"Image": models, code, and papers
Self-supervised driven consistency training for annotation efficient histopathology image analysis
Feb 09, 2021
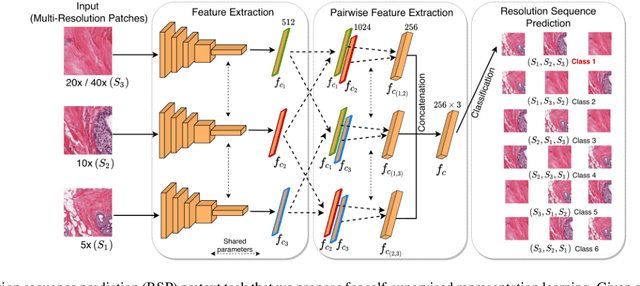
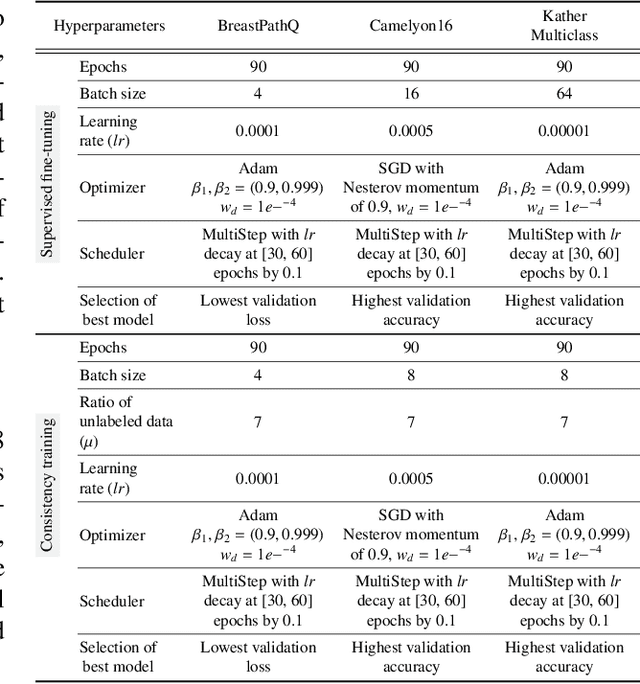
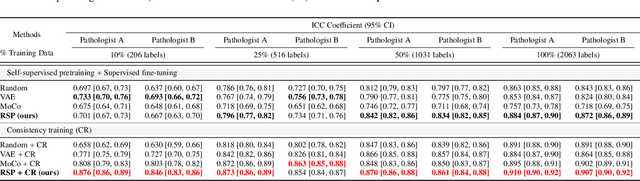
Training a neural network with a large labeled dataset is still a dominant paradigm in computational histopathology. However, obtaining such exhaustive manual annotations is often expensive, laborious, and prone to inter and Intra-observer variability. While recent self-supervised and semi-supervised methods can alleviate this need by learn-ing unsupervised feature representations, they still struggle to generalize well to downstream tasks when the number of labeled instances is small. In this work, we overcome this challenge by leveraging both task-agnostic and task-specific unlabeled data based on two novel strategies: i) a self-supervised pretext task that harnesses the underlying multi-resolution contextual cues in histology whole-slide images to learn a powerful supervisory signal for unsupervised representation learning; ii) a new teacher-student semi-supervised consistency paradigm that learns to effectively transfer the pretrained representations to downstream tasks based on prediction consistency with the task-specific un-labeled data. We carry out extensive validation experiments on three histopathology benchmark datasets across two classification and one regression-based tasks, i.e., tumor metastasis detection, tissue type classification, and tumor cellularity quantification. Under limited-label data, the proposed method yields tangible improvements, which is close or even outperforming other state-of-the-art self-supervised and supervised baselines. Furthermore, we empirically show that the idea of bootstrapping the self-supervised pretrained features is an effective way to improve the task-specific semi-supervised learning on standard benchmarks. Code and pretrained models will be made available at: https://github.com/srinidhiPY/SSL_CR_Histo
Object Localization Through a Single Multiple-Model Convolutional Neural Network with a Specific Training Approach
Mar 24, 2021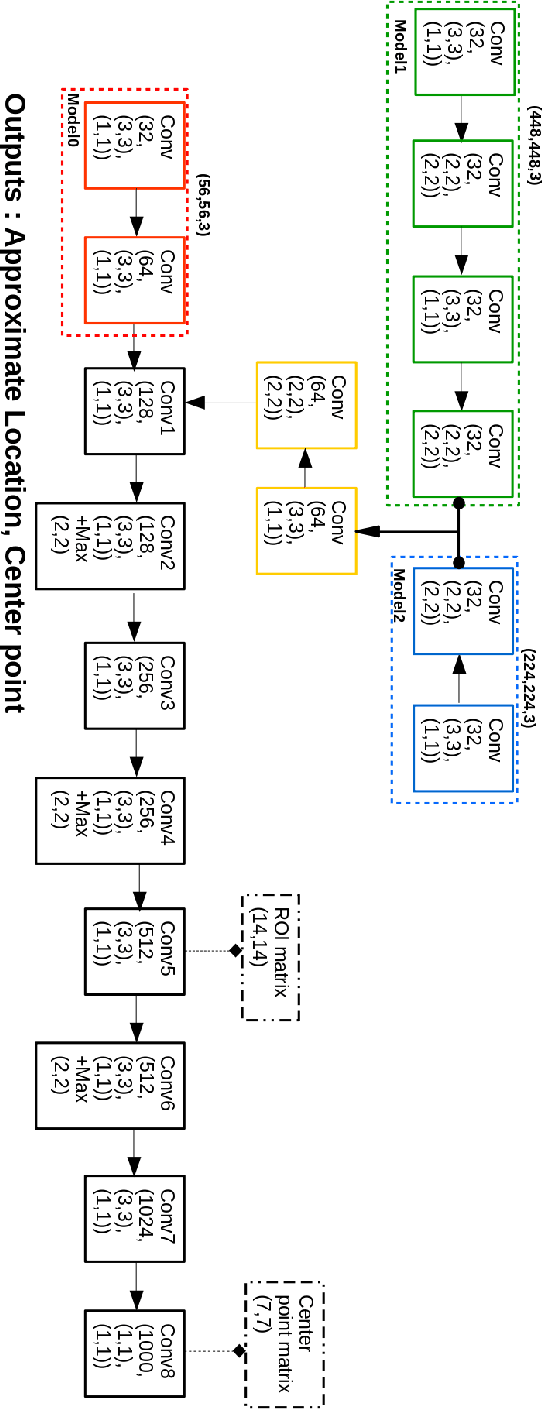
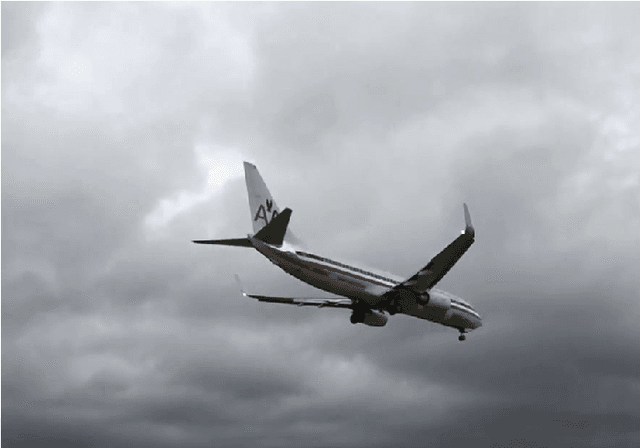


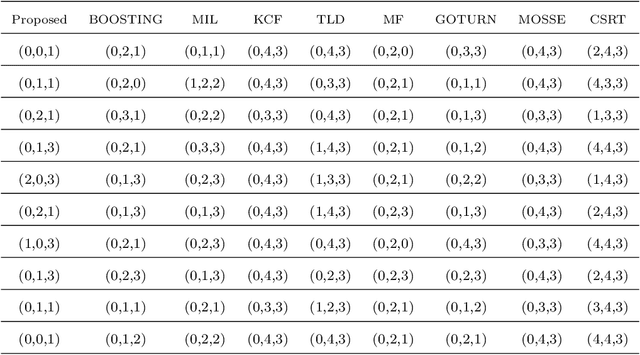
Object localization has a vital role in any object detector, and therefore, has been the focus of attention by many researchers. In this article, a special training approach is proposed for a light convolutional neural network (CNN) to determine the region of interest (ROI) in an image while effectively reducing the number of probable anchor boxes. Almost all CNN-based detectors utilize a fixed input size image, which may yield poor performance when dealing with various object sizes. In this paper, a different CNN structure is proposed taking three different input sizes, to enhance the performance. In order to demonstrate the effectiveness of the proposed method, two common data set are used for training while tracking by localization application is considered to demonstrate its final performance. The promising results indicate the applicability of the presented structure and the training method in practice.
Training Data Independent Image Registration With GANs Using Transfer Learning And Segmentation Information
Apr 11, 2019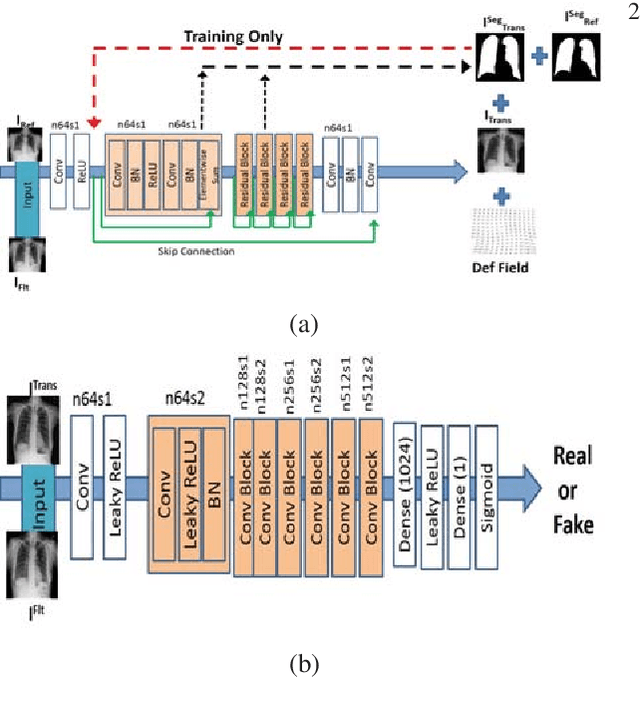
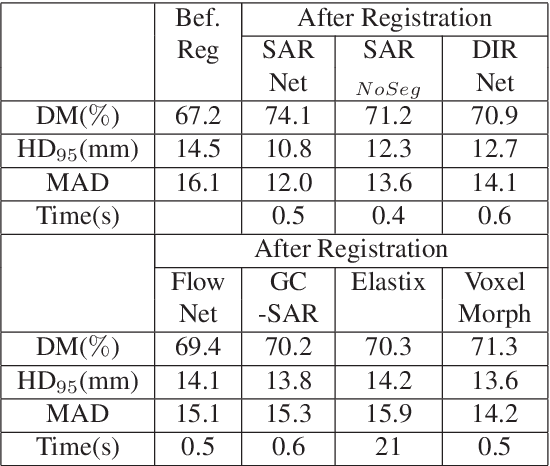
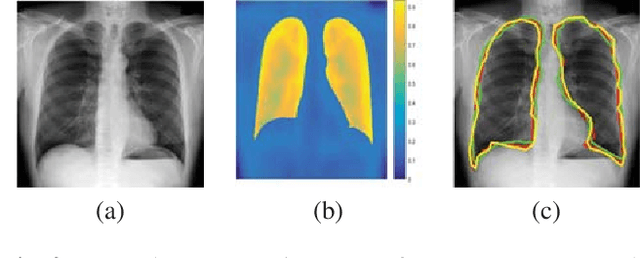
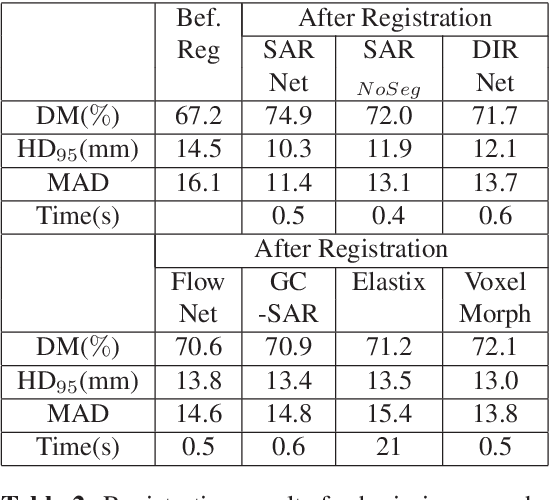
Registration is an important task in automated medical image analysis. Although deep learning (DL) based image registration methods out perform time consuming conventional approaches, they are heavily dependent on training data and do not generalize well for new images types. We present a DL based approach that can register an image pair which is different from the training images. This is achieved by training generative adversarial networks (GANs) in combination with segmentation information and transfer learning. Experiments on chest Xray and brain MR images show that our method gives better registration performance over conventional methods.
Efficient and Generic Interactive Segmentation Framework to Correct Mispredictions during Clinical Evaluation of Medical Images
Aug 06, 2021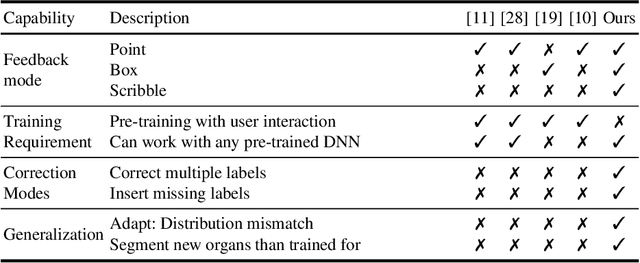
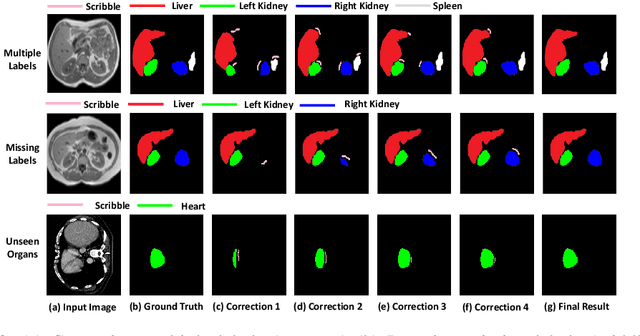
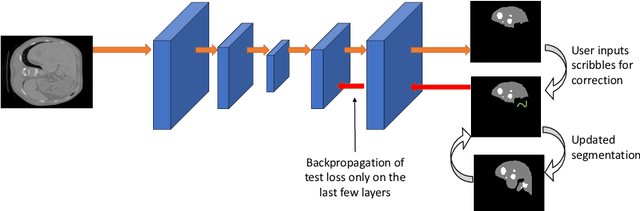
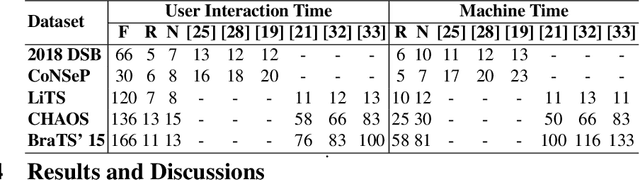

Semantic segmentation of medical images is an essential first step in computer-aided diagnosis systems for many applications. However, given many disparate imaging modalities and inherent variations in the patient data, it is difficult to consistently achieve high accuracy using modern deep neural networks (DNNs). This has led researchers to propose interactive image segmentation techniques where a medical expert can interactively correct the output of a DNN to the desired accuracy. However, these techniques often need separate training data with the associated human interactions, and do not generalize to various diseases, and types of medical images. In this paper, we suggest a novel conditional inference technique for DNNs which takes the intervention by a medical expert as test time constraints and performs inference conditioned upon these constraints. Our technique is generic can be used for medical images from any modality. Unlike other methods, our approach can correct multiple structures simultaneously and add structures missed at initial segmentation. We report an improvement of 13.3, 12.5, 17.8, 10.2, and 12.4 times in user annotation time than full human annotation for the nucleus, multiple cells, liver and tumor, organ, and brain segmentation respectively. We report a time saving of 2.8, 3.0, 1.9, 4.4, and 8.6 fold compared to other interactive segmentation techniques. Our method can be useful to clinicians for diagnosis and post-surgical follow-up with minimal intervention from the medical expert. The source-code and the detailed results are available here [1].
UU-Nets Connecting Discriminator and Generator for Image to Image Translation
Apr 04, 2019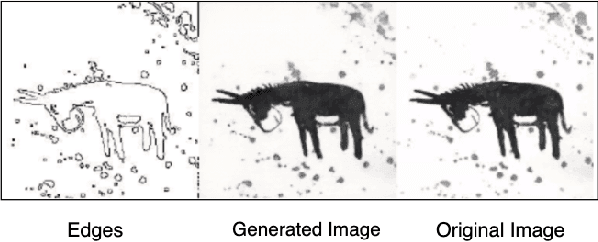
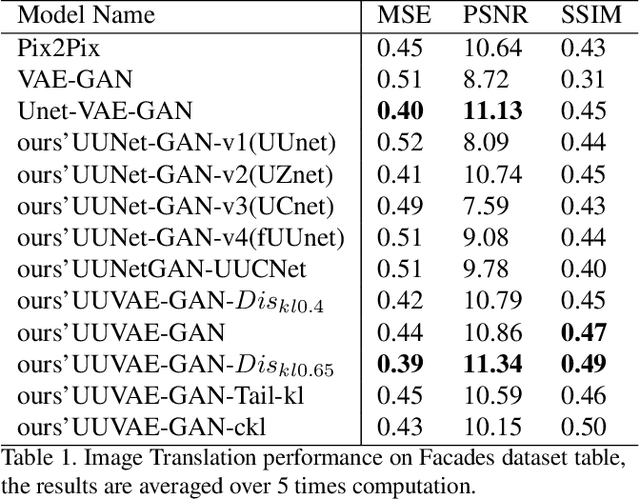
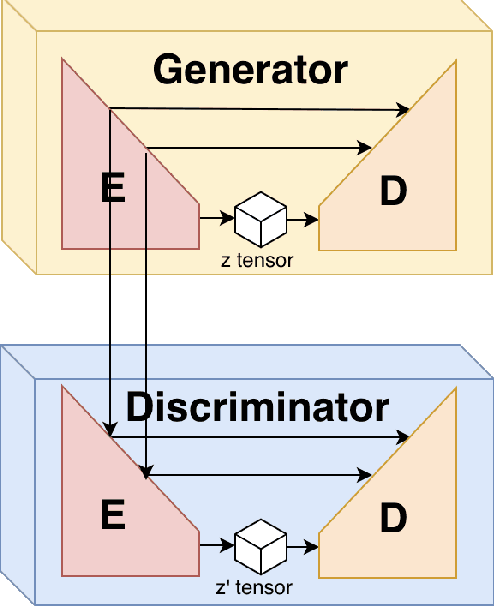
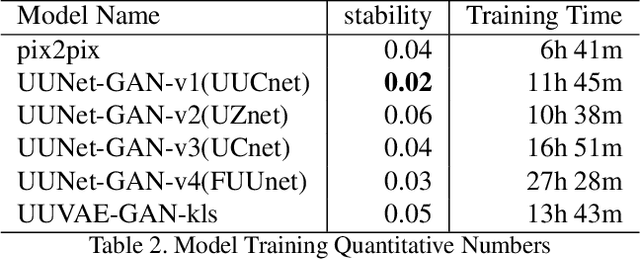
Adversarial generative model have successfully manifest itself in image synthesis. However, the performance deteriorate and unstable, because discriminator is far stable than generator, and it is hard to control the game between the two modules. Various methods have been introduced to tackle the problem such as WGAN, Relativistic GAN and their successors by adding or restricting the loss function, which certainly help balance the min-max game, but they all focused on the loss function ignoring the intrinsic structure limitation. We present a UU-Net architecture inspired by U-net bridging the encoder and the decoder, UU-Net composed by two U-Net liked modules respectively served as generator and discriminator. Because the modules in U-net are symmetrical, therefore it shares weights easily between all four components. Thanks to UU-net's modules identical and symmetric property, we could not only carried the features from inner generator's encoder to its decoder, but also to the discriminator's encoder and decoder. By this design, it give us more control and condition flexibility to intervene the process between the generator and the discriminator.
Rapid Detection of Aircrafts in Satellite Imagery based on Deep Neural Networks
Apr 21, 2021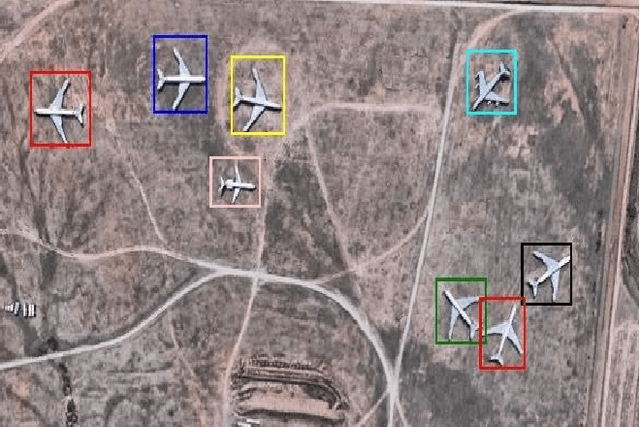
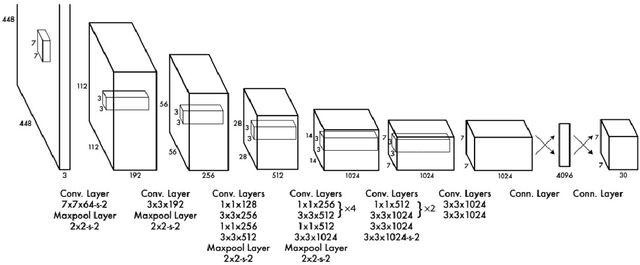
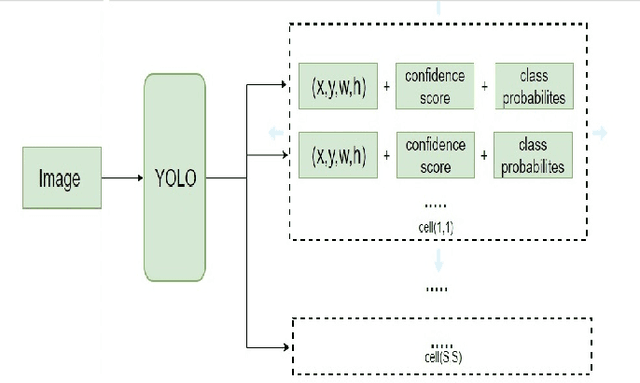
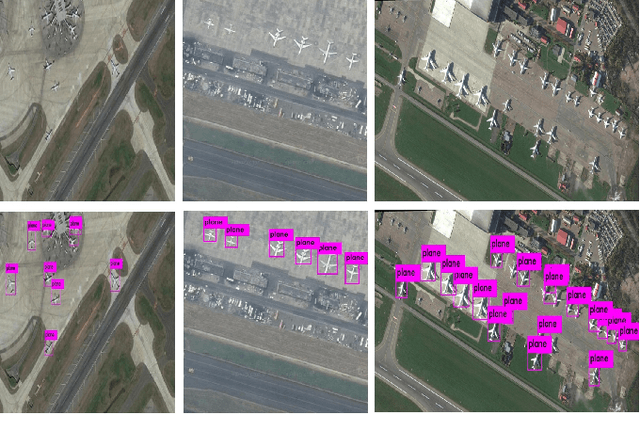
Object detection is one of the fundamental objectives in Applied Computer Vision. In some of the applications, object detection becomes very challenging such as in the case of satellite image processing. Satellite image processing has remained the focus of researchers in domains of Precision Agriculture, Climate Change, Disaster Management, etc. Therefore, object detection in satellite imagery is one of the most researched problems in this domain. This paper focuses on aircraft detection. in satellite imagery using deep learning techniques. In this paper, we used YOLO deep learning framework for aircraft detection. This method uses satellite images collected by different sources as learning for the model to perform detection. Object detection in satellite images is mostly complex because objects have many variations, types, poses, sizes, complex and dense background. YOLO has some limitations for small size objects (less than$\sim$32 pixels per object), therefore we upsample the prediction grid to reduce the coarseness of the model and to accurately detect the densely clustered objects. The improved model shows good accuracy and performance on different unknown images having small, rotating, and dense objects to meet the requirements in real-time.
Align Yourself: Self-supervised Pre-training for Fine-grained Recognition via Saliency Alignment
Jun 30, 2021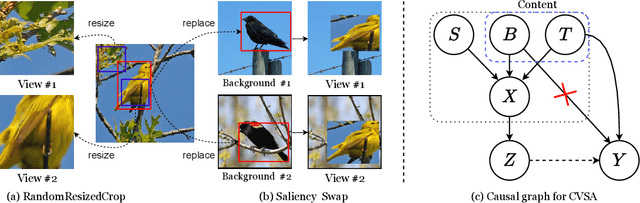
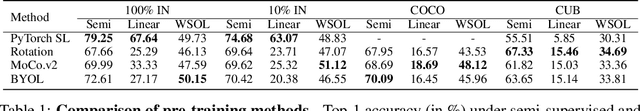
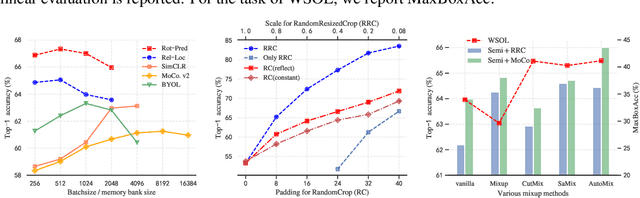
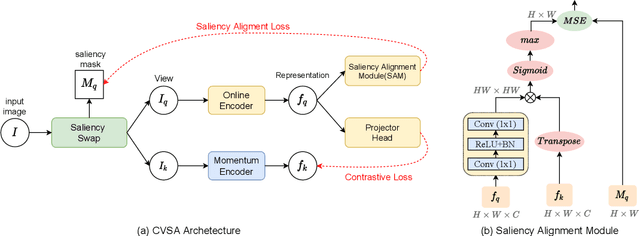
Self-supervised contrastive learning has demonstrated great potential in learning visual representations. Despite their success on various downstream tasks such as image classification and object detection, self-supervised pre-training for fine-grained scenarios is not fully explored. In this paper, we first point out that current contrastive methods are prone to memorizing background/foreground texture and therefore have a limitation in localizing the foreground object. Analysis suggests that learning to extract discriminative texture information and localization are equally crucial for self-supervised pre-training under fine-grained scenarios. Based on our findings, we introduce Cross-view Saliency Alignment (CVSA), a contrastive learning framework that first crops and swaps saliency regions of images as a novel view generation and then guides the model to localize on the foreground object via a cross-view alignment loss. Extensive experiments on four popular fine-grained classification benchmarks show that CVSA significantly improves the learned representation.
Dynamic Imaging using Deep Bi-linear Unsupervised Regularization (DEBLUR)
Jun 30, 2021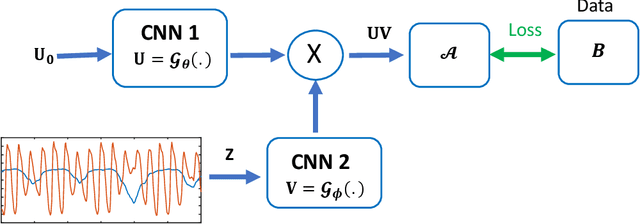
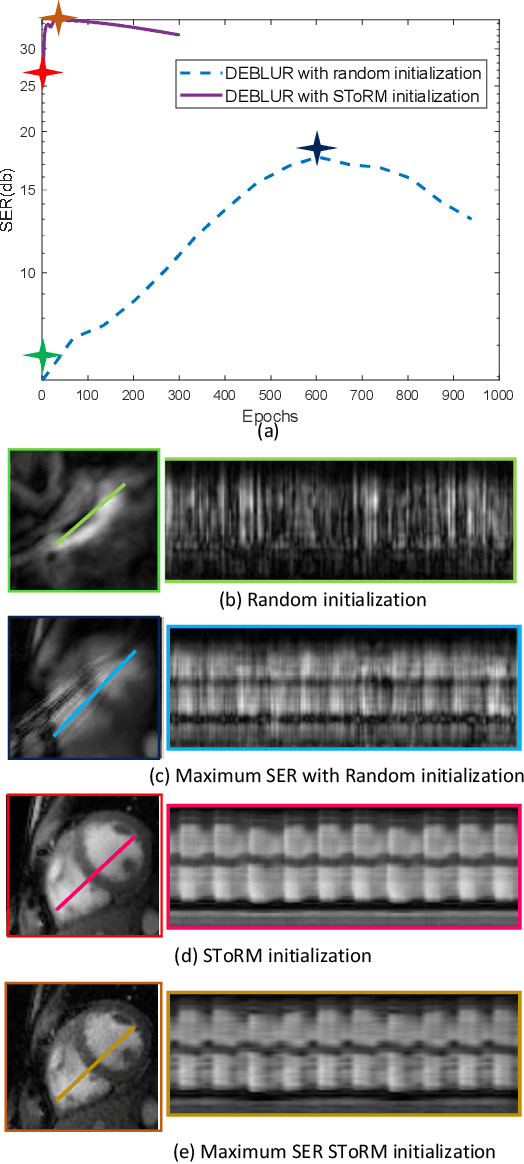
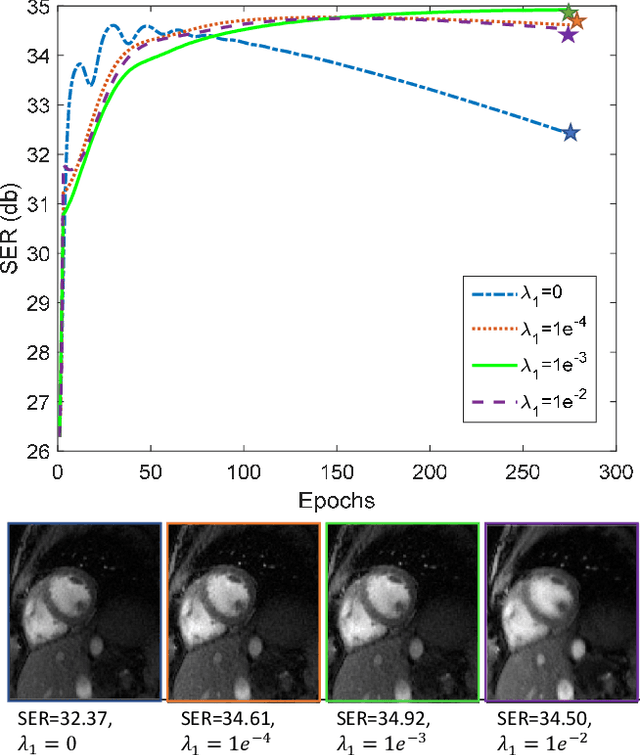
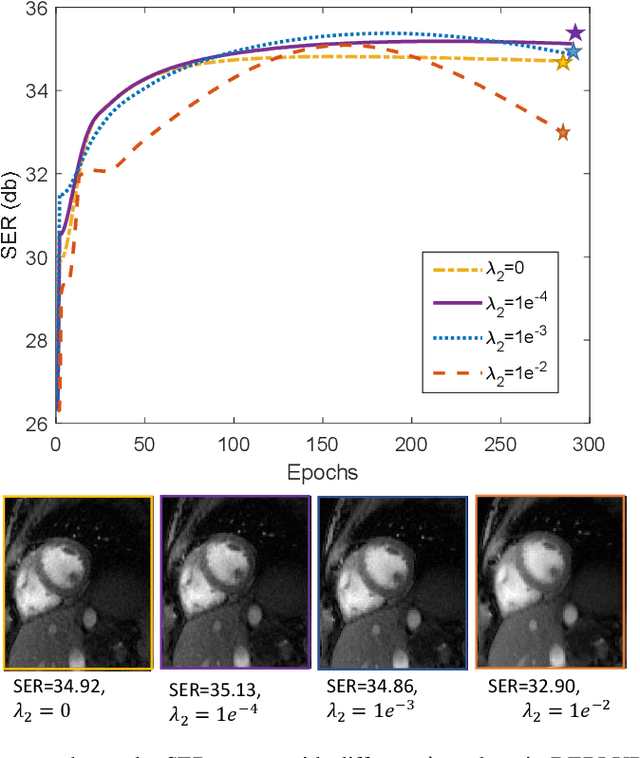
Bilinear models that decompose dynamic data to spatial and temporal factors are powerful and memory-efficient tools for the recovery of dynamic MRI data. These methods rely on sparsity and energy compaction priors on the factors to regularize the recovery. The quality of the recovered images depend on the specific priors. Motivated by deep image prior, we introduce a novel bilinear model whose factors are represented using convolutional neural networks (CNNs). The CNN parameters are learned from the undersampled data off the same subject. To reduce the run time and to improve performance, we initialize the CNN parameters. We use sparsity regularization of the network parameters to minimize the overfitting of the network to measurement noise. Our experiments on free breathing and ungated cardiac cine data acquired using a navigated golden-angle gradient-echo radial sequence show the ability of our method to provide reduced spatial blurring as compared to low-rank and SToRM reconstructions.
Analyzing an Imitation Learning Network for Fundus Image Registration Using a Divide-and-Conquer Approach
Dec 19, 2019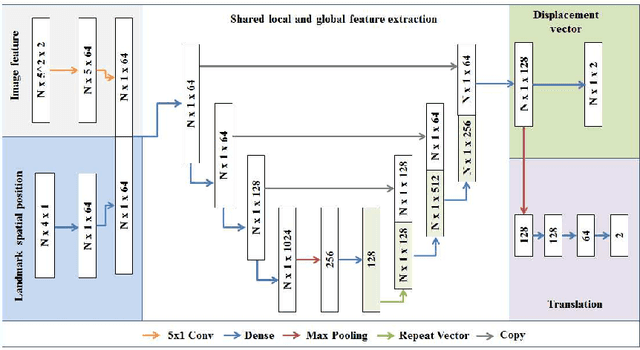
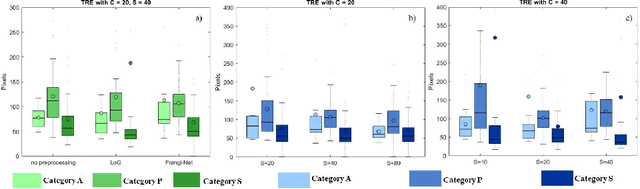
Comparison of microvascular circulation on fundoscopic images is a non-invasive clinical indication for the diagnosis and monitoring of diseases, such as diabetes and hypertensions. The differences between intra-patient images can be assessed quantitatively by registering serial acquisitions. Due to the variability of the images (i.e. contrast, luminosity) and the anatomical changes of the retina, the registration of fundus images remains a challenging task. Recently, several deep learning approaches have been proposed to register fundus images in an end-to-end fashion, achieving remarkable results. However, the results are difficult to interpret and analyze. In this work, we propose an imitation learning framework for the registration of 2D color funduscopic images for a wide range of applications such as disease monitoring, image stitching and super-resolution. We follow a divide-and-conquer approach to improve the interpretability of the proposed network, and analyze both the influence of the input image and the hyperparameters on the registration result. The results show that the proposed registration network reduces the initial target registration error up to 95\%.
GENESIS-V2: Inferring Unordered Object Representations without Iterative Refinement
Apr 21, 2021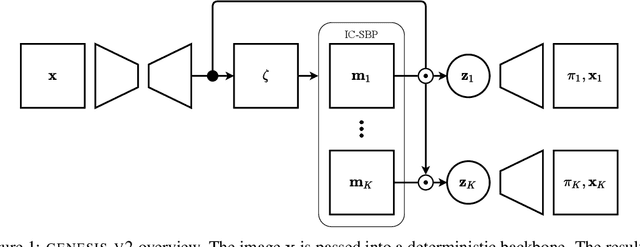

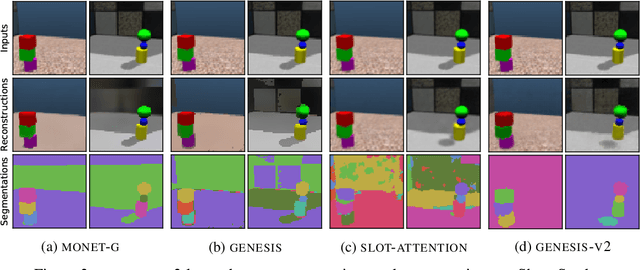
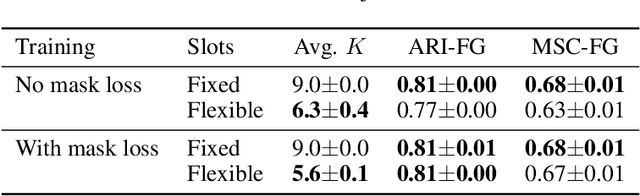
Advances in object-centric generative models (OCGMs) have culminated in the development of a broad range of methods for unsupervised object segmentation and interpretable object-centric scene generation. These methods, however, are limited to simulated and real-world datasets with limited visual complexity. Moreover, object representations are often inferred using RNNs which do not scale well to large images or iterative refinement which avoids imposing an unnatural ordering on objects in an image but requires the a priori initialisation of a fixed number of object representations. In contrast to established paradigms, this work proposes an embedding-based approach in which embeddings of pixels are clustered in a differentiable fashion using a stochastic, non-parametric stick-breaking process. Similar to iterative refinement, this clustering procedure also leads to randomly ordered object representations, but without the need of initialising a fixed number of clusters a priori. This is used to develop a new model, GENESIS-V2, which can infer a variable number of object representations without using RNNs or iterative refinement. We show that GENESIS-V2 outperforms previous methods for unsupervised image segmentation and object-centric scene generation on established synthetic datasets as well as more complex real-world datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge