"Image": models, code, and papers
Towards localisation of keywords in speech using weak supervision
Dec 14, 2020Developments in weakly supervised and self-supervised models could enable speech technology in low-resource settings where full transcriptions are not available. We consider whether keyword localisation is possible using two forms of weak supervision where location information is not provided explicitly. In the first, only the presence or absence of a word is indicated, i.e. a bag-of-words (BoW) labelling. In the second, visual context is provided in the form of an image paired with an unlabelled utterance; a model then needs to be trained in a self-supervised fashion using the paired data. For keyword localisation, we adapt a saliency-based method typically used in the vision domain. We compare this to an existing technique that performs localisation as a part of the network architecture. While the saliency-based method is more flexible (it can be applied without architectural restrictions), we identify a critical limitation when using it for keyword localisation. Of the two forms of supervision, the visually trained model performs worse than the BoW-trained model. We show qualitatively that the visually trained model sometimes locate semantically related words, but this is not consistent. While our results show that there is some signal allowing for localisation, it also calls for other localisation methods better matched to these forms of weak supervision.
Born Identity Network: Multi-way Counterfactual Map Generation to Explain a Classifier's Decision
Nov 24, 2020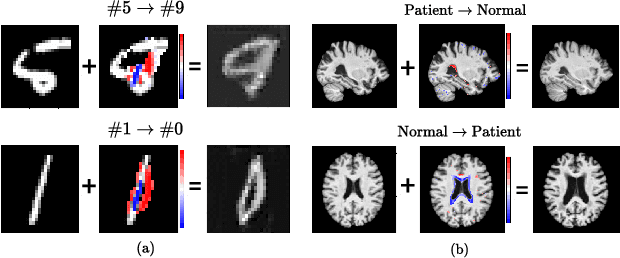
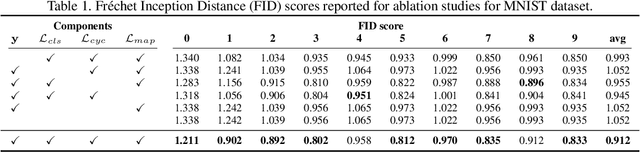
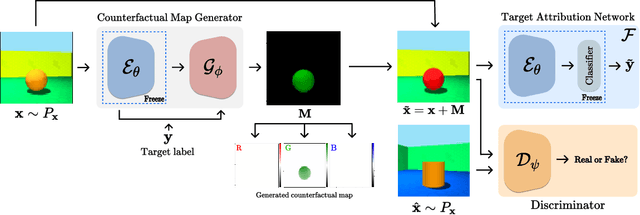
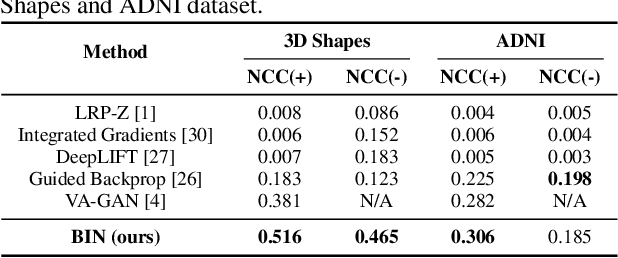
There exists an apparent negative correlation between performance and interpretability of deep learning models. In an effort to reduce this negative correlation, we propose Born Identity Network (BIN), which is a post-hoc approach for producing multi-way counterfactual maps. A counterfactual map transforms an input sample to be classified as a target label, which is similar to how humans process knowledge through counterfactual thinking. Thus, producing a better counterfactual map may be a step towards explanation at the level of human knowledge. For example, a counterfactual map can localize hypothetical abnormalities from a normal brain image that may cause it to be diagnosed with a disease. Specifically, our proposed BIN consists of two core components: Counterfactual Map Generator and Target Attribution Network. The Counterfactual Map Generator is a variation of conditional GAN which can synthesize a counterfactual map conditioned on an arbitrary target label. The Target Attribution Network works in a complementary manner to enforce target label attribution to the synthesized map. We have validated our proposed BIN in qualitative, quantitative analysis on MNIST, 3D Shapes, and ADNI datasets, and show the comprehensibility and fidelity of our method from various ablation studies.
Phase Retrieval with Holography and Untrained Priors: Tackling the Challenges of Low-Photon Nanoscale Imaging
Dec 14, 2020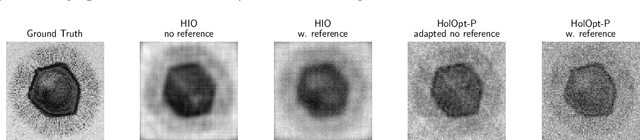
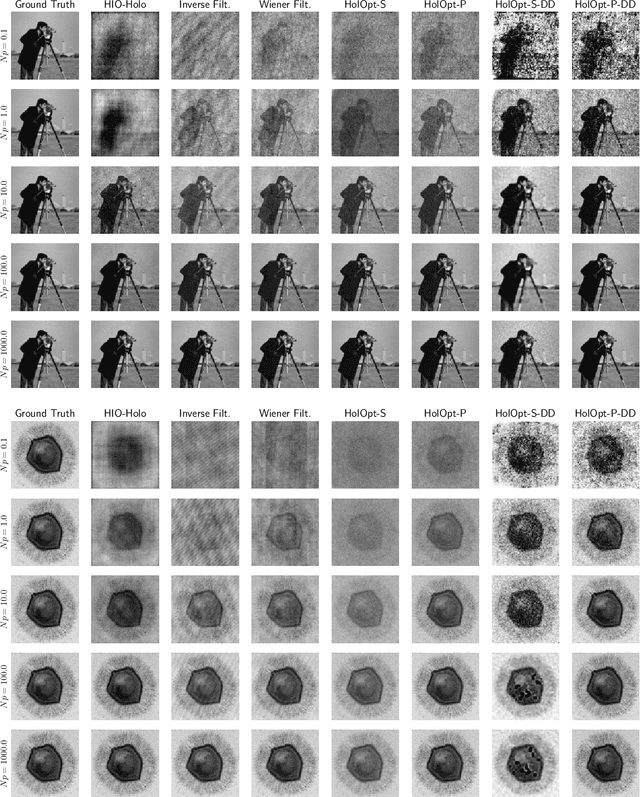
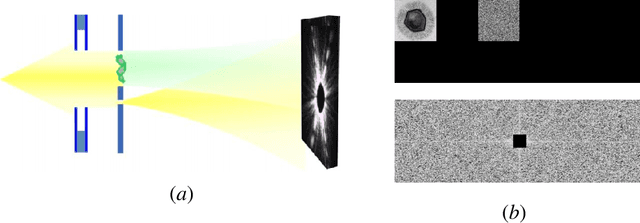
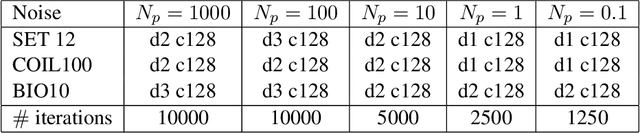


Phase retrieval is the inverse problem of recovering a signal from magnitude-only Fourier measurements, and underlies numerous imaging modalities, such as Coherent Diffraction Imaging (CDI). A variant of this setup, known as holography, includes a reference object that is placed adjacent to the specimen of interest before measurements are collected. The resulting inverse problem, known as holographic phase retrieval, is well-known to have improved problem conditioning relative to the original. This innovation, i.e. Holographic CDI, becomes crucial at the nanoscale, where imaging specimens such as viruses, proteins, and crystals require low-photon measurements. This data is highly corrupted by Poisson shot noise, and often lacks low-frequency content as well. In this work, we introduce a dataset-free deep learning framework for holographic phase retrieval adapted to these challenges. The key ingredients of our approach are the explicit and flexible incorporation of the physical forward model into an automatic differentiation procedure, the Poisson log-likelihood objective function, and an optional untrained deep image prior. We perform extensive evaluation under realistic conditions. Compared to competing classical methods, our method recovers signal from higher noise levels and is more resilient to suboptimal reference design, as well as to large missing regions of low frequencies in the observations. To the best of our knowledge, this is the first work to consider a dataset-free machine learning approach for holographic phase retrieval.
Dense Pixel-wise Micro-motion Estimation of Object Surface by using Low Dimensional Embedding of Laser Speckle Pattern
Oct 31, 2020
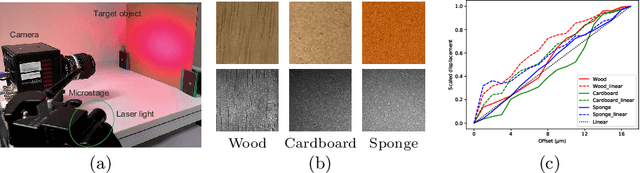
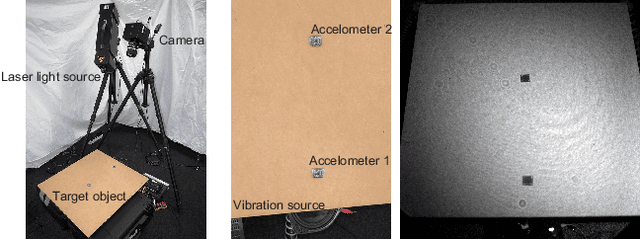
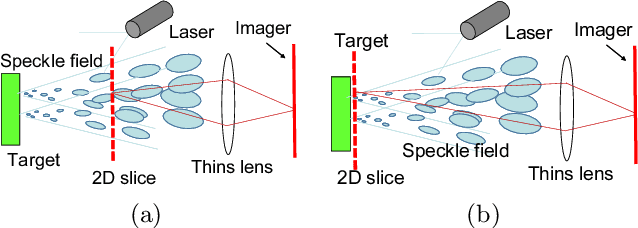


This paper proposes a method of estimating micro-motion of an object at each pixel that is too small to detect under a common setup of camera and illumination. The method introduces an active-lighting approach to make the motion visually detectable. The approach is based on speckle pattern, which is produced by the mutual interference of laser light on object's surface and continuously changes its appearance according to the out-of-plane motion of the surface. In addition, speckle pattern becomes uncorrelated with large motion. To compensate such micro- and large motion, the method estimates the motion parameters up to scale at each pixel by nonlinear embedding of the speckle pattern into low-dimensional space. The out-of-plane motion is calculated by making the motion parameters spatially consistent across the image. In the experiments, the proposed method is compared with other measuring devices to prove the effectiveness of the method.
HR-Depth: High Resolution Self-Supervised Monocular Depth Estimation
Dec 14, 2020
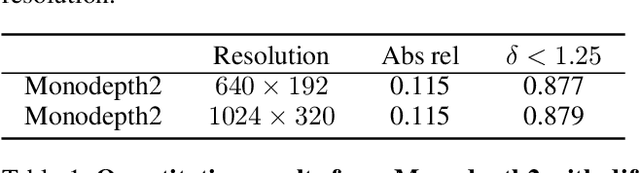
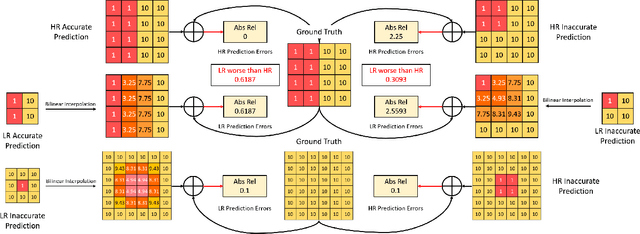
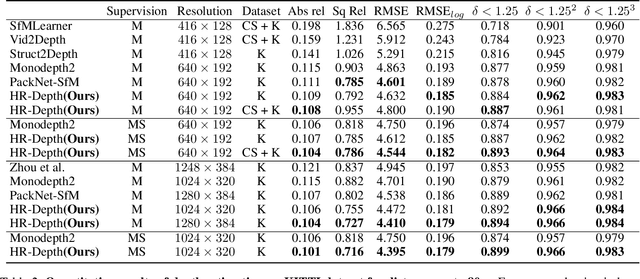
Self-supervised learning shows great potential in monoculardepth estimation, using image sequences as the only source ofsupervision. Although people try to use the high-resolutionimage for depth estimation, the accuracy of prediction hasnot been significantly improved. In this work, we find thecore reason comes from the inaccurate depth estimation inlarge gradient regions, making the bilinear interpolation er-ror gradually disappear as the resolution increases. To obtainmore accurate depth estimation in large gradient regions, itis necessary to obtain high-resolution features with spatialand semantic information. Therefore, we present an improvedDepthNet, HR-Depth, with two effective strategies: (1) re-design the skip-connection in DepthNet to get better high-resolution features and (2) propose feature fusion Squeeze-and-Excitation(fSE) module to fuse feature more efficiently.Using Resnet-18 as the encoder, HR-Depth surpasses all pre-vious state-of-the-art(SoTA) methods with the least param-eters at both high and low resolution. Moreover, previousstate-of-the-art methods are based on fairly complex and deepnetworks with a mass of parameters which limits their realapplications. Thus we also construct a lightweight networkwhich uses MobileNetV3 as encoder. Experiments show thatthe lightweight network can perform on par with many largemodels like Monodepth2 at high-resolution with only20%parameters. All codes and models will be available at https://github.com/shawLyu/HR-Depth.
Hyperspectral Unmixing via Nonnegative Matrix Factorization with Handcrafted and Learnt Priors
Oct 09, 2020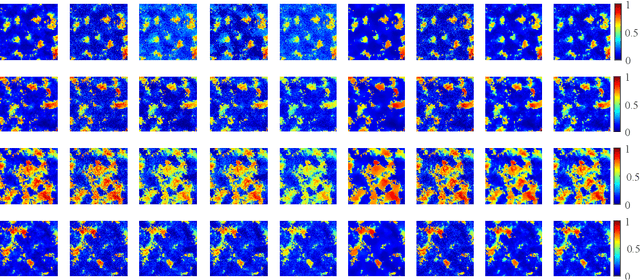
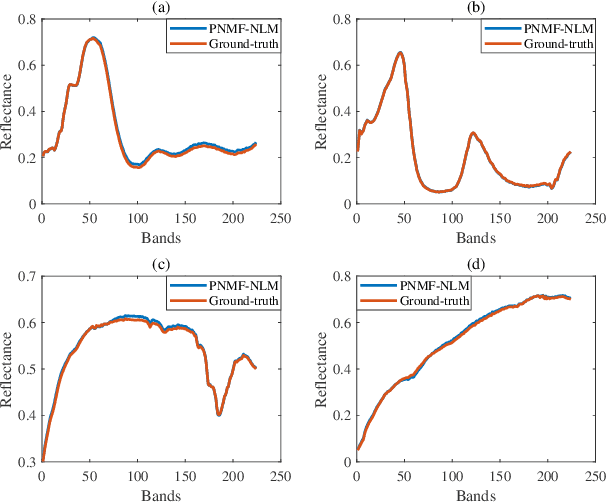
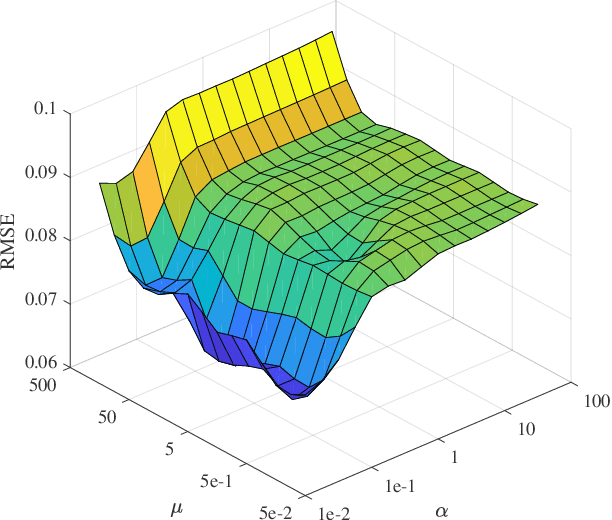
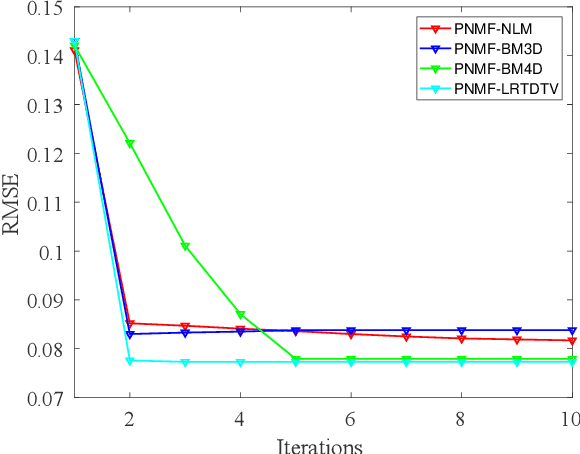
Nowadays, nonnegative matrix factorization (NMF) based methods have been widely applied to blind spectral unmixing. Introducing proper regularizers to NMF is crucial for mathematically constraining the solutions and physically exploiting spectral and spatial properties of images. Generally, properly handcrafting regularizers and solving the associated complex optimization problem are non-trivial tasks. In our work, we propose an NMF based unmixing framework which jointly uses a handcrafting regularizer and a learnt regularizer from data. we plug learnt priors of abundances where the associated subproblem can be addressed using various image denoisers, and we consider an l_2,1-norm regularizer to the abundance matrix to promote sparse unmixing results. The proposed framework is flexible and extendable. Both synthetic data and real airborne data are conducted to confirm the effectiveness of our method.
Neural Star Domain as Primitive Representation
Nov 12, 2020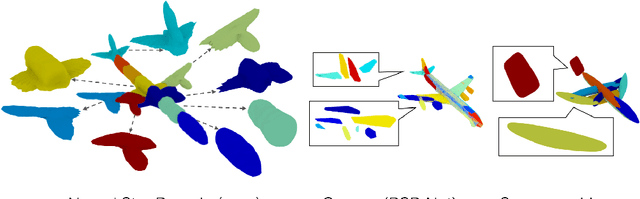
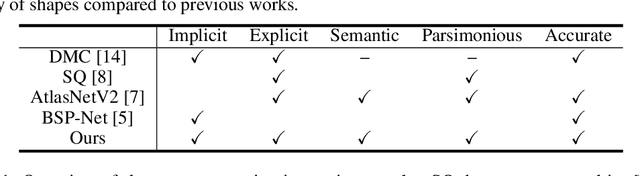

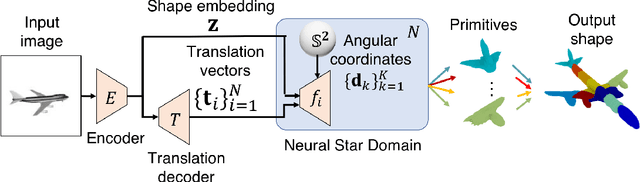
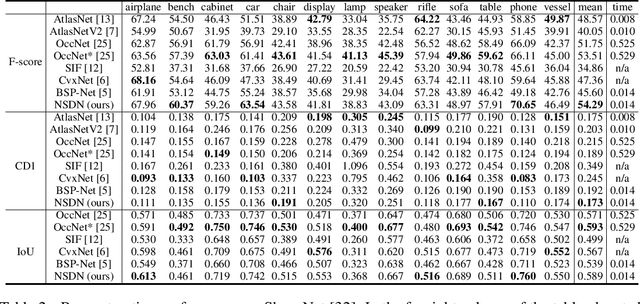
Reconstructing 3D objects from 2D images is a fundamental task in computer vision. Accurate structured reconstruction by parsimonious and semantic primitive representation further broadens its application. When reconstructing a target shape with multiple primitives, it is preferable that one can instantly access the union of basic properties of the shape such as collective volume and surface, treating the primitives as if they are one single shape. This becomes possible by primitive representation with unified implicit and explicit representations. However, primitive representations in current approaches do not satisfy all of the above requirements at the same time. To solve this problem, we propose a novel primitive representation named neural star domain (NSD) that learns primitive shapes in the star domain. We show that NSD is a universal approximator of the star domain and is not only parsimonious and semantic but also an implicit and explicit shape representation. We demonstrate that our approach outperforms existing methods in image reconstruction tasks, semantic capabilities, and speed and quality of sampling high-resolution meshes.
From Boundaries to Bumps: when closed (extremal) contours are critical
May 16, 2020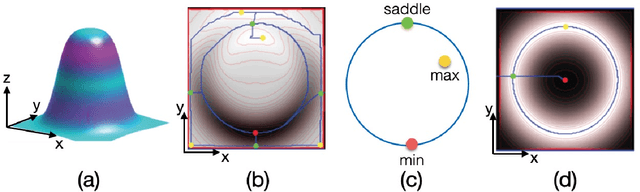
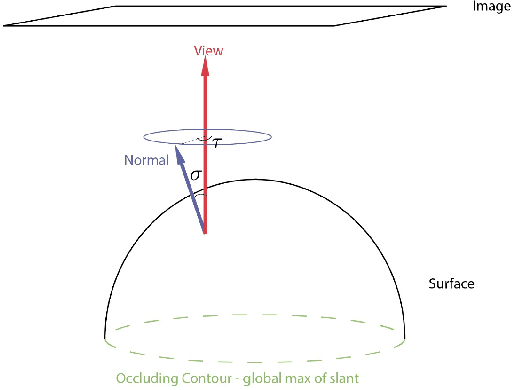
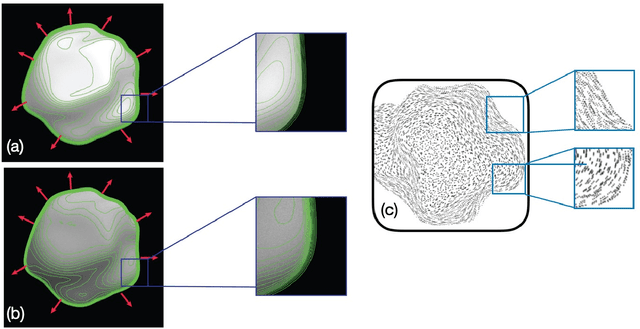
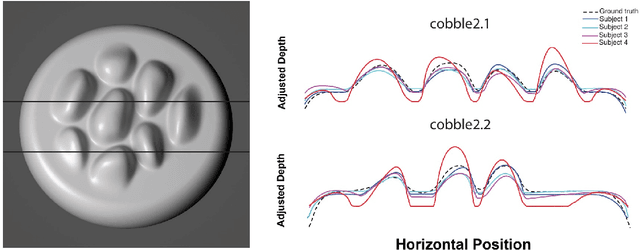
Invariants underlying shape inference are elusive: a variety of shapes can give rise to the same image, and a variety of images can be rendered from the same shape. The occluding contour is a rare exception: it has both image salience, in terms of isophotes, and surface meaning, in terms of surface normal. We relax the notion of occluding contour to define closed extremal curves, a new shape invariant that exists at the topological level. They surround bumps, a common but ill-specified interior shape component, and formalize the qualitative nature of bump perception. Extremal curves are biologically computable, unify shape inferences from shading, texture, and specular materials, and predict new phenomena in bump perception.
Comparisons among different stochastic selection of activation layers for convolutional neural networks for healthcare
Nov 24, 2020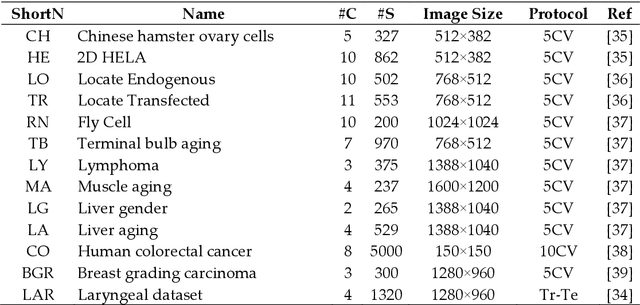

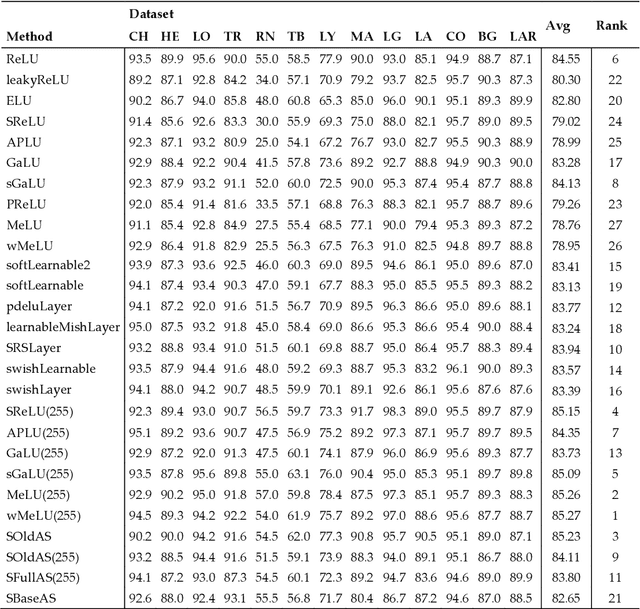
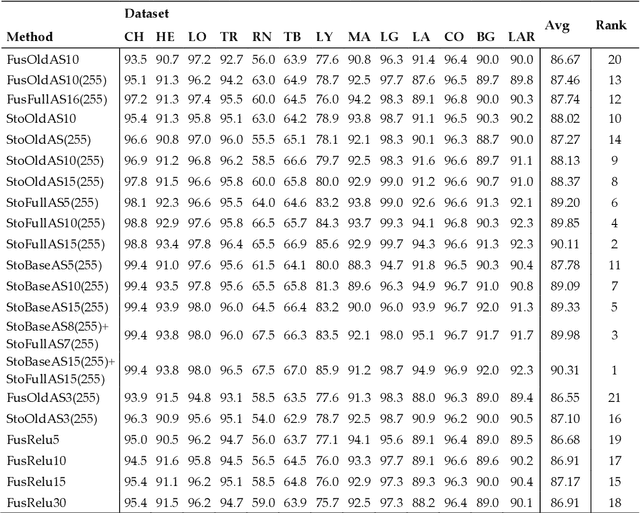
Classification of biological images is an important task with crucial application in many fields, such as cell phenotypes recognition, detection of cell organelles and histopathological classification, and it might help in early medical diagnosis, allowing automatic disease classification without the need of a human expert. In this paper we classify biomedical images using ensembles of neural networks. We create this ensemble using a ResNet50 architecture and modifying its activation layers by substituting ReLUs with other functions. We select our activations among the following ones: ReLU, leaky ReLU, Parametric ReLU, ELU, Adaptive Piecewice Linear Unit, S-Shaped ReLU, Swish , Mish, Mexican Linear Unit, Gaussian Linear Unit, Parametric Deformable Linear Unit, Soft Root Sign (SRS) and others. As a baseline, we used an ensemble of neural networks that only use ReLU activations. We tested our networks on several small and medium sized biomedical image datasets. Our results prove that our best ensemble obtains a better performance than the ones of the naive approaches. In order to encourage the reproducibility of this work, the MATLAB code of all the experiments will be shared at https://github.com/LorisNanni.
CPR-GCN: Conditional Partial-Residual Graph Convolutional Network in Automated Anatomical Labeling of Coronary Arteries
Apr 18, 2020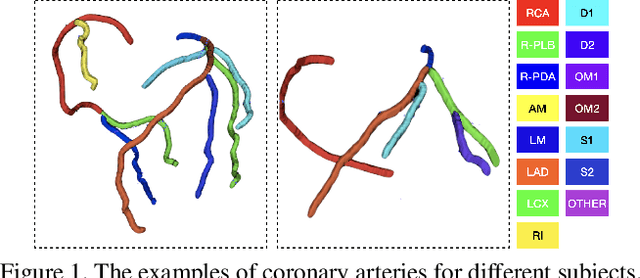
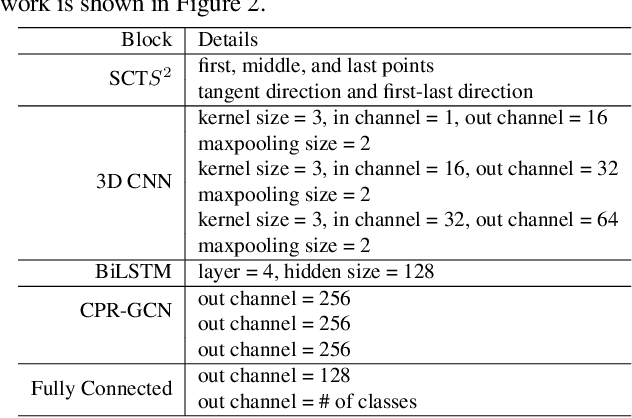
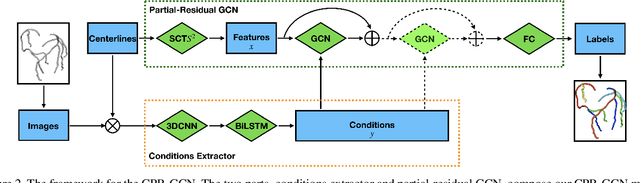
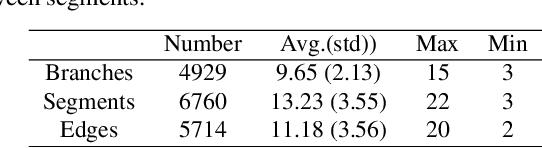
Automated anatomical labeling plays a vital role in coronary artery disease diagnosing procedure. The main challenge in this problem is the large individual variability inherited in human anatomy. Existing methods usually rely on the position information and the prior knowledge of the topology of the coronary artery tree, which may lead to unsatisfactory performance when the main branches are confusing. Motivated by the wide application of the graph neural network in structured data, in this paper, we propose a conditional partial-residual graph convolutional network (CPR-GCN), which takes both position and CT image into consideration, since CT image contains abundant information such as branch size and spanning direction. Two majority parts, a Partial-Residual GCN and a conditions extractor, are included in CPR-GCN. The conditions extractor is a hybrid model containing the 3D CNN and the LSTM, which can extract 3D spatial image features along the branches. On the technical side, the Partial-Residual GCN takes the position features of the branches, with the 3D spatial image features as conditions, to predict the label for each branches. While on the mathematical side, our approach twists the partial differential equation (PDE) into the graph modeling. A dataset with 511 subjects is collected from the clinic and annotated by two experts with a two-phase annotation process. According to the five-fold cross-validation, our CPR-GCN yields 95.8% meanRecall, 95.4% meanPrecision and 0.955 meanF1, which outperforms state-of-the-art approaches.
* This work is done by Xingjian Zhen during internship in Alibaba Damo Academy
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge