"Image": models, code, and papers
Classifying Organisms and Artefacts By Their Shapes
Aug 27, 2021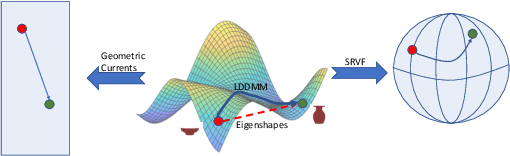
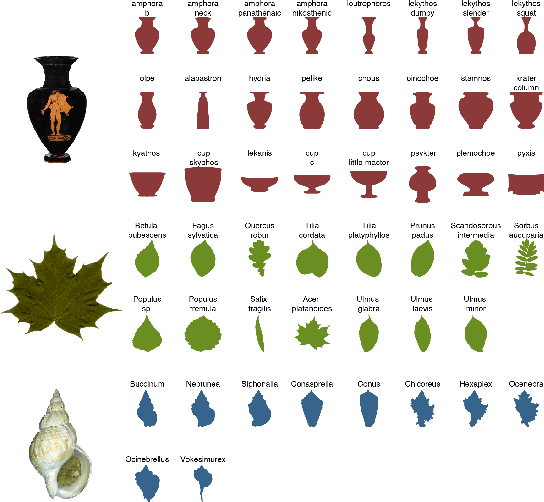
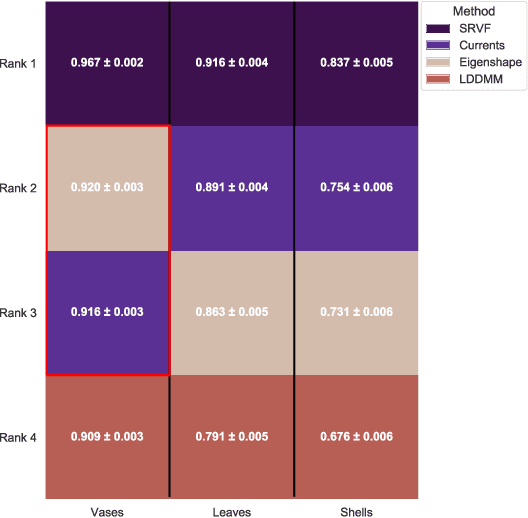
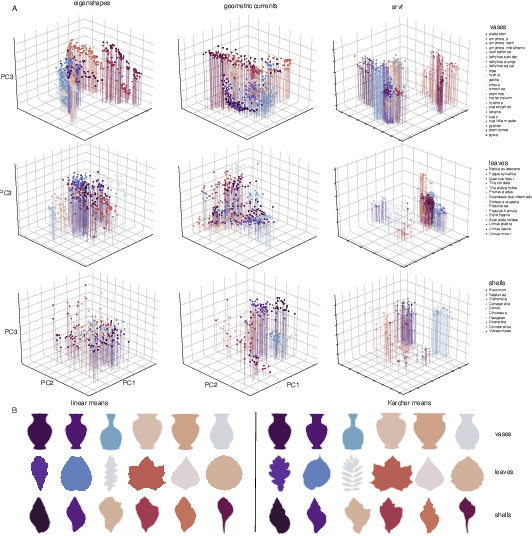
We often wish to classify objects by their shapes. Indeed, the study of shapes is an important part of many scientific fields such as evolutionary biology, structural biology, image processing, and archaeology. The most widely-used method of shape analysis, Geometric Morphometrics, assumes that that the mathematical space in which shapes are represented is linear. However, it has long been known that shape space is, in fact, rather more complicated, and certainly non-linear. Diffeomorphic methods that take this non-linearity into account, and so give more accurate estimates of the distances among shapes, exist but have rarely been applied to real-world problems. Using a machine classifier, we tested the ability of several of these methods to describe and classify the shapes of a variety of organic and man-made objects. We find that one method, the Square-Root Velocity Function (SRVF), is superior to all others, including a standard Geometric Morphometric method (eigenshapes). We also show that computational shape classifiers outperform human experts, and that the SRVF shortest-path between shapes can be used to estimate the shapes of intermediate steps in evolutionary series. Diffeomorphic shape analysis methods, we conclude, now provide practical and effective solutions to many shape description and classification problems in the natural and human sciences.
Progressive Transformer-Based Generation of Radiology Reports
Feb 19, 2021
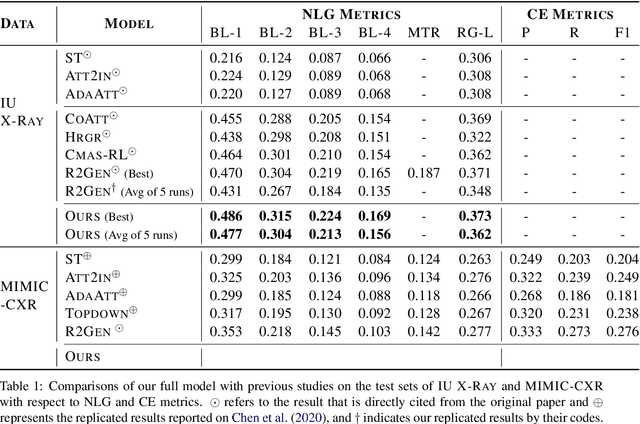
Inspired by Curriculum Learning, we propose a consecutive (i.e. image-to-text-to-text) generation framework where we divide the problem of radiology report generation into two steps. Contrary to generating the full radiology report from the image at once, the model generates global concepts from the image in the first step and then reforms them into finer and coherent texts using transformer-based architecture. We follow the transformer-based sequence-to-sequence paradigm at each step. We improve upon the state-of-the-art on two benchmark datasets.
Computed Tomography Reconstruction Using Deep Image Prior and Learned Reconstruction Methods
Mar 12, 2020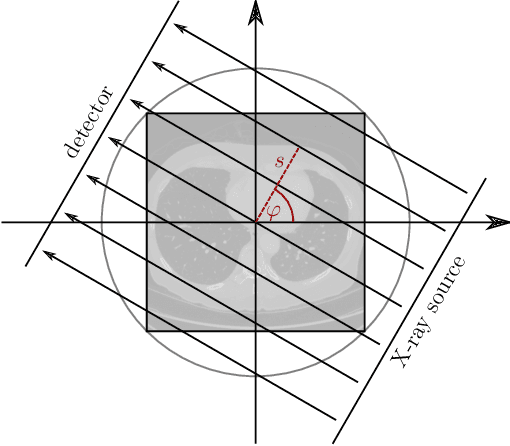


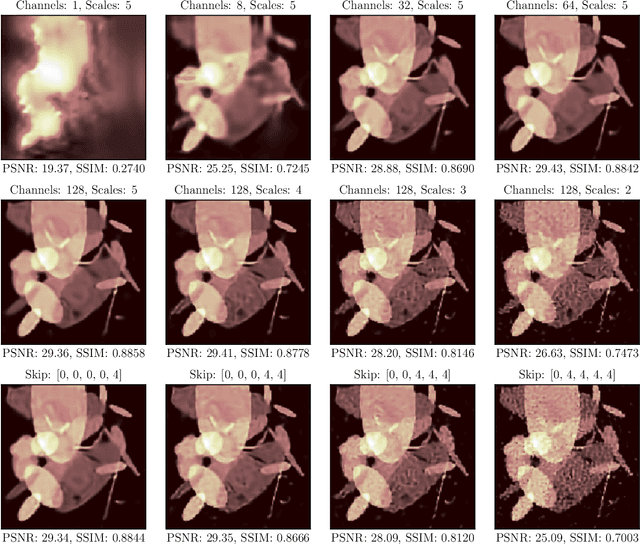
In this work, we investigate the application of deep learning methods for computed tomography in the context of having a low-data regime. As motivation, we review some of the existing approaches and obtain quantitative results after training them with different amounts of data. We find that the learned primal-dual has an outstanding performance in terms of reconstruction quality and data efficiency. However, in general, end-to-end learned methods have two issues: a) lack of classical guarantees in inverse problems and b) lack of generalization when not trained with enough data. To overcome these issues, we bring in the deep image prior approach in combination with classical regularization. The proposed methods improve the state-of-the-art results in the low data-regime.
Terabyte-scale supervised 3D training and benchmarking dataset of the mouse kidney
Aug 04, 2021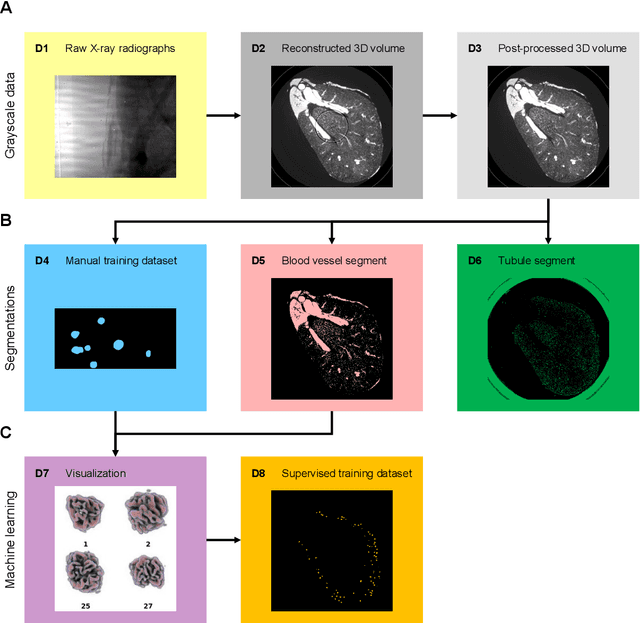
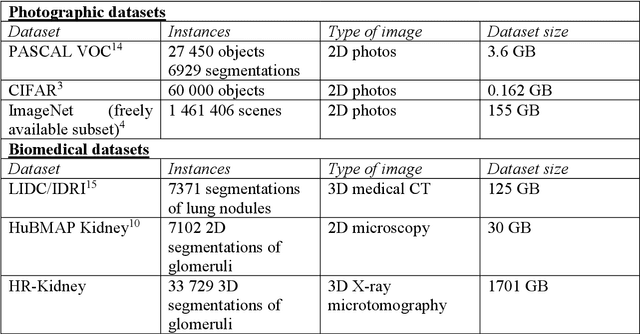
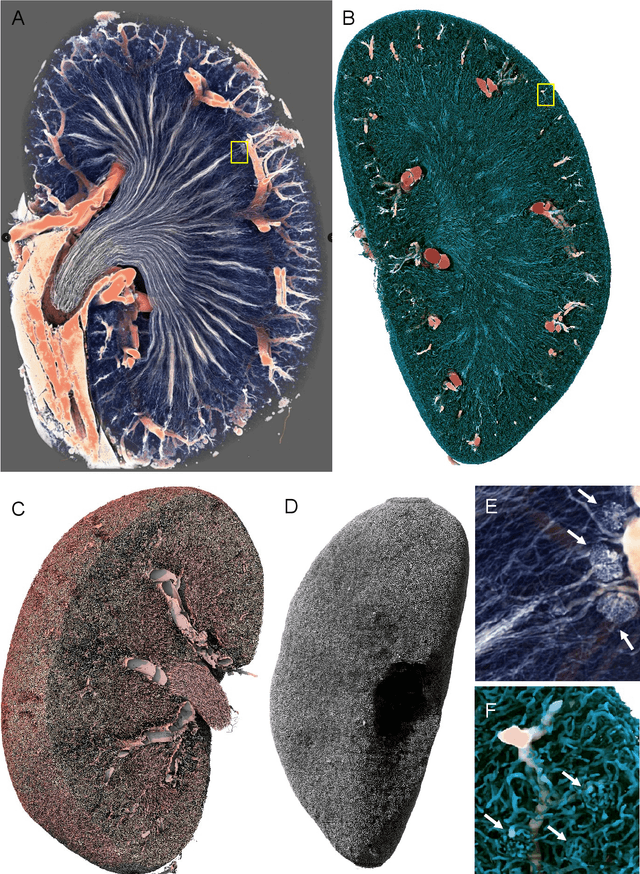
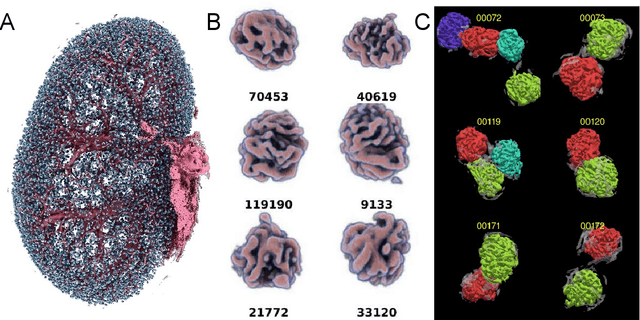
The performance of machine learning algorithms used for the segmentation of 3D biomedical images lags behind that of the algorithms employed in the classification of 2D photos. This may be explained by the comparative lack of high-volume, high-quality training datasets, which require state-of-the art imaging facilities, domain experts for annotation and large computational and personal resources to create. The HR-Kidney dataset presented in this work bridges this gap by providing 1.7 TB of artefact-corrected synchrotron radiation-based X-ray phase-contrast microtomography images of whole mouse kidneys and validated segmentations of 33 729 glomeruli, which represents a 1-2 orders of magnitude increase over currently available biomedical datasets. The dataset further contains the underlying raw data, classical segmentations of renal vasculature and uriniferous tubules, as well as true 3D manual annotations. By removing limits currently imposed by small training datasets, the provided data open up the possibility for disruptions in machine learning for biomedical image analysis.
Deep Learning for Hyperspectral Image Classification: An Overview
Oct 26, 2019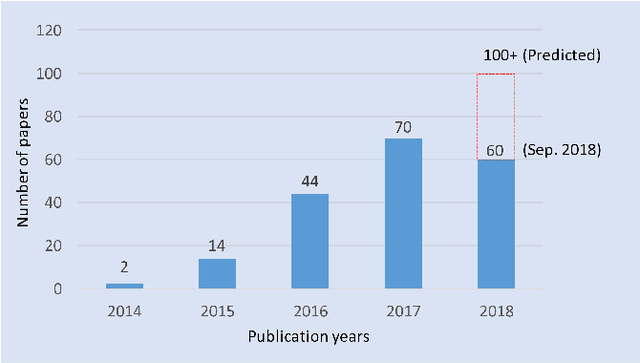
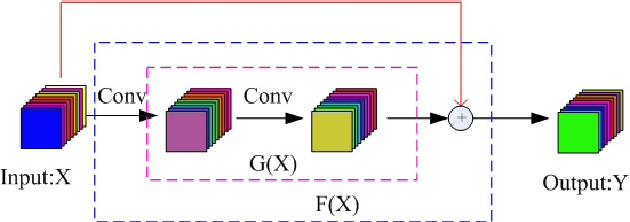
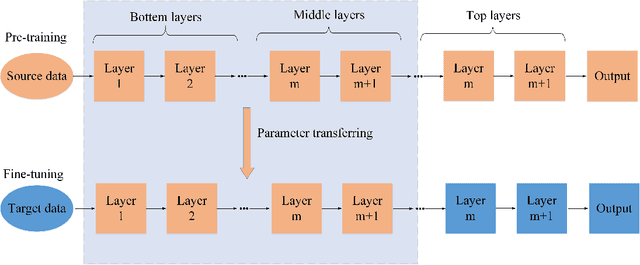
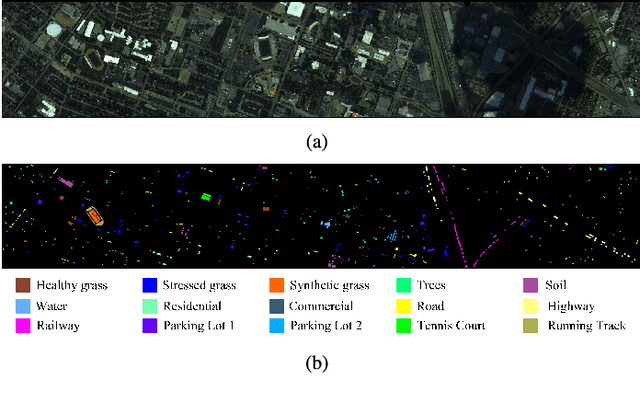
Hyperspectral image (HSI) classification has become a hot topic in the field of remote sensing. In general, the complex characteristics of hyperspectral data make the accurate classification of such data challenging for traditional machine learning methods. In addition, hyperspectral imaging often deals with an inherently nonlinear relation between the captured spectral information and the corresponding materials. In recent years, deep learning has been recognized as a powerful feature-extraction tool to effectively address nonlinear problems and widely used in a number of image processing tasks. Motivated by those successful applications, deep learning has also been introduced to classify HSIs and demonstrated good performance. This survey paper presents a systematic review of deep learning-based HSI classification literatures and compares several strategies for this topic. Specifically, we first summarize the main challenges of HSI classification which cannot be effectively overcome by traditional machine learning methods, and also introduce the advantages of deep learning to handle these problems. Then, we build a framework which divides the corresponding works into spectral-feature networks, spatial-feature networks, and spectral-spatial-feature networks to systematically review the recent achievements in deep learning-based HSI classification. In addition, considering the fact that available training samples in the remote sensing field are usually very limited and training deep networks require a large number of samples, we include some strategies to improve classification performance, which can provide some guidelines for future studies on this topic. Finally, several representative deep learning-based classification methods are conducted on real HSIs in our experiments.
Tiled sparse coding in eigenspaces for the COVID-19 diagnosis in chest X-ray images
Jun 28, 2021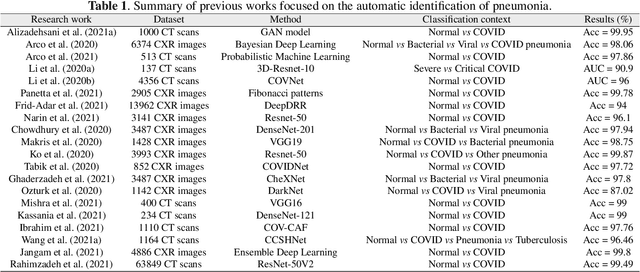
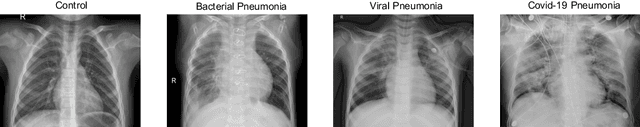
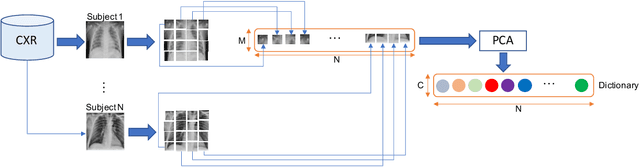
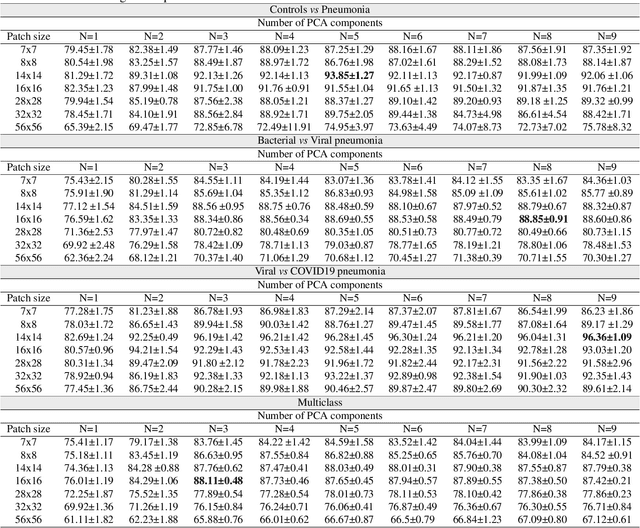
The ongoing crisis of the COVID-19 (Coronavirus disease 2019) pandemic has changed the world. According to the World Health Organization (WHO), 4 million people have died due to this disease, whereas there have been more than 180 million confirmed cases of COVID-19. The collapse of the health system in many countries has demonstrated the need of developing tools to automatize the diagnosis of the disease from medical imaging. Previous studies have used deep learning for this purpose. However, the performance of this alternative highly depends on the size of the dataset employed for training the algorithm. In this work, we propose a classification framework based on sparse coding in order to identify the pneumonia patterns associated with different pathologies. Specifically, each chest X-ray (CXR) image is partitioned into different tiles. The most relevant features extracted from PCA are then used to build the dictionary within the sparse coding procedure. Once images are transformed and reconstructed from the elements of the dictionary, classification is performed from the reconstruction errors of individual patches associated with each image. Performance is evaluated in a real scenario where simultaneously differentiation between four different pathologies: control vs bacterial pneumonia vs viral pneumonia vs COVID-19. The accuracy when identifying the presence of pneumonia is 93.85%, whereas 88.11% is obtained in the 4-class classification context. The excellent results and the pioneering use of sparse coding in this scenario evidence the applicability of this approach as an aid for clinicians in a real-world environment.
To What Extent Does Downsampling, Compression, and Data Scarcity Impact Renal Image Analysis?
Sep 22, 2019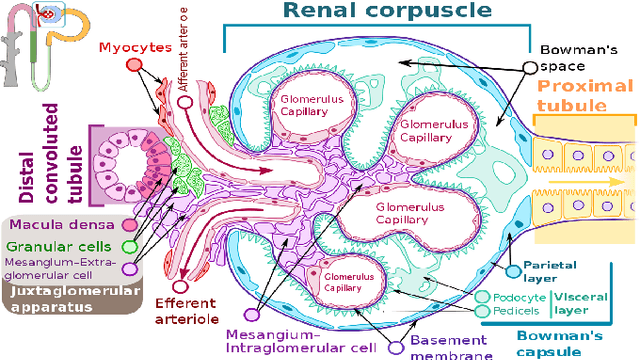
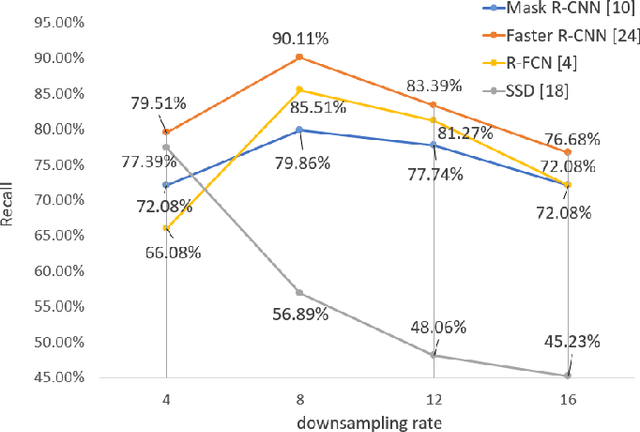
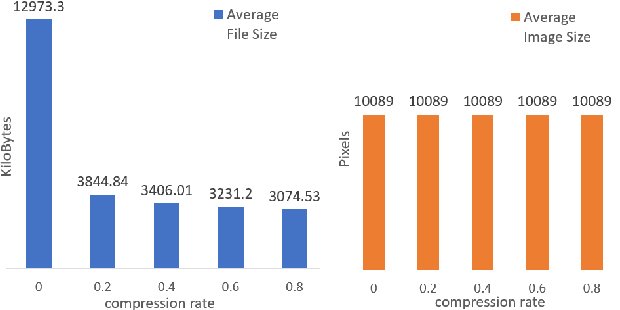
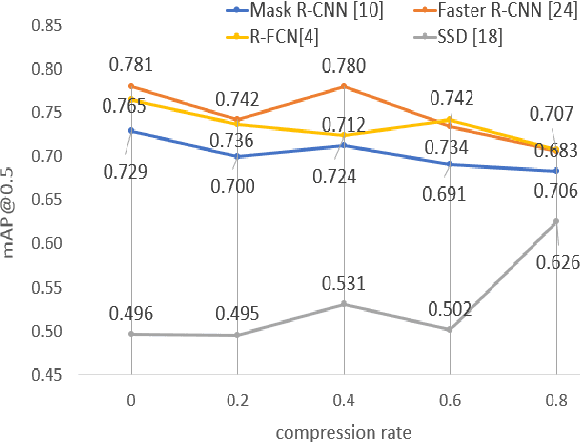
The condition of the Glomeruli, or filter sacks, in renal Direct Immunofluorescence (DIF) specimens is a critical indicator for diagnosing kidney diseases. A digital pathology system which digitizes a glass histology slide into a Whole Slide Image (WSI) and then automatically detects and zooms in on the glomeruli with a higher magnification objective will be extremely helpful for pathologists. In this paper, using glomerulus detection as the study case, we provide analysis and observations on several important issues to help with the development of Computer Aided Diagnostic (CAD) systems to process WSIs. Large image resolution, large file size, and data scarcity are always challenging to deal with. To this end, we first examine image downsampling rates in terms of their effect on detection accuracy. Second, we examine the impact of image compression. Third, we examine the relationship between the size of the training set and detection accuracy. To understand the above issues, experiments are performed on the state-of-the-art detectors: Faster R-CNN, R-FCN, Mask R-CNN and SSD. Critical findings are observed: (1) The best balance between detection accuracy, detection speed and file size is achieved at 8 times downsampling captured with a $40\times$ objective; (2) compression which reduces the file size dramatically, does not necessarily have an adverse effect on overall accuracy; (3) reducing the amount of training data to some extents causes a drop in precision but has a negligible impact on the recall; (4) in most cases, Faster R-CNN achieves the best accuracy in the glomerulus detection task. We show that the image file size of $40\times$ WSI images can be reduced by a factor of over 6000 with negligible loss of glomerulus detection accuracy.
SDL: New data generation tools for full-level annotated document layout
Jun 29, 2021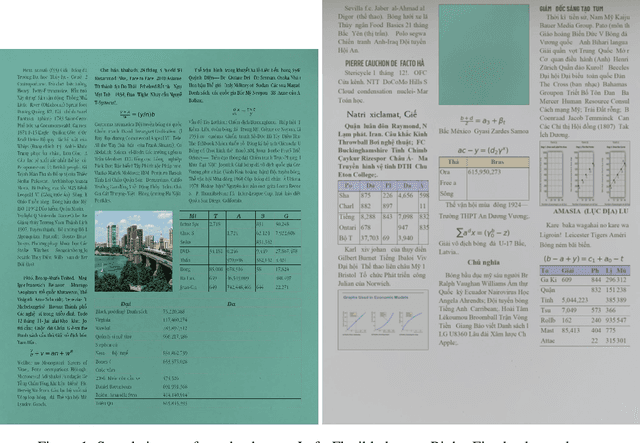
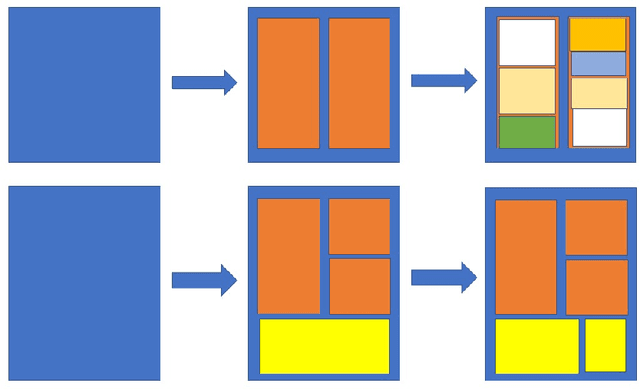
We present a novel data generation tool for document processing. The tool focuses on providing a maximal level of visual information in a normal type document, ranging from character position to paragraph-level position. It also enables working with a large dataset on low-resource languages as well as providing a mean of processing thorough full-level information of the documented text. The data generation tools come with a dataset of 320000 Vietnamese synthetic document images and an instruction to generate a dataset of similar size in other languages. The repository can be found at: https://github.com/tson1997/SDL-Document-Image-Generation
IUPUI Driving Videos and Images in All Weather and Illumination Conditions
Apr 17, 2021This document describes an image and video dataset of driving views captured in all weather and illumination conditions. The data set has been submitted to CDVL.
Mitigating Information Leakage in Image Representations: A Maximum Entropy Approach
Apr 11, 2019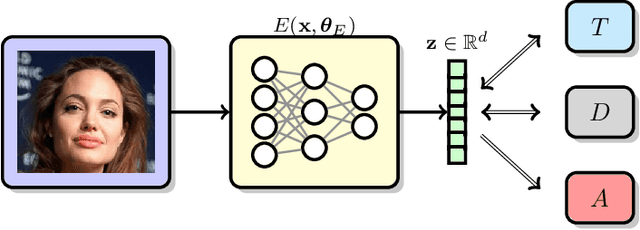
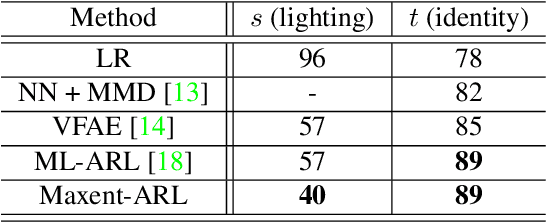
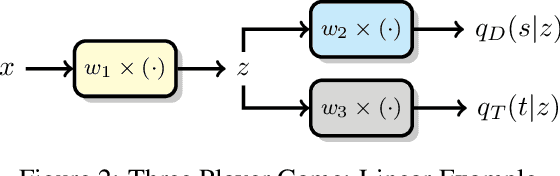
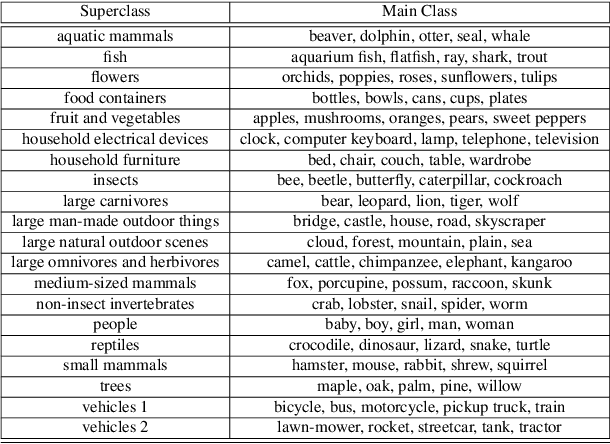
Image recognition systems have demonstrated tremendous progress over the past few decades thanks, in part, to our ability of learning compact and robust representations of images. As we witness the wide spread adoption of these systems, it is imperative to consider the problem of unintended leakage of information from an image representation, which might compromise the privacy of the data owner. This paper investigates the problem of learning an image representation that minimizes such leakage of user information. We formulate the problem as an adversarial non-zero sum game of finding a good embedding function with two competing goals: to retain as much task dependent discriminative image information as possible, while simultaneously minimizing the amount of information, as measured by entropy, about other sensitive attributes of the user. We analyze the stability and convergence dynamics of the proposed formulation using tools from non-linear systems theory and compare to that of the corresponding adversarial zero-sum game formulation that optimizes likelihood as a measure of information content. Numerical experiments on UCI, Extended Yale B, CIFAR-10 and CIFAR-100 datasets indicate that our proposed approach is able to learn image representations that exhibit high task performance while mitigating leakage of predefined sensitive information.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge