"Image": models, code, and papers
Isometric Propagation Network for Generalized Zero-shot Learning
Feb 03, 2021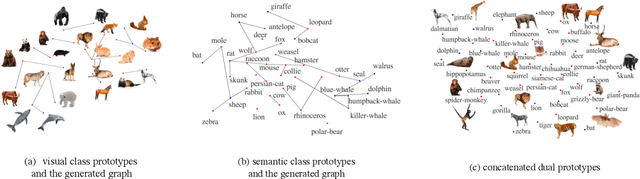
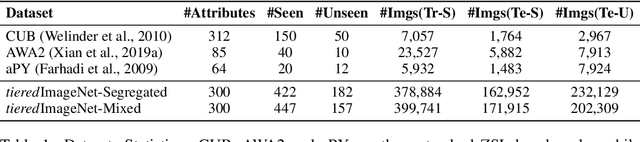
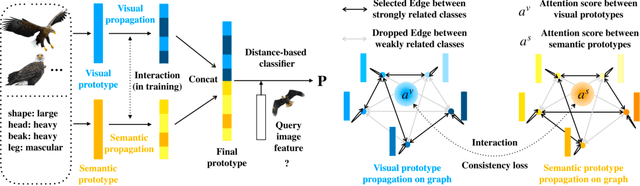
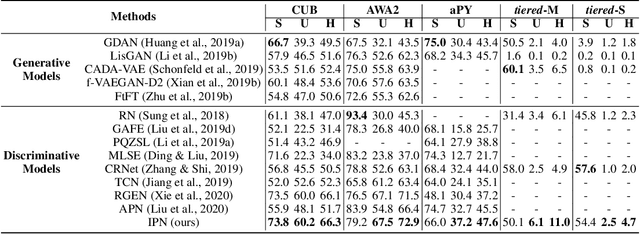
Zero-shot learning (ZSL) aims to classify images of an unseen class only based on a few attributes describing that class but no access to any training sample. A popular strategy is to learn a mapping between the semantic space of class attributes and the visual space of images based on the seen classes and their data. Thus, an unseen class image can be ideally mapped to its corresponding class attributes. The key challenge is how to align the representations in the two spaces. For most ZSL settings, the attributes for each seen/unseen class are only represented by a vector while the seen-class data provide much more information. Thus, the imbalanced supervision from the semantic and the visual space can make the learned mapping easily overfitting to the seen classes. To resolve this problem, we propose Isometric Propagation Network (IPN), which learns to strengthen the relation between classes within each space and align the class dependency in the two spaces. Specifically, IPN learns to propagate the class representations on an auto-generated graph within each space. In contrast to only aligning the resulted static representation, we regularize the two dynamic propagation procedures to be isometric in terms of the two graphs' edge weights per step by minimizing a consistency loss between them. IPN achieves state-of-the-art performance on three popular ZSL benchmarks. To evaluate the generalization capability of IPN, we further build two larger benchmarks with more diverse unseen classes and demonstrate the advantages of IPN on them.
Graph-Based Social Relation Reasoning
Jul 17, 2020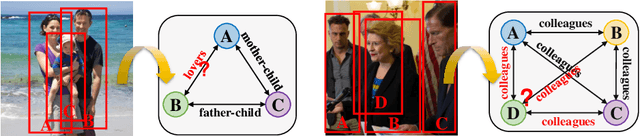
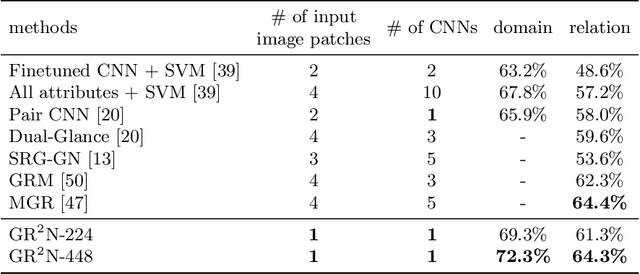
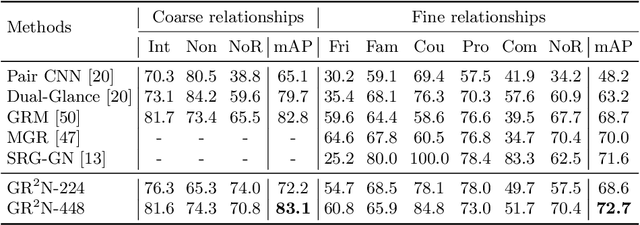
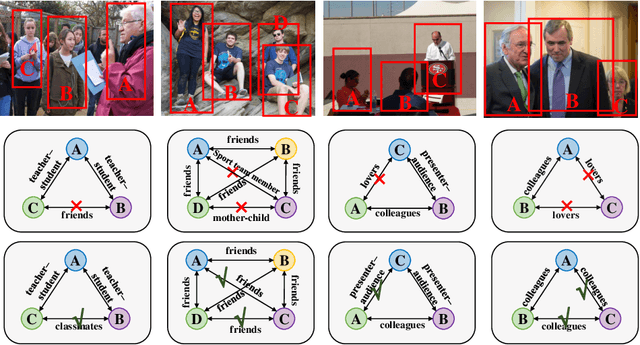
Human beings are fundamentally sociable -- that we generally organize our social lives in terms of relations with other people. Understanding social relations from an image has great potential for intelligent systems such as social chatbots and personal assistants. In this paper, we propose a simpler, faster, and more accurate method named graph relational reasoning network (GR2N) for social relation recognition. Different from existing methods which process all social relations on an image independently, our method considers the paradigm of jointly inferring the relations by constructing a social relation graph. Furthermore, the proposed GR2N constructs several virtual relation graphs to explicitly grasp the strong logical constraints among different types of social relations. Experimental results illustrate that our method generates a reasonable and consistent social relation graph and improves the performance in both accuracy and efficiency.
Selecting Regions of Interest in Large Multi-Scale Images for Cancer Pathology
Jul 03, 2020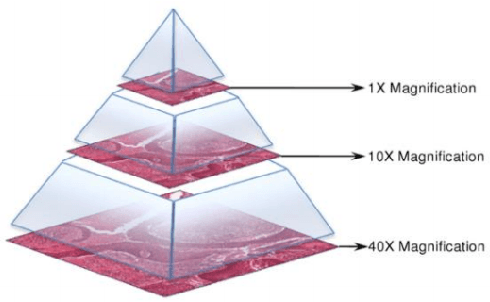
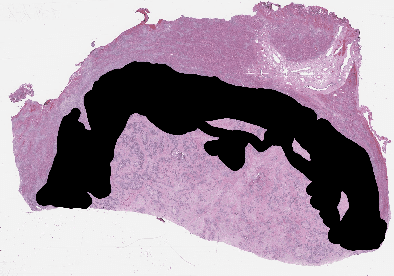
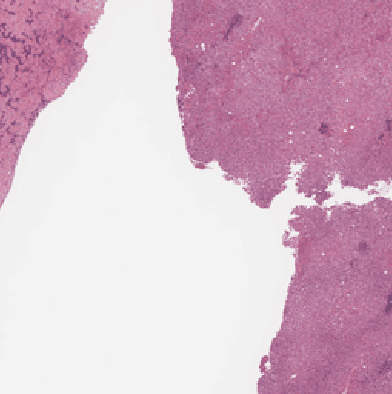
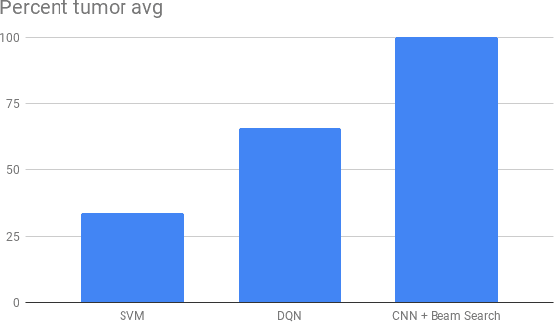
Recent breakthroughs in object detection and image classification using Convolutional Neural Networks (CNNs) are revolutionizing the state of the art in medical imaging, and microscopy in particular presents abundant opportunities for computer vision algorithms to assist medical professionals in diagnosis of diseases ranging from malaria to cancer. High resolution scans of microscopy slides called Whole Slide Images (WSIs) offer enough information for a cancer pathologist to come to a conclusion regarding cancer presence, subtype, and severity based on measurements of features within the slide image at multiple scales and resolutions. WSIs' extremely high resolutions and feature scales ranging from gross anatomical structures down to cell nuclei preclude the use of standard CNN models for object detection and classification, which have typically been designed for images with dimensions in the hundreds of pixels and with objects on the order of the size of the image itself. We explore parallel approaches based on Reinforcement Learning and Beam Search to learn to progressively zoom into the WSI to detect Regions of Interest (ROIs) in liver pathology slides containing one of two types of liver cancer, namely Hepatocellular Carcinoma (HCC) and Cholangiocarcinoma (CC). These ROIs can then be presented directly to the pathologist to aid in measurement and diagnosis or be used for automated classification of tumor subtype.
On Deep Learning Techniques to Boost Monocular Depth Estimation for Autonomous Navigation
Oct 13, 2020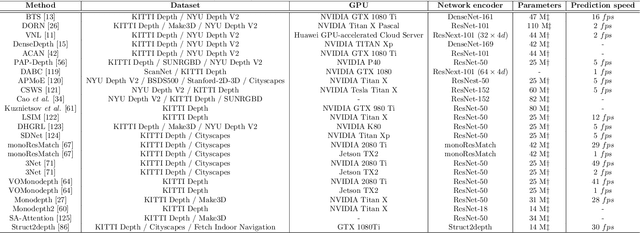
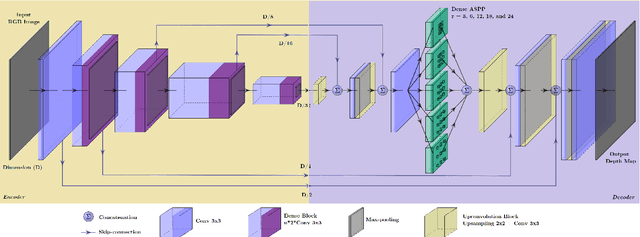
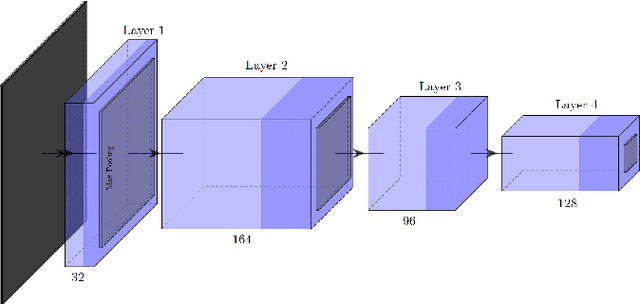
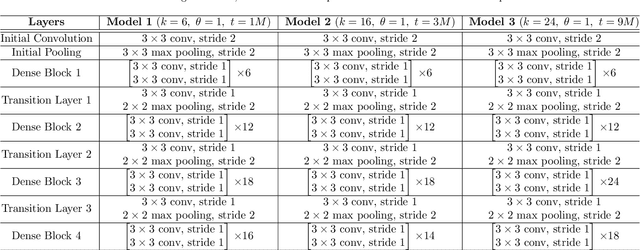
Inferring the depth of images is a fundamental inverse problem within the field of Computer Vision since depth information is obtained through 2D images, which can be generated from infinite possibilities of observed real scenes. Benefiting from the progress of Convolutional Neural Networks (CNNs) to explore structural features and spatial image information, Single Image Depth Estimation (SIDE) is often highlighted in scopes of scientific and technological innovation, as this concept provides advantages related to its low implementation cost and robustness to environmental conditions. In the context of autonomous vehicles, state-of-the-art CNNs optimize the SIDE task by producing high-quality depth maps, which are essential during the autonomous navigation process in different locations. However, such networks are usually supervised by sparse and noisy depth data, from Light Detection and Ranging (LiDAR) laser scans, and are carried out at high computational cost, requiring high-performance Graphic Processing Units (GPUs). Therefore, we propose a new lightweight and fast supervised CNN architecture combined with novel feature extraction models which are designed for real-world autonomous navigation. We also introduce an efficient surface normals module, jointly with a simple geometric 2.5D loss function, to solve SIDE problems. We also innovate by incorporating multiple Deep Learning techniques, such as the use of densification algorithms and additional semantic, surface normals and depth information to train our framework. The method introduced in this work focuses on robotic applications in indoor and outdoor environments and its results are evaluated on the competitive and publicly available NYU Depth V2 and KITTI Depth datasets.
MeDaS: An open-source platform as service to help break the walls between medicine and informatics
Jul 14, 2020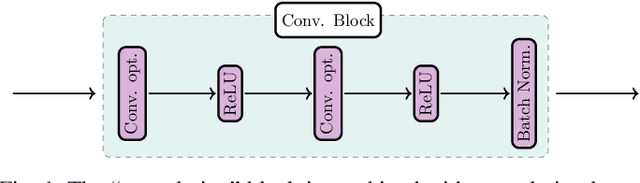
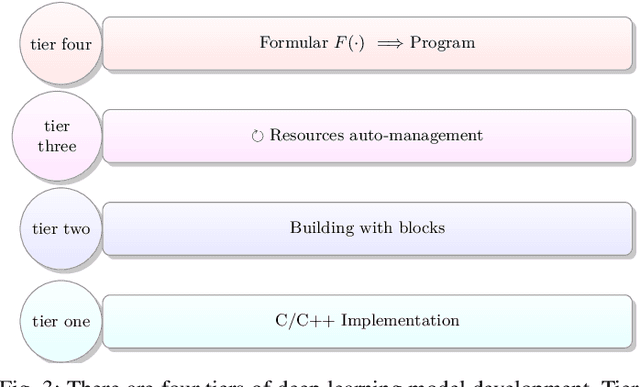


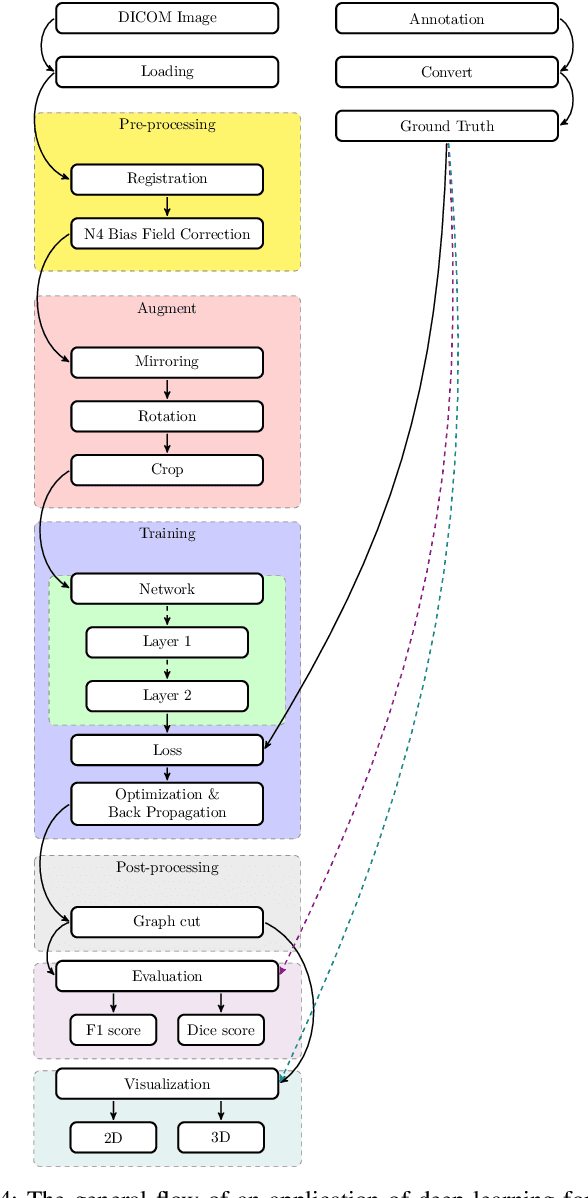
In the past decade, deep learning (DL) has achieved unprecedented success in numerous fields including computer vision, natural language processing, and healthcare. In particular, DL is experiencing an increasing development in applications for advanced medical image analysis in terms of analysis, segmentation, classification, and furthermore. On the one hand, tremendous needs that leverage the power of DL for medical image analysis are arising from the research community of a medical, clinical, and informatics background to jointly share their expertise, knowledge, skills, and experience. On the other hand, barriers between disciplines are on the road for them often hampering a full and efficient collaboration. To this end, we propose our novel open-source platform, i.e., MeDaS -- the MeDical open-source platform as Service. To the best of our knowledge, MeDaS is the first open-source platform proving a collaborative and interactive service for researchers from a medical background easily using DL related toolkits, and at the same time for scientists or engineers from information sciences to understand the medical knowledge side. Based on a series of toolkits and utilities from the idea of RINV (Rapid Implementation aNd Verification), our proposed MeDaS platform can implement pre-processing, post-processing, augmentation, visualization, and other phases needed in medical image analysis. Five tasks including the subjects of lung, liver, brain, chest, and pathology, are validated and demonstrated to be efficiently realisable by using MeDaS.
Image Generation and Editing with Variational Info Generative AdversarialNetworks
Jan 17, 2017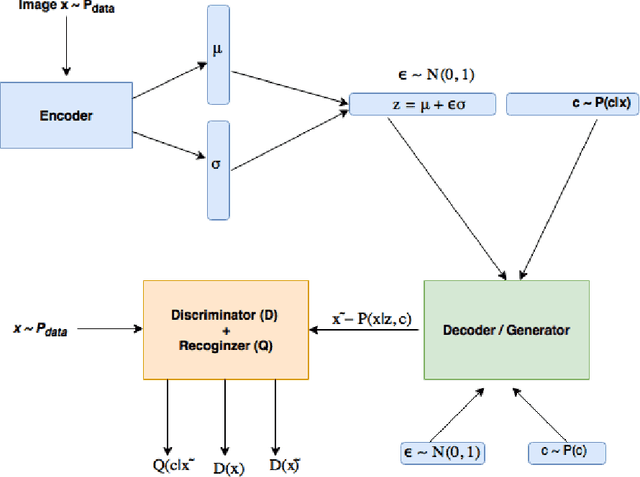
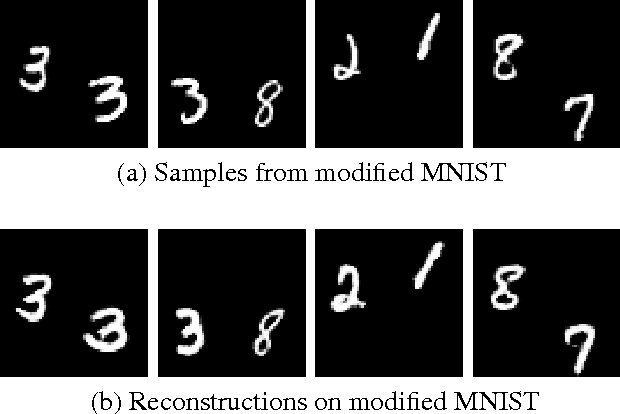

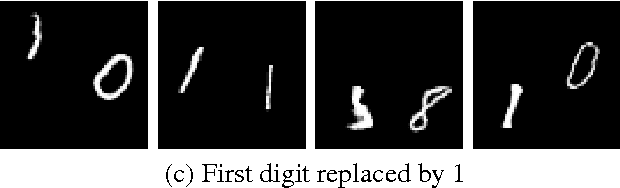
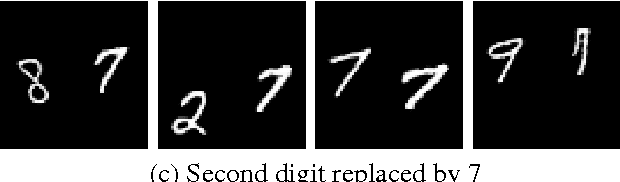
Recently there has been an enormous interest in generative models for images in deep learning. In pursuit of this, Generative Adversarial Networks (GAN) and Variational Auto-Encoder (VAE) have surfaced as two most prominent and popular models. While VAEs tend to produce excellent reconstructions but blurry samples, GANs generate sharp but slightly distorted images. In this paper we propose a new model called Variational InfoGAN (ViGAN). Our aim is two fold: (i) To generated new images conditioned on visual descriptions, and (ii) modify the image, by fixing the latent representation of image and varying the visual description. We evaluate our model on Labeled Faces in the Wild (LFW), celebA and a modified version of MNIST datasets and demonstrate the ability of our model to generate new images as well as to modify a given image by changing attributes.
Rethinking the Form of Latent States in Image Captioning
Jul 26, 2018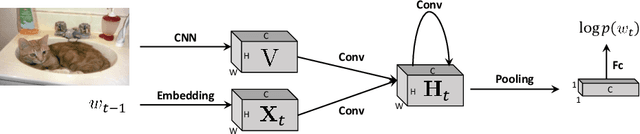
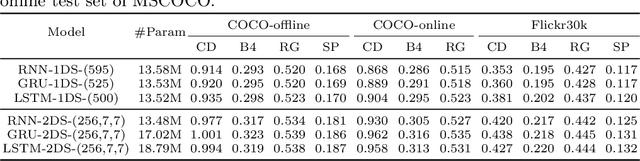


RNNs and their variants have been widely adopted for image captioning. In RNNs, the production of a caption is driven by a sequence of latent states. Existing captioning models usually represent latent states as vectors, taking this practice for granted. We rethink this choice and study an alternative formulation, namely using two-dimensional maps to encode latent states. This is motivated by the curiosity about a question: how the spatial structures in the latent states affect the resultant captions? Our study on MSCOCO and Flickr30k leads to two significant observations. First, the formulation with 2D states is generally more effective in captioning, consistently achieving higher performance with comparable parameter sizes. Second, 2D states preserve spatial locality. Taking advantage of this, we visually reveal the internal dynamics in the process of caption generation, as well as the connections between input visual domain and output linguistic domain.
Unified Depth Prediction and Intrinsic Image Decomposition from a Single Image via Joint Convolutional Neural Fields
Mar 21, 2016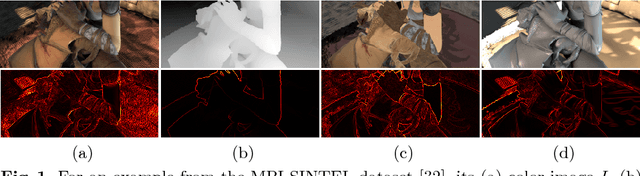
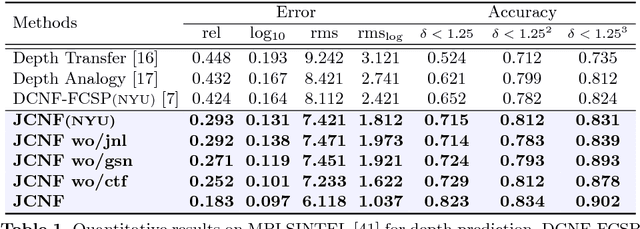
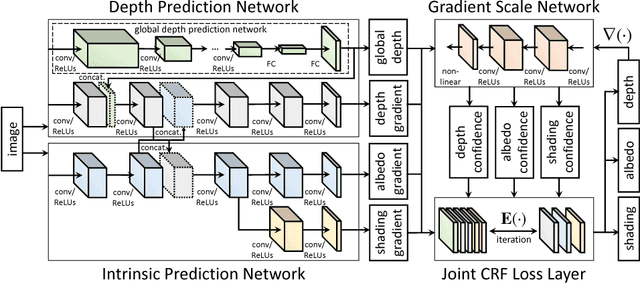
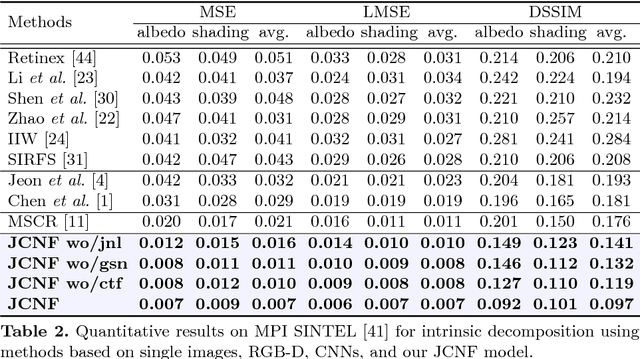
We present a method for jointly predicting a depth map and intrinsic images from single-image input. The two tasks are formulated in a synergistic manner through a joint conditional random field (CRF) that is solved using a novel convolutional neural network (CNN) architecture, called the joint convolutional neural field (JCNF) model. Tailored to our joint estimation problem, JCNF differs from previous CNNs in its sharing of convolutional activations and layers between networks for each task, its inference in the gradient domain where there exists greater correlation between depth and intrinsic images, and the incorporation of a gradient scale network that learns the confidence of estimated gradients in order to effectively balance them in the solution. This approach is shown to surpass state-of-the-art methods both on single-image depth estimation and on intrinsic image decomposition.
RAILS: A Robust Adversarial Immune-inspired Learning System
Dec 18, 2020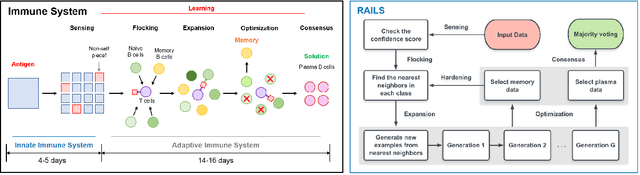
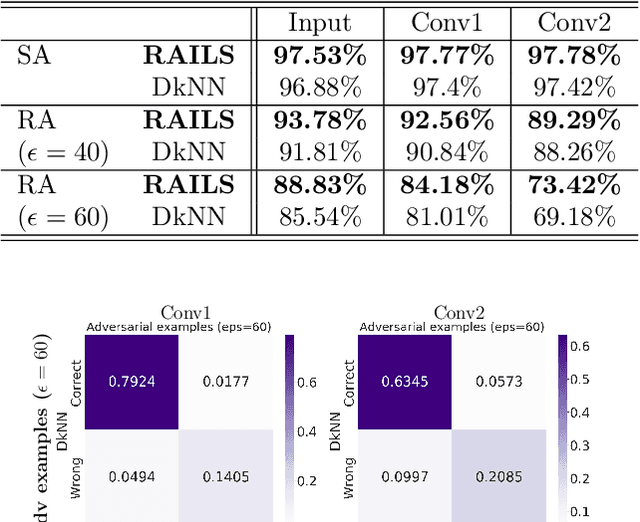
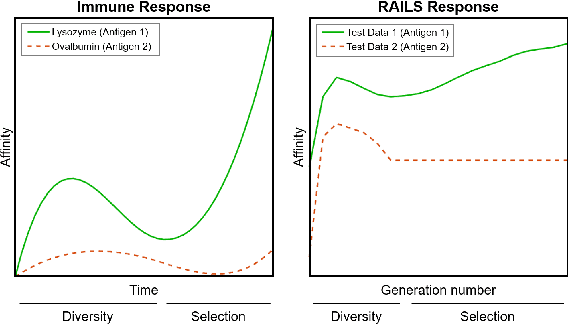
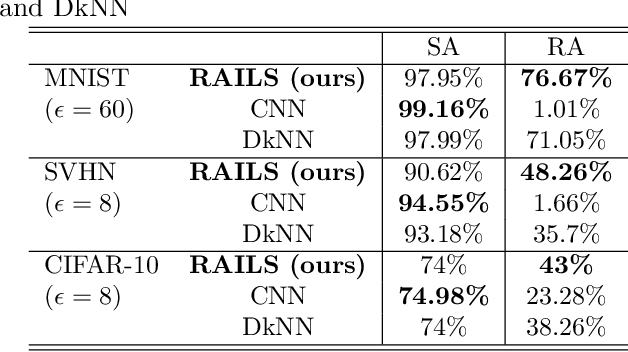
Adversarial attacks against deep neural networks are continuously evolving. Without effective defenses, they can lead to catastrophic failure. The long-standing and arguably most powerful natural defense system is the mammalian immune system, which has successfully defended against attacks by novel pathogens for millions of years. In this paper, we propose a new adversarial defense framework, called the Robust Adversarial Immune-inspired Learning System (RAILS). RAILS incorporates an Adaptive Immune System Emulation (AISE), which emulates in silico the biological mechanisms that are used to defend the host against attacks by pathogens. We use RAILS to harden Deep k-Nearest Neighbor (DkNN) architectures against evasion attacks. Evolutionary programming is used to simulate processes in the natural immune system: B-cell flocking, clonal expansion, and affinity maturation. We show that the RAILS learning curve exhibits similar diversity-selection learning phases as observed in our in vitro biological experiments. When applied to adversarial image classification on three different datasets, RAILS delivers an additional 5.62%/12.56%/4.74% robustness improvement as compared to applying DkNN alone, without appreciable loss of accuracy on clean data.
Minimax Active Learning
Dec 18, 2020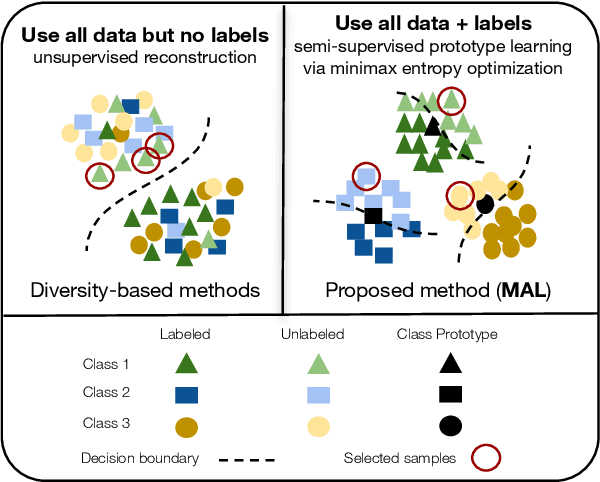
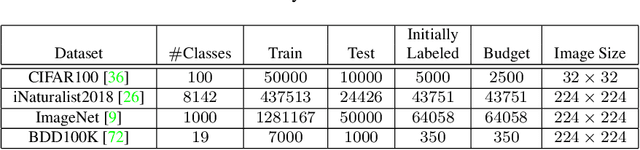
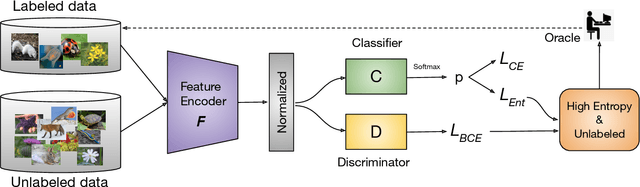

Active learning aims to develop label-efficient algorithms by querying the most representative samples to be labeled by a human annotator. Current active learning techniques either rely on model uncertainty to select the most uncertain samples or use clustering or reconstruction to choose the most diverse set of unlabeled examples. While uncertainty-based strategies are susceptible to outliers, solely relying on sample diversity does not capture the information available on the main task. In this work, we develop a semi-supervised minimax entropy-based active learning algorithm that leverages both uncertainty and diversity in an adversarial manner. Our model consists of an entropy minimizing feature encoding network followed by an entropy maximizing classification layer. This minimax formulation reduces the distribution gap between the labeled/unlabeled data, while a discriminator is simultaneously trained to distinguish the labeled/unlabeled data. The highest entropy samples from the classifier that the discriminator predicts as unlabeled are selected for labeling. We extensively evaluate our method on various image classification and semantic segmentation benchmark datasets and show superior performance over the state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge