"Image": models, code, and papers
Explain Me the Painting: Multi-Topic Knowledgeable Art Description Generation
Sep 13, 2021
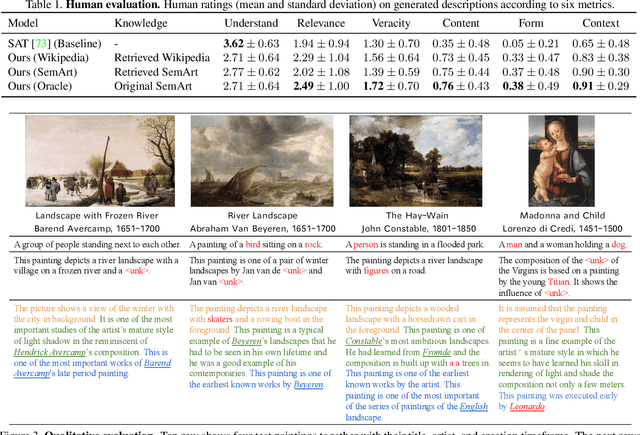
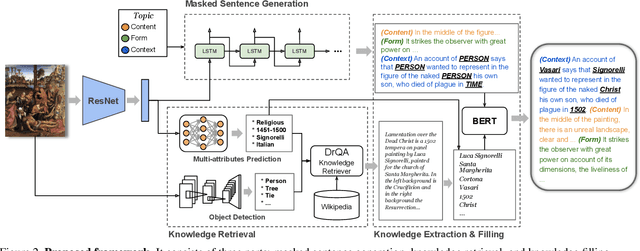
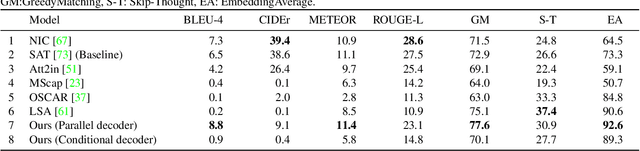
Have you ever looked at a painting and wondered what is the story behind it? This work presents a framework to bring art closer to people by generating comprehensive descriptions of fine-art paintings. Generating informative descriptions for artworks, however, is extremely challenging, as it requires to 1) describe multiple aspects of the image such as its style, content, or composition, and 2) provide background and contextual knowledge about the artist, their influences, or the historical period. To address these challenges, we introduce a multi-topic and knowledgeable art description framework, which modules the generated sentences according to three artistic topics and, additionally, enhances each description with external knowledge. The framework is validated through an exhaustive analysis, both quantitative and qualitative, as well as a comparative human evaluation, demonstrating outstanding results in terms of both topic diversity and information veracity.
All-Optical Image Identification with Programmable Matrix Transformation
Apr 01, 2021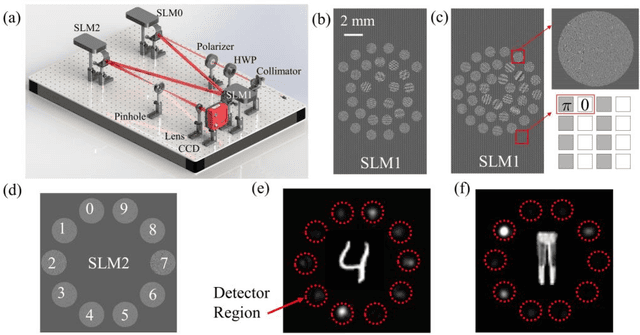
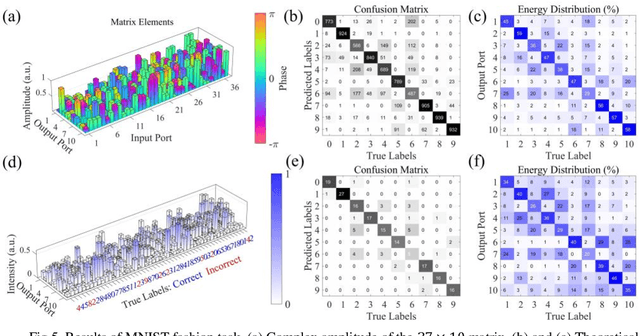
An optical neural network is proposed and demonstrated with programmable matrix transformation and nonlinear activation function of photodetection (square-law detection). Based on discrete phase-coherent spatial modes, the dimensionality of programmable optical matrix operations is 30~37, which is implemented by spatial light modulators. With this architecture, all-optical classification tasks of handwritten digits, objects and depth images are performed on the same platform with high accuracy. Due to the parallel nature of matrix multiplication, the processing speed of our proposed architecture is potentially as high as7.4T~74T FLOPs per second (with 10~100GHz detector)
E1D3 U-Net for Brain Tumor Segmentation: Submission to the RSNA-ASNR-MICCAI BraTS 2021 Challenge
Oct 06, 2021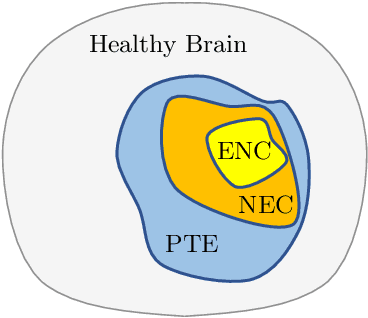
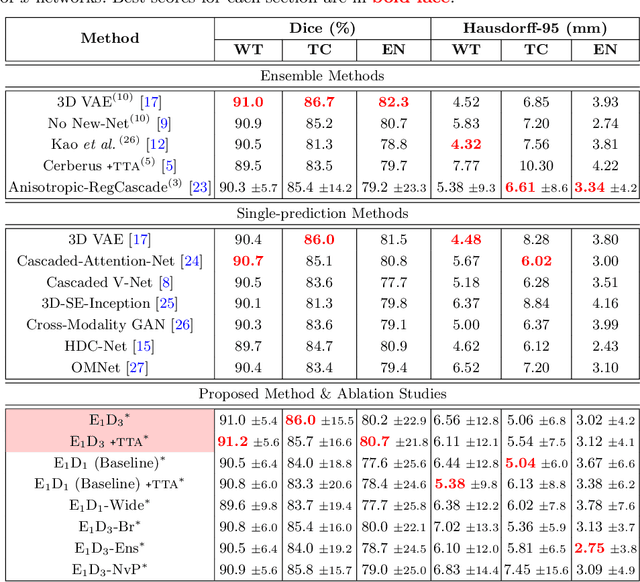
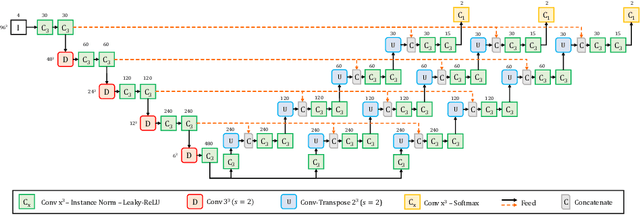
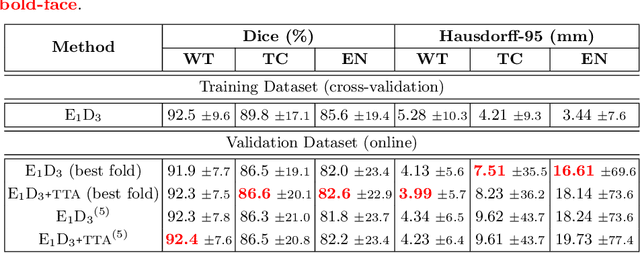
Convolutional Neural Networks (CNNs) have demonstrated state-of-the-art performance in medical image segmentation tasks. A common feature in most top-performing CNNs is an encoder-decoder architecture inspired by the U-Net. For multi-region brain tumor segmentation, 3D U-Net architecture and its variants provide the most competitive segmentation performances. In this work, we propose an interesting extension of the standard 3D U-Net architecture, specialized for brain tumor segmentation. The proposed network, called E1D3 U-Net, is a one-encoder, three-decoder fully-convolutional neural network architecture where each decoder segments one of the hierarchical regions of interest: whole tumor, tumor core, and enhancing core. On the BraTS 2018 validation (unseen) dataset, E1D3 U-Net demonstrates single-prediction performance comparable with most state-of-the-art networks in brain tumor segmentation, with reasonable computational requirements and without ensembling. As a submission to the RSNA-ASNR-MICCAI BraTS 2021 challenge, we also evaluate our proposal on the BraTS 2021 dataset. E1D3 U-Net showcases the flexibility in the standard 3D U-Net architecture which we exploit for the task of brain tumor segmentation.
Does Vision-and-Language Pretraining Improve Lexical Grounding?
Sep 21, 2021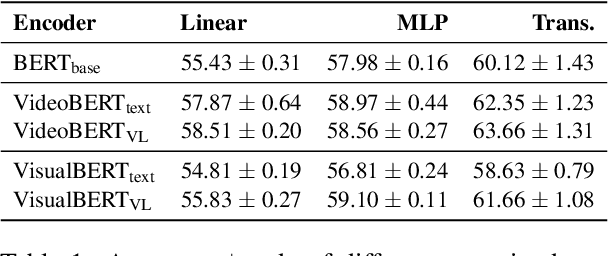
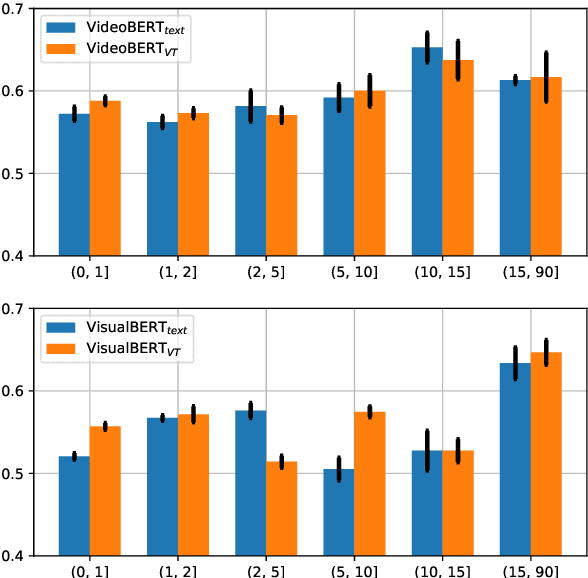
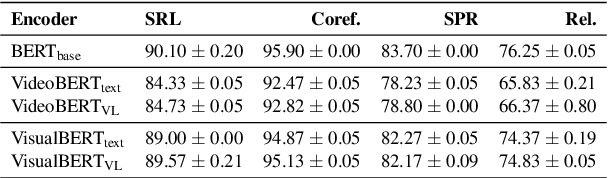
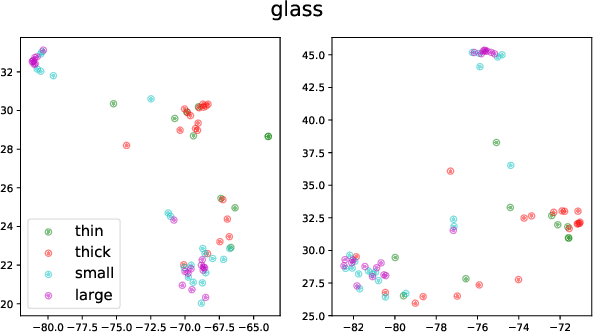
Linguistic representations derived from text alone have been criticized for their lack of grounding, i.e., connecting words to their meanings in the physical world. Vision-and-Language (VL) models, trained jointly on text and image or video data, have been offered as a response to such criticisms. However, while VL pretraining has shown success on multimodal tasks such as visual question answering, it is not yet known how the internal linguistic representations themselves compare to their text-only counterparts. This paper compares the semantic representations learned via VL vs. text-only pretraining for two recent VL models using a suite of analyses (clustering, probing, and performance on a commonsense question answering task) in a language-only setting. We find that the multimodal models fail to significantly outperform the text-only variants, suggesting that future work is required if multimodal pretraining is to be pursued as a means of improving NLP in general.
SuperCaptioning: Image Captioning Using Two-dimensional Word Embedding
Jun 04, 2019
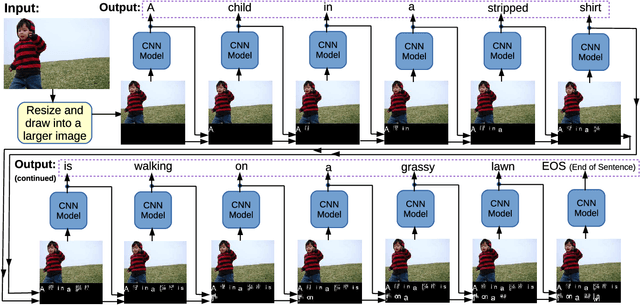
Language and vision are processed as two different modal in current work for image captioning. However, recent work on Super Characters method shows the effectiveness of two-dimensional word embedding, which converts text classification problem into image classification problem. In this paper, we propose the SuperCaptioning method, which borrows the idea of two-dimensional word embedding from Super Characters method, and processes the information of language and vision together in one single CNN model. The experimental results on Flickr30k data shows the proposed method gives high quality image captions. An interactive demo is ready to show at the workshop.
Identity-Expression Ambiguity in 3D Morphable Face Models
Sep 29, 2021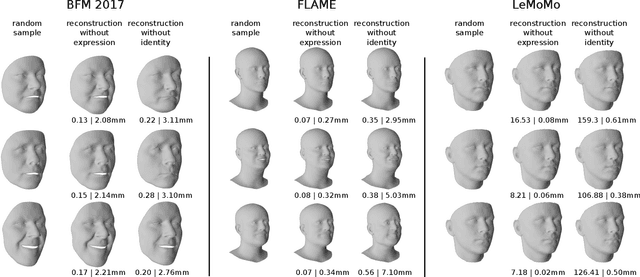
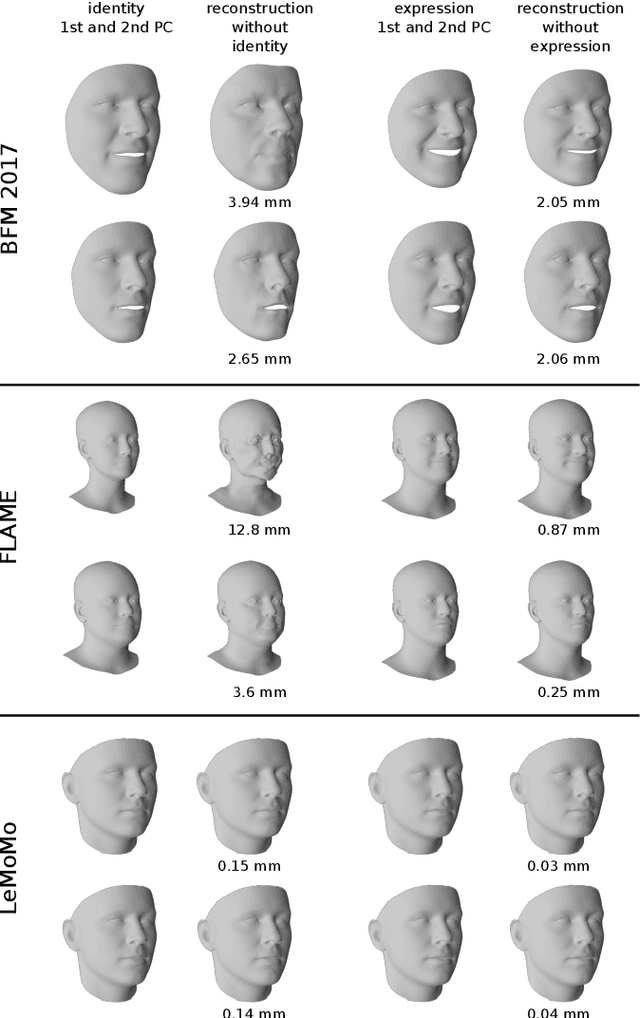

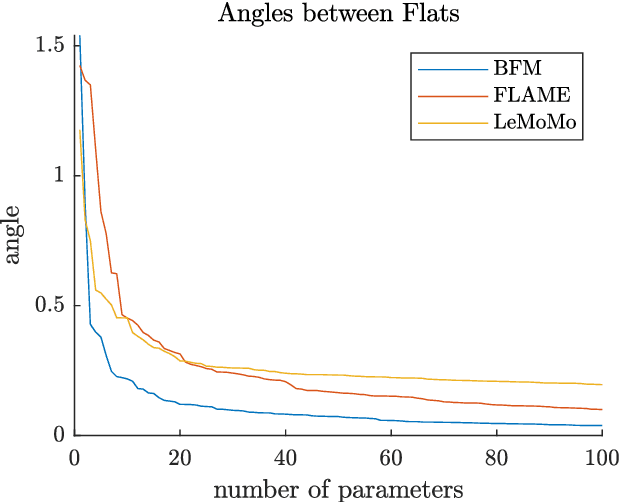
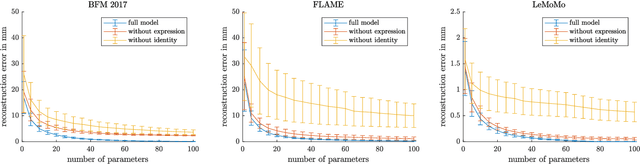
3D Morphable Models are a class of generative models commonly used to model faces. They are typically applied to ill-posed problems such as 3D reconstruction from 2D data. Several ambiguities in this problem's image formation process have been studied explicitly. We demonstrate that non-orthogonality of the variation in identity and expression can cause identity-expression ambiguity in 3D Morphable Models, and that in practice expression and identity are far from orthogonal and can explain each other surprisingly well. Whilst previously reported ambiguities only arise in an inverse rendering setting, identity-expression ambiguity emerges in the 3D shape generation process itself. We demonstrate this effect with 3D shapes directly as well as through an inverse rendering task, and use two popular models built from high quality 3D scans as well as a model built from a large collection of 2D images and videos. We explore this issue's implications for inverse rendering and observe that it cannot be resolved by a purely statistical prior on identity and expression deformations.
Transform-Invariant Convolutional Neural Networks for Image Classification and Search
Nov 28, 2019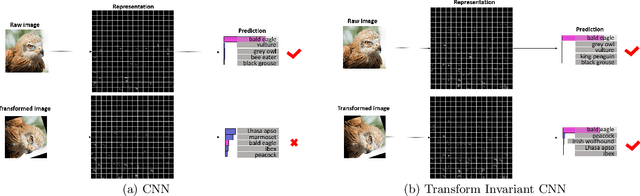
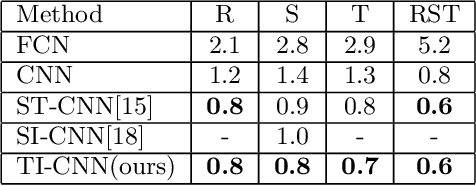
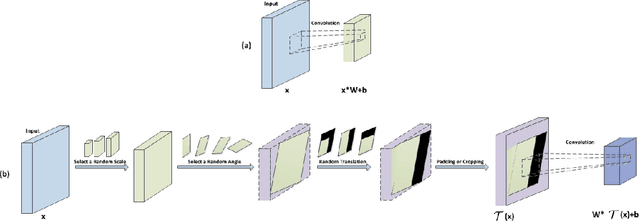
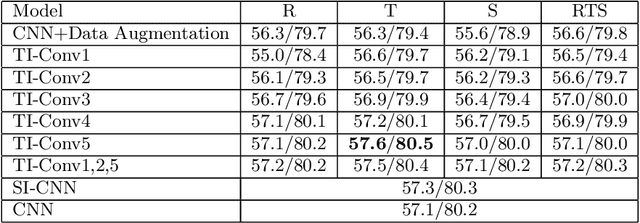
Convolutional neural networks (CNNs) have achieved state-of-the-art results on many visual recognition tasks. However, current CNN models still exhibit a poor ability to be invariant to spatial transformations of images. Intuitively, with sufficient layers and parameters, hierarchical combinations of convolution (matrix multiplication and non-linear activation) and pooling operations should be able to learn a robust mapping from transformed input images to transform-invariant representations. In this paper, we propose randomly transforming (rotation, scale, and translation) feature maps of CNNs during the training stage. This prevents complex dependencies of specific rotation, scale, and translation levels of training images in CNN models. Rather, each convolutional kernel learns to detect a feature that is generally helpful for producing the transform-invariant answer given the combinatorially large variety of transform levels of its input feature maps. In this way, we do not require any extra training supervision or modification to the optimization process and training images. We show that random transformation provides significant improvements of CNNs on many benchmark tasks, including small-scale image recognition, large-scale image recognition, and image retrieval. The code is available at https://github.com/jasonustc/caffe-multigpu/tree/TICNN.
Joint Triplet Autoencoder for Histopathological Colon Cancer Nuclei Retrieval
May 24, 2021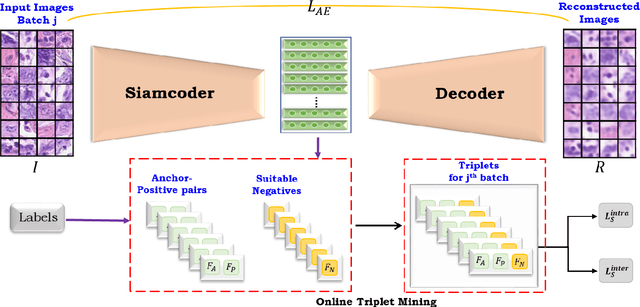
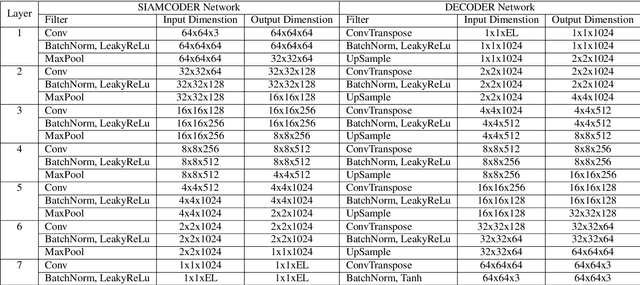
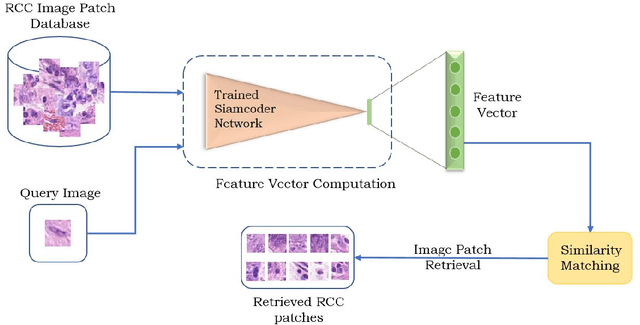
Deep learning has shown a great improvement in the performance of visual tasks. Image retrieval is the task of extracting the visually similar images from a database for a query image. The feature matching is performed to rank the images. Various hand-designed features have been derived in past to represent the images. Nowadays, the power of deep learning is being utilized for automatic feature learning from data in the field of biomedical image analysis. Autoencoder and Siamese networks are two deep learning models to learn the latent space (i.e., features or embedding). Autoencoder works based on the reconstruction of the image from latent space. Siamese network utilizes the triplets to learn the intra-class similarity and inter-class dissimilarity. Moreover, Autoencoder is unsupervised, whereas Siamese network is supervised. We propose a Joint Triplet Autoencoder Network (JTANet) by facilitating the triplet learning in autoencoder framework. A joint supervised learning for Siamese network and unsupervised learning for Autoencoder is performed. Moreover, the Encoder network of Autoencoder is shared with Siamese network and referred as the Siamcoder network. The features are extracted by using the trained Siamcoder network for retrieval purpose. The experiments are performed over Histopathological Routine Colon Cancer dataset. We have observed the promising performance using the proposed JTANet model against the Autoencoder and Siamese models for colon cancer nuclei retrieval in histopathological images.
Second-Order Neural ODE Optimizer
Sep 29, 2021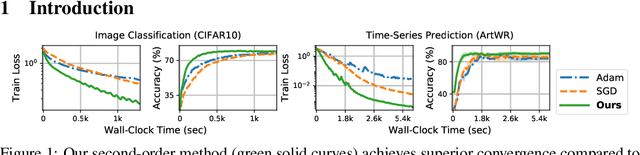
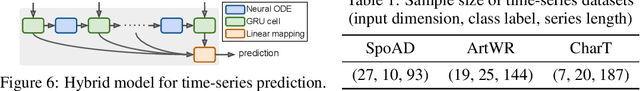
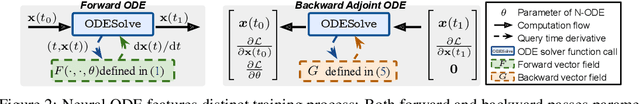
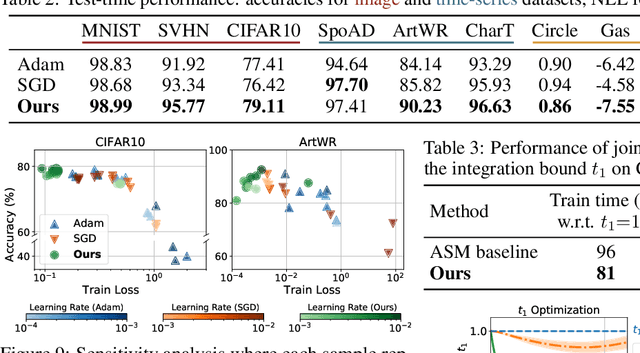
We propose a novel second-order optimization framework for training the emerging deep continuous-time models, specifically the Neural Ordinary Differential Equations (Neural ODEs). Since their training already involves expensive gradient computation by solving a backward ODE, deriving efficient second-order methods becomes highly nontrivial. Nevertheless, inspired by the recent Optimal Control (OC) interpretation of training deep networks, we show that a specific continuous-time OC methodology, called Differential Programming, can be adopted to derive backward ODEs for higher-order derivatives at the same O(1) memory cost. We further explore a low-rank representation of the second-order derivatives and show that it leads to efficient preconditioned updates with the aid of Kronecker-based factorization. The resulting method converges much faster than first-order baselines in wall-clock time, and the improvement remains consistent across various applications, e.g. image classification, generative flow, and time-series prediction. Our framework also enables direct architecture optimization, such as the integration time of Neural ODEs, with second-order feedback policies, strengthening the OC perspective as a principled tool of analyzing optimization in deep learning.
Hyperspectral-Multispectral Image Fusion with Weighted LASSO
Mar 15, 2020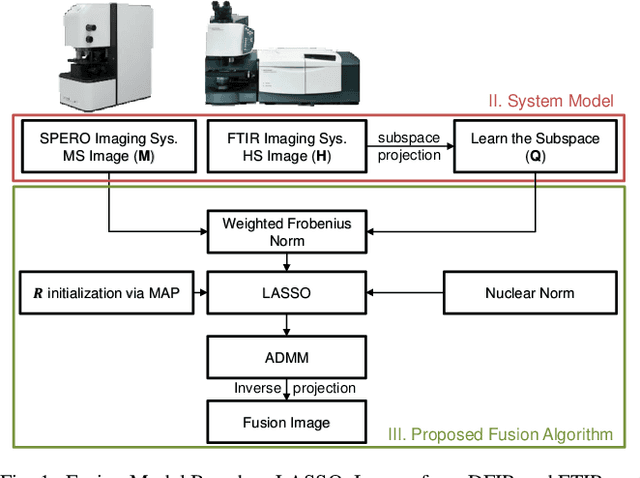
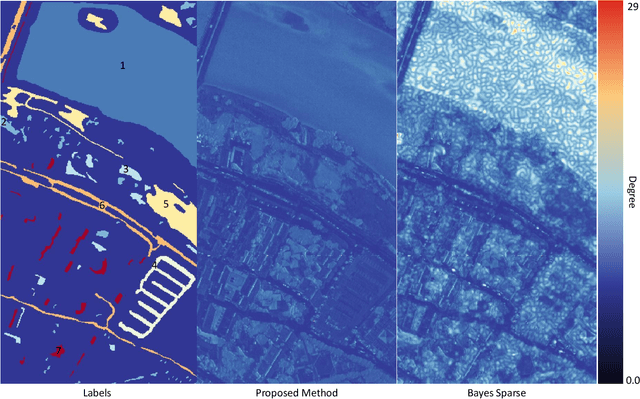

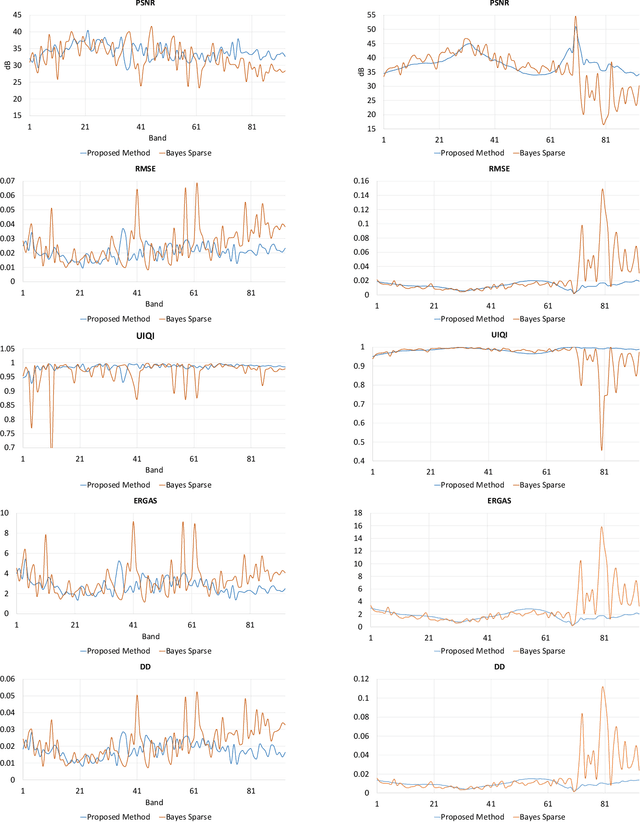
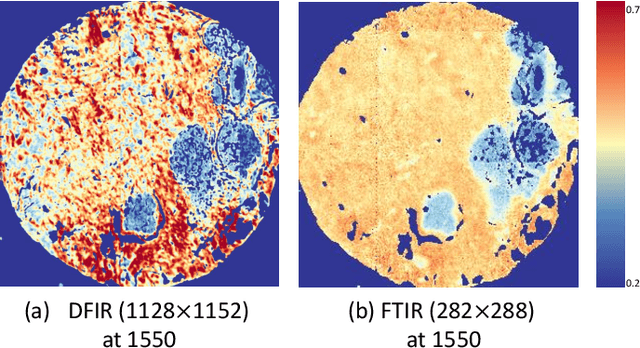
Spectral imaging enables spatially-resolved identification of materials in remote sensing, biomedicine, and astronomy. However, acquisition times require balancing spectral and spatial resolution with signal-to-noise. Hyperspectral imaging provides superior material specificity, while multispectral images are faster to collect at greater fidelity. We propose an approach for fusing hyperspectral and multispectral images to provide high-quality hyperspectral output. The proposed optimization leverages the least absolute shrinkage and selection operator (LASSO) to perform variable selection and regularization. Computational time is reduced by applying the alternating direction method of multipliers (ADMM), as well as initializing the fusion image by estimating it using maximum a posteriori (MAP) based on Hardie's method. We demonstrate that the proposed sparse fusion and reconstruction provides quantitatively superior results when compared to existing methods on publicly available images. Finally, we show how the proposed method can be practically applied in biomedical infrared spectroscopic microscopy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge