"Image": models, code, and papers
Image Inpainting for Irregular Holes Using Partial Convolutions
Apr 20, 2018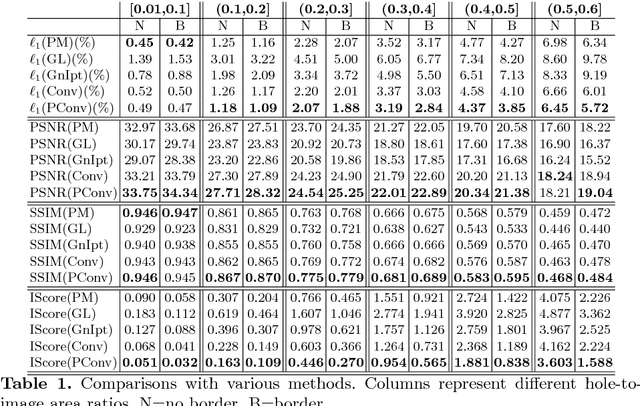
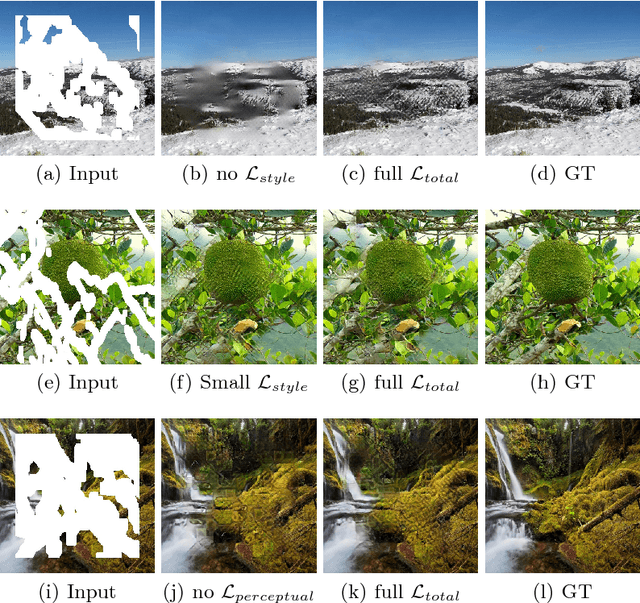
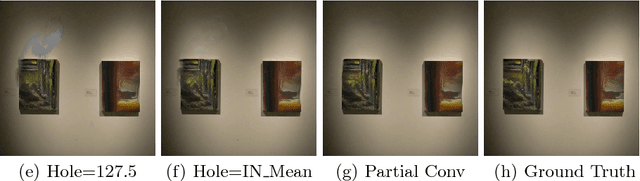

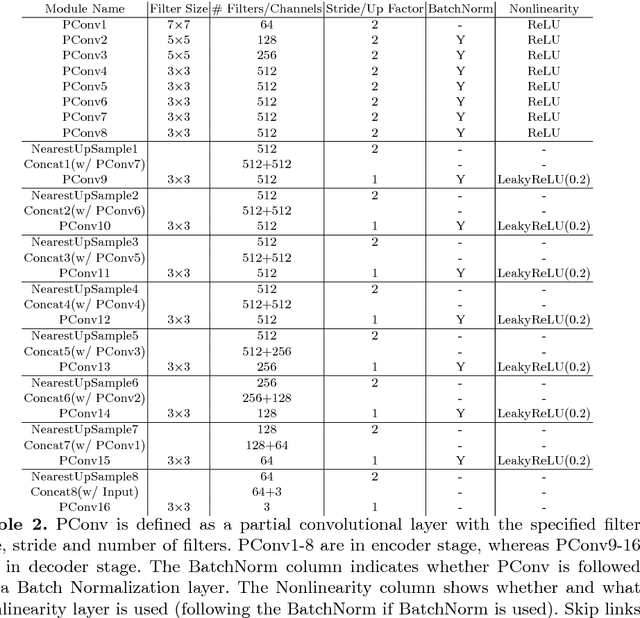

Existing deep learning based image inpainting methods use a standard convolutional network over the corrupted image, using convolutional filter responses conditioned on both valid pixels as well as the substitute values in the masked holes (typically the mean value). This often leads to artifacts such as color discrepancy and blurriness. Post-processing is usually used to reduce such artifacts, but are expensive and may fail. We propose the use of partial convolutions, where the convolution is masked and renormalized to be conditioned on only valid pixels. We further include a mechanism to automatically generate an updated mask for the next layer as part of the forward pass. Our model outperforms other methods for irregular masks. We show qualitative and quantitative comparisons with other methods to validate our approach.
Causal Intervention for Weakly-Supervised Semantic Segmentation
Oct 07, 2020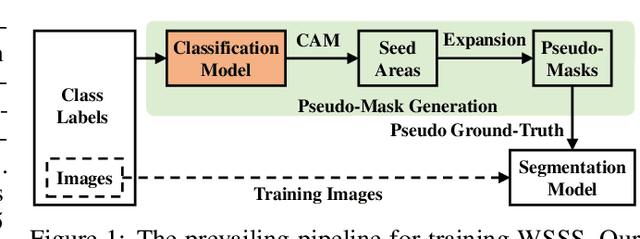
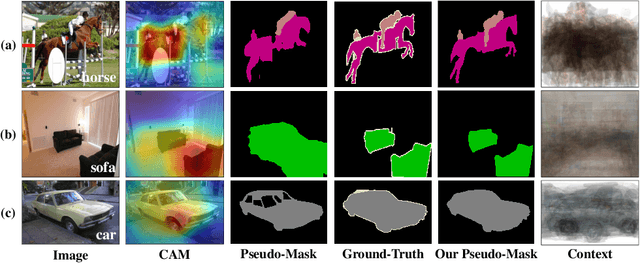
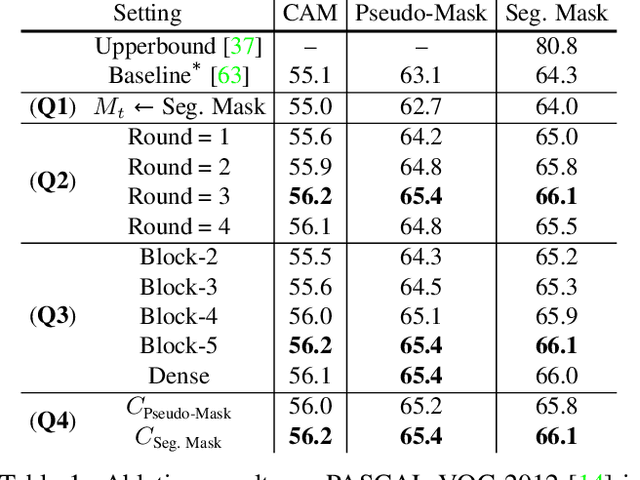

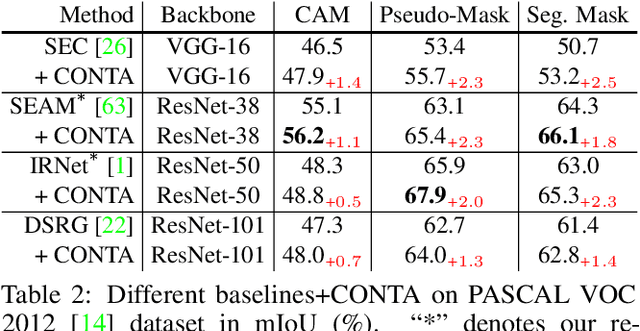
We present a causal inference framework to improve Weakly-Supervised Semantic Segmentation (WSSS). Specifically, we aim to generate better pixel-level pseudo-masks by using only image-level labels -- the most crucial step in WSSS. We attribute the cause of the ambiguous boundaries of pseudo-masks to the confounding context, e.g., the correct image-level classification of "horse" and "person" may be not only due to the recognition of each instance, but also their co-occurrence context, making the model inspection (e.g., CAM) hard to distinguish between the boundaries. Inspired by this, we propose a structural causal model to analyze the causalities among images, contexts, and class labels. Based on it, we develop a new method: Context Adjustment (CONTA), to remove the confounding bias in image-level classification and thus provide better pseudo-masks as ground-truth for the subsequent segmentation model. On PASCAL VOC 2012 and MS-COCO, we show that CONTA boosts various popular WSSS methods to new state-of-the-arts.
ChallenCap: Monocular 3D Capture of Challenging Human Performances using Multi-Modal References
Mar 29, 2021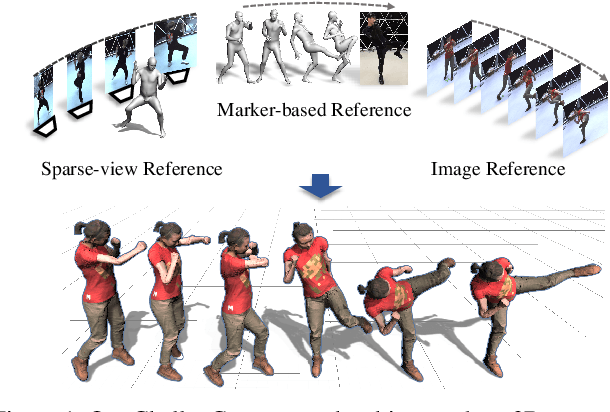
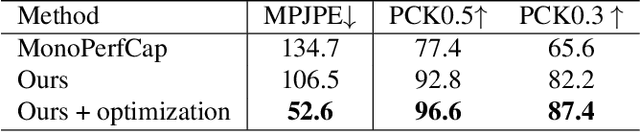
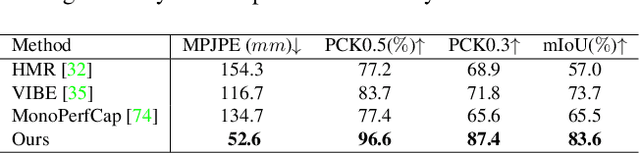
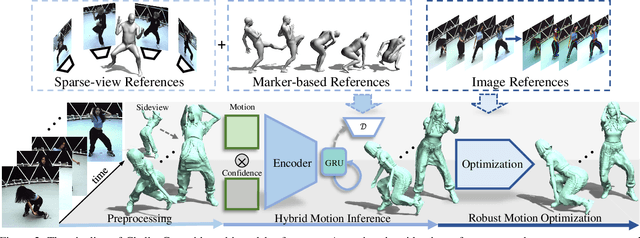

Capturing challenging human motions is critical for numerous applications, but it suffers from complex motion patterns and severe self-occlusion under the monocular setting. In this paper, we propose ChallenCap -- a template-based approach to capture challenging 3D human motions using a single RGB camera in a novel learning-and-optimization framework, with the aid of multi-modal references. We propose a hybrid motion inference stage with a generation network, which utilizes a temporal encoder-decoder to extract the motion details from the pair-wise sparse-view reference, as well as a motion discriminator to utilize the unpaired marker-based references to extract specific challenging motion characteristics in a data-driven manner. We further adopt a robust motion optimization stage to increase the tracking accuracy, by jointly utilizing the learned motion details from the supervised multi-modal references as well as the reliable motion hints from the input image reference. Extensive experiments on our new challenging motion dataset demonstrate the effectiveness and robustness of our approach to capture challenging human motions.
Lung Nodule Classification Using Biomarkers, Volumetric Radiomics and 3D CNNs
Oct 19, 2020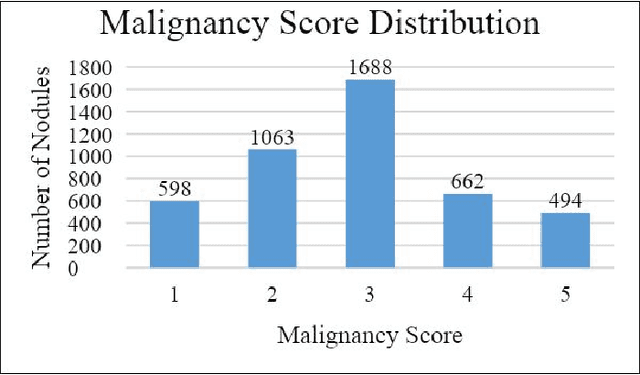
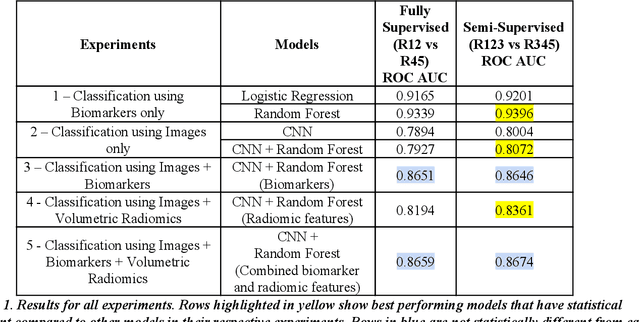
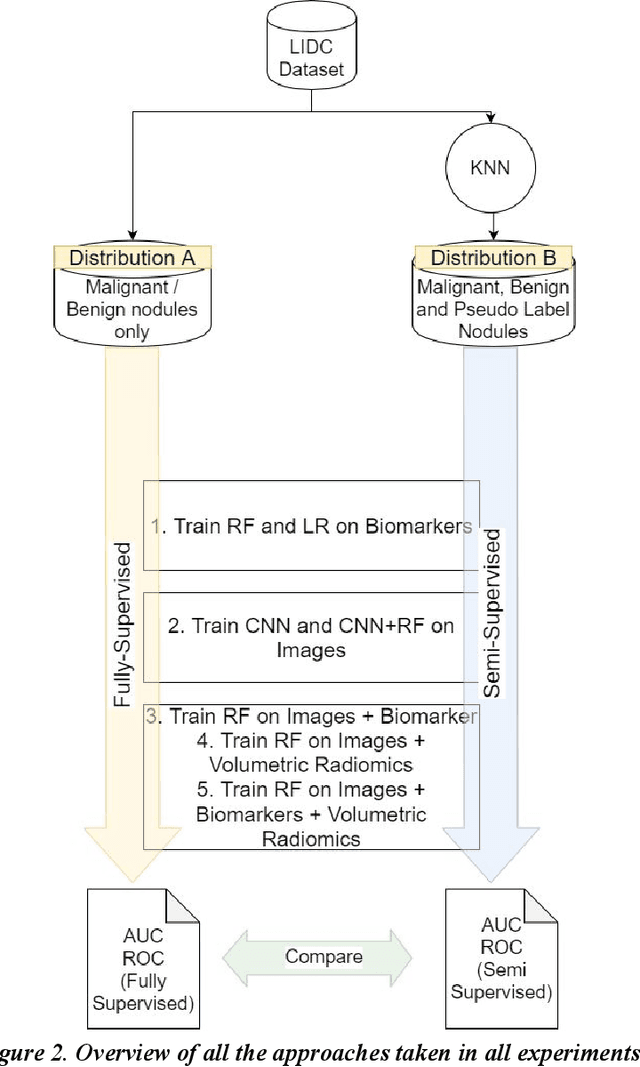
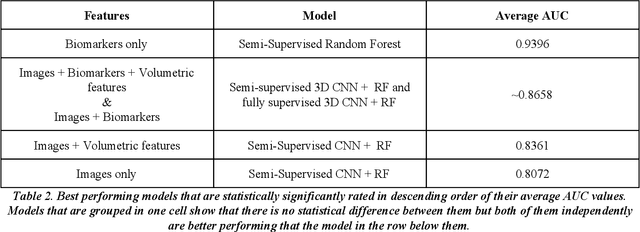
We present a hybrid algorithm to estimate lung nodule malignancy that combines imaging biomarkers from Radiologist's annotation with image classification of CT scans. Our algorithm employs a 3D Convolutional Neural Network (CNN) as well as a Random Forest in order to combine CT imagery with biomarker annotation and volumetric radiomic features. We analyze and compare the performance of the algorithm using only imagery, only biomarkers, combined imagery + biomarkers, combined imagery + volumetric radiomic features and finally the combination of imagery + biomarkers + volumetric features in order to classify the suspicion level of nodule malignancy. The National Cancer Institute (NCI) Lung Image Database Consortium (LIDC) IDRI dataset is used to train and evaluate the classification task. We show that the incorporation of semi-supervised learning by means of K-Nearest-Neighbors (KNN) can increase the available training sample size of the LIDC-IDRI thereby further improving the accuracy of malignancy estimation of most of the models tested although there is no significant improvement with the use of KNN semi-supervised learning if image classification with CNNs and volumetric features are combined with descriptive biomarkers. Unexpectedly, we also show that a model using image biomarkers alone is more accurate than one that combines biomarkers with volumetric radiomics, 3D CNNs, and semi-supervised learning. We discuss the possibility that this result may be influenced by cognitive bias in LIDC-IDRI because malignancy estimates were recorded by the same radiologist panel as biomarkers, as well as future work to incorporate pathology information over a subset of study participants.
Diverse Branch Block: Building a Convolution as an Inception-like Unit
Mar 29, 2021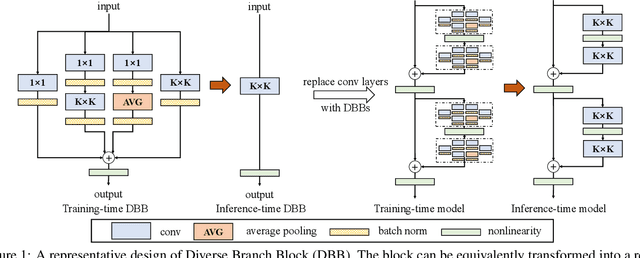
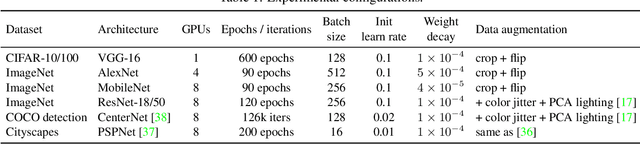
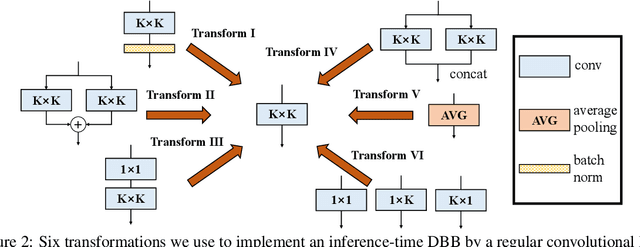
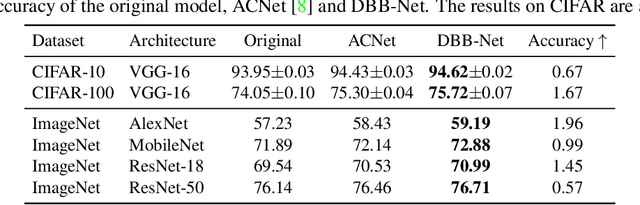
We propose a universal building block of Convolutional Neural Network (ConvNet) to improve the performance without any inference-time costs. The block is named Diverse Branch Block (DBB), which enhances the representational capacity of a single convolution by combining diverse branches of different scales and complexities to enrich the feature space, including sequences of convolutions, multi-scale convolutions, and average pooling. After training, a DBB can be equivalently converted into a single conv layer for deployment. Unlike the advancements of novel ConvNet architectures, DBB complicates the training-time microstructure while maintaining the macro architecture, so that it can be used as a drop-in replacement for regular conv layers of any architecture. In this way, the model can be trained to reach a higher level of performance and then transformed into the original inference-time structure for inference. DBB improves ConvNets on image classification (up to 1.9% higher top-1 accuracy on ImageNet), object detection and semantic segmentation. The PyTorch code and models are released at https://github.com/DingXiaoH/DiverseBranchBlock.
EXSCLAIM! -- An automated pipeline for the construction of labeled materials imaging datasets from literature
Mar 19, 2021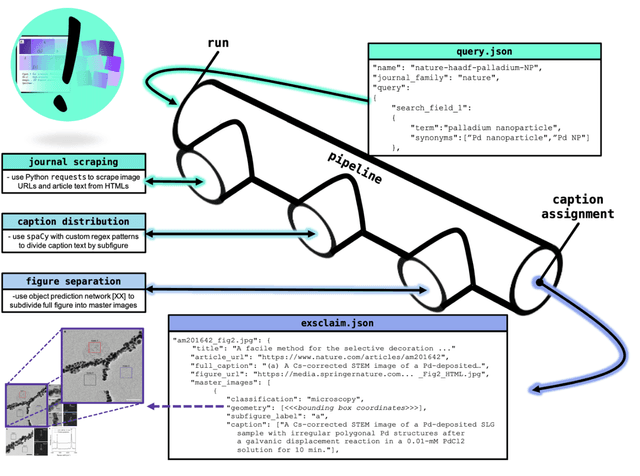
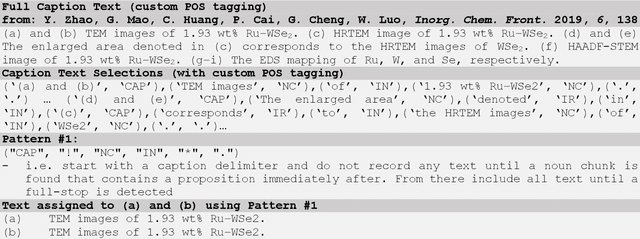
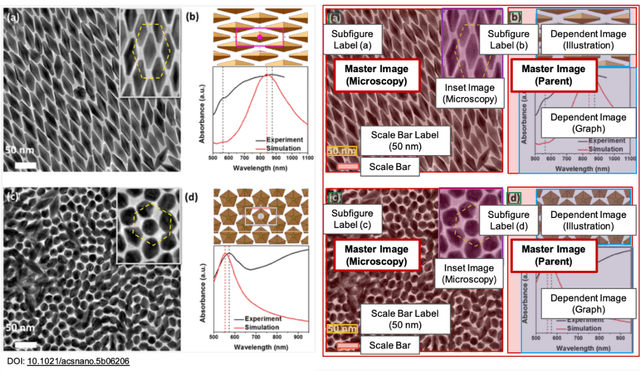
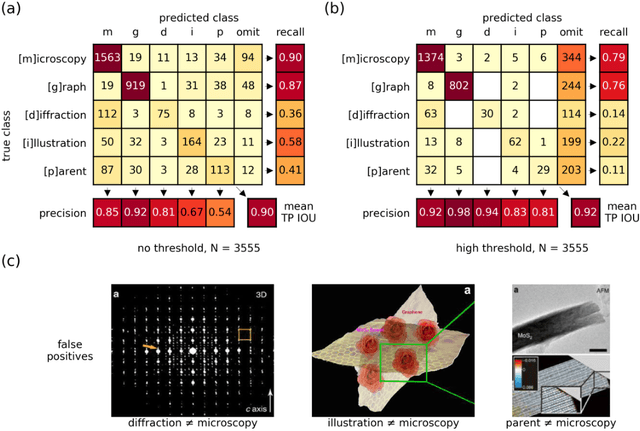
Due to recent improvements in image resolution and acquisition speed, materials microscopy is experiencing an explosion of published imaging data. The standard publication format, while sufficient for traditional data ingestion scenarios where a select number of images can be critically examined and curated manually, is not conducive to large-scale data aggregation or analysis, hindering data sharing and reuse. Most images in publications are presented as components of a larger figure with their explicit context buried in the main body or caption text, so even if aggregated, collections of images with weak or no digitized contextual labels have limited value. To solve the problem of curating labeled microscopy data from literature, this work introduces the EXSCLAIM! Python toolkit for the automatic EXtraction, Separation, and Caption-based natural Language Annotation of IMages from scientific literature. We highlight the methodology behind the construction of EXSCLAIM! and demonstrate its ability to extract and label open-source scientific images at high volume.
Genetic Algorithms for the Optimization of Diffusion Parameters in Content-Based Image Retrieval
Aug 19, 2019
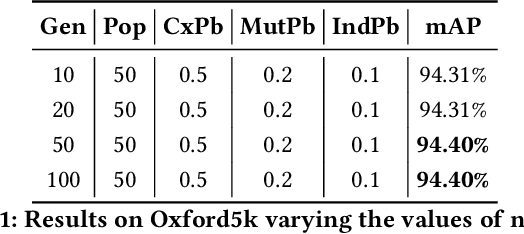
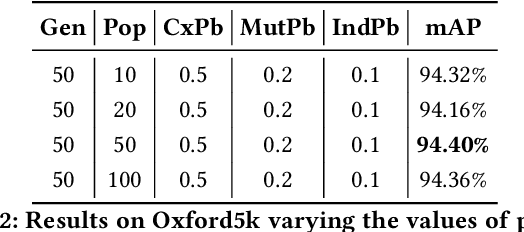
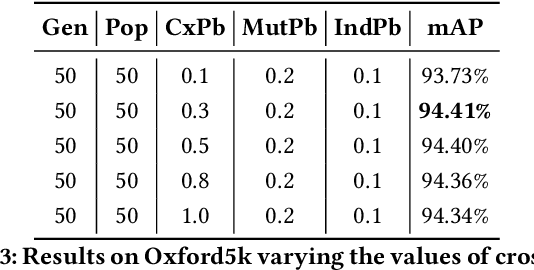
Several computer vision and artificial intelligence projects are nowadays exploiting the manifold data distribution using, e.g., the diffusion process. This approach has produced dramatic improvements on the final performance thanks to the application of such algorithms to the kNN graph. Unfortunately, this recent technique needs a manual configuration of several parameters, thus it is not straightforward to find the best configuration for each dataset. Moreover, the brute-force approach is computationally very demanding when used to optimally set the parameters of the diffusion approach. We propose to use genetic algorithms to find the optimal setting of all the diffusion parameters with respect to retrieval performance for each different dataset. Our approach is faster than others used as references (brute-force, random-search and PSO). A comparison with these methods has been made on three public image datasets: Oxford5k, Paris6k and Oxford105k.
Adaptive Local Structure Consistency based Heterogeneous Remote Sensing Change Detection
Aug 29, 2020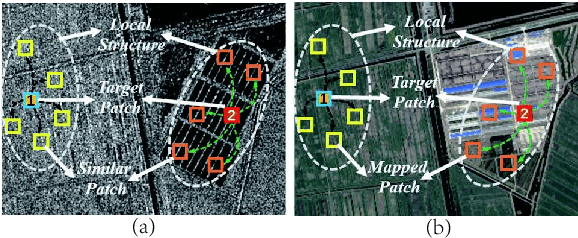
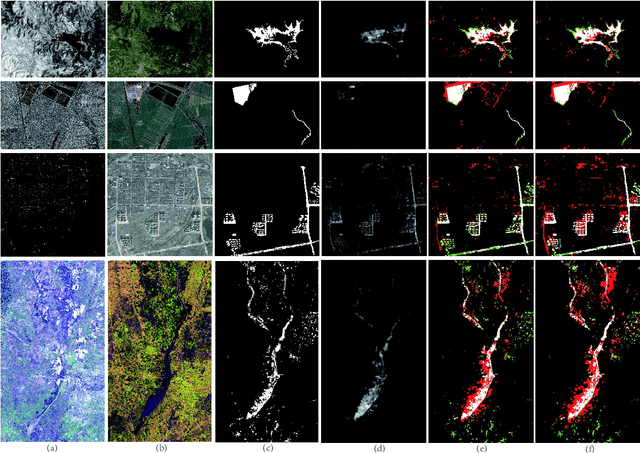
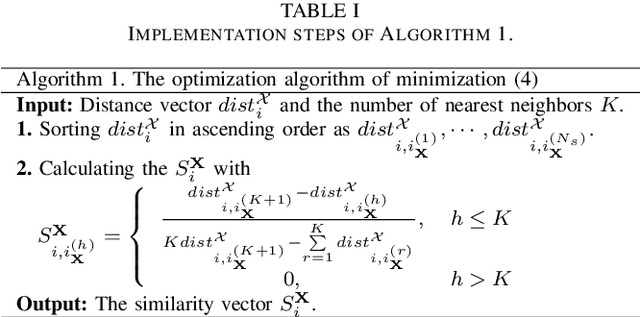

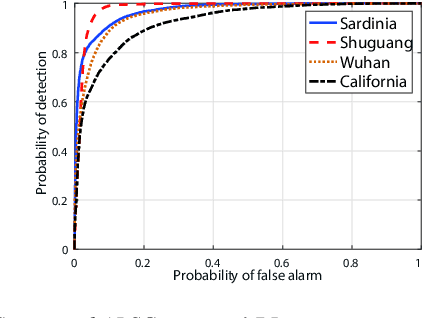

Change detection of heterogeneous remote sensing images is an important and challenging topic in remote sensing for emergency situation resulting from nature disaster. Due to the different imaging mechanisms of heterogeneous sensors, it is difficult to directly compare the images. To address this challenge, we explore an unsupervised change detection method based on adaptive local structure consistency (ALSC) between heterogeneous images in this letter, which constructs an adaptive graph representing the local structure for each patch in one image domain and then projects this graph to the other image domain to measure the change level. This local structure consistency exploits the fact that the heterogeneous images share the same structure information for the same ground object, which is imaging modality-invariant. To avoid the leakage of heterogeneous data, the pixelwise change image is calculated in the same image domain by graph projection. Experiment results demonstrate the effectiveness of the proposed ALSC based change detection method by comparing with some state-of-the-art methods.
SREdgeNet: Edge Enhanced Single Image Super Resolution using Dense Edge Detection Network and Feature Merge Network
Dec 18, 2018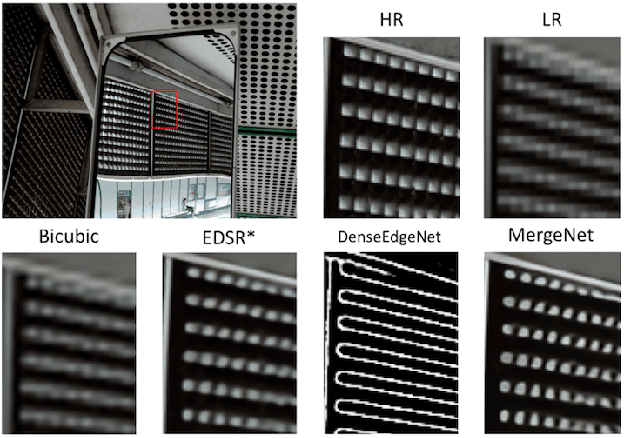
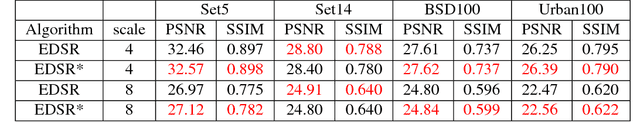
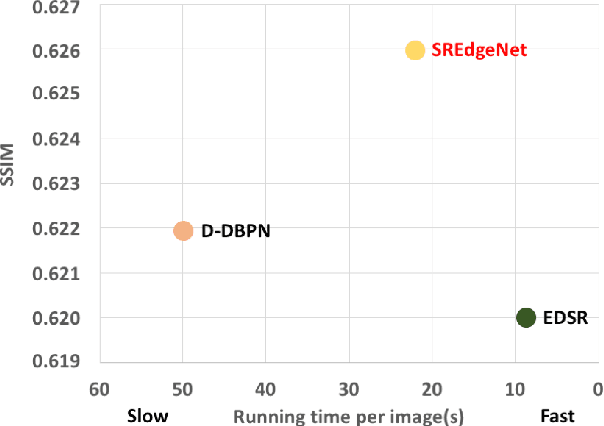
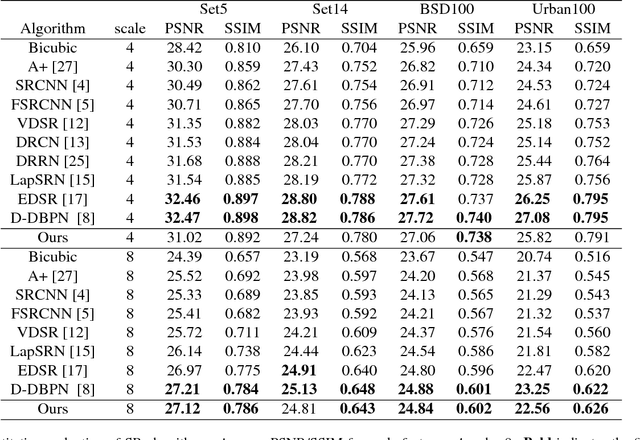
Deep learning based single image super-resolution (SR) methods have been rapidly evolved over the past few years and have yielded state-of-the-art performances over conventional methods. Since these methods usually minimized l1 loss between the output SR image and the ground truth image, they yielded very high peak signal-to-noise ratio (PSNR) that is inversely proportional to these losses. Unfortunately, minimizing these losses inevitably lead to blurred edges due to averaging of plausible solutions. Recently, SRGAN was proposed to avoid this average effect by minimizing perceptual losses instead of l1 loss and it yielded perceptually better SR images (or images with sharp edges) at the price of lowering PSNR. In this paper, we propose SREdgeNet, edge enhanced single image SR network, that was inspired by conventional SR theories so that average effect could be avoided not by changing the loss, but by changing the SR network property with the same l1 loss. Our SREdgeNet consists of 3 sequential deep neural network modules: the first module is any state-of-the-art SR network and we selected a variant of EDSR. The second module is any edge detection network taking the output of the first SR module as an input and we propose DenseEdgeNet for this module. Lastly, the third module is merging the outputs of the first and second modules to yield edge enhanced SR image and we propose MergeNet for this module. Qualitatively, our proposed method yielded images with sharp edges compared to other state-of-the-art SR methods. Quantitatively, our SREdgeNet yielded state-of-the-art performance in terms of structural similarity (SSIM) while maintained comparable PSNR for x8 enlargement.
Real-Time COVID-19 Diagnosis from X-Ray Images Using Deep CNN and Extreme Learning Machines Stabilized by Chimp Optimization Algorithm
May 14, 2021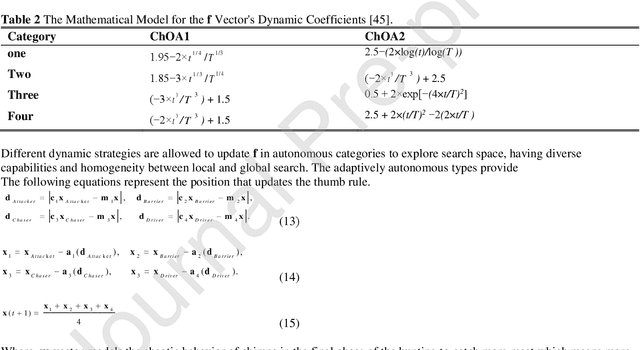
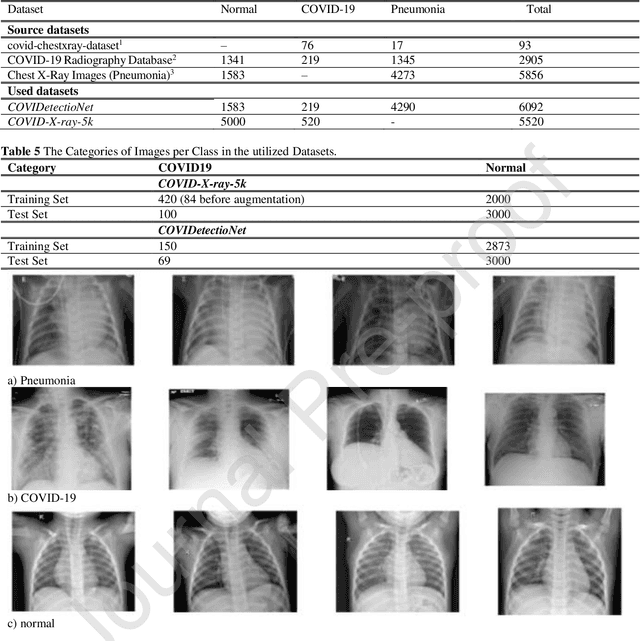
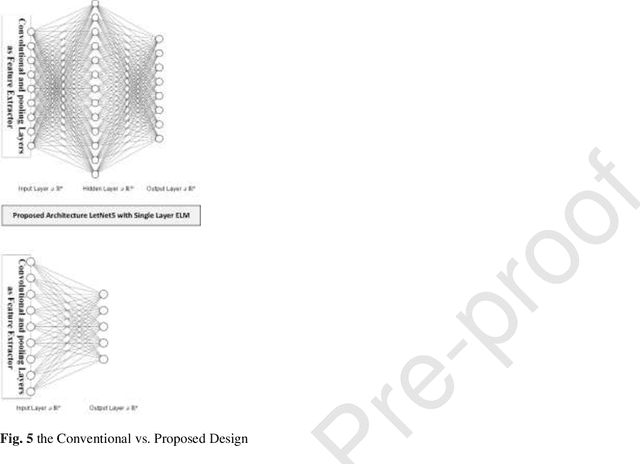
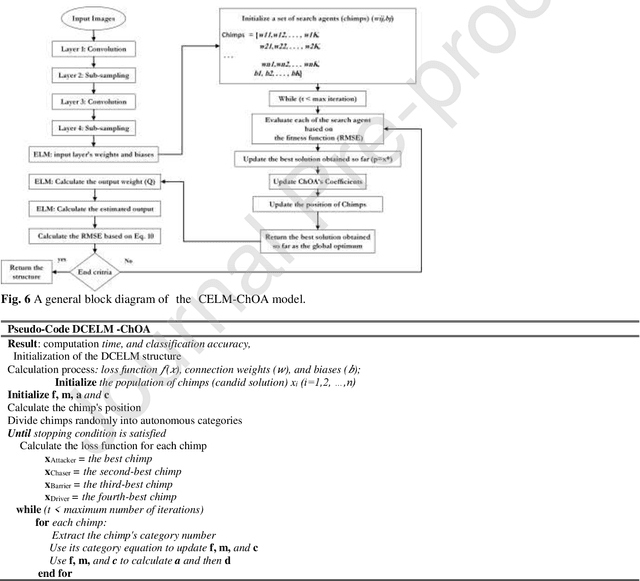
Real-time detection of COVID-19 using radiological images has gained priority due to the increasing demand for fast diagnosis of COVID-19 cases. This paper introduces a novel two-phase approach for classifying chest X-ray images. Deep Learning (DL) methods fail to cover these aspects since training and fine-tuning the model's parameters consume much time. In this approach, the first phase comes to train a deep CNN working as a feature extractor, and the second phase comes to use Extreme Learning Machines (ELMs) for real-time detection. The main drawback of ELMs is to meet the need of a large number of hidden-layer nodes to gain a reliable and accurate detector in applying image processing since the detective performance remarkably depends on the setting of initial weights and biases. Therefore, this paper uses Chimp Optimization Algorithm (ChOA) to improve results and increase the reliability of the network while maintaining real-time capability. The designed detector is to be benchmarked on the COVID-Xray-5k and COVIDetectioNet datasets, and the results are verified by comparing it with the classic DCNN, Genetic Algorithm optimized ELM (GA-ELM), Cuckoo Search optimized ELM (CS-ELM), and Whale Optimization Algorithm optimized ELM (WOA-ELM). The proposed approach outperforms other comparative benchmarks with 98.25% and 99.11% as ultimate accuracy on the COVID-Xray-5k and COVIDetectioNet datasets, respectively, and it led relative error to reduce as the amount of 1.75% and 1.01% as compared to a convolutional CNN. More importantly, the time needed for training deep ChOA-ELM is only 0.9474 milliseconds, and the overall testing time for 3100 images is 2.937 seconds.
* 17 pages. arXiv admin note: text overlap with arXiv:2105.14192
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge