"Image": models, code, and papers
Semi-supervised Left Atrium Segmentation with Mutual Consistency Training
Mar 04, 2021
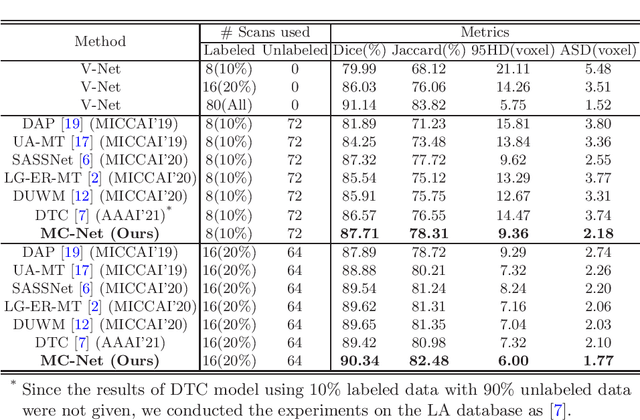
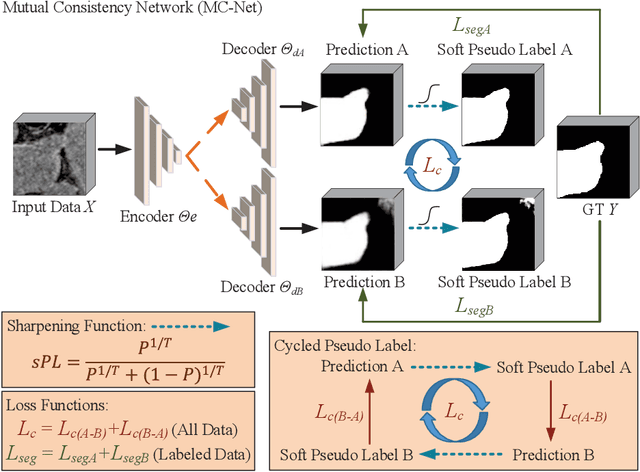
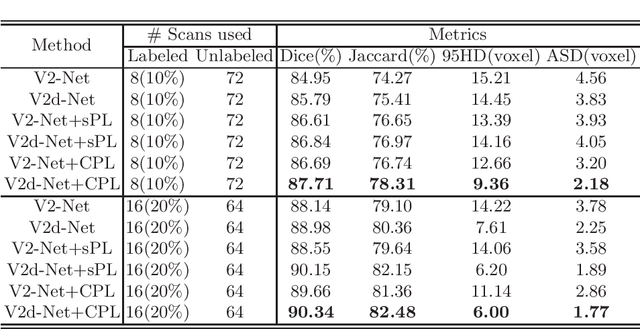
Semi-supervised learning has attracted great attention in the field of machine learning, especially for medical image segmentation tasks, since it alleviates the heavy burden of collecting abundant densely annotated data for training. However, most of existing methods underestimate the importance of challenging regions (e.g. small branches or blurred edges) during training. We believe that these unlabeled regions may contain more crucial information to minimize the uncertainty prediction for the model and should be emphasized in the training process. Therefore, in this paper, we propose a novel Mutual Consistency Network (MC-Net) for semi-supervised left atrium segmentation from 3D MR images. Particularly, our MC-Net consists of one encoder and two slightly different decoders, and the prediction discrepancies of two decoders are transformed as an unsupervised loss by our designed cycled pseudo label scheme to encourage mutual consistency. Such mutual consistency encourages the two decoders to have consistent and low-entropy predictions and enables the model to gradually capture generalized features from these unlabeled challenging regions. We evaluate our MC-Net on the public Left Atrium (LA) database and it obtains impressive performance gains by exploiting the unlabeled data effectively. Our MC-Net outperforms six recent semi-supervised methods for left atrium segmentation, and sets the new state-of-the-art performance on the LA database.
Incident Light Frequency-based Image Defogging Algorithm
Mar 03, 2017
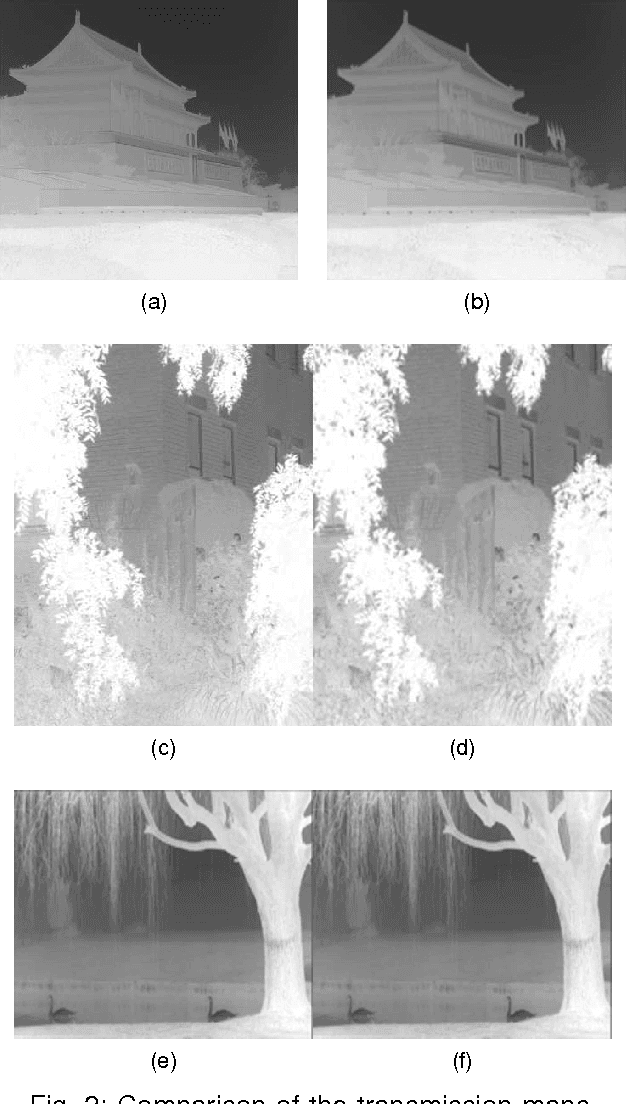



Considering the problem of color distortion caused by the defogging algorithm based on dark channel prior, an improved algorithm was proposed to calculate the transmittance of all channels respectively. First, incident light frequency's effect on the transmittance of various color channels was analyzed according to the Beer-Lambert's Law, from which a proportion among various channel transmittances was derived; afterwards, images were preprocessed by down-sampling to refine transmittance, and then the original size was restored to enhance the operational efficiency of the algorithm; finally, the transmittance of all color channels was acquired in accordance with the proportion, and then the corresponding transmittance was used for image restoration in each channel. The experimental results show that compared with the existing algorithm, this improved image defogging algorithm could make image colors more natural, solve the problem of slightly higher color saturation caused by the existing algorithm, and shorten the operation time by four to nine times.
Chaotic-to-Fine Clustering for Unlabeled Plant Disease Images
Jan 18, 2021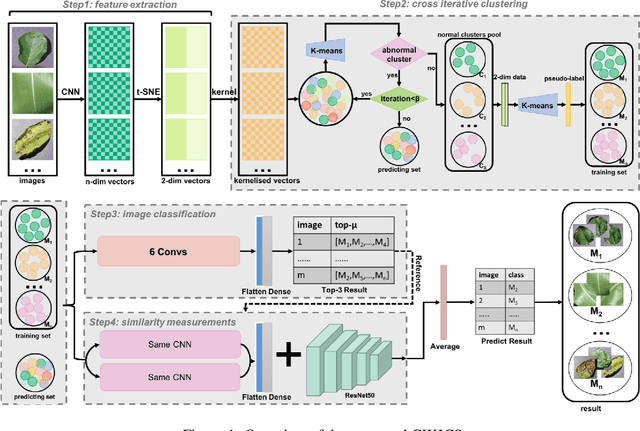
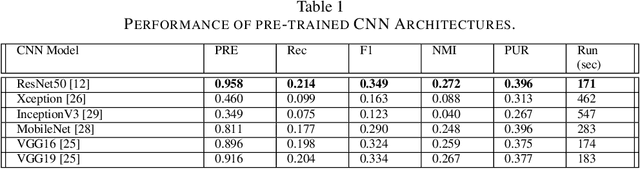
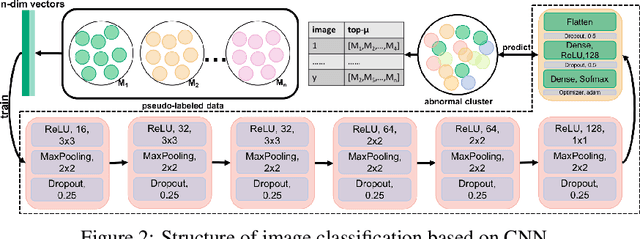
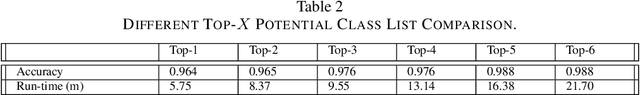
Current annotation for plant disease images depends on manual sorting and handcrafted features by agricultural experts, which is time-consuming and labour-intensive. In this paper, we propose a self-supervised clustering framework for grouping plant disease images based on the vulnerability of Kernel K-means. The main idea is to establish a cross iterative under-clustering algorithm based on Kernel K-means to produce the pseudo-labeled training set and a chaotic cluster to be further classified by a deep learning module. In order to verify the effectiveness of our proposed framework, we conduct extensive experiments on three different plant disease datatsets with five plants and 17 plant diseases. The experimental results show the high superiority of our method to do image-based plant disease classification over balanced and unbalanced datasets by comparing with five state-of-the-art existing works in terms of different metrics.
VDM-DA: Virtual Domain Modeling for Source Data-free Domain Adaptation
Mar 26, 2021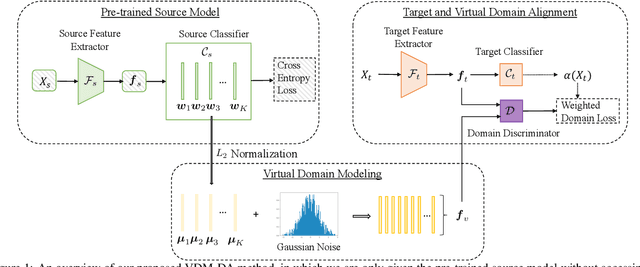
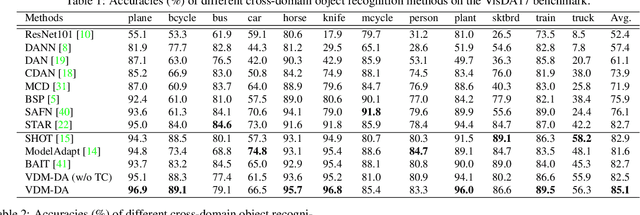
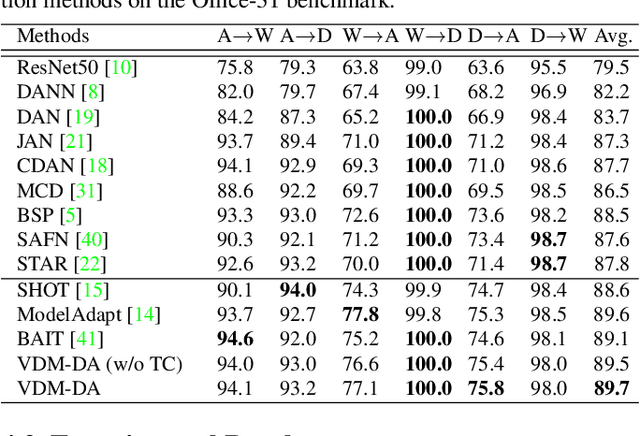
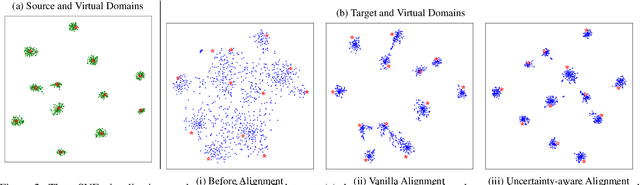
Domain adaptation aims to leverage a label-rich domain (the source domain) to help model learning in a label-scarce domain (the target domain). Most domain adaptation methods require the co-existence of source and target domain samples to reduce the distribution mismatch, however, access to the source domain samples may not always be feasible in the real world applications due to different problems (e.g., storage, transmission, and privacy issues). In this work, we deal with the source data-free unsupervised domain adaptation problem, and propose a novel approach referred to as Virtual Domain Modeling (VDM-DA). The virtual domain acts as a bridge between the source and target domains. On one hand, we generate virtual domain samples based on an approximated Gaussian Mixture Model (GMM) in the feature space with the pre-trained source model, such that the virtual domain maintains a similar distribution with the source domain without accessing to the original source data. On the other hand, we also design an effective distribution alignment method to reduce the distribution divergence between the virtual domain and the target domain by gradually improving the compactness of the target domain distribution through model learning. In this way, we successfully achieve the goal of distribution alignment between the source and target domains by training deep networks without accessing to the source domain data. We conduct extensive experiments on benchmark datasets for both 2D image-based and 3D point cloud-based cross-domain object recognition tasks, where the proposed method referred to Domain Adaptation with Virtual Domain Modeling (VDM-DA) achieves the state-of-the-art performances on all datasets.
FSOCO: The Formula Student Objects in Context Dataset
Dec 13, 2020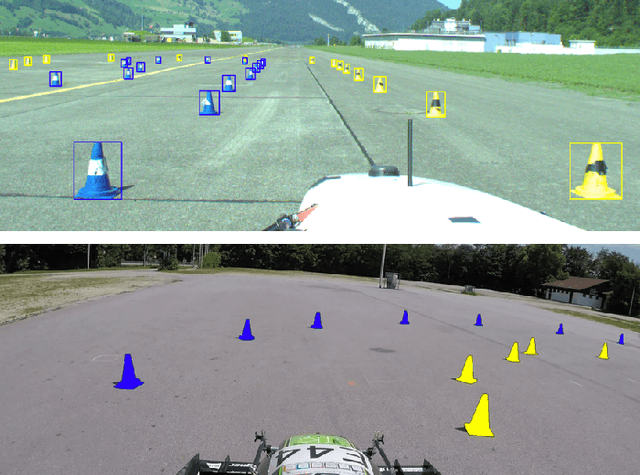
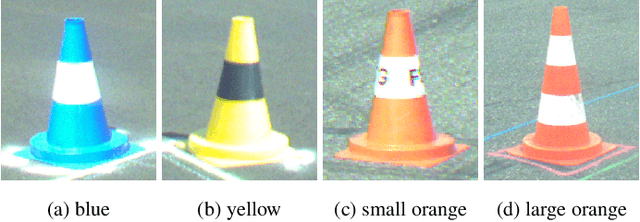
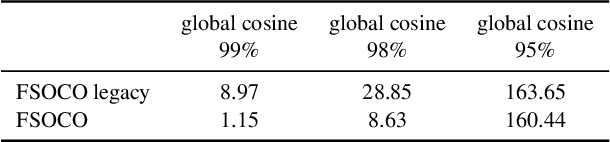

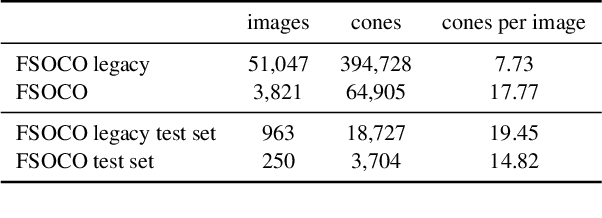
This paper presents the FSOCO dataset, a collaborative dataset for vision-based cone detection systems in Formula Student Driverless competitions. It contains human annotated ground truth labels for both bounding boxes and instance-wise segmentation masks. The data buy-in philosophy of FSOCO asks student teams to contribute to the database first before being granted access ensuring continuous growth. By providing clear labeling guidelines and tools for a sophisticated raw image selection, new annotations are guaranteed to meet the desired quality. The effectiveness of the approach is shown by comparing prediction results of a network trained on FSOCO and its unregulated predecessor. The FSOCO dataset can be found at fsoco-dataset.com.
Universal Denoising Networks : A Novel CNN Architecture for Image Denoising
Mar 24, 2018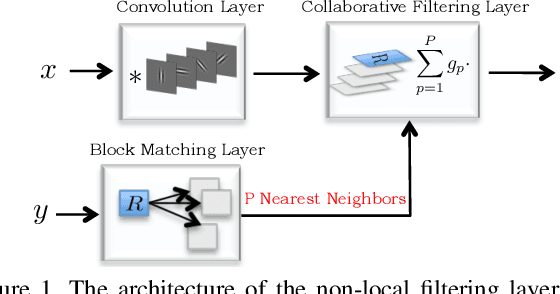
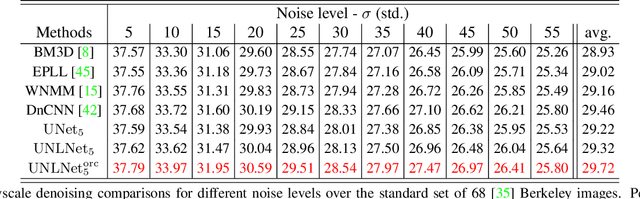
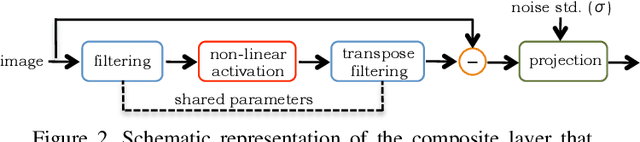
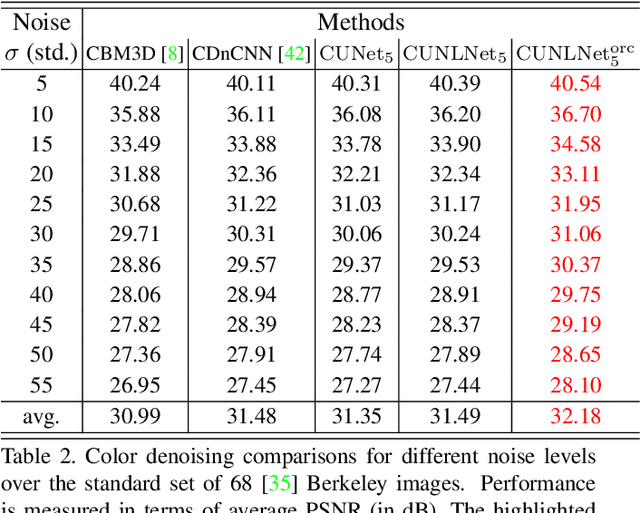
We design a novel network architecture for learning discriminative image models that are employed to efficiently tackle the problem of grayscale and color image denoising. Based on the proposed architecture, we introduce two different variants. The first network involves convolutional layers as a core component, while the second one relies instead on non-local filtering layers and thus it is able to exploit the inherent non-local self-similarity property of natural images. As opposed to most of the existing deep network approaches, which require the training of a specific model for each considered noise level, the proposed models are able to handle a wide range of noise levels using a single set of learned parameters, while they are very robust when the noise degrading the latent image does not match the statistics of the noise used during training. The latter argument is supported by results that we report on publicly available images corrupted by unknown noise and which we compare against solutions obtained by competing methods. At the same time the introduced networks achieve excellent results under additive white Gaussian noise (AWGN), which are comparable to those of the current state-of-the-art network, while they depend on a more shallow architecture with the number of trained parameters being one order of magnitude smaller. These properties make the proposed networks ideal candidates to serve as sub-solvers on restoration methods that deal with general inverse imaging problems such as deblurring, demosaicking, superresolution, etc.
Plexus Convolutional Neural Network (PlexusNet): A novel neural network architecture for histologic image analysis
Aug 24, 2019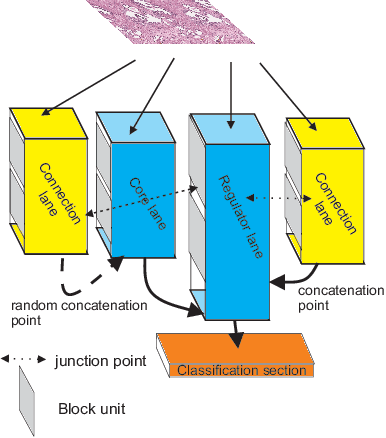
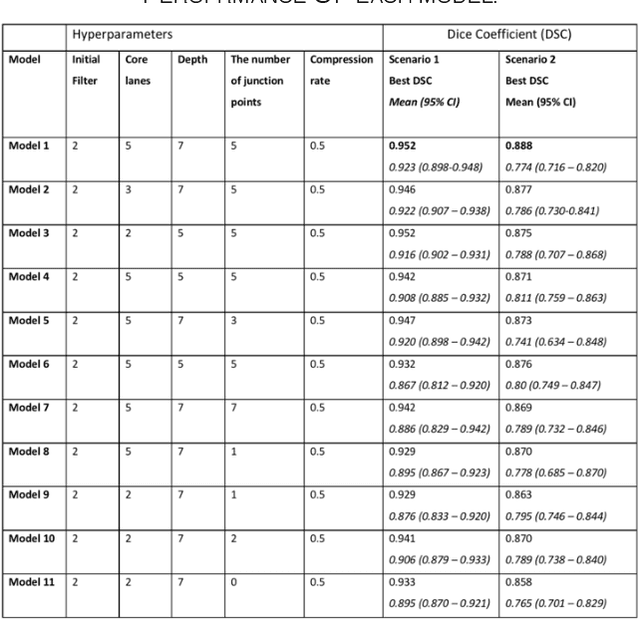
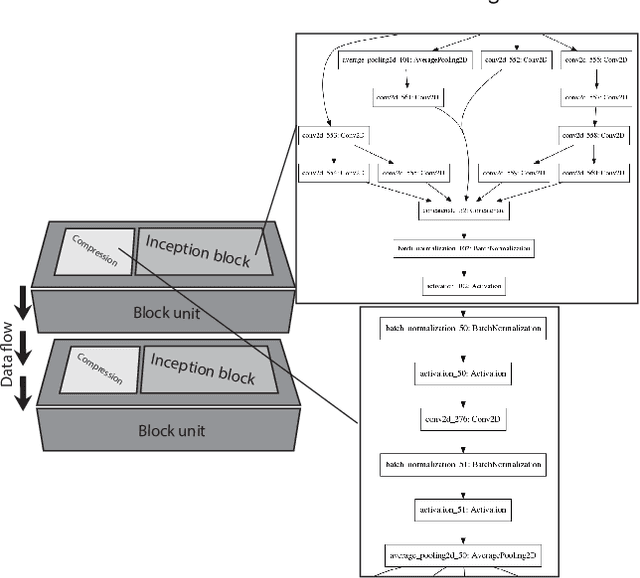
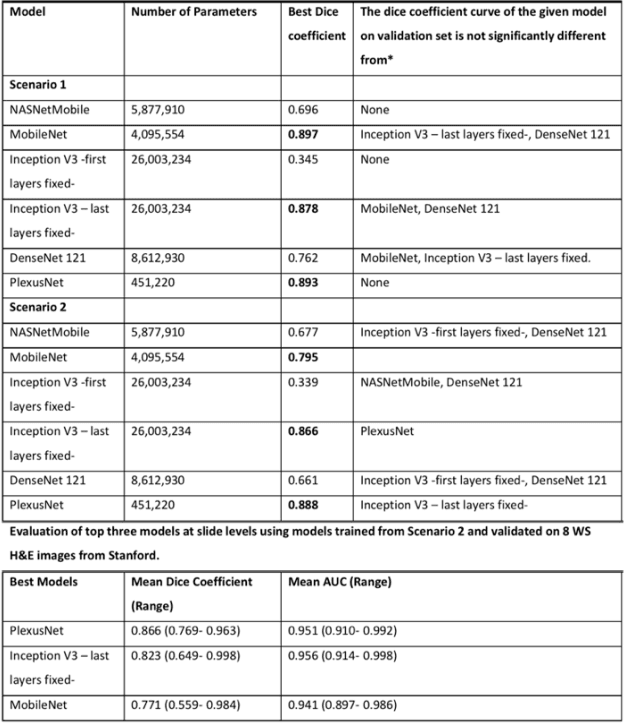
Different convolutional neural network (CNN) models have been tested for their application in histologic imaging analyses. However, these models are prone to overfitting due to their large parameter capacity, requiring more data and expensive computational resources for model training. Given these limitations, we developed and tested PlexusNet for histologic evaluation using a single GPU by a batch dimension of 16x512x512x3. We utilized 62 Hematoxylin and eosin stain (H&E) annotated histological images of radical prostatectomy cases from TCGA-PRAD and Stanford University, and 24 H&E whole-slide images with hepatocellular carcinoma from TCGA-LIHC diagnostic histology images. Base models were DenseNet, Inception V3, and MobileNet and compared with PlexusNet. The dice coefficient (DSC) was evaluated for each model. PlexusNet delivered comparable classification performance (DSC at patch level: 0.89) for H&E whole-slice images in distinguishing prostate cancer from normal tissues. The parameter capacity of PlexusNet is 9 times smaller than MobileNet or 58 times smaller than Inception V3, respectively. Similar findings were observed in distinguishing hepatocellular carcinoma from non-cancerous liver histologies (DSC at patch level: 0.85). As conclusion, PlexusNet represents a novel model architecture for histological image analysis that achieves classification performance comparable to the base models while providing orders-of-magnitude memory savings.
Unsupervised Shadow Removal Using Target Consistency Generative Adversarial Network
Oct 03, 2020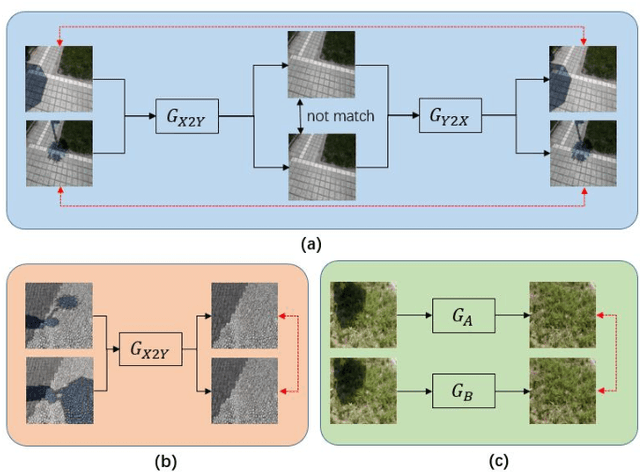
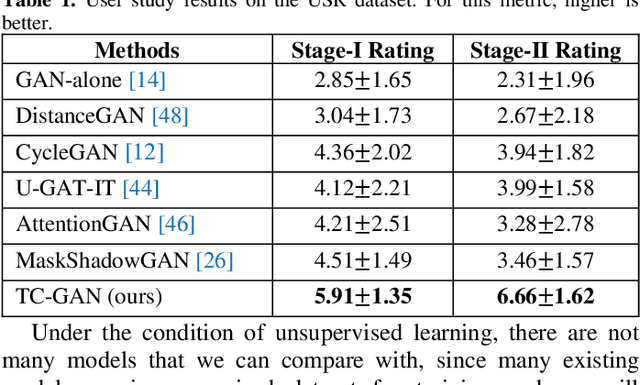
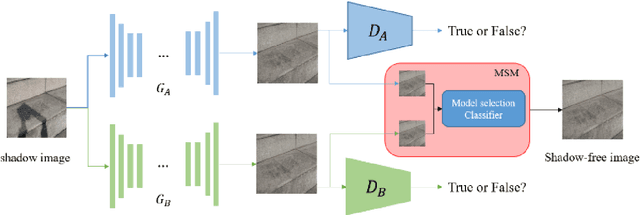
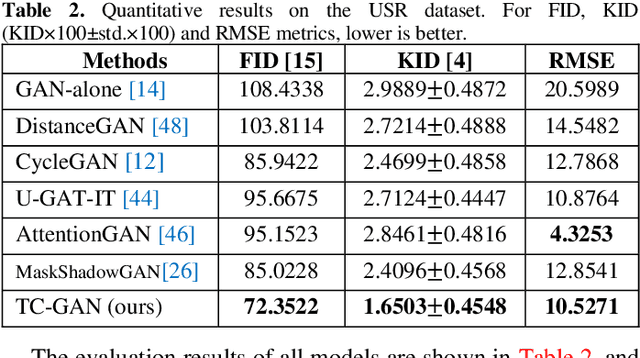
Unsupervised shadow removal aims to learn a non-linear function to map the original image from shadow domain to non-shadow domain in the absence of paired shadow and non-shadow data. In this paper, we develop a simple yet efficient target-consistency generative adversarial network (TC-GAN) for the shadow removal task in the unsupervised manner. Compared with the bidirectional mapping in cycle-consistency GAN based methods for shadow removal, TC-GAN tries to learn a one-sided mapping to cast shadow images into shadow-free ones. With the proposed target-consistency constraint, the correlations between shadow images and the output shadow-free image are strictly confined. Extensive comparison experiments results show that TC-GAN outperforms the state-of-the-art unsupervised shadow removal methods by 14.9% in terms of FID and 31.5% in terms of KID. It is rather remarkable that TC-GAN achieves comparable performance with supervised shadow removal methods.
Crop Classification under Varying Cloud Cover with Neural Ordinary Differential Equations
Dec 04, 2020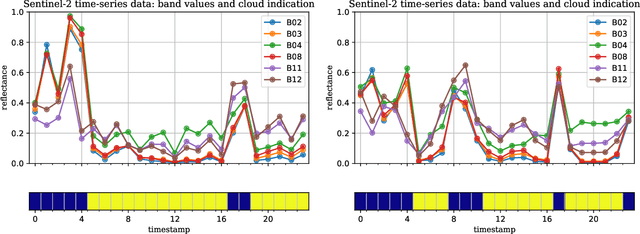
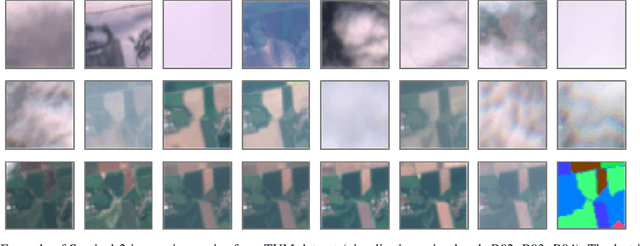

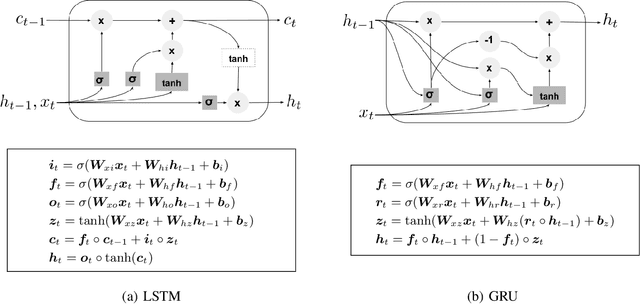
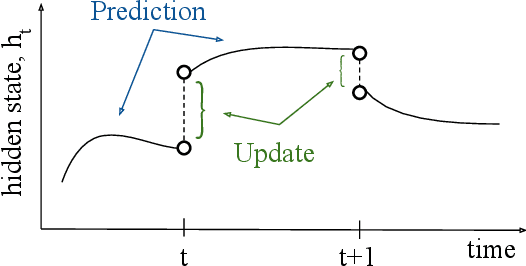
Optical satellite sensors cannot see the Earth's surface through clouds. Despite the periodic revisit cycle, image sequences acquired by Earth observation satellites are therefore irregularly sampled in time. State-of-the-art methods for crop classification (and other time series analysis tasks) rely on techniques that implicitly assume regular temporal spacing between observations, such as recurrent neural networks (RNNs). We propose to use neural ordinary differential equations (NODEs) in combination with RNNs to classify crop types in irregularly spaced image sequences. The resulting ODE-RNN models consist of two steps: an update step, where a recurrent unit assimilates new input data into the model's hidden state; and a prediction step, in which NODE propagates the hidden state until the next observation arrives. The prediction step is based on a continuous representation of the latent dynamics, which has several advantages. At the conceptual level, it is a more natural way to describe the mechanisms that govern the phenological cycle. From a practical point of view, it makes it possible to sample the system state at arbitrary points in time, such that one can integrate observations whenever they are available, and extrapolate beyond the last observation. Our experiments show that ODE-RNN indeed improves classification accuracy over common baselines such as LSTM, GRU, and temporal convolution. The gains are most prominent in the challenging scenario where only few observations are available (i.e., frequent cloud cover). Moreover, we show that the ability to extrapolate translates to better classification performance early in the season, which is important for forecasting.
Classifying high-dimensional Gaussian mixtures: Where kernel methods fail and neural networks succeed
Feb 23, 2021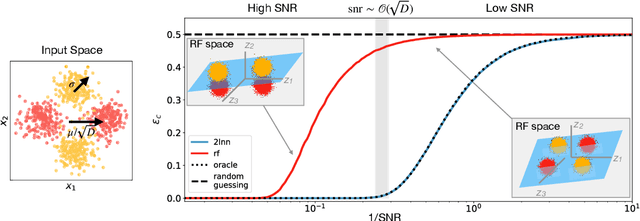
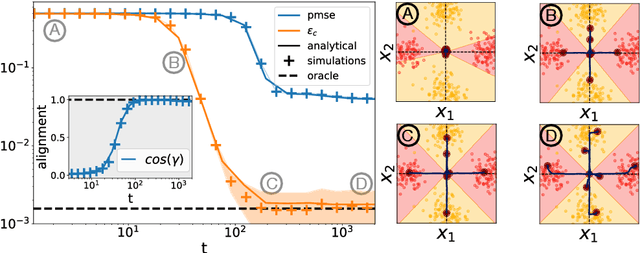
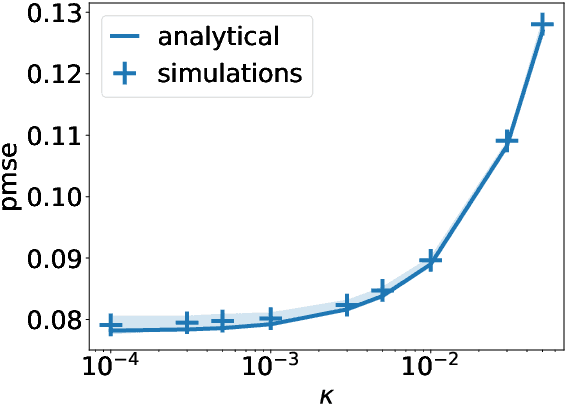
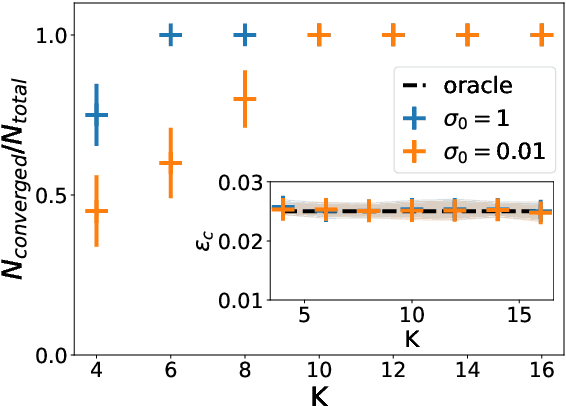
A recent series of theoretical works showed that the dynamics of neural networks with a certain initialisation are well-captured by kernel methods. Concurrent empirical work demonstrated that kernel methods can come close to the performance of neural networks on some image classification tasks. These results raise the question of whether neural networks only learn successfully if kernels also learn successfully, despite neural networks being more expressive. Here, we show theoretically that two-layer neural networks (2LNN) with only a few hidden neurons can beat the performance of kernel learning on a simple Gaussian mixture classification task. We study the high-dimensional limit where the number of samples is linearly proportional to the input dimension, and show that while small 2LNN achieve near-optimal performance on this task, lazy training approaches such as random features and kernel methods do not. Our analysis is based on the derivation of a closed set of equations that track the learning dynamics of the 2LNN and thus allow to extract the asymptotic performance of the network as a function of signal-to-noise ratio and other hyperparameters. We finally illustrate how over-parametrising the neural network leads to faster convergence, but does not improve its final performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge