"Image": models, code, and papers
MeDaS: An open-source platform as service to help break the walls between medicine and informatics
Jul 12, 2020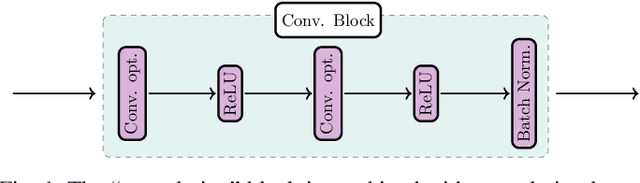
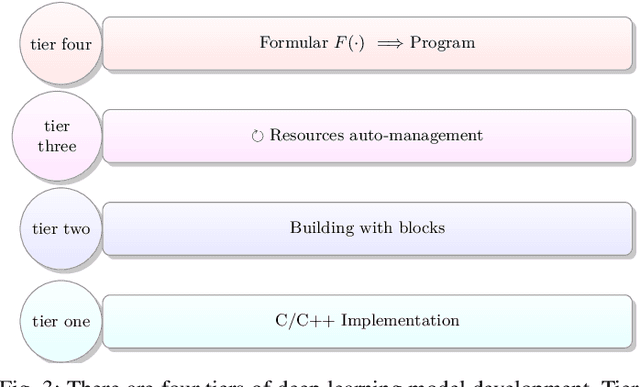


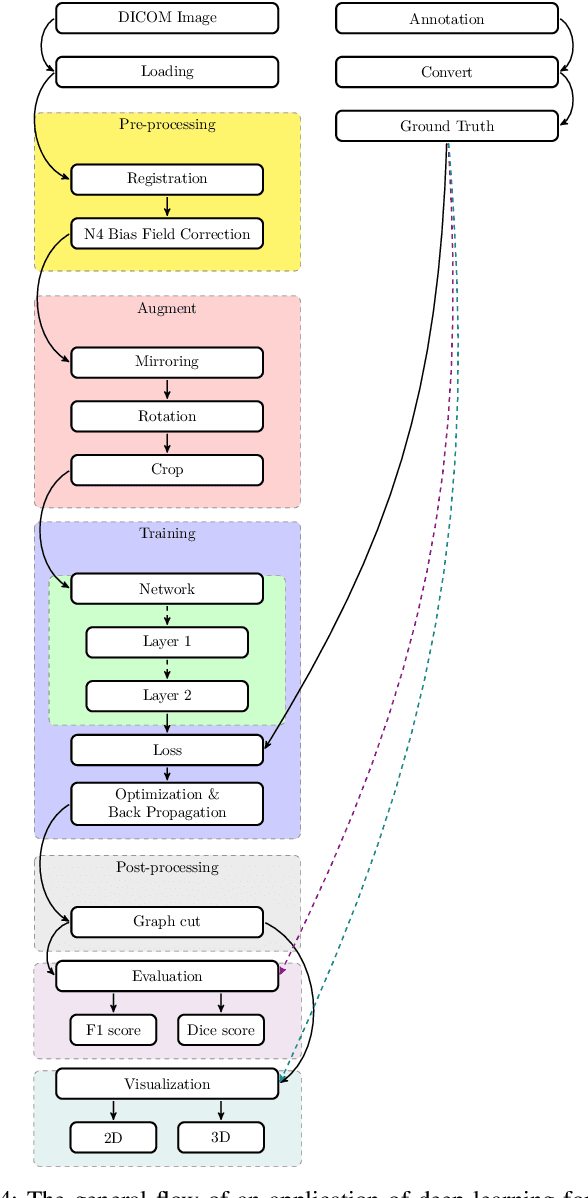
In the past decade, deep learning (DL) has achieved unprecedented success in numerous fields including computer vision, natural language processing, and healthcare. In particular, DL is experiencing an increasing development in applications for advanced medical image analysis in terms of analysis, segmentation, classification, and furthermore. On the one hand, tremendous needs that leverage the power of DL for medical image analysis are arising from the research community of a medical, clinical, and informatics background to jointly share their expertise, knowledge, skills, and experience. On the other hand, barriers between disciplines are on the road for them often hampering a full and efficient collaboration. To this end, we propose our novel open-source platform, i.e., MeDaS -- the MeDical open-source platform as Service. To the best of our knowledge, MeDaS is the first open-source platform proving a collaborative and interactive service for researchers from a medical background easily using DL related toolkits, and at the same time for scientists or engineers from information sciences to understand the medical knowledge side. Based on a series of toolkits and utilities from the idea of RINV (Rapid Implementation aNd Verification), our proposed MeDaS platform can implement pre-processing, post-processing, augmentation, visualization, and other phases needed in medical image analysis. Five tasks including the subjects of lung, liver, brain, chest, and pathology, are validated and demonstrated to be efficiently realisable by using MeDaS.
A Generalization of Otsu's Method and Minimum Error Thresholding
Jul 23, 2020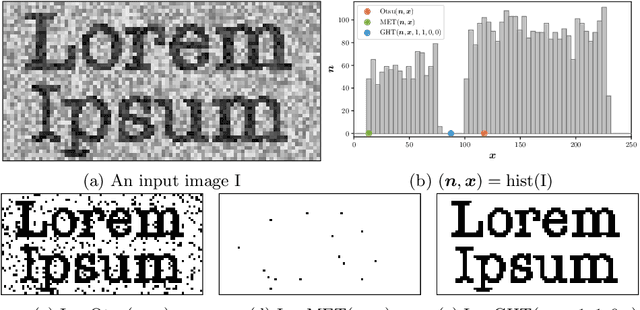
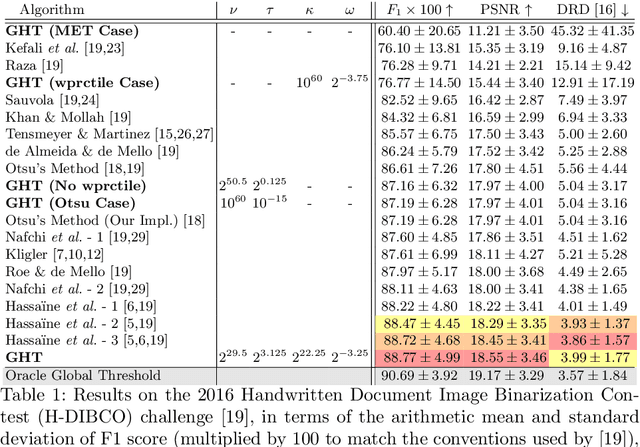
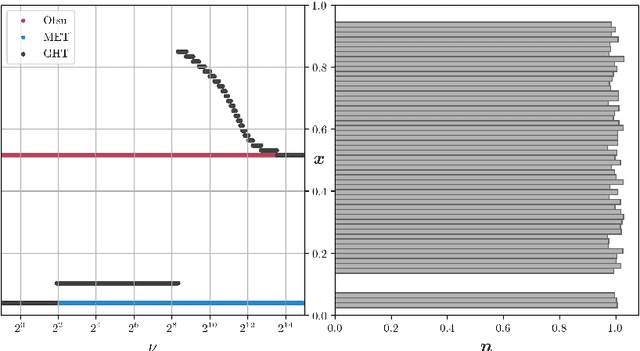
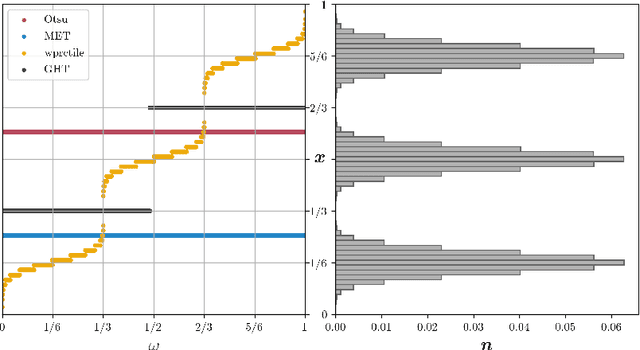
We present Generalized Histogram Thresholding (GHT), a simple, fast, and effective technique for histogram-based image thresholding. GHT works by performing approximate maximum a posteriori estimation of a mixture of Gaussians with appropriate priors. We demonstrate that GHT subsumes three classic thresholding techniques as special cases: Otsu's method, Minimum Error Thresholding (MET), and weighted percentile thresholding. GHT thereby enables the continuous interpolation between those three algorithms, which allows thresholding accuracy to be improved significantly. GHT also provides a clarifying interpretation of the common practice of coarsening a histogram's bin width during thresholding. We show that GHT outperforms or matches the performance of all algorithms on a recent challenge for handwritten document image binarization (including deep neural networks trained to produce per-pixel binarizations), and can be implemented in a dozen lines of code or as a trivial modification to Otsu's method or MET.
Bridging the gap between Natural and Medical Images through Deep Colorization
May 21, 2020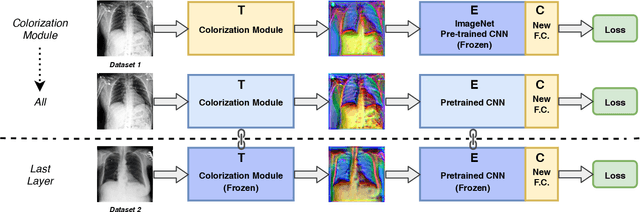
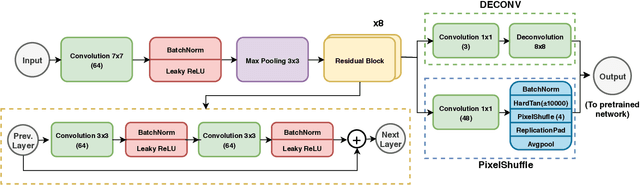
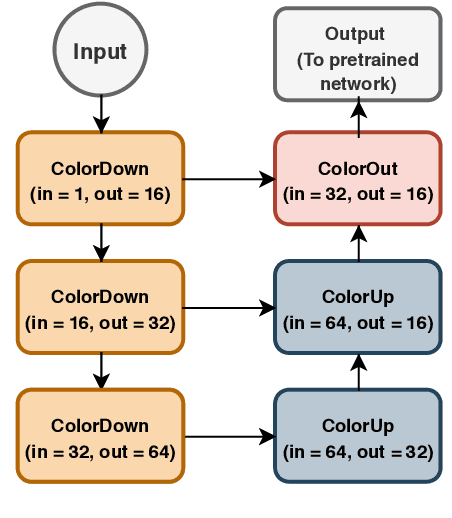
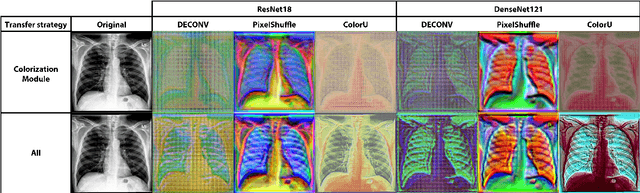
Deep learning has thrived by training on large-scale datasets. However, in many applications, as for medical image diagnosis, getting massive amount of data is still prohibitive due to privacy, lack of acquisition homogeneity and annotation cost. In this scenario, transfer learning from natural image collections is a standard practice that attempts to tackle shape, texture and color discrepancies all at once through pretrained model fine-tuning. In this work, we propose to disentangle those challenges and design a dedicated network module that focuses on color adaptation. We combine learning from scratch of the color module with transfer learning of different classification backbones, obtaining an end-to-end, easy-to-train architecture for diagnostic image recognition on X-ray images. Extensive experiments showed how our approach is particularly efficient in case of data scarcity and provides a new path for further transferring the learned color information across multiple medical datasets.
Weighted Truncated Nuclear Norm Regularization for Low-Rank Quaternion Matrix Completion
Jan 07, 2021
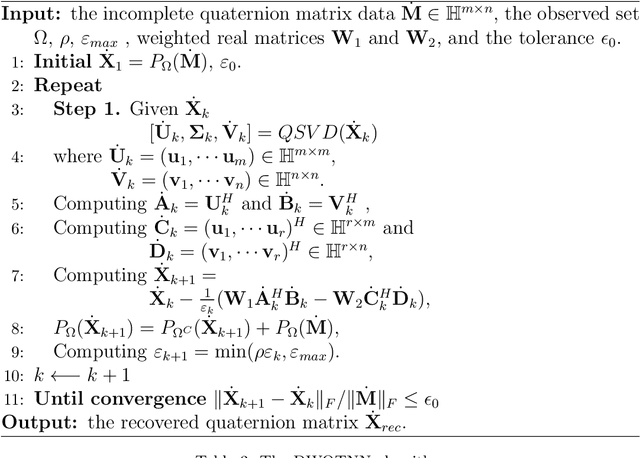



In recent years, quaternion matrix completion (QMC) based on low-rank regularization has been gradually used in image de-noising and de-blurring.Unlike low-rank matrix completion (LRMC) which handles RGB images by recovering each color channel separately, the QMC models utilize the connection of three channels by processing them as a whole. Most of the existing quaternion-based methods formulate low-rank QMC (LRQMC) as a quaternion nuclear norm (a convex relaxation of the rank) minimization problem.The main limitation of these approaches is that the singular values being minimized simultaneously so that the low-rank property could not be approximated well and efficiently. To achieve a more accurate low-rank approximation, the matrix-based truncated nuclear norm has been proposed and also been proved to have the superiority. In this paper, we introduce a quaternion truncated nuclear norm (QTNN) for LRQMC and utilize the alternating direction method of multipliers (ADMM) to get the optimization.We further propose weights to the residual error quaternion matrix during the update process for accelerating the convergence of the QTNN method with admissible performance. The weighted method utilizes a concise gradient descent strategy which has a theoretical guarantee in optimization. The effectiveness of our method is illustrated by experiments on real visual datasets.
Dairy Cow rumination detection: A deep learning approach
Jan 07, 2021
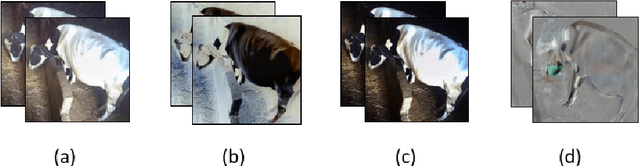

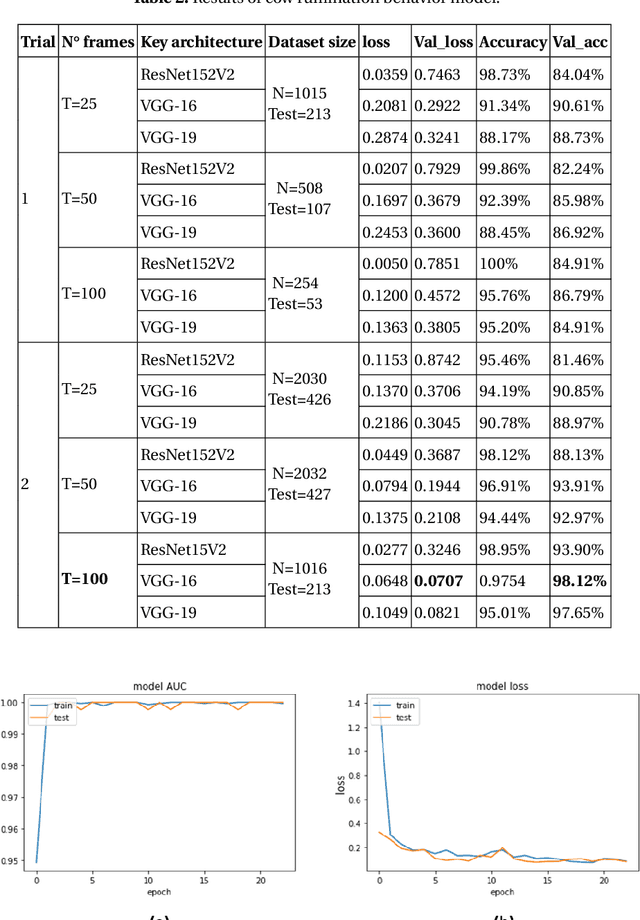

Cattle activity is an essential index for monitoring health and welfare of the ruminants. Thus, changes in the livestock behavior are a critical indicator for early detection and prevention of several diseases. Rumination behavior is a significant variable for tracking the development and yield of animal husbandry. Therefore, various monitoring methods and measurement equipment have been used to assess cattle behavior. However, these modern attached devices are invasive, stressful and uncomfortable for the cattle and can influence negatively the welfare and diurnal behavior of the animal. Multiple research efforts addressed the problem of rumination detection by adopting new methods by relying on visual features. However, they only use few postures of the dairy cow to recognize the rumination or feeding behavior. In this study, we introduce an innovative monitoring method using Convolution Neural Network (CNN)-based deep learning models. The classification process is conducted under two main labels: ruminating and other, using all cow postures captured by the monitoring camera. Our proposed system is simple and easy-to-use which is able to capture long-term dynamics using a compacted representation of a video in a single 2D image. This method proved efficiency in recognizing the rumination behavior with 95%, 98% and 98% of average accuracy, recall and precision, respectively.
* 17 pages, 6 figures, 4 tables
Saliency maps on image hierarchies
Aug 19, 2015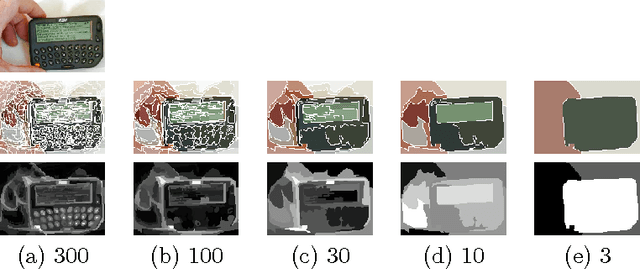
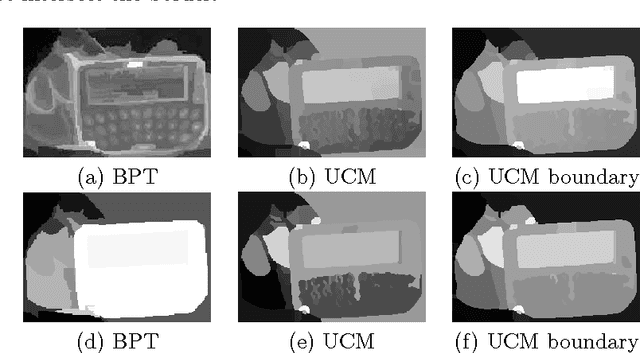
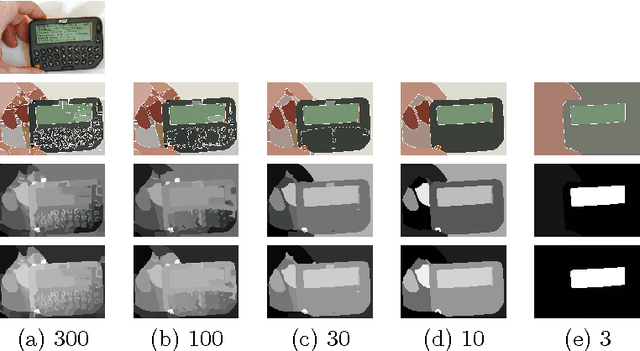

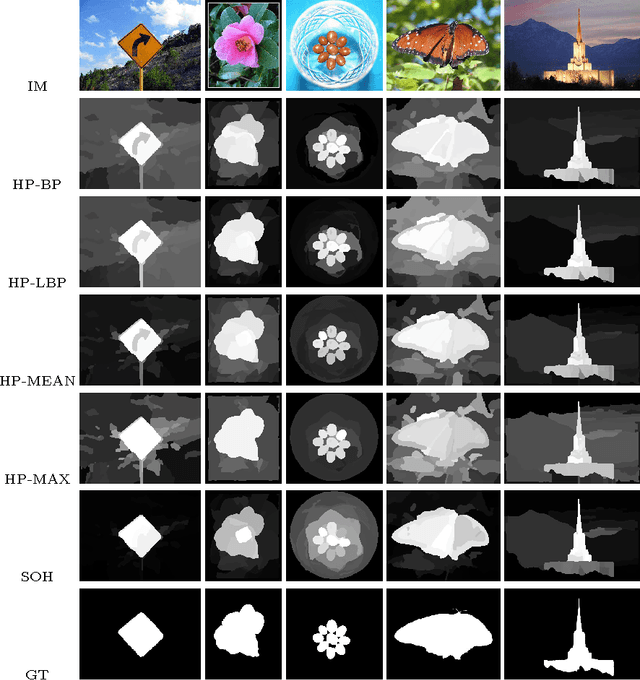
In this paper we propose two saliency models for salient object segmentation based on a hierarchical image segmentation, a tree-like structure that represents regions at different scales from the details to the whole image (e.g. gPb-UCM, BPT). The first model is based on a hierarchy of image partitions. The saliency at each level is computed on a region basis, taking into account the contrast between regions. The maps obtained for the different partitions are then integrated into a final saliency map. The second model directly works on the structure created by the segmentation algorithm, computing saliency at each node and integrating these cues in a straightforward manner into a single saliency map. We show that the proposed models produce high quality saliency maps. Objective evaluation demonstrates that the two methods achieve state-of-the-art performance in several benchmark datasets.
Open Source Iris Recognition Hardware and Software with Presentation Attack Detection
Aug 19, 2020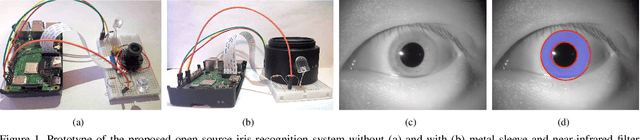
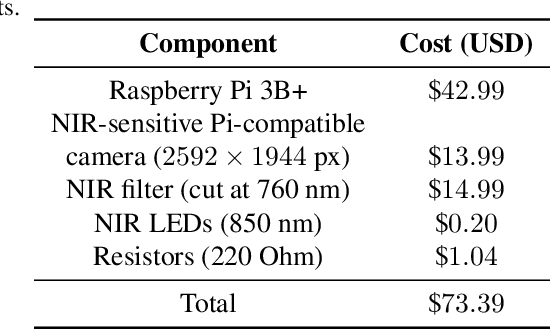
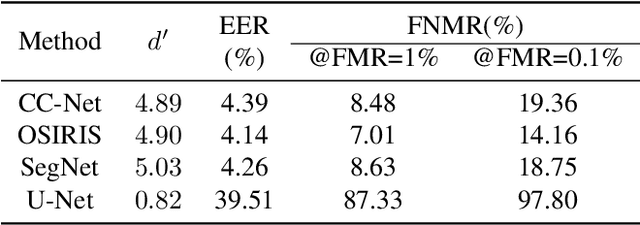
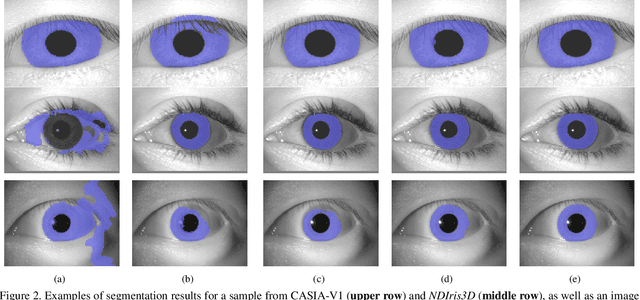
This paper proposes the first known to us open source hardware and software iris recognition system with presentation attack detection (PAD), which can be easily assembled for about 75 USD using Raspberry Pi board and a few peripherals. The primary goal of this work is to offer a low-cost baseline for spoof-resistant iris recognition, which may (a) stimulate research in iris PAD and allow for easy prototyping of secure iris recognition systems, (b) offer a low-cost secure iris recognition alternative to more sophisticated systems, and (c) serve as an educational platform. We propose a lightweight image complexity-guided convolutional network for fast and accurate iris segmentation, domain-specific human-inspired Binarized Statistical Image Features (BSIF) to build an iris template, and to combine 2D (iris texture) and 3D (photometric stereo-based) features for PAD. The proposed iris recognition runs in about 3.2 seconds and the proposed PAD runs in about 4.5 seconds on Raspberry Pi 3B+. The hardware specifications and all source codes of the entire pipeline are made available along with this paper.
Stereo Plane SLAM Based on Intersecting Lines
Aug 19, 2020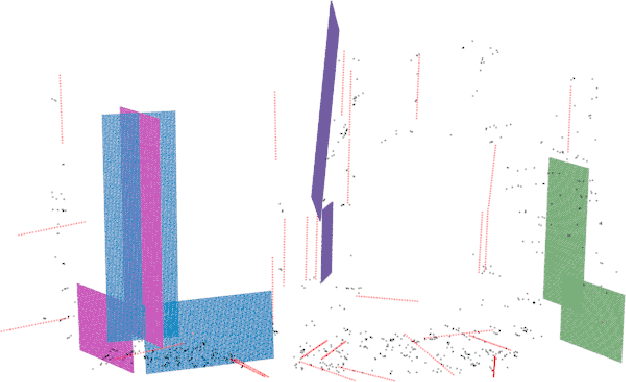
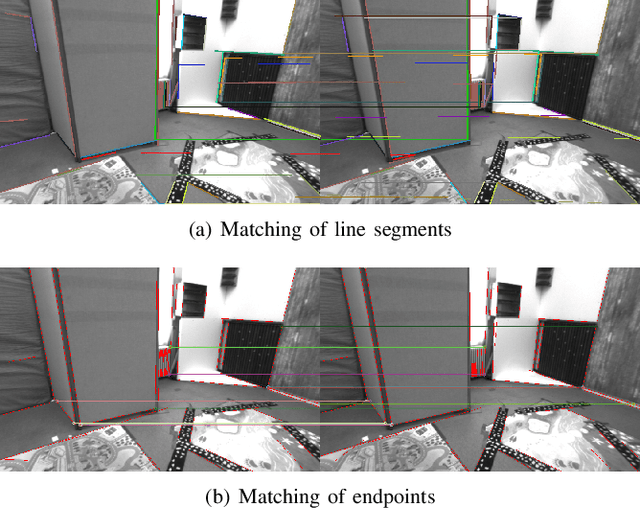
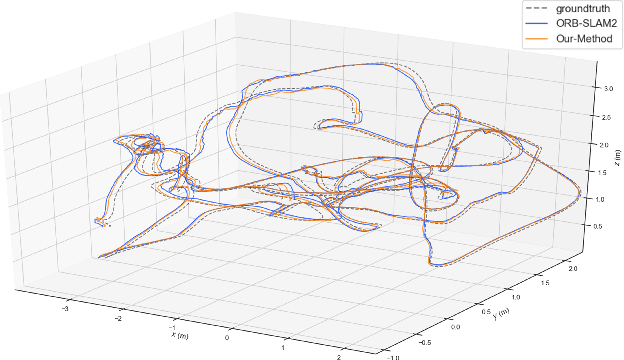

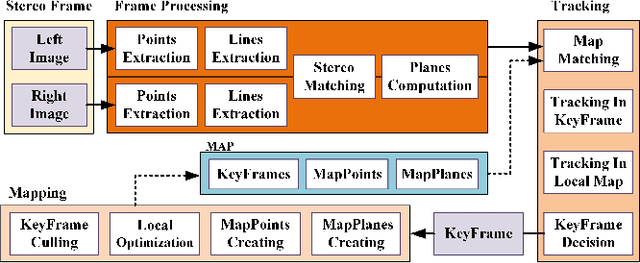

Plane feature is a kind of stable landmark to reduce drift error in SLAM system. It is easy and fast to extract planes from dense point cloud, which is commonly acquired from RGB-D camera or lidar. But for stereo camera, it is hard to compute dense point cloud accurately and efficiently. In this paper, we propose a novel method to compute plane parameters from intersecting lines extracted for stereo image. The plane features commonly exist on the surface of man-made objects and structure, which have regular shape and straight edge lines. In 3D space, two intersecting lines can determine such a plane. Thus we extract line segments from both stereo left and right image. By stereo matching, we compute the endpoints and line directions in 3D space, and then the planes can be computed. Adding such computed plane features in stereo SLAM system reduces the drift error and refines the performance. We test our proposed system on public datasets and demonstrate its robust and accurate estimation results, compared with state-of-the-art SLAM systems.
Deep Co-Attention Network for Multi-View Subspace Learning
Feb 15, 2021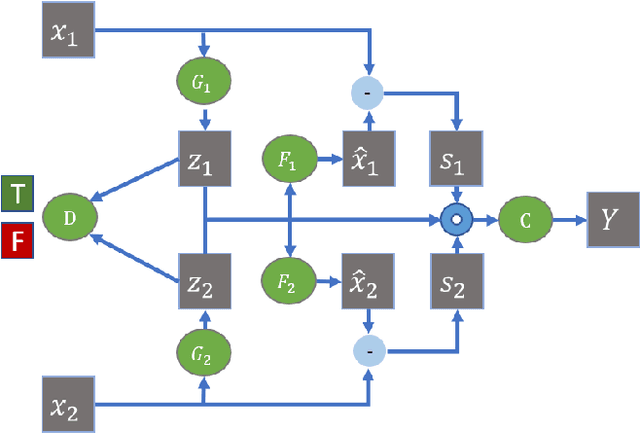
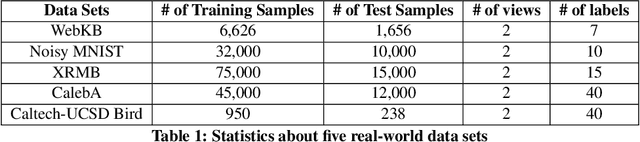
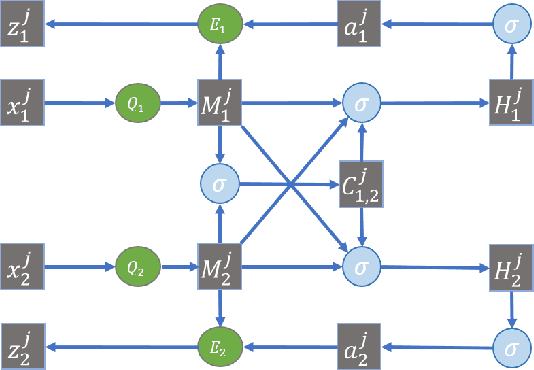
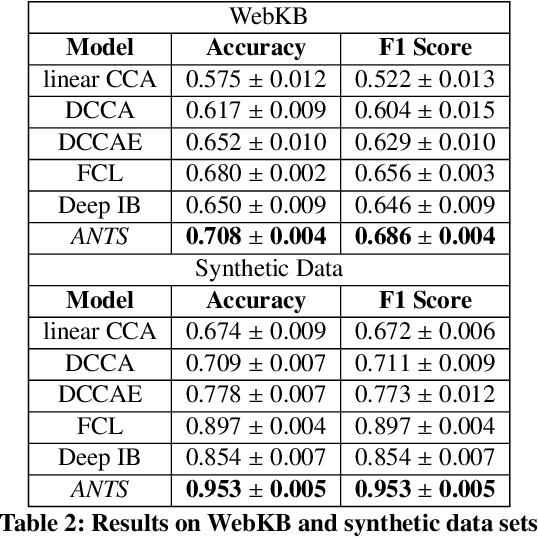
Many real-world applications involve data from multiple modalities and thus exhibit the view heterogeneity. For example, user modeling on social media might leverage both the topology of the underlying social network and the content of the users' posts; in the medical domain, multiple views could be X-ray images taken at different poses. To date, various techniques have been proposed to achieve promising results, such as canonical correlation analysis based methods, etc. In the meanwhile, it is critical for decision-makers to be able to understand the prediction results from these methods. For example, given the diagnostic result that a model provided based on the X-ray images of a patient at different poses, the doctor needs to know why the model made such a prediction. However, state-of-the-art techniques usually suffer from the inability to utilize the complementary information of each view and to explain the predictions in an interpretable manner. To address these issues, in this paper, we propose a deep co-attention network for multi-view subspace learning, which aims to extract both the common information and the complementary information in an adversarial setting and provide robust interpretations behind the prediction to the end-users via the co-attention mechanism. In particular, it uses a novel cross reconstruction loss and leverages the label information to guide the construction of the latent representation by incorporating the classifier into our model. This improves the quality of latent representation and accelerates the convergence speed. Finally, we develop an efficient iterative algorithm to find the optimal encoders and discriminator, which are evaluated extensively on synthetic and real-world data sets. We also conduct a case study to demonstrate how the proposed method robustly interprets the predictions on an image data set.
Image Segmentation and Classification for Sickle Cell Disease using Deformable U-Net
Oct 29, 2017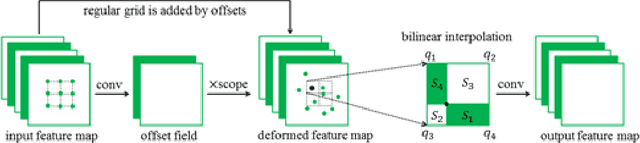
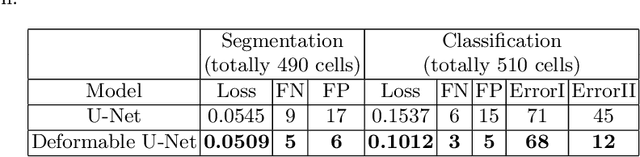
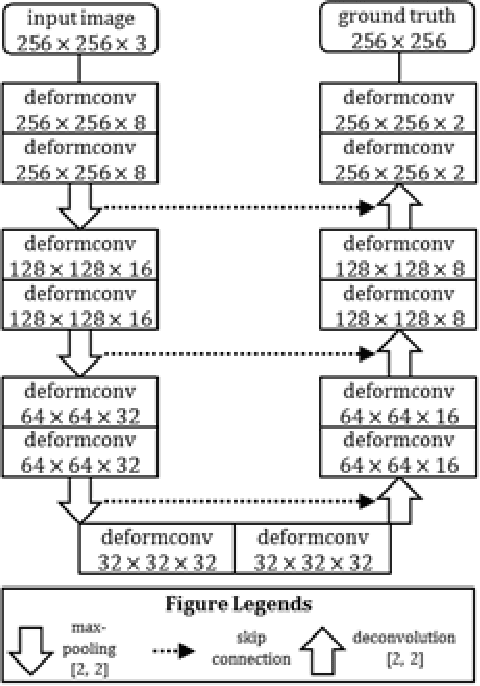
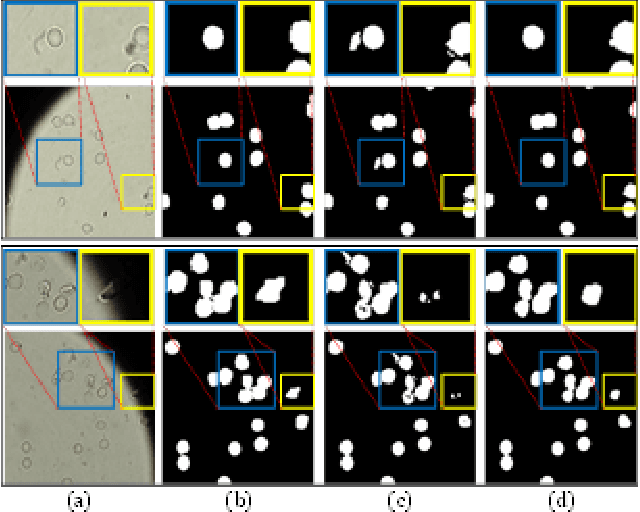
Reliable cell segmentation and classification from biomedical images is a crucial step for both scientific research and clinical practice. A major challenge for more robust segmentation and classification methods is the large variations in the size, shape and viewpoint of the cells, combining with the low image quality caused by noise and artifacts. To address this issue, in this work we propose a learning-based, simultaneous cell segmentation and classification method based on the deep U-Net structure with deformable convolution layers. The U-Net architecture for deep learning has been shown to offer a precise localization for image semantic segmentation. Moreover, deformable convolution layer enables the free form deformation of the feature learning process, thus makes the whole network more robust to various cell morphologies and image settings. The proposed method is tested on microscopic red blood cell images from patients with sickle cell disease. The results show that U-Net with deformable convolution achieves the highest accuracy for segmentation and classification, comparing with original U-Net structure.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge