"Image": models, code, and papers
Sentinel-1 Additive Noise Removal from Cross-Polarization Extra-Wide TOPSAR with Dynamic Least-Squares
Jul 12, 2021
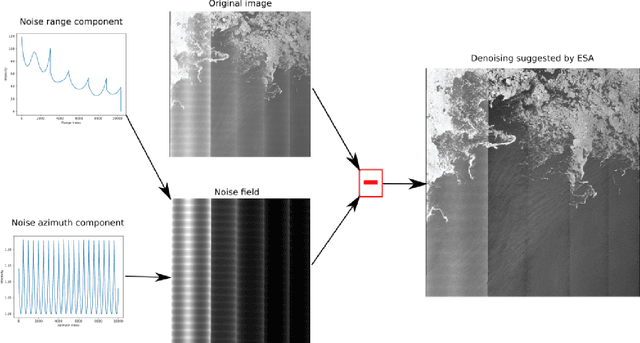


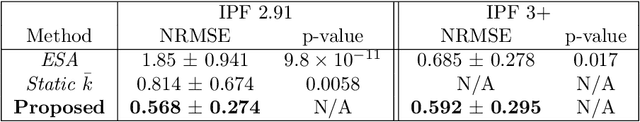
Sentinel-1 is a synthetic aperture radar (SAR) platform with an operational mode called extra wide (EW) that offers large regions of ocean areas to be observed. A major issue with EW images is that the cross-polarized HV and VH channels have prominent additive noise patterns relative to low backscatter intensity, which disrupts tasks that require manual or automated interpretation. The European Space Agency (ESA) provides a method for removing the additive noise pattern by means of lookup tables, but applying them directly produces unsatisfactory results because characteristics of the noise still remain. Furthermore, evidence suggests that the magnitude of the additive noise dynamically depends on factors that are not considered by the ESA estimated noise field. To address these issues we propose a quadratic objective function to model the mis-scale of the provided noise field on an image. We consider a linear denoising model that re-scales the noise field for each subswath, whose parameters are found from a least-squares solution over the objective function. This method greatly reduces the presence of additive noise while not requiring a set of training images, is robust to heterogeneity in images, dynamically estimates parameters for each image, and finds parameters using a closed-form solution. Two experiments were performed to validate the proposed method. The first experiment simulated noise removal on a set of RADARSAT-2 images with noise fields artificially imposed on them. The second experiment conducted noise removal on a set of Sentinel-1 images taken over the five oceans. Afterwards, quality of the noise removal was evaluated based on the appearance of open-water. The two experiments indicate that the proposed method marks an improvement both visually and through numerical measures.
* 22 pages, 14 figures
Partial 3D Object Retrieval using Local Binary QUICCI Descriptors and Dissimilarity Tree Indexing
Jul 07, 2021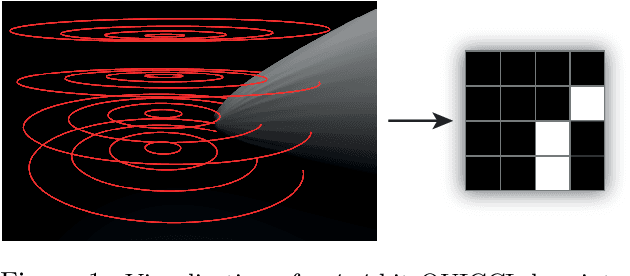

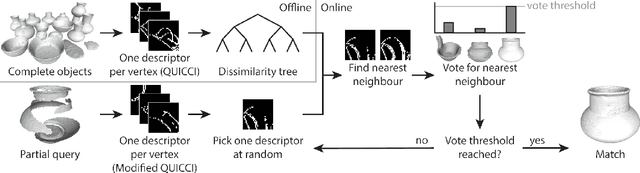
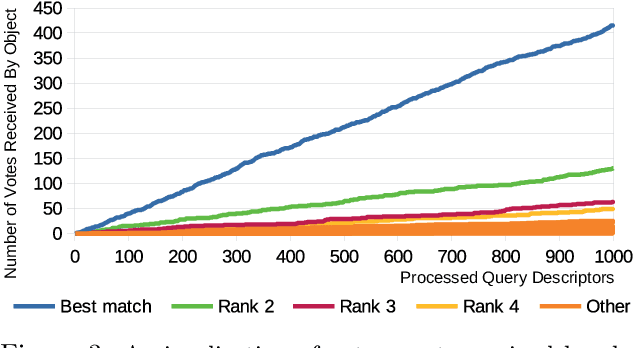
A complete pipeline is presented for accurate and efficient partial 3D object retrieval based on Quick Intersection Count Change Image (QUICCI) binary local descriptors and a novel indexing tree. It is shown how a modification to the QUICCI query descriptor makes it ideal for partial retrieval. An indexing structure called Dissimilarity Tree is proposed which can significantly accelerate searching the large space of local descriptors; this is applicable to QUICCI and other binary descriptors. The index exploits the distribution of bits within descriptors for efficient retrieval. The retrieval pipeline is tested on the artificial part of SHREC'16 dataset with near-ideal retrieval results.
Compressing Weight-updates for Image Artifacts Removal Neural Networks
May 10, 2019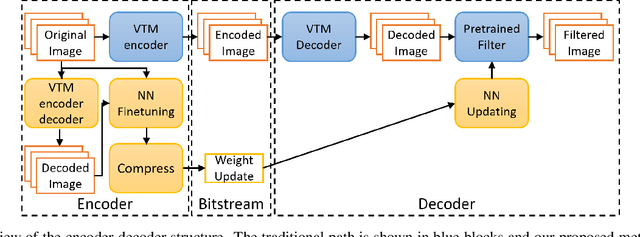
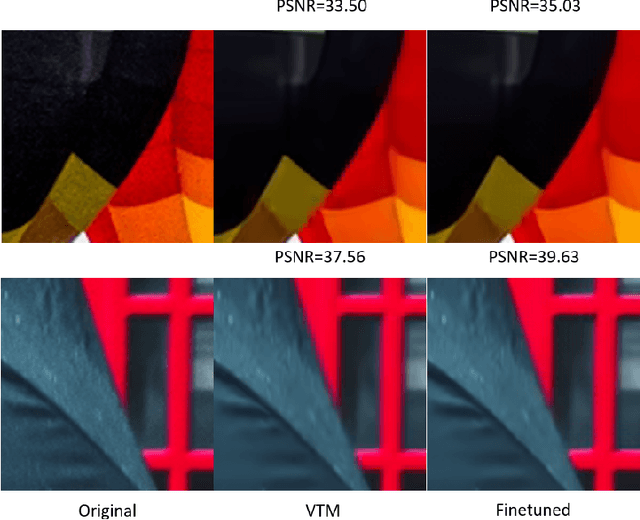
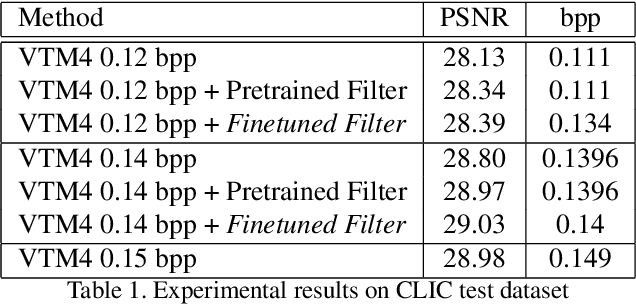

In this paper, we present a novel approach for fine-tuning a decoder-side neural network in the context of image compression, such that the weight-updates are better compressible. At encoder side, we fine-tune a pre-trained artifact removal network on target data by using a compression objective applied on the weight-update. In particular, the compression objective encourages weight-updates which are sparse and closer to quantized values. This way, the final weight-update can be compressed more efficiently by pruning and quantization, and can be included into the encoded bitstream together with the image bitstream of a traditional codec. We show that this approach achieves reconstruction quality which is on-par or slightly superior to a traditional codec, at comparable bitrates. To our knowledge, this is the first attempt to combine image compression and neural network's weight update compression.
X-Ray Image Compression Using Convolutional Recurrent Neural Networks
May 09, 2019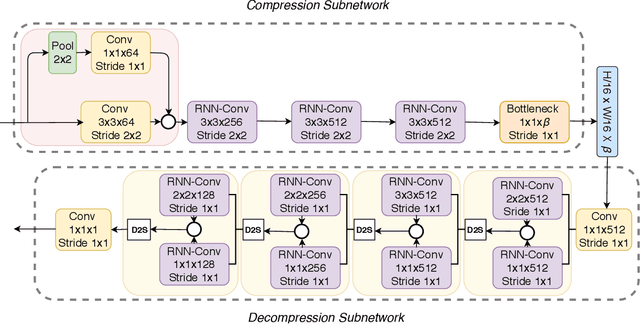
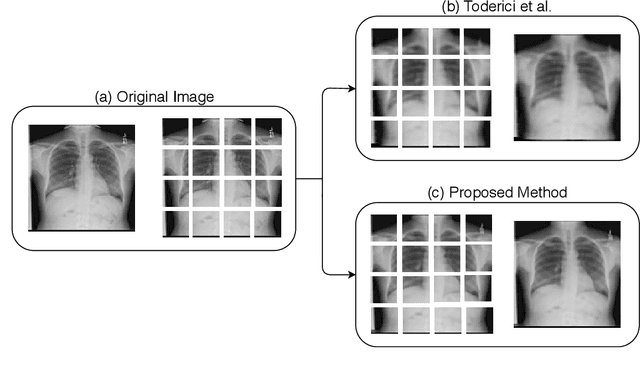
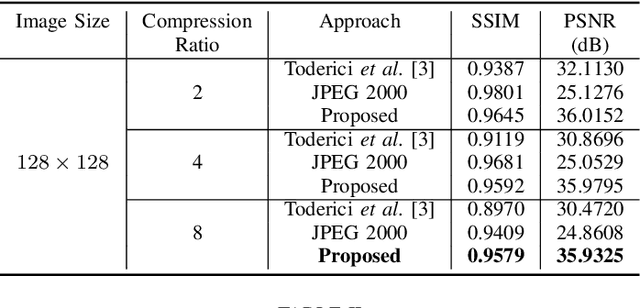
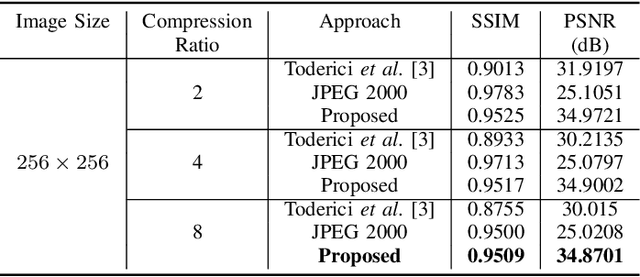
In the advent of a digital health revolution, vast amounts of clinical data are being generated, stored and processed on a daily basis. This has made the storage and retrieval of large volumes of health-care data, especially, high-resolution medical images, particularly challenging. Effective image compression for medical images thus plays a vital role in today's healthcare information system, particularly in teleradiology. In this work, an X-ray image compression method based on a Convolutional Recurrent Neural Networks RNN-Conv is presented. The proposed architecture can provide variable compression rates during deployment while it requires each network to be trained only once for a specific dimension of X-ray images. The model uses a multi-level pooling scheme that learns contextualized features for effective compression. We perform our image compression experiments on the National Institute of Health (NIH) ChestX-ray8 dataset and compare the performance of the proposed architecture with a state-of-the-art RNN based technique and JPEG 2000. The experimental results depict improved compression performance achieved by the proposed method in terms of Structural Similarity Index (SSIM) and Peak Signal-to-Noise Ratio (PSNR) metrics. To the best of our knowledge, this is the first reported evaluation on using a deep convolutional RNN for medical image compression.
Constrained Linear Data-feature Mapping for Image Classification
Nov 23, 2019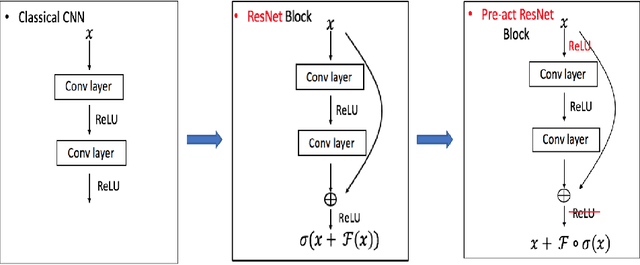
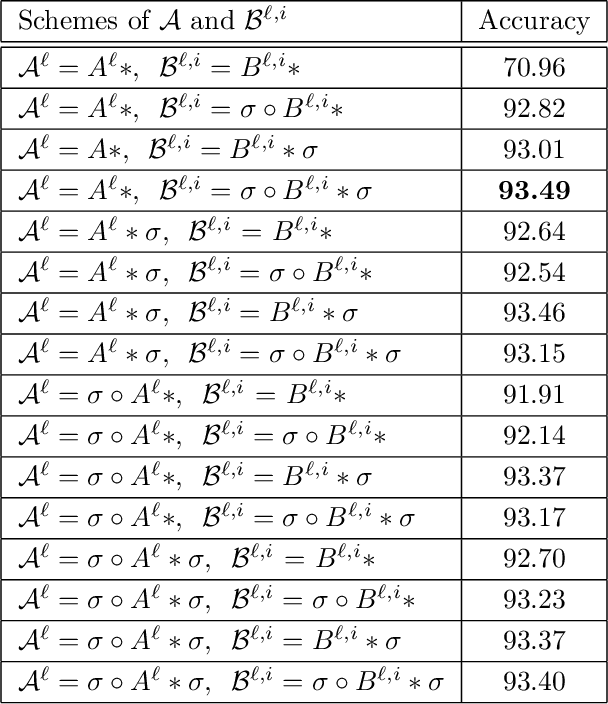
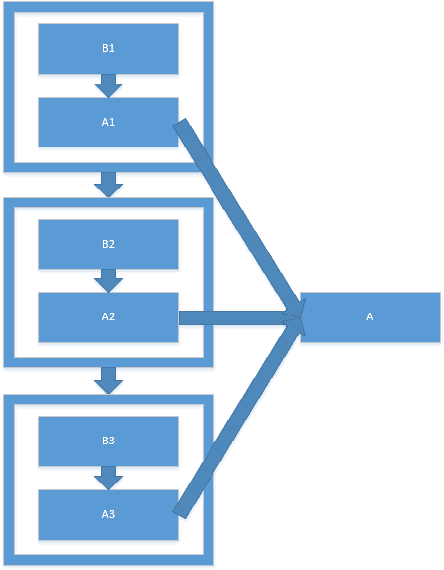
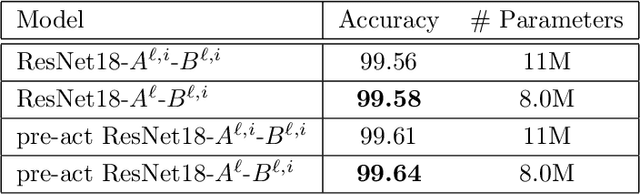
In this paper, we propose a constrained linear data-feature mapping model as an interpretable mathematical model for image classification using convolutional neural network (CNN) such as the ResNet. From this viewpoint, we establish the detailed connections in a technical level between the traditional iterative schemes for constrained linear system and the architecture for the basic block of ResNet. Under these connections, we propose some natural modifications of ResNet type models which will have less parameters but can keep almost the same accuracy as these original models. Some numerical experiments are shown to demonstrate the validity of this constrained learning data-feature mapping assumption.
Multi-Stage Pathological Image Classification using Semantic Segmentation
Oct 10, 2019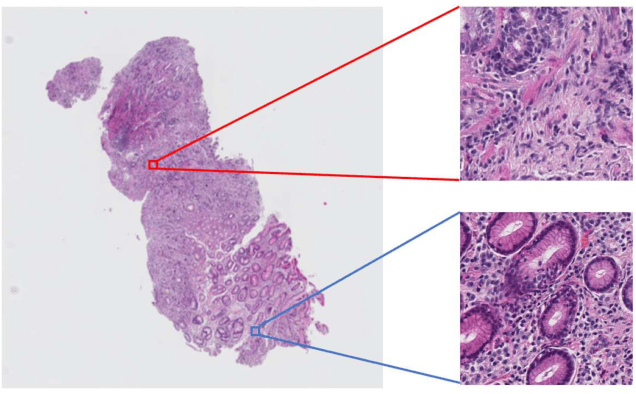
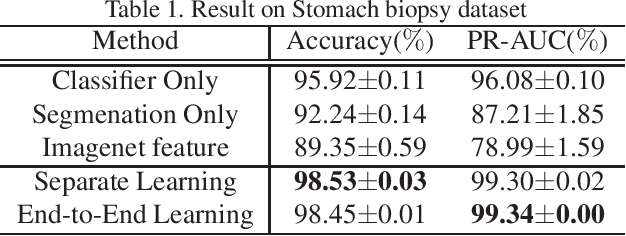
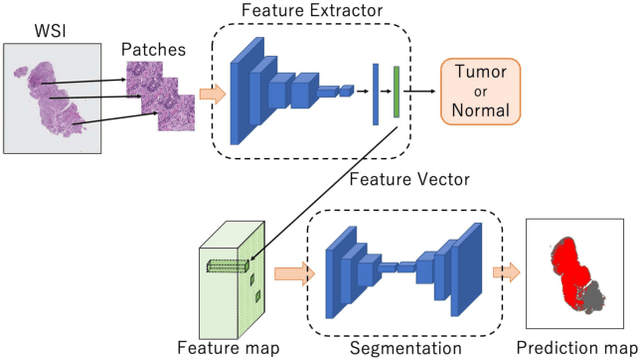
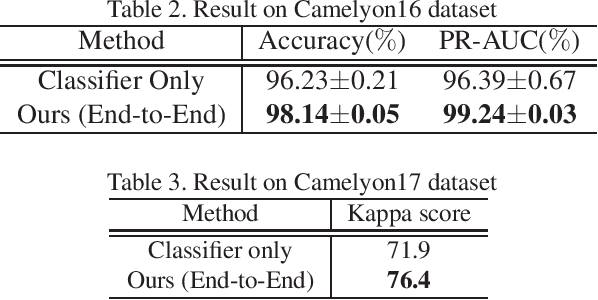
Histopathological image analysis is an essential process for the discovery of diseases such as cancer. However, it is challenging to train CNN on whole slide images (WSIs) of gigapixel resolution considering the available memory capacity. Most of the previous works divide high resolution WSIs into small image patches and separately input them into the model to classify it as a tumor or a normal tissue. However, patch-based classification uses only patch-scale local information but ignores the relationship between neighboring patches. If we consider the relationship of neighboring patches and global features, we can improve the classification performance. In this paper, we propose a new model structure combining the patch-based classification model and whole slide-scale segmentation model in order to improve the prediction performance of automatic pathological diagnosis. We extract patch features from the classification model and input them into the segmentation model to obtain a whole slide tumor probability heatmap. The classification model considers patch-scale local features, and the segmentation model can take global information into account. We also propose a new optimization method that retains gradient information and trains the model partially for end-to-end learning with limited GPU memory capacity. We apply our method to the tumor/normal prediction on WSIs and the classification performance is improved compared with the conventional patch-based method.
Fair Conformal Predictors for Applications in Medical Imaging
Sep 09, 2021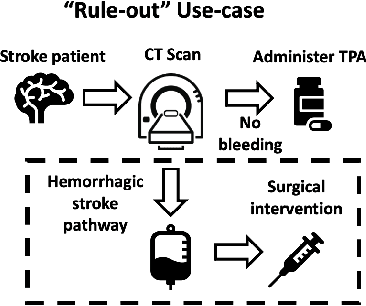
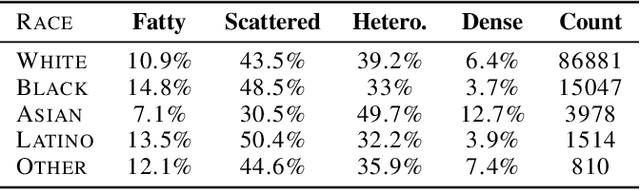
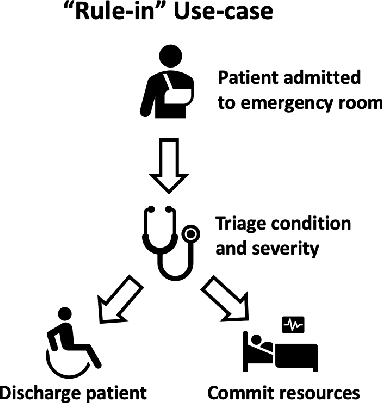
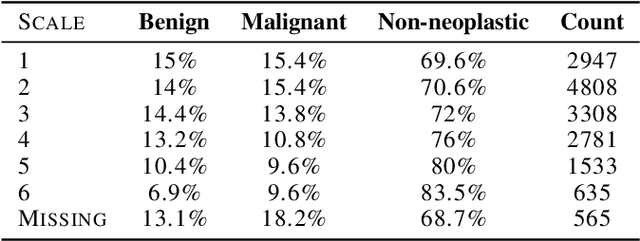
Deep learning has the potential to augment many components of the clinical workflow, such as medical image interpretation. However, the translation of these black box algorithms into clinical practice has been marred by the relative lack of transparency compared to conventional machine learning methods, hindering in clinician trust in the systems for critical medical decision-making. Specifically, common deep learning approaches do not have intuitive ways of expressing uncertainty with respect to cases that might require further human review. Furthermore, the possibility of algorithmic bias has caused hesitancy regarding the use of developed algorithms in clinical settings. To these ends, we explore how conformal methods can complement deep learning models by providing both clinically intuitive way (by means of confidence prediction sets) of expressing model uncertainty as well as facilitating model transparency in clinical workflows. In this paper, we conduct a field survey with clinicians to assess clinical use-cases of conformal predictions. Next, we conduct experiments with a mammographic breast density and dermatology photography datasets to demonstrate the utility of conformal predictions in "rule-in" and "rule-out" disease scenarios. Further, we show that conformal predictors can be used to equalize coverage with respect to patient demographics such as race and skin tone. We find that a conformal predictions to be a promising framework with potential to increase clinical usability and transparency for better collaboration between deep learning algorithms and clinicians.
Spatially Consistent Representation Learning
Mar 10, 2021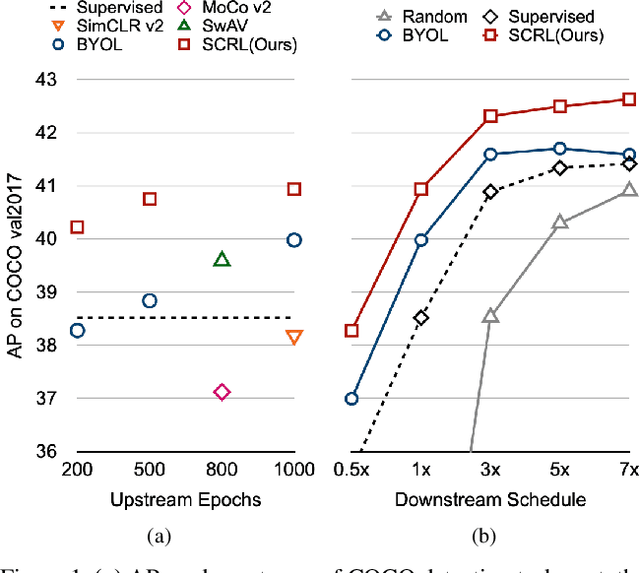

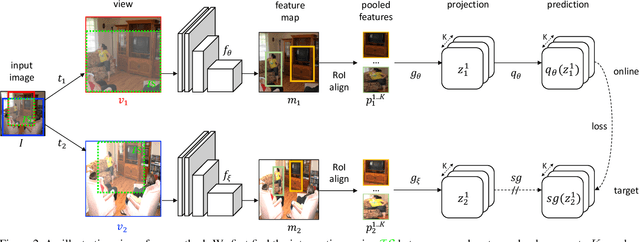
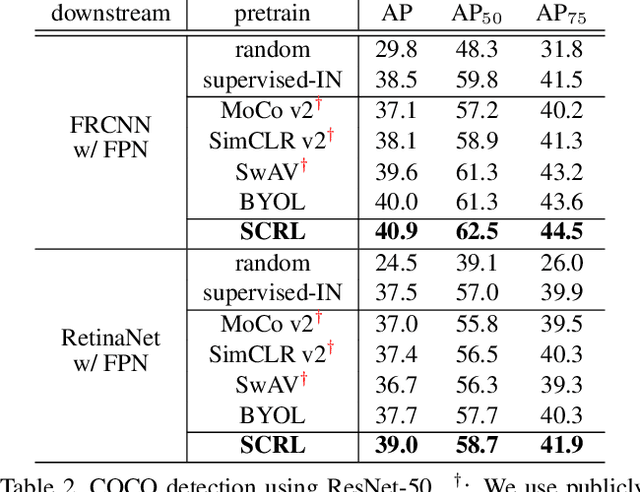
Self-supervised learning has been widely used to obtain transferrable representations from unlabeled images. Especially, recent contrastive learning methods have shown impressive performances on downstream image classification tasks. While these contrastive methods mainly focus on generating invariant global representations at the image-level under semantic-preserving transformations, they are prone to overlook spatial consistency of local representations and therefore have a limitation in pretraining for localization tasks such as object detection and instance segmentation. Moreover, aggressively cropped views used in existing contrastive methods can minimize representation distances between the semantically different regions of a single image. In this paper, we propose a spatially consistent representation learning algorithm (SCRL) for multi-object and location-specific tasks. In particular, we devise a novel self-supervised objective that tries to produce coherent spatial representations of a randomly cropped local region according to geometric translations and zooming operations. On various downstream localization tasks with benchmark datasets, the proposed SCRL shows significant performance improvements over the image-level supervised pretraining as well as the state-of-the-art self-supervised learning methods.
Global Guidance Network for Breast Lesion Segmentation in Ultrasound Images
Apr 05, 2021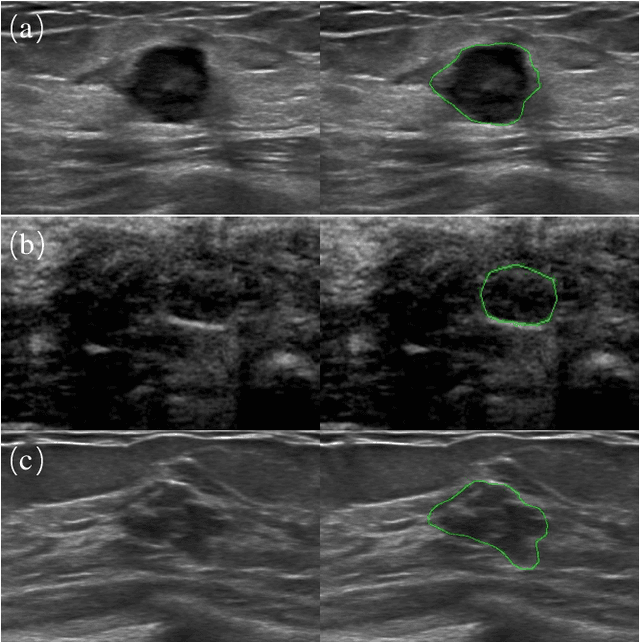
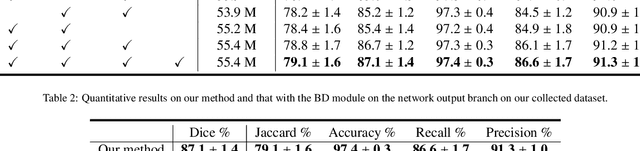
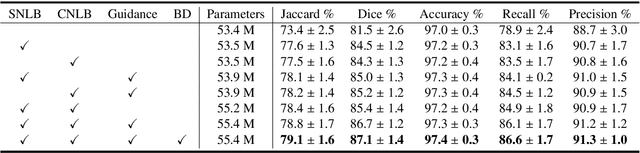
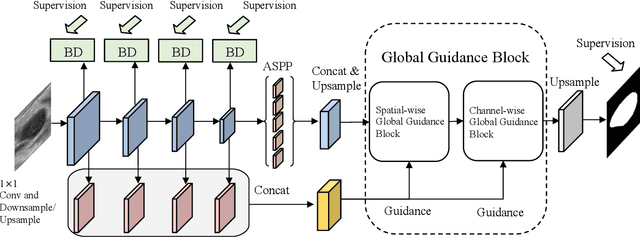

Automatic breast lesion segmentation in ultrasound helps to diagnose breast cancer, which is one of the dreadful diseases that affect women globally. Segmenting breast regions accurately from ultrasound image is a challenging task due to the inherent speckle artifacts, blurry breast lesion boundaries, and inhomogeneous intensity distributions inside the breast lesion regions. Recently, convolutional neural networks (CNNs) have demonstrated remarkable results in medical image segmentation tasks. However, the convolutional operations in a CNN often focus on local regions, which suffer from limited capabilities in capturing long-range dependencies of the input ultrasound image, resulting in degraded breast lesion segmentation accuracy. In this paper, we develop a deep convolutional neural network equipped with a global guidance block (GGB) and breast lesion boundary detection (BD) modules for boosting the breast ultrasound lesion segmentation. The GGB utilizes the multi-layer integrated feature map as a guidance information to learn the long-range non-local dependencies from both spatial and channel domains. The BD modules learn additional breast lesion boundary map to enhance the boundary quality of a segmentation result refinement. Experimental results on a public dataset and a collected dataset show that our network outperforms other medical image segmentation methods and the recent semantic segmentation methods on breast ultrasound lesion segmentation. Moreover, we also show the application of our network on the ultrasound prostate segmentation, in which our method better identifies prostate regions than state-of-the-art networks.
WenLan: Bridging Vision and Language by Large-Scale Multi-Modal Pre-Training
Mar 17, 2021
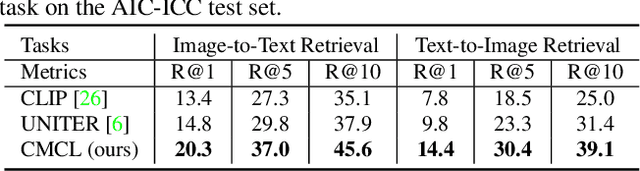
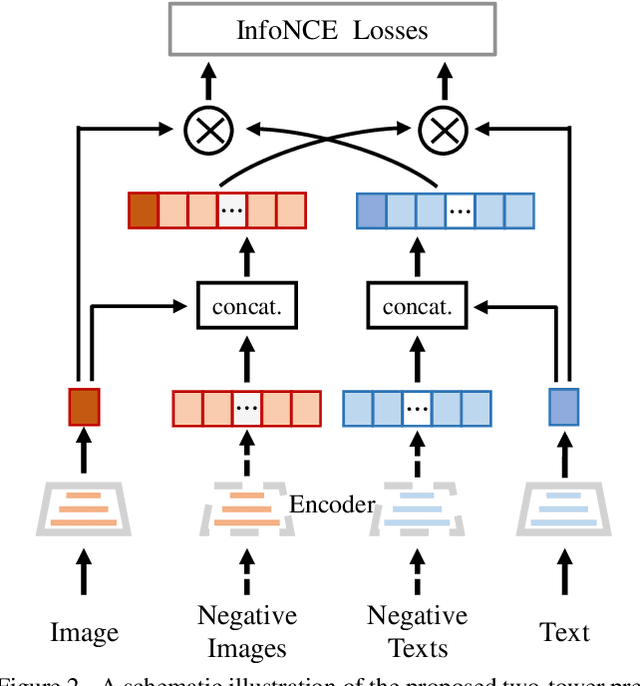
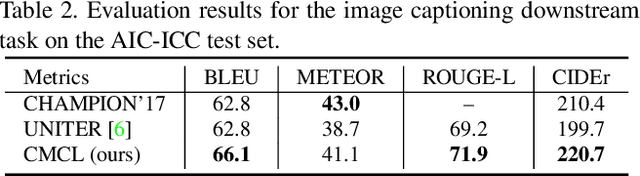
Multi-modal pre-training models have been intensively explored to bridge vision and language in recent years. However, most of them explicitly model the cross-modal interaction between image-text pairs, by assuming that there exists strong semantic correlation between the text and image modalities. Since this strong assumption is often invalid in real-world scenarios, we choose to implicitly model the cross-modal correlation for large-scale multi-modal pre-training, which is the focus of the Chinese project `WenLan' led by our team. Specifically, with the weak correlation assumption over image-text pairs, we propose a two-tower pre-training model called BriVL within the cross-modal contrastive learning framework. Unlike OpenAI CLIP that adopts a simple contrastive learning method, we devise a more advanced algorithm by adapting the latest method MoCo into the cross-modal scenario. By building a large queue-based dictionary, our BriVL can incorporate more negative samples in limited GPU resources. We further construct a large Chinese multi-source image-text dataset called RUC-CAS-WenLan for pre-training our BriVL model. Extensive experiments demonstrate that the pre-trained BriVL model outperforms both UNITER and OpenAI CLIP on various downstream tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge