"Image": models, code, and papers
SHARP: Shape-Aware Reconstruction of People In Loose Clothing
Jun 09, 2021
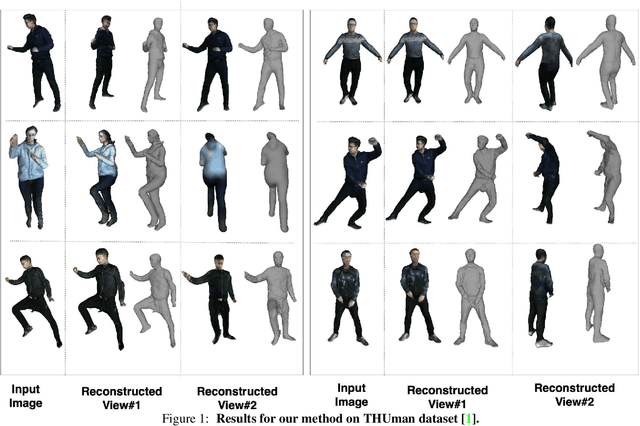
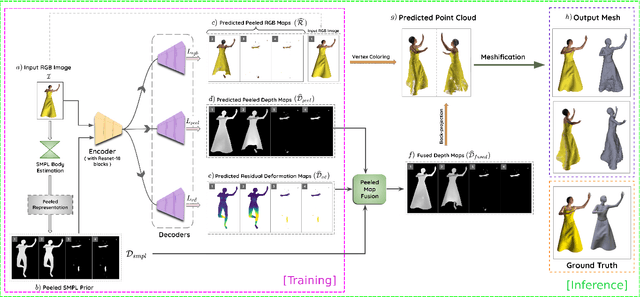
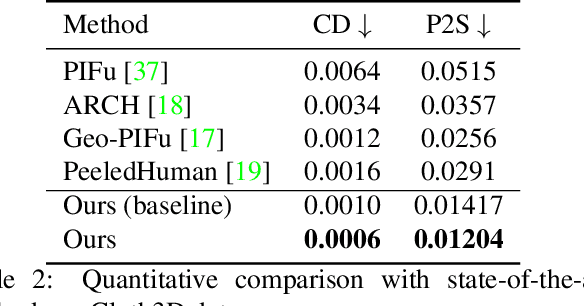
3D human body reconstruction from monocular images is an interesting and ill-posed problem in computer vision with wider applications in multiple domains. In this paper, we propose SHARP, a novel end-to-end trainable network that accurately recovers the detailed geometry and appearance of 3D people in loose clothing from a monocular image. We propose a sparse and efficient fusion of a parametric body prior with a non-parametric peeled depth map representation of clothed models. The parametric body prior constraints our model in two ways: first, the network retains geometrically consistent body parts that are not occluded by clothing, and second, it provides a body shape context that improves prediction of the peeled depth maps. This enables SHARP to recover fine-grained 3D geometrical details with just L1 losses on the 2D maps, given an input image. We evaluate SHARP on publicly available Cloth3D and THuman datasets and report superior performance to state-of-the-art approaches.
LB-CNN: An Open Source Framework for Fast Training of Light Binary Convolutional Neural Networks using Chainer and Cupy
Jun 25, 2021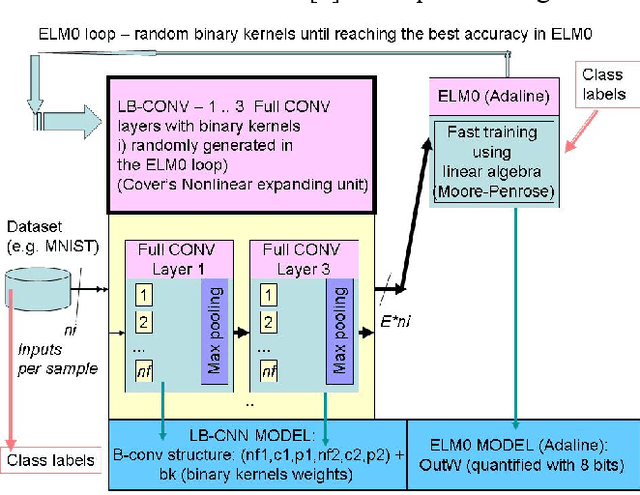
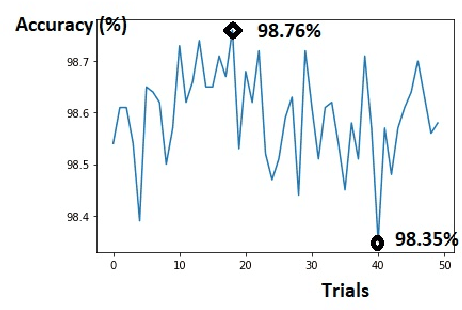
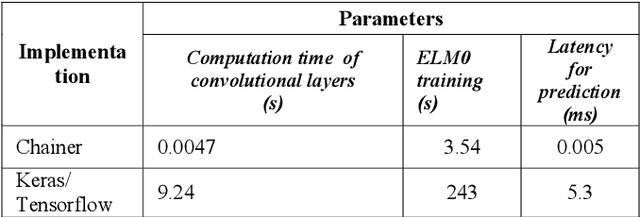
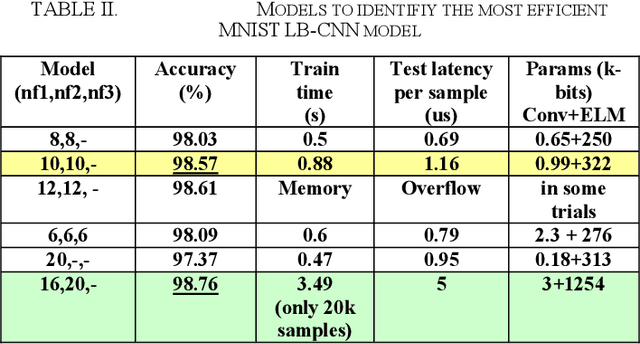
Light binary convolutional neural networks (LB-CNN) are particularly useful when implemented in low-energy computing platforms as required in many industrial applications. Herein, a framework for optimizing compact LB-CNN is introduced and its effectiveness is evaluated. The framework is freely available and may run on free-access cloud platforms, thus requiring no major investments. The optimized model is saved in the standardized .h5 format and can be used as input to specialized tools for further deployment into specific technologies, thus enabling the rapid development of various intelligent image sensors. The main ingredient in accelerating the optimization of our model, particularly the selection of binary convolution kernels, is the Chainer/Cupy machine learning library offering significant speed-ups for training the output layer as an extreme-learning machine. Additional training of the output layer using Keras/Tensorflow is included, as it allows an increase in accuracy. Results for widely used datasets including MNIST, GTSRB, ORL, VGG show very good compromise between accuracy and complexity. Particularly, for face recognition problems a carefully optimized LB-CNN model provides up to 100% accuracies. Such TinyML solutions are well suited for industrial applications requiring image recognition with low energy consumption.
Disentangle, align and fuse for multimodal and zero-shot image segmentation
Nov 12, 2019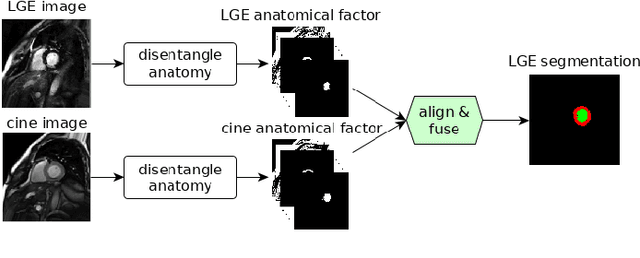
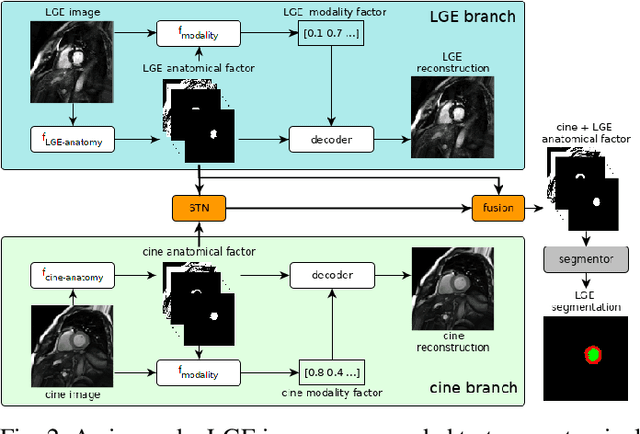
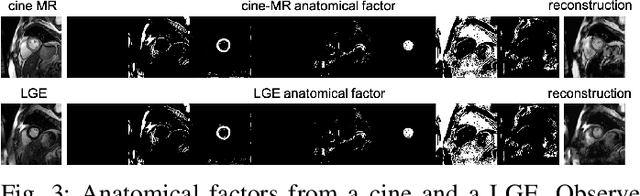

Magnetic resonance (MR) protocols rely on several sequences to properly assess pathology and organ status. Yet, despite advances in image analysis we tend to treat each sequence, here termed modality, in isolation. Taking advantage of the information shared between modalities (largely an organ's anatomy) is beneficial for multi-modality multi-input processing and learning. However, we must overcome inherent anatomical misregistrations and disparities in signal intensity across the modalities to claim this benefit. We present a method that offers improved segmentation accuracy of the modality of interest (over a single input model), by learning to leverage information present in other modalities, enabling semi-supervised and zero shot learning. Core to our method is learning a disentangled decomposition into anatomical and imaging factors. Shared anatomical factors from the different inputs are jointly processed and fused to extract more accurate segmentation masks. Image misregistrations are corrected with a Spatial Transformer Network, that non-linearly aligns the anatomical factors. The imaging factor captures signal intensity characteristics across different modality data, and is used for image reconstruction, enabling semi-supervised learning. Temporal and slice pairing between inputs are learned dynamically. We demonstrate applications in Late Gadolinium Enhanced (LGE) and Blood Oxygenation Level Dependent (BOLD) cardiac segmentation, as well as in T2 abdominal segmentation.
Learning of Image Dehazing Models for Segmentation Tasks
Mar 04, 2019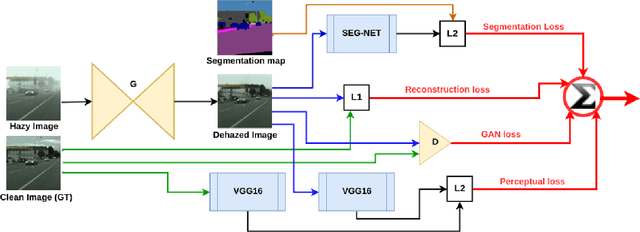
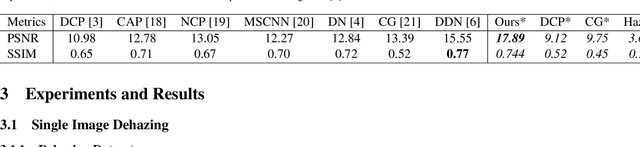

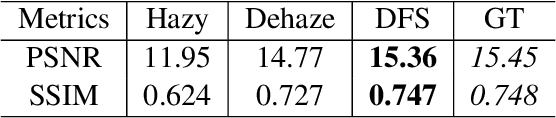
To evaluate their performance, existing dehazing approaches generally rely on distance measures between the generated image and its corresponding ground truth. Despite its ability to produce visually good images, using pixel-based or even perceptual metrics do not guarantee, in general, that the produced image is fit for being used as input for low-level computer vision tasks such as segmentation. To overcome this weakness, we are proposing a novel end-to-end approach for image dehazing, fit for being used as input to an image segmentation procedure, while maintaining the visual quality of the generated images. Inspired by the success of Generative Adversarial Networks (GAN), we propose to optimize the generator by introducing a discriminator network and a loss function that evaluates segmentation quality of dehazed images. In addition, we make use of a supplementary loss function that verifies that the visual and the perceptual quality of the generated image are preserved in hazy conditions. Results obtained using the proposed technique are appealing, with a favorable comparison to state-of-the-art approaches when considering the performance of segmentation algorithms on the hazy images.
NeMI: Unifying Neural Radiance Fields with Multiplane Images for Novel View Synthesis
Mar 27, 2021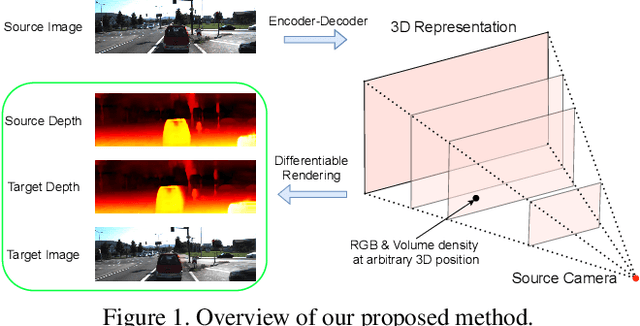
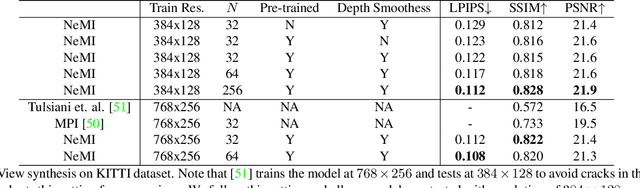
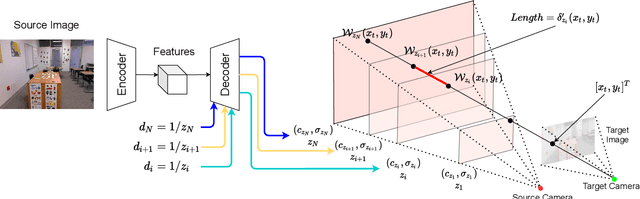
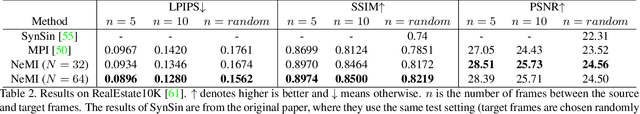
In this paper, we propose an approach to perform novel view synthesis and depth estimation via dense 3D reconstruction from a single image. Our NeMI unifies Neural radiance fields (NeRF) with Multiplane Images (MPI). Specifically, our NeMI is a general two-dimensional and image-conditioned extension of NeRF, and a continuous depth generalization of MPI. Given a single image as input, our method predicts a 4-channel image (RGB and volume density) at arbitrary depth values to jointly reconstruct the camera frustum and fill in occluded contents. The reconstructed and inpainted frustum can then be easily rendered into novel RGB or depth views using differentiable rendering. Extensive experiments on RealEstate10K, KITTI and Flowers Light Fields show that our NeMI outperforms state-of-the-art by a large margin in novel view synthesis. We also achieve competitive results in depth estimation on iBims-1 and NYU-v2 without annotated depth supervision. Project page available at https://vincentfung13.github.io/projects/nemi/
Training Automatic View Planner for Cardiac MR Imaging via Self-Supervision by Spatial Relationship between Views
Sep 24, 2021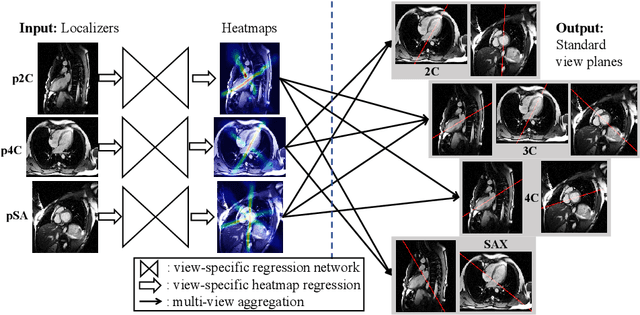
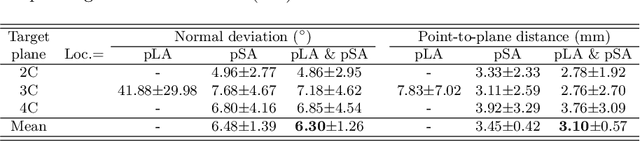
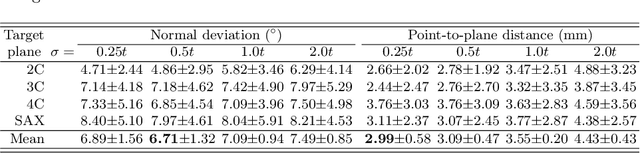
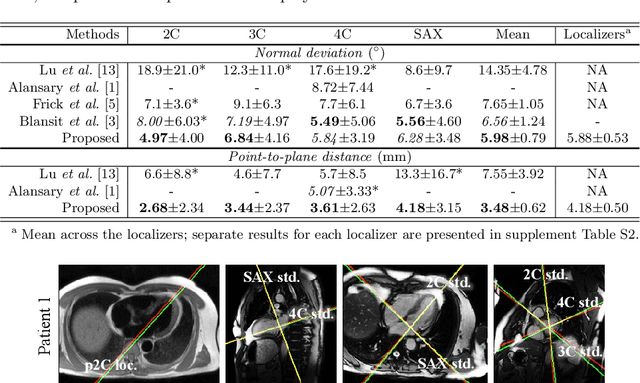
View planning for the acquisition of cardiac magnetic resonance imaging (CMR) requires acquaintance with the cardiac anatomy and remains a challenging task in clinical practice. Existing approaches to its automation relied either on an additional volumetric image not typically acquired in clinic routine, or on laborious manual annotations of cardiac structural landmarks. This work presents a clinic-compatible and annotation-free system for automatic CMR view planning. The system mines the spatial relationship -- more specifically, locates and exploits the intersecting lines -- between the source and target views, and trains deep networks to regress heatmaps defined by these intersecting lines. As the spatial relationship is self-contained in properly stored data, e.g., in the DICOM format, the need for manual annotation is eliminated. Then, a multi-view planning strategy is proposed to aggregate information from the predicted heatmaps for all the source views of a target view, for a globally optimal prescription. The multi-view aggregation mimics the similar strategy practiced by skilled human prescribers. Experimental results on 181 clinical CMR exams show that our system achieves superior accuracy to existing approaches including conventional atlas-based and newer deep learning based ones, in prescribing four standard CMR views. The mean angle difference and point-to-plane distance evaluated against the ground truth planes are 5.98 degrees and 3.48 mm, respectively.
e-UDA: Efficient Unsupervised Domain Adaptation for Cross-Site Medical Image Segmentation
Jan 25, 2020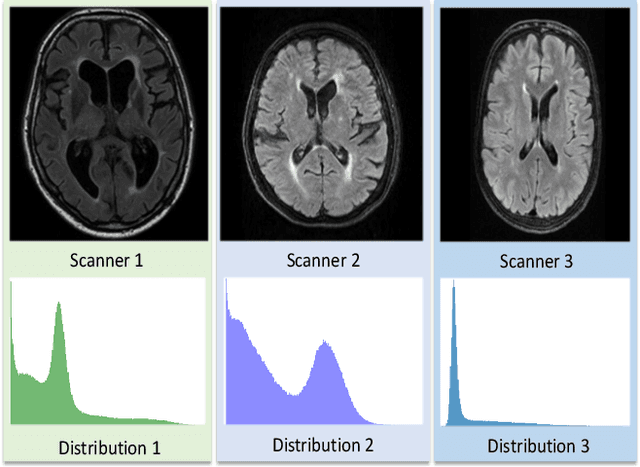
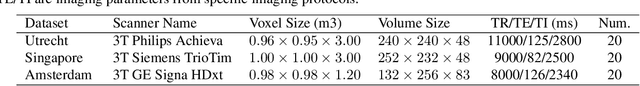
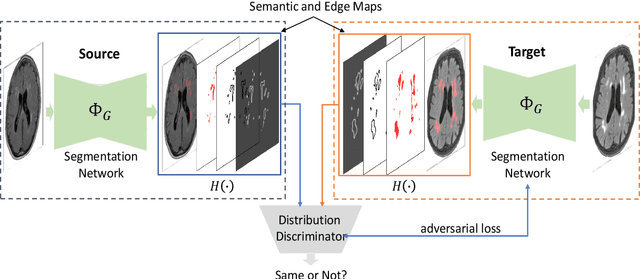
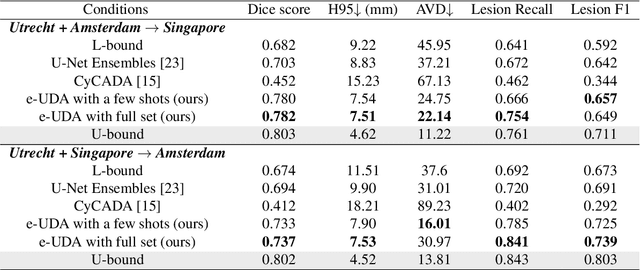
Domain adaptation in healthcare data is a potentially critical component in making computer-aided diagnostic systems applicable cross multiple sites and imaging scanners. In this paper, we propose an efficient unsupervised domain adaptation framework for robust image segmentation cross multiple similar domains. We enforce our algorithm to not only adapt to the new domains via an adversarial optimization, rejecting unlikely segmentation patterns, but also to maintain its performance on the source training data, by incorporating both semantic and boundary information into the data distributions. Further, as we do not have labels for the transfer domain, we propose a new quality score for the adaptation process, and strategies to retrain the diagnostic algorithm in a stable fashion. Using multi-centric data from a public benchmark for brain lesion segmentation, we demonstrate that recalibrating on just few unlabeled image sets from the target domain improves segmentation accuracies drastically, with performances almost similar to those from algorithms trained on fresh and fully annotated data from the test domain.
U-Net Using Stacked Dilated Convolutions for Medical Image Segmentation
Apr 10, 2020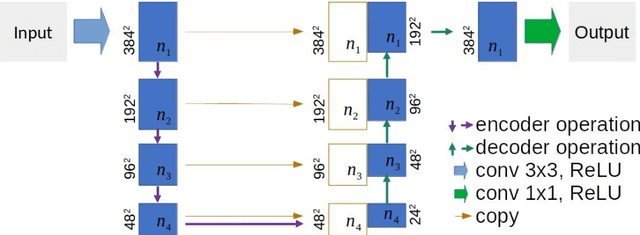

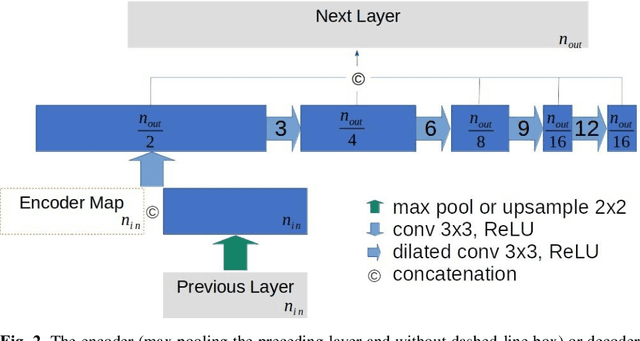
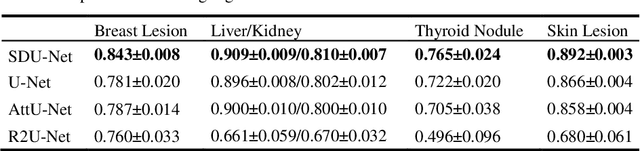
This paper proposes a novel U-Net variant using stacked dilated convolutions for medical image segmentation (SDU-Net). SDU-Net adopts the architecture of vanilla U-Net with modifications in the encoder and decoder operations (an operation indicates all the processing for feature maps of the same resolution). Unlike vanilla U-Net which incorporates two standard convolutions in each encoder/decoder operation, SDU-Net uses one standard convolution followed by multiple dilated convolutions and concatenates all dilated convolution outputs as input to the next operation. Experiments showed that SDU-Net outperformed vanilla U-Net, attention U-Net (AttU-Net), and recurrent residual U-Net (R2U-Net) in all four tested segmentation tasks while using parameters around 40% of vanilla U-Net's, 17% of AttU-Net's, and 15% of R2U-Net's.
High Resolution Medical Image Analysis with Spatial Partitioning
Sep 12, 2019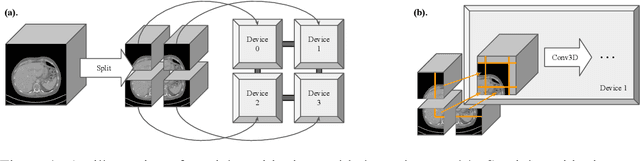
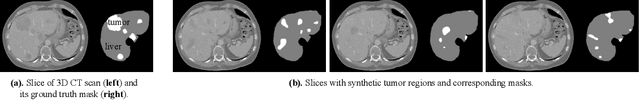
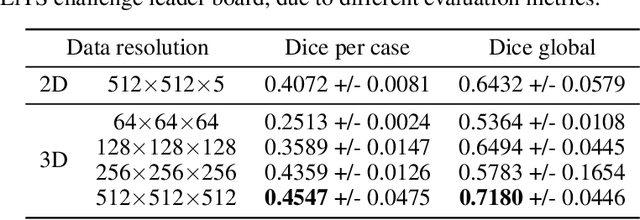

Medical images such as 3D computerized tomography (CT) scans and pathology images, have hundreds of millions or billions of voxels/pixels. It is infeasible to train CNN models directly on such high resolution images, because neural activations of a single image do not fit in the memory of a single GPU/TPU, and naive data and model parallelism approaches do not work. Existing image analysis approaches alleviate this problem by cropping or down-sampling input images, which leads to complicated implementation and sub-optimal performance due to information loss. In this paper, we implement spatial partitioning, which internally distributes the input and output of convolutional layers across GPUs/TPUs. Our implementation is based on the Mesh-TensorFlow framework and the computation distribution is transparent to end users. With this technique, we train a 3D Unet on up to 512 by 512 by 512 resolution data. To the best of our knowledge, this is the first work for handling such high resolution images end-to-end.
Image search using multilingual texts: a cross-modal learning approach between image and text
Apr 02, 2019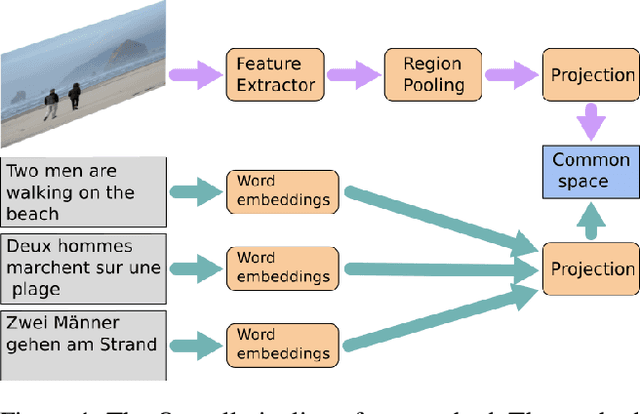
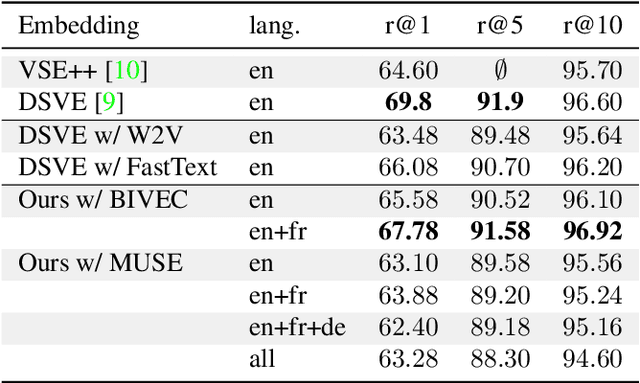
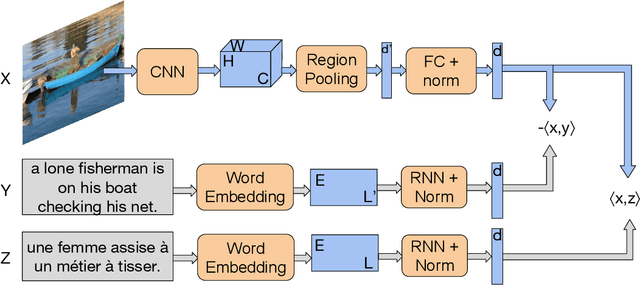
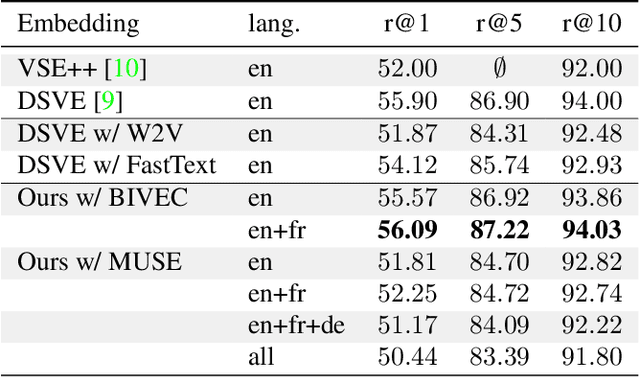
Multilingual (or cross-lingual) embeddings represent several languages in a unique vector space. Using a common embedding space enables for a shared semantic between words from different languages. In this paper, we propose to embed images and texts into a unique distributional vector space, enabling to search images by using text queries expressing information needs related to the (visual) content of images, as well as using image similarity. Our framework forces the representation of an image to be similar to the representation of the text that describes it. Moreover, by using multilingual embeddings we ensure that words from two different languages have close descriptors and thus are attached to similar images. We provide experimental evidence of the efficiency of our approach by experimenting it on two datasets: Common Objects in COntext (COCO) [19] and Multi30K [7].
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge