"Image": models, code, and papers
A Review on Image Texture Analysis Methods
Mar 13, 2018Texture classification is an active topic in image processing which plays an important role in many applications such as image retrieval, inspection systems, face recognition, medical image processing, etc. There are many approaches extracting texture features in gray-level images such as local binary patterns, gray level co-occurrence matrices, statistical features, skeleton, scale invariant feature transform, etc. The texture analysis methods can be categorized in 4 groups titles: statistical methods, structural methods, filter-based and model based approaches. In many related researches, authors have tried to extract color and texture features jointly. In this respect, combined methods are considered as efficient image analysis descriptors. Mostly important challenges in image texture analysis are rotation sensitivity, gray scale variations, noise sensitivity, illumination and brightness conditions, etc. In this paper, we review most efficient and state-of-the-art image texture analysis methods. Also, some texture classification approaches are survived.
* in Persian
Exploiting Raw Images for Real-Scene Super-Resolution
Feb 02, 2021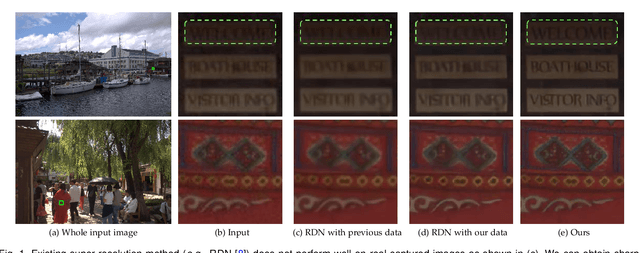
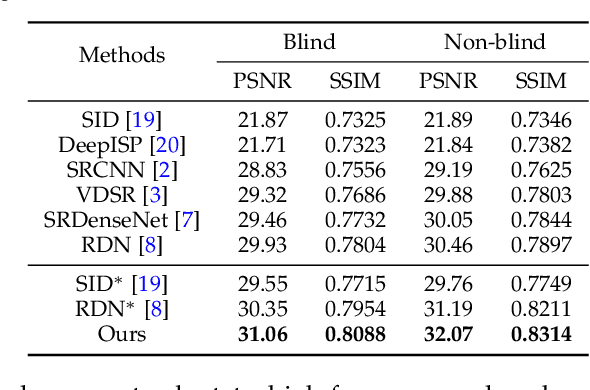
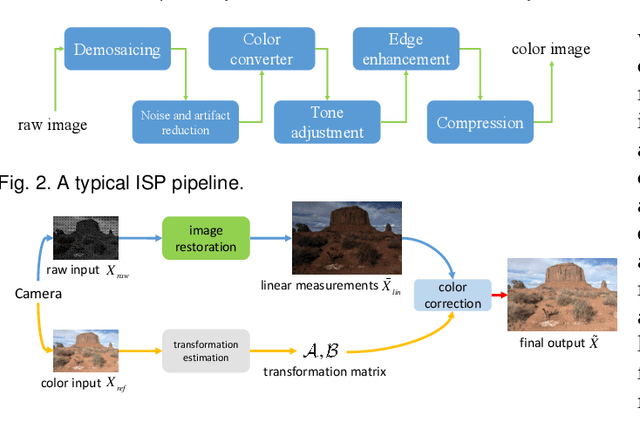
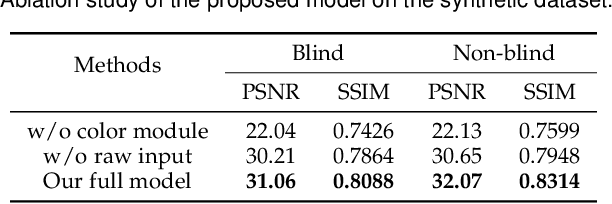
Super-resolution is a fundamental problem in computer vision which aims to overcome the spatial limitation of camera sensors. While significant progress has been made in single image super-resolution, most algorithms only perform well on synthetic data, which limits their applications in real scenarios. In this paper, we study the problem of real-scene single image super-resolution to bridge the gap between synthetic data and real captured images. We focus on two issues of existing super-resolution algorithms: lack of realistic training data and insufficient utilization of visual information obtained from cameras. To address the first issue, we propose a method to generate more realistic training data by mimicking the imaging process of digital cameras. For the second issue, we develop a two-branch convolutional neural network to exploit the radiance information originally-recorded in raw images. In addition, we propose a dense channel-attention block for better image restoration as well as a learning-based guided filter network for effective color correction. Our model is able to generalize to different cameras without deliberately training on images from specific camera types. Extensive experiments demonstrate that the proposed algorithm can recover fine details and clear structures, and achieve high-quality results for single image super-resolution in real scenes.
* A larger version with higher-resolution figures is available at: https://sites.google.com/view/xiangyuxu. arXiv admin note: text overlap with arXiv:1905.12156
ICMSC: Intra- and Cross-modality Semantic Consistency for Unsupervised Domain Adaptation on Hip Joint Bone Segmentation
Dec 23, 2020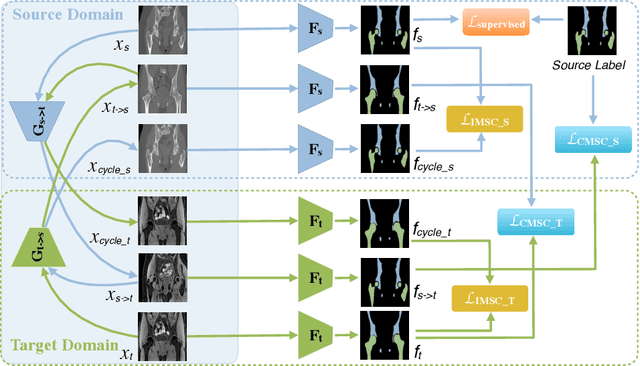
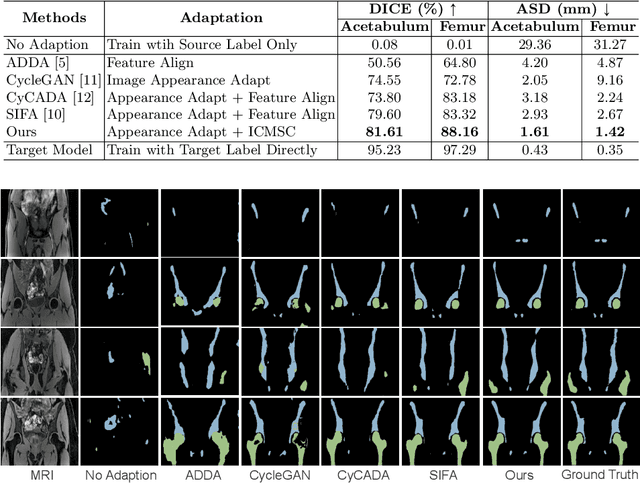
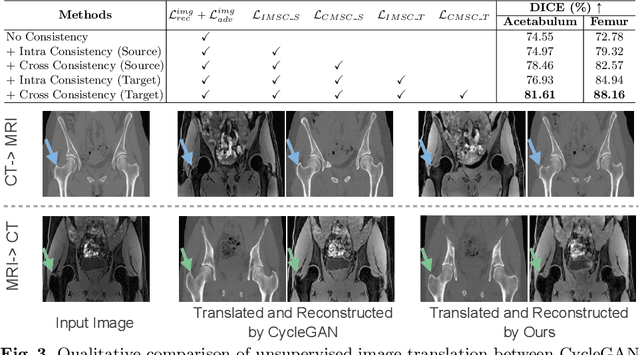
Unsupervised domain adaptation (UDA) for cross-modality medical image segmentation has shown great progress by domain-invariant feature learning or image appearance translation. Adapted feature learning usually cannot detect domain shifts at the pixel level and is not able to achieve good results in dense semantic segmentation tasks. Image appearance translation, e.g. CycleGAN, translates images into different styles with good appearance, despite its population, its semantic consistency is hardly to maintain and results in poor cross-modality segmentation. In this paper, we propose intra- and cross-modality semantic consistency (ICMSC) for UDA and our key insight is that the segmentation of synthesised images in different styles should be consistent. Specifically, our model consists of an image translation module and a domain-specific segmentation module. The image translation module is a standard CycleGAN, while the segmentation module contains two domain-specific segmentation networks. The intra-modality semantic consistency (IMSC) forces the reconstructed image after a cycle to be segmented in the same way as the original input image, while the cross-modality semantic consistency (CMSC) encourages the synthesized images after translation to be segmented exactly the same as before translation. Comprehensive experimental results on cross-modality hip joint bone segmentation show the effectiveness of our proposed method, which achieves an average DICE of 81.61% on the acetabulum and 88.16% on the proximal femur, outperforming other state-of-the-art methods. It is worth to note that without UDA, a model trained on CT for hip joint bone segmentation is non-transferable to MRI and has almost zero-DICE segmentation.
PC-DAN: Point Cloud based Deep Affinity Network for 3D Multi-Object Tracking (Accepted as an extended abstract in JRDB-ACT Workshop at CVPR21)
Jun 03, 2021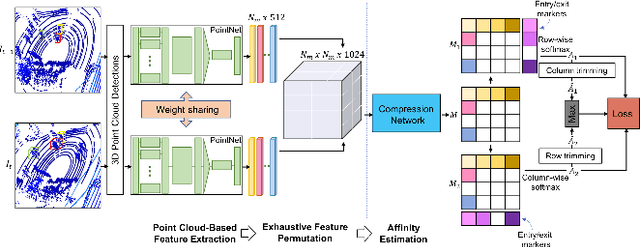
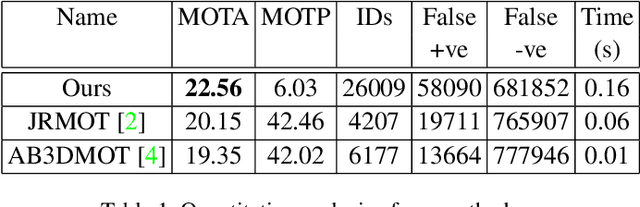
In recent times, the scope of LIDAR (Light Detection and Ranging) sensor-based technology has spread across numerous fields. It is popularly used to map terrain and navigation information into reliable 3D point cloud data, potentially revolutionizing the autonomous vehicles and assistive robotic industry. A point cloud is a dense compilation of spatial data in 3D coordinates. It plays a vital role in modeling complex real-world scenes since it preserves structural information and avoids perspective distortion, unlike image data, which is the projection of a 3D structure on a 2D plane. In order to leverage the intrinsic capabilities of the LIDAR data, we propose a PointNet-based approach for 3D Multi-Object Tracking (MOT).
A Pseudo Multi-Exposure Fusion Method Using Single Image
Aug 01, 2018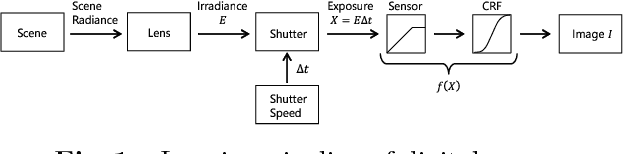
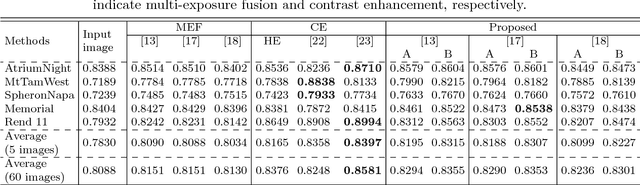
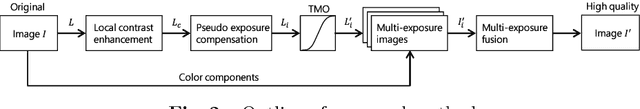
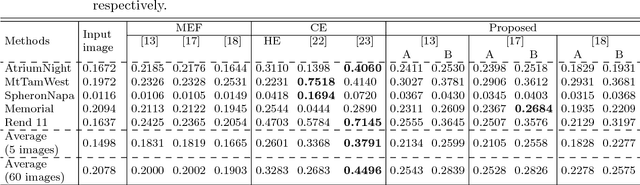
This paper proposes a novel pseudo multi-exposure image fusion method based on a single image. Multi-exposure image fusion is used to produce images without saturation regions, by using photos with different exposures. However, it is difficult to take photos suited for the multi-exposure image fusion when we take a photo of dynamic scenes or record a video. In addition, the multi-exposure image fusion cannot be applied to existing images with a single exposure or videos. The proposed method enables us to produce pseudo multi-exposure images from a single image. To produce multi-exposure images, the proposed method utilizes the relationship between the exposure values and pixel values, which is obtained by assuming that a digital camera has a linear response function. Moreover, it is shown that the use of a local contrast enhancement method allows us to produce pseudo multi-exposure images with higher quality. Most of conventional multi-exposure image fusion methods are also applicable to the proposed multi-exposure images. Experimental results show the effectiveness of the proposed method by comparing the proposed one with conventional ones.
Extraction of physically meaningful endmembers from STEM spectrum-images combining geometrical and statistical approaches
May 21, 2021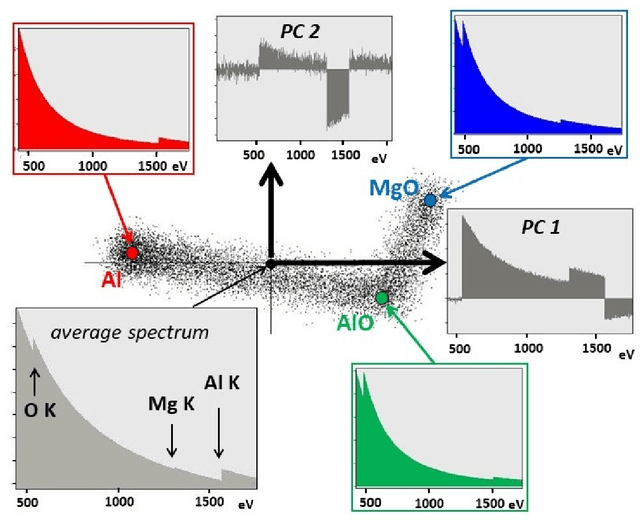
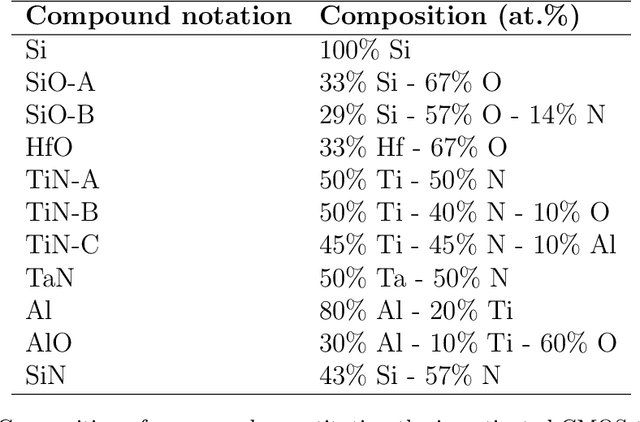
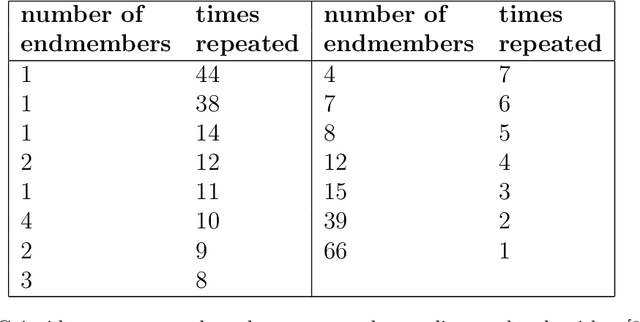

This article addresses extraction of physically meaningful information from STEM EELS and EDX spectrum-images using methods of Multivariate Statistical Analysis. The problem is interpreted in terms of data distribution in a multi-dimensional factor space, which allows for a straightforward and intuitively clear comparison of various approaches. A new computationally efficient and robust method for finding physically meaningful endmembers in spectrum-image datasets is presented. The method combines the geometrical approach of Vertex Component Analysis with the statistical approach of Bayesian inference. The algorithm is described in detail at an example of EELS spectrum-imaging of a multi-compound CMOS transistor.
LeViT: a Vision Transformer in ConvNet's Clothing for Faster Inference
Apr 02, 2021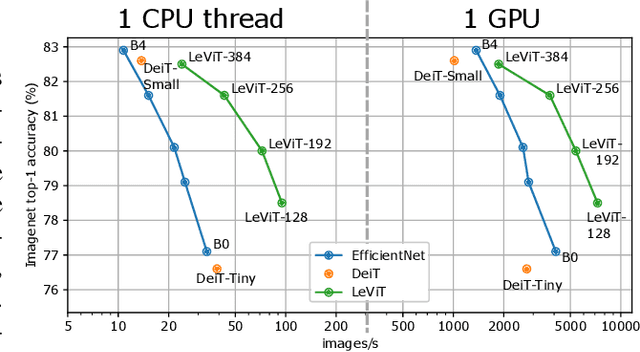
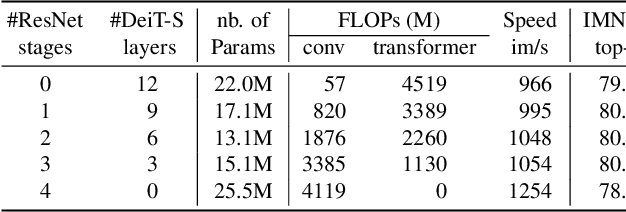

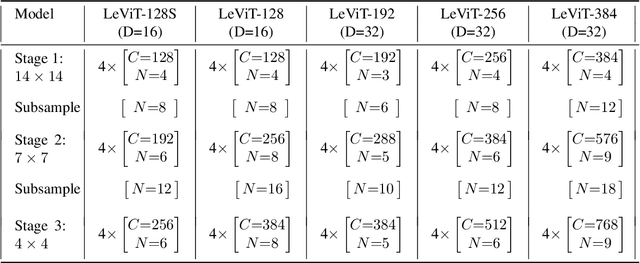
We design a family of image classification architectures that optimize the trade-off between accuracy and efficiency in a high-speed regime. Our work exploits recent findings in attention-based architectures, which are competitive on highly parallel processing hardware. We re-evaluated principles from the extensive literature on convolutional neural networks to apply them to transformers, in particular activation maps with decreasing resolutions. We also introduce the attention bias, a new way to integrate positional information in vision transformers. As a result, we propose LeVIT: a hybrid neural network for fast inference image classification. We consider different measures of efficiency on different hardware platforms, so as to best reflect a wide range of application scenarios. Our extensive experiments empirically validate our technical choices and show they are suitable to most architectures. Overall, LeViT significantly outperforms existing convnets and vision transformers with respect to the speed/accuracy tradeoff. For example, at 80\% ImageNet top-1 accuracy, LeViT is 3.3 times faster than EfficientNet on the CPU.
Spline Positional Encoding for Learning 3D Implicit Signed Distance Fields
Jun 03, 2021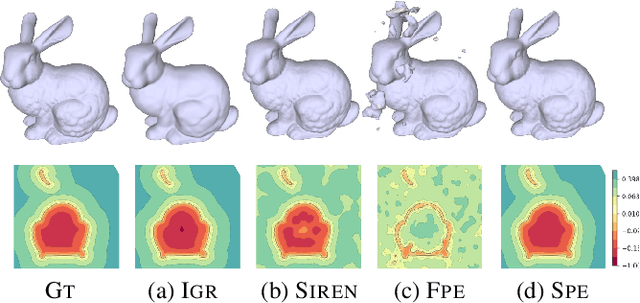
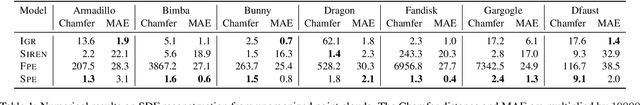
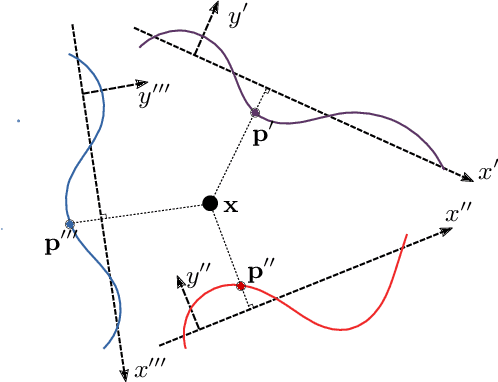
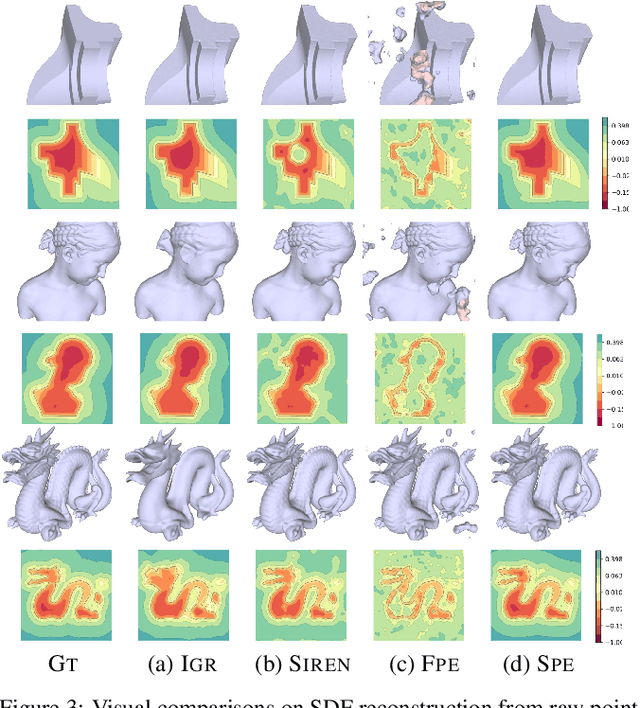
Multilayer perceptrons (MLPs) have been successfully used to represent 3D shapes implicitly and compactly, by mapping 3D coordinates to the corresponding signed distance values or occupancy values. In this paper, we propose a novel positional encoding scheme, called Spline Positional Encoding, to map the input coordinates to a high dimensional space before passing them to MLPs, for helping to recover 3D signed distance fields with fine-scale geometric details from unorganized 3D point clouds. We verified the superiority of our approach over other positional encoding schemes on tasks of 3D shape reconstruction from input point clouds and shape space learning. The efficacy of our approach extended to image reconstruction is also demonstrated and evaluated.
Transfer learning approach to Classify the X-ray image that corresponds to corona disease Using ResNet50 pretrained by ChexNet
May 18, 2021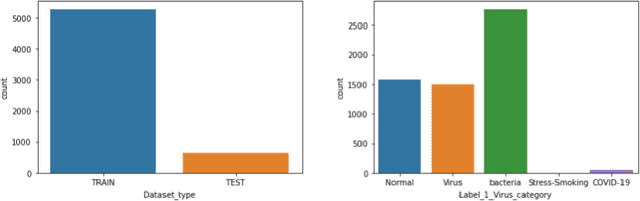
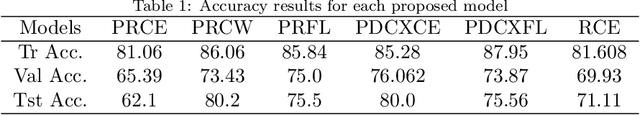
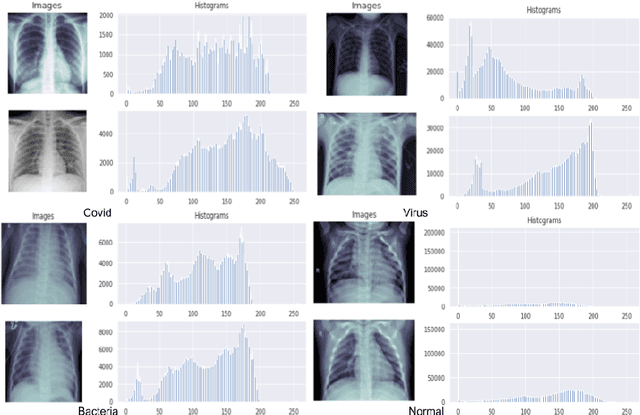
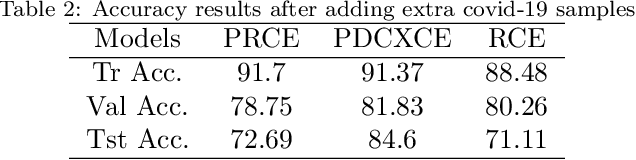
Coronavirus adversely has affected people worldwide. There are common symptoms between the Covid19 virus disease and other respiratory diseases like pneumonia or Influenza. Therefore, diagnosing it fast is crucial not only to save patients but also to prevent it from spreading. One of the most reliant methods of diagnosis is through X-ray images of a lung. With the help of deep learning approaches, we can teach the deep model to learn the condition of an affected lung. Therefore, it can classify the new sample as if it is a Covid19 infected patient or not. In this project, we train a deep model based on ResNet50 pretrained by ImageNet dataset and CheXNet dataset. Based on the imbalanced CoronaHack Chest X-Ray dataset introducing by Kaggle we applied both binary and multi-class classification. Also, we compare the results when using Focal loss and Cross entropy loss.
Leaf Image-based Plant Disease Identification using Color and Texture Features
Feb 08, 2021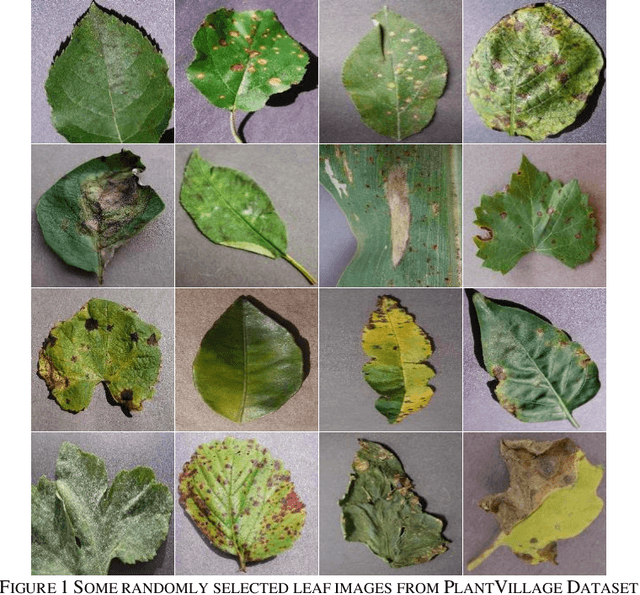
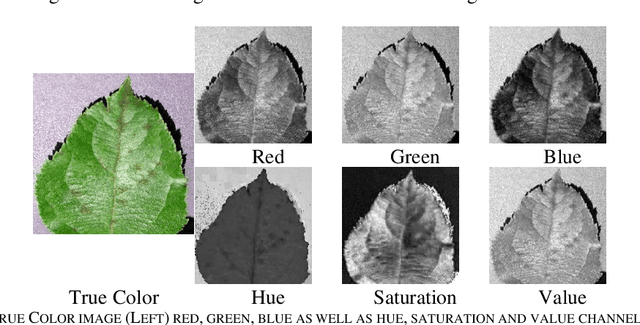
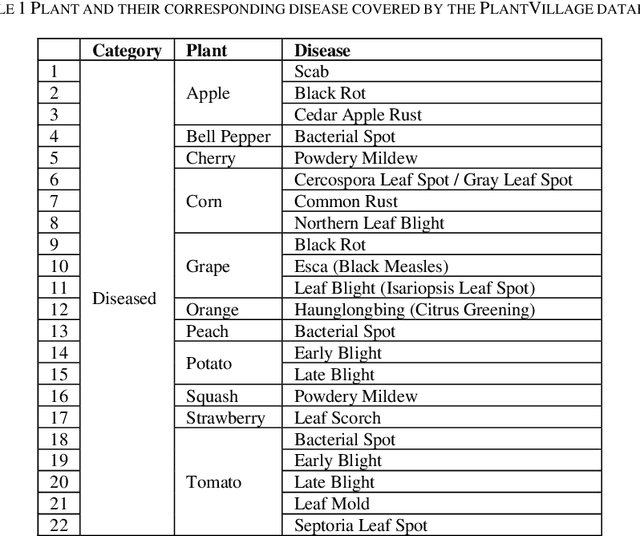

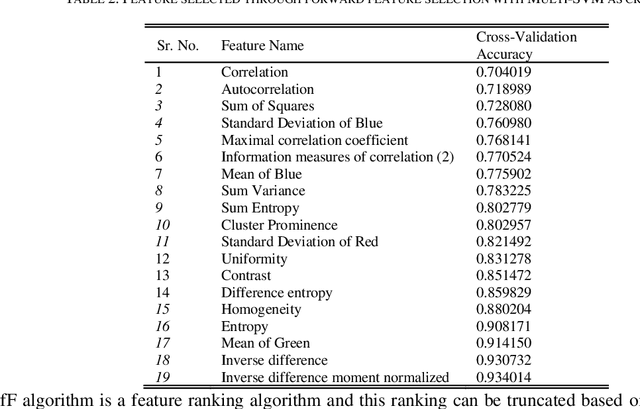
Identification of plant disease is usually done through visual inspection or during laboratory examination which causes delays resulting in yield loss by the time identification is complete. On the other hand, complex deep learning models perform the task with reasonable performance but due to their large size and high computational requirements, they are not suited to mobile and handheld devices. Our proposed approach contributes automated identification of plant diseases which follows a sequence of steps involving pre-processing, segmentation of diseased leaf area, calculation of features based on the Gray-Level Co-occurrence Matrix (GLCM), feature selection and classification. In this study, six color features and twenty-two texture features have been calculated. Support vector machines is used to perform one-vs-one classification of plant disease. The proposed model of disease identification provides an accuracy of 98.79% with a standard deviation of 0.57 on 10-fold cross-validation. The accuracy on a self-collected dataset is 82.47% for disease identification and 91.40% for healthy and diseased classification. The reported performance measures are better or comparable to the existing approaches and highest among the feature-based methods, presenting it as the most suitable method to automated leaf-based plant disease identification. This prototype system can be extended by adding more disease categories or targeting specific crop or disease categories.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge