"Image": models, code, and papers
Dataset on Bi- and Multi-Nucleated Tumor Cells in Canine Cutaneous Mast Cell Tumors
Jan 05, 2021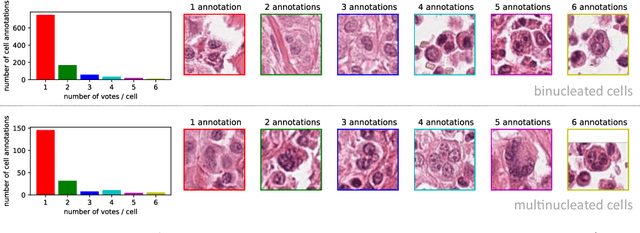
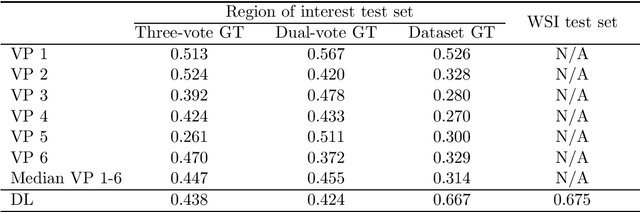
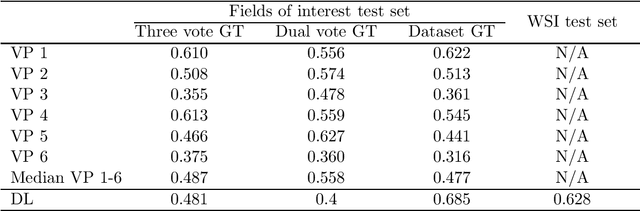
Tumor cells with two nuclei (binucleated cells, BiNC) or more nuclei (multinucleated cells, MuNC) indicate an increased amount of cellular genetic material which is thought to facilitate oncogenesis, tumor progression and treatment resistance. In canine cutaneous mast cell tumors (ccMCT), binucleation and multinucleation are parameters used in cytologic and histologic grading schemes (respectively) which correlate with poor patient outcome. For this study, we created the first open source data-set with 19,983 annotations of BiNC and 1,416 annotations of MuNC in 32 histological whole slide images of ccMCT. Labels were created by a pathologist and an algorithmic-aided labeling approach with expert review of each generated candidate. A state-of-the-art deep learning-based model yielded an $F_1$ score of 0.675 for BiNC and 0.623 for MuNC on 11 test whole slide images. In regions of interest ($2.37 mm^2$) extracted from these test images, 6 pathologists had an object detection performance between 0.270 - 0.526 for BiNC and 0.316 - 0.622 for MuNC, while our model archived an $F_1$ score of 0.667 for BiNC and 0.685 for MuNC. This open dataset can facilitate development of automated image analysis for this task and may thereby help to promote standardization of this facet of histologic tumor prognostication.
Automated cardiovascular magnetic resonance image analysis with fully convolutional networks
May 22, 2018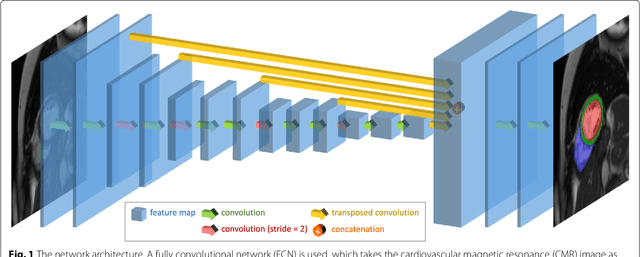

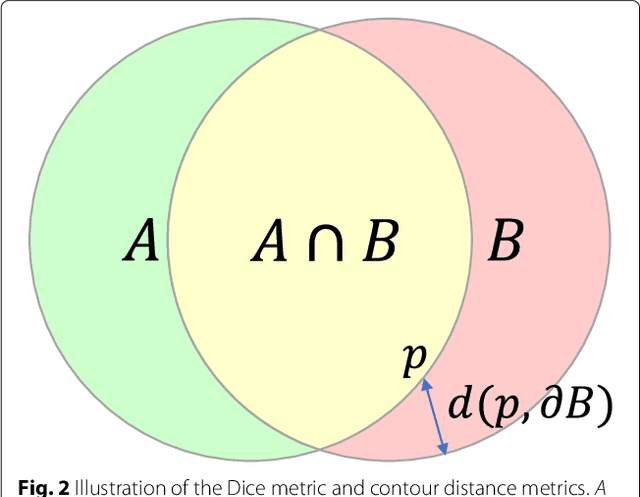
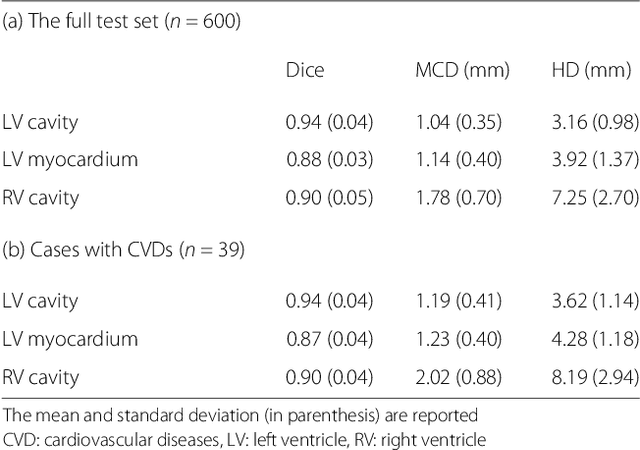
Cardiovascular magnetic resonance (CMR) imaging is a standard imaging modality for assessing cardiovascular diseases (CVDs), the leading cause of death globally. CMR enables accurate quantification of the cardiac chamber volume, ejection fraction and myocardial mass, providing information for diagnosis and monitoring of CVDs. However, for years, clinicians have been relying on manual approaches for CMR image analysis, which is time consuming and prone to subjective errors. It is a major clinical challenge to automatically derive quantitative and clinically relevant information from CMR images. Deep neural networks have shown a great potential in image pattern recognition and segmentation for a variety of tasks. Here we demonstrate an automated analysis method for CMR images, which is based on a fully convolutional network (FCN). The network is trained and evaluated on a large-scale dataset from the UK Biobank, consisting of 4,875 subjects with 93,500 pixelwise annotated images. The performance of the method has been evaluated using a number of technical metrics, including the Dice metric, mean contour distance and Hausdorff distance, as well as clinically relevant measures, including left ventricle (LV) end-diastolic volume (LVEDV) and end-systolic volume (LVESV), LV mass (LVM); right ventricle (RV) end-diastolic volume (RVEDV) and end-systolic volume (RVESV). By combining FCN with a large-scale annotated dataset, the proposed automated method achieves a high performance on par with human experts in segmenting the LV and RV on short-axis CMR images and the left atrium (LA) and right atrium (RA) on long-axis CMR images.
Going Deeper with Contextual CNN for Hyperspectral Image Classification
May 09, 2017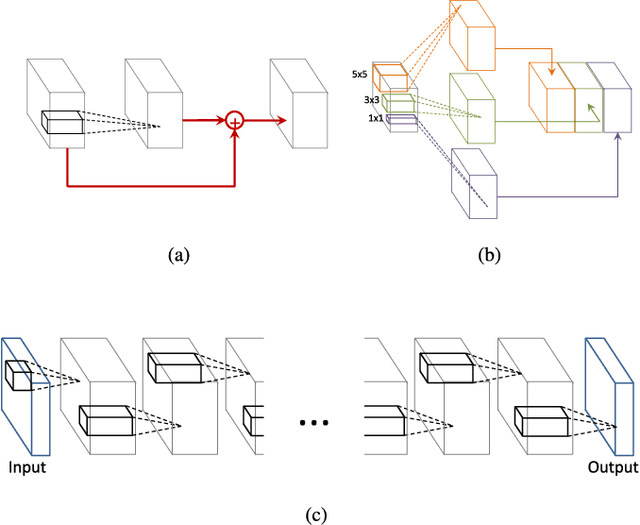
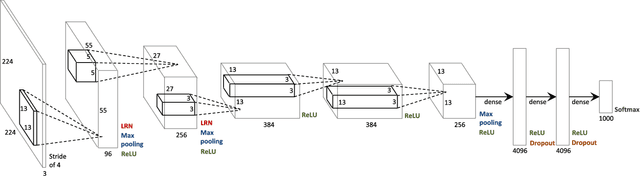
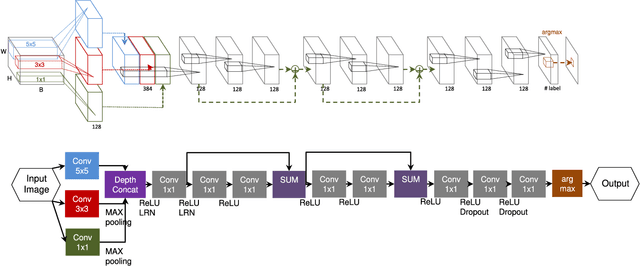
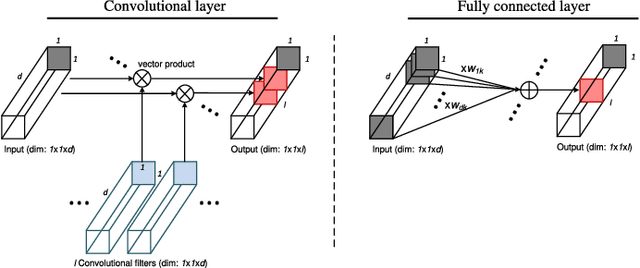
In this paper, we describe a novel deep convolutional neural network (CNN) that is deeper and wider than other existing deep networks for hyperspectral image classification. Unlike current state-of-the-art approaches in CNN-based hyperspectral image classification, the proposed network, called contextual deep CNN, can optimally explore local contextual interactions by jointly exploiting local spatio-spectral relationships of neighboring individual pixel vectors. The joint exploitation of the spatio-spectral information is achieved by a multi-scale convolutional filter bank used as an initial component of the proposed CNN pipeline. The initial spatial and spectral feature maps obtained from the multi-scale filter bank are then combined together to form a joint spatio-spectral feature map. The joint feature map representing rich spectral and spatial properties of the hyperspectral image is then fed through a fully convolutional network that eventually predicts the corresponding label of each pixel vector. The proposed approach is tested on three benchmark datasets: the Indian Pines dataset, the Salinas dataset and the University of Pavia dataset. Performance comparison shows enhanced classification performance of the proposed approach over the current state-of-the-art on the three datasets.
Revisiting Stereo Depth Estimation From a Sequence-to-Sequence Perspective with Transformers
Nov 05, 2020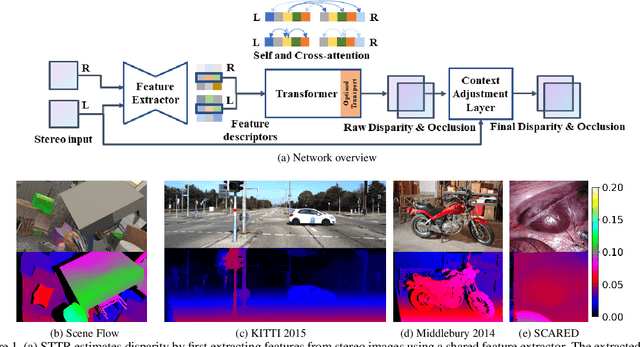
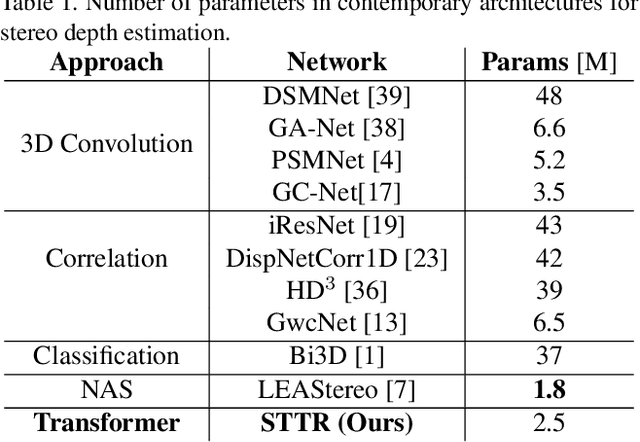
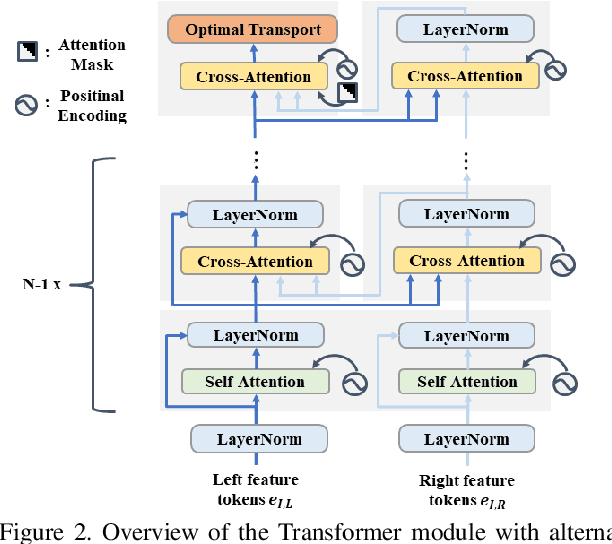
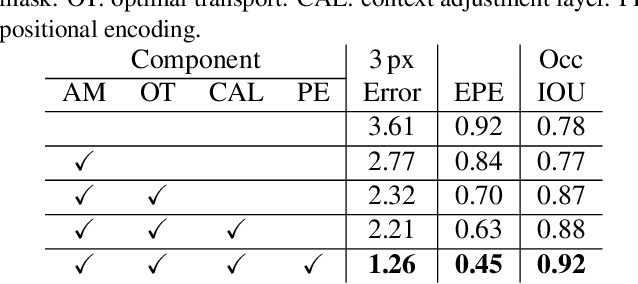
Stereo depth estimation relies on optimal correspondence matching between pixels on epipolar lines in the left and right image to infer depth. Rather than matching individual pixels, in this work, we revisit the problem from a sequence-to-sequence correspondence perspective to replace cost volume construction with dense pixel matching using position information and attention. This approach, named STereo TRansformer (STTR), has several advantages: It 1) relaxes the limitation of a fixed disparity range, 2) identifies occluded regions and provides confidence of estimation, and 3) imposes uniqueness constraints during the matching process. We report promising results on both synthetic and real-world datasets and demonstrate that STTR generalizes well across different domains, even without fine-tuning. Our code is publicly available at https://github.com/mli0603/stereo-transformer.
Rethinking CNN-Based Pansharpening: Guided Colorization of Panchromatic Images via GANs
Jun 30, 2020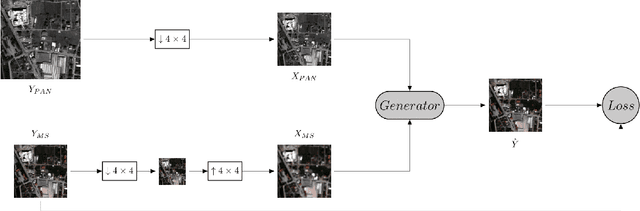
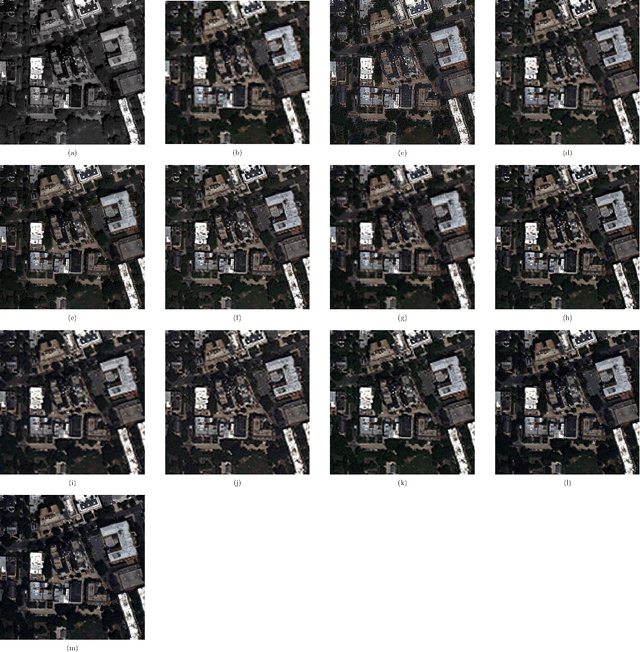
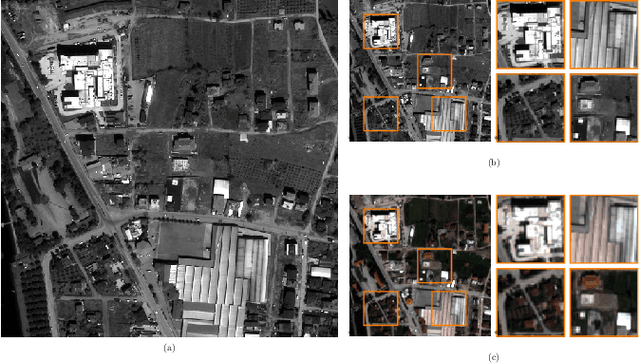


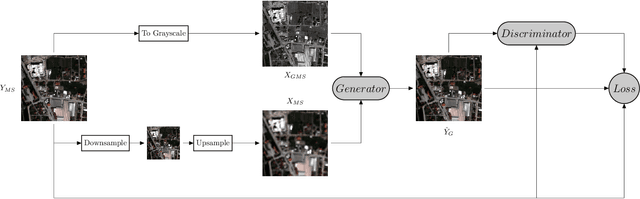
Convolutional Neural Networks (CNN)-based approaches have shown promising results in pansharpening of satellite images in recent years. However, they still exhibit limitations in producing high-quality pansharpening outputs. To that end, we propose a new self-supervised learning framework, where we treat pansharpening as a colorization problem, which brings an entirely novel perspective and solution to the problem compared to existing methods that base their solution solely on producing a super-resolution version of the multispectral image. Whereas CNN-based methods provide a reduced resolution panchromatic image as input to their model along with reduced resolution multispectral images, hence learn to increase their resolution together, we instead provide the grayscale transformed multispectral image as input, and train our model to learn the colorization of the grayscale input. We further address the fixed downscale ratio assumption during training, which does not generalize well to the full-resolution scenario. We introduce a noise injection into the training by randomly varying the downsampling ratios. Those two critical changes, along with the addition of adversarial training in the proposed PanColorization Generative Adversarial Networks (PanColorGAN) framework, help overcome the spatial detail loss and blur problems that are observed in CNN-based pansharpening. The proposed approach outperforms the previous CNN-based and traditional methods as demonstrated in our experiments.
New Techniques for Preserving Global Structure and Denoising with Low Information Loss in Single-Image Super-Resolution
Jun 16, 2018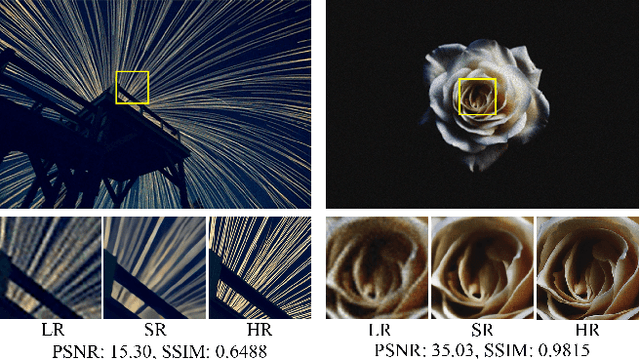
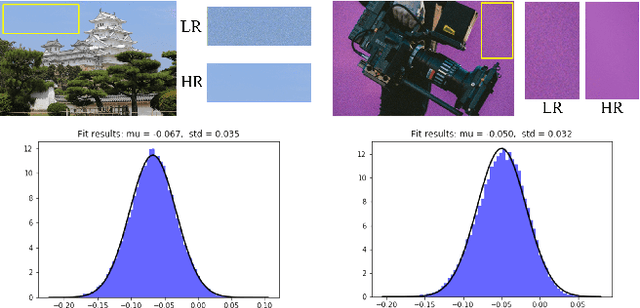
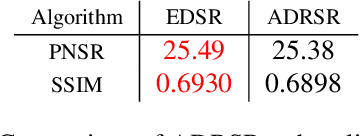
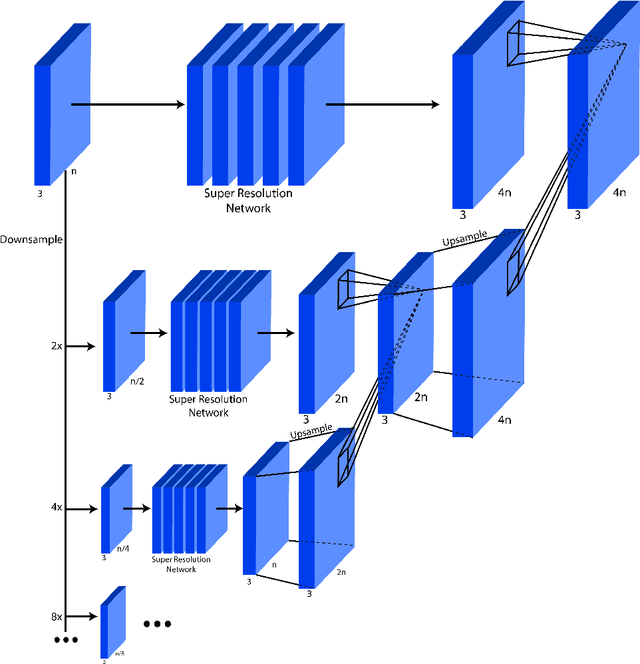
This work identifies and addresses two important technical challenges in single-image super-resolution: (1) how to upsample an image without magnifying noise and (2) how to preserve large scale structure when upsampling. We summarize the techniques we developed for our second place entry in Track 1 (Bicubic Downsampling), seventh place entry in Track 2 (Realistic Adverse Conditions), and seventh place entry in Track 3 (Realistic difficult) in the 2018 NTIRE Super-Resolution Challenge. Furthermore, we present new neural network architectures that specifically address the two challenges listed above: denoising and preservation of large-scale structure.
Attend to You: Personalized Image Captioning with Context Sequence Memory Networks
Apr 25, 2017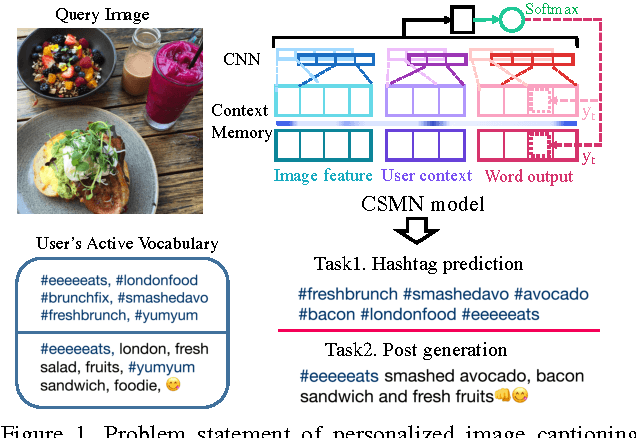
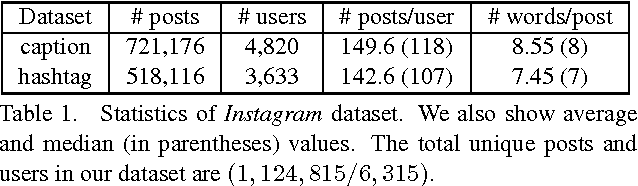
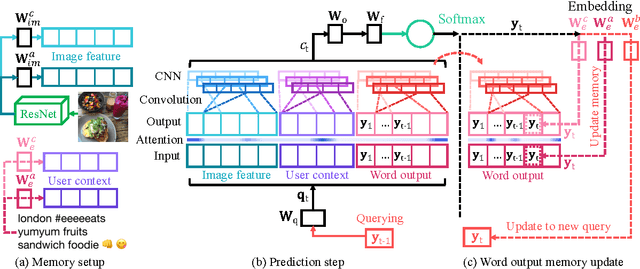

We address personalization issues of image captioning, which have not been discussed yet in previous research. For a query image, we aim to generate a descriptive sentence, accounting for prior knowledge such as the user's active vocabularies in previous documents. As applications of personalized image captioning, we tackle two post automation tasks: hashtag prediction and post generation, on our newly collected Instagram dataset, consisting of 1.1M posts from 6.3K users. We propose a novel captioning model named Context Sequence Memory Network (CSMN). Its unique updates over previous memory network models include (i) exploiting memory as a repository for multiple types of context information, (ii) appending previously generated words into memory to capture long-term information without suffering from the vanishing gradient problem, and (iii) adopting CNN memory structure to jointly represent nearby ordered memory slots for better context understanding. With quantitative evaluation and user studies via Amazon Mechanical Turk, we show the effectiveness of the three novel features of CSMN and its performance enhancement for personalized image captioning over state-of-the-art captioning models.
Deepzzle: Solving Visual Jigsaw Puzzles with Deep Learning andShortest Path Optimization
May 26, 2020
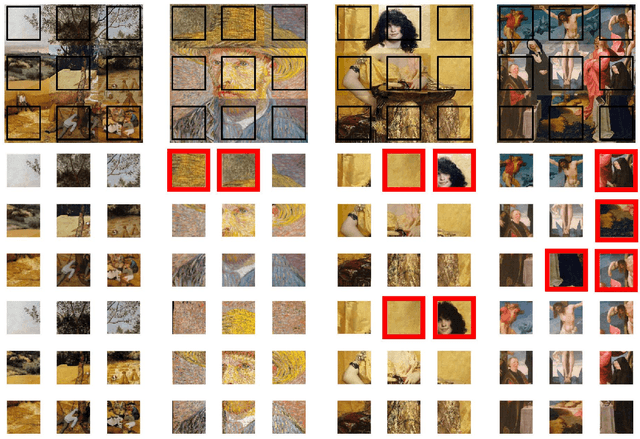
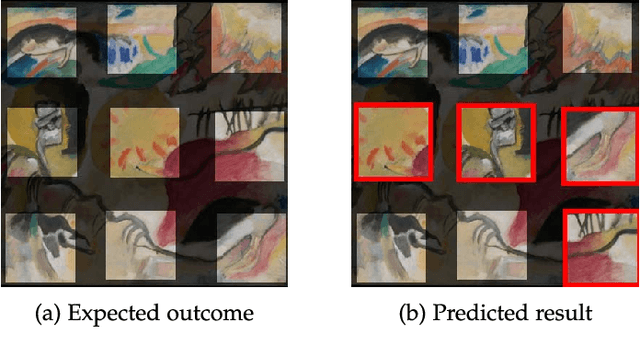

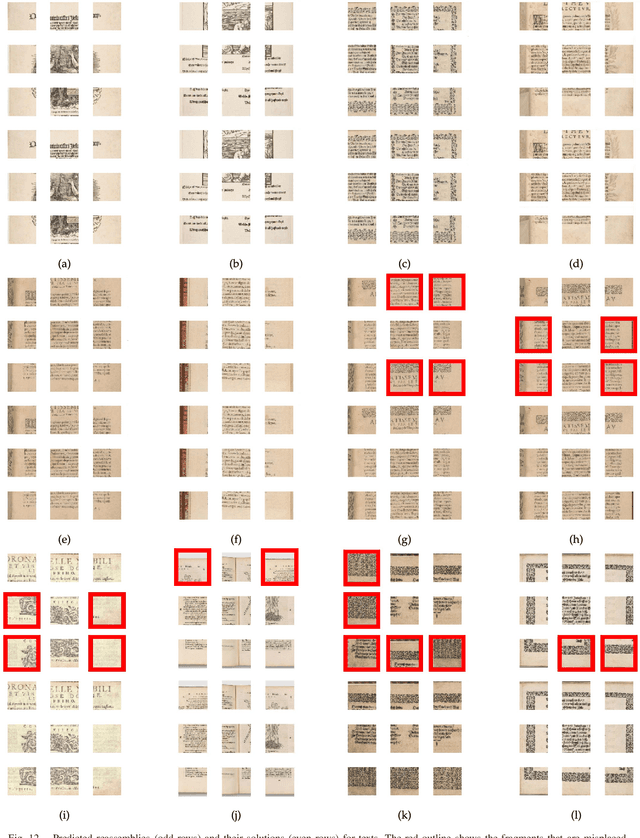
We tackle the image reassembly problem with wide space between the fragments, in such a way that the patterns and colors continuity is mostly unusable. The spacing emulates the erosion of which the archaeological fragments suffer. We crop-square the fragments borders to compel our algorithm to learn from the content of the fragments. We also complicate the image reassembly by removing fragments and adding pieces from other sources. We use a two-step method to obtain the reassemblies: 1) a neural network predicts the positions of the fragments despite the gaps between them; 2) a graph that leads to the best reassemblies is made from these predictions. In this paper, we notably investigate the effect of branch-cut in the graph of reassemblies. We also provide a comparison with the literature, solve complex images reassemblies, explore at length the dataset, and propose a new metric that suits its specificities. Keywords: image reassembly, jigsaw puzzle, deep learning, graph, branch-cut, cultural heritage
Radon-Gabor Barcodes for Medical Image Retrieval
Sep 16, 2016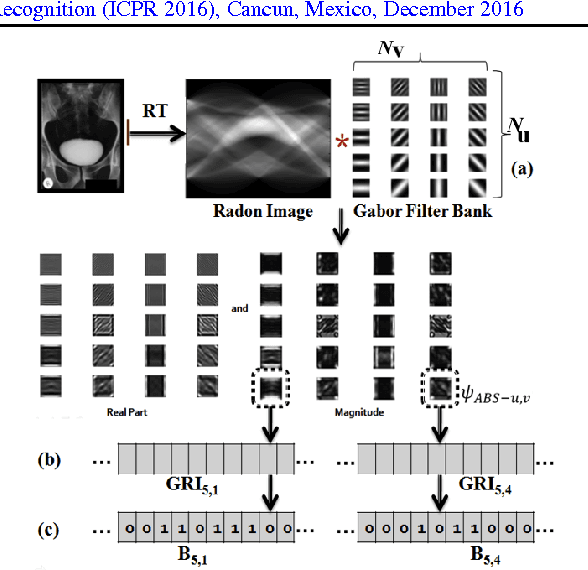
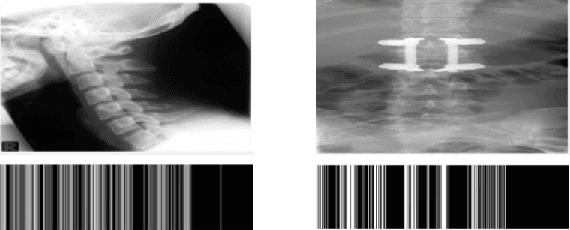
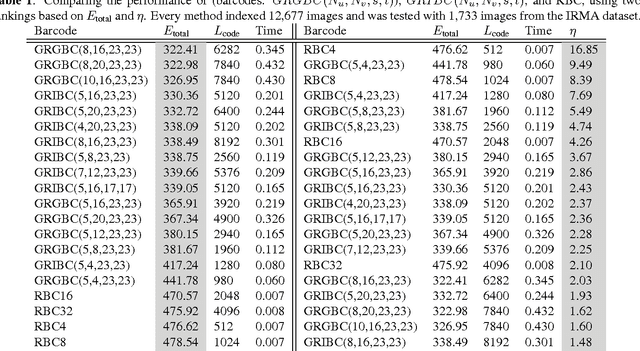

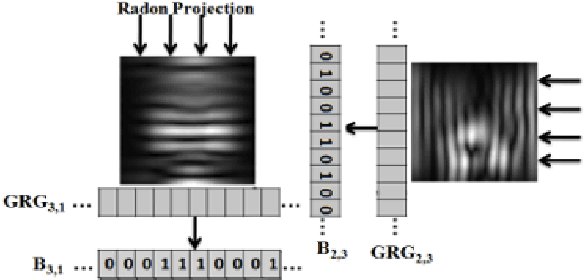
In recent years, with the explosion of digital images on the Web, content-based retrieval has emerged as a significant research area. Shapes, textures, edges and segments may play a key role in describing the content of an image. Radon and Gabor transforms are both powerful techniques that have been widely studied to extract shape-texture-based information. The combined Radon-Gabor features may be more robust against scale/rotation variations, presence of noise, and illumination changes. The objective of this paper is to harness the potentials of both Gabor and Radon transforms in order to introduce expressive binary features, called barcodes, for image annotation/tagging tasks. We propose two different techniques: Gabor-of-Radon-Image Barcodes (GRIBCs), and Guided-Radon-of-Gabor Barcodes (GRGBCs). For validation, we employ the IRMA x-ray dataset with 193 classes, containing 12,677 training images and 1,733 test images. A total error score as low as 322 and 330 were achieved for GRGBCs and GRIBCs, respectively. This corresponds to $\approx 81\%$ retrieval accuracy for the first hit.
EC-GAN: Low-Sample Classification using Semi-Supervised Algorithms and GANs
Dec 26, 2020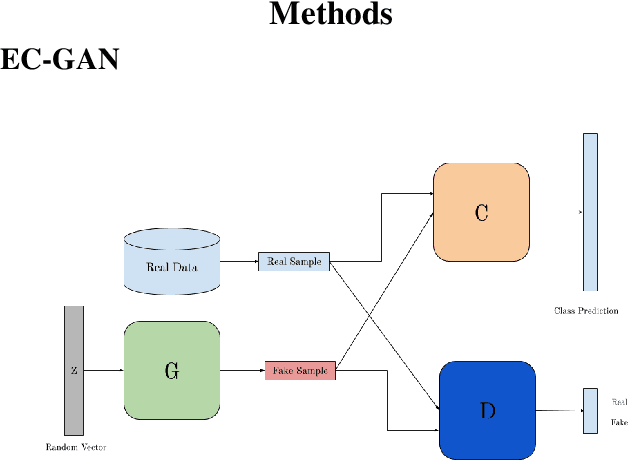
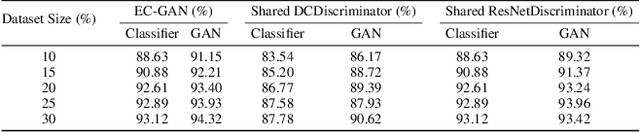
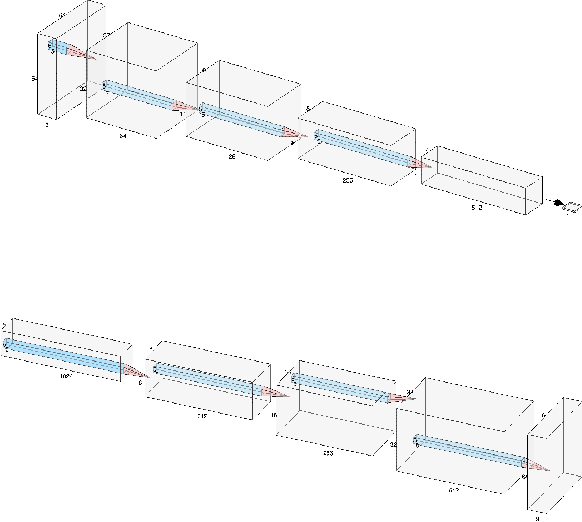
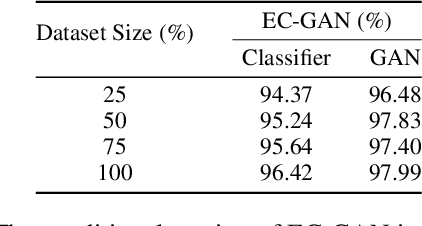
Semi-supervised learning has been gaining attention as it allows for performing image analysis tasks such as classification with limited labeled data. Some popular algorithms using Generative Adversarial Networks (GANs) for semi-supervised classification share a single architecture for classification and discrimination. However, this may require a model to converge to a separate data distribution for each task, which may reduce overall performance. While progress in semi-supervised learning has been made, less addressed are small-scale, fully-supervised tasks where even unlabeled data is unavailable and unattainable. We therefore, propose a novel GAN model namely External Classifier GAN (EC-GAN), that utilizes GANs and semi-supervised algorithms to improve classification in fully-supervised regimes. Our method leverages a GAN to generate artificial data used to supplement supervised classification. More specifically, we attach an external classifier, hence the name EC-GAN, to the GAN's generator, as opposed to sharing an architecture with the discriminator. Our experiments demonstrate that EC-GAN's performance is comparable to the shared architecture method, far superior to the standard data augmentation and regularization-based approach, and effective on a small, realistic dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge