"Image": models, code, and papers
Convolutional Neural Network (CNN) vs Visual Transformer (ViT) for Digital Holography
Aug 20, 2021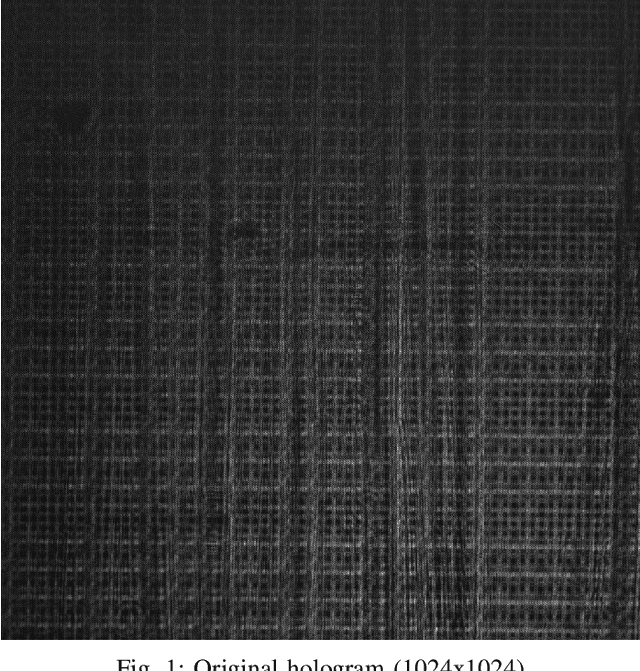
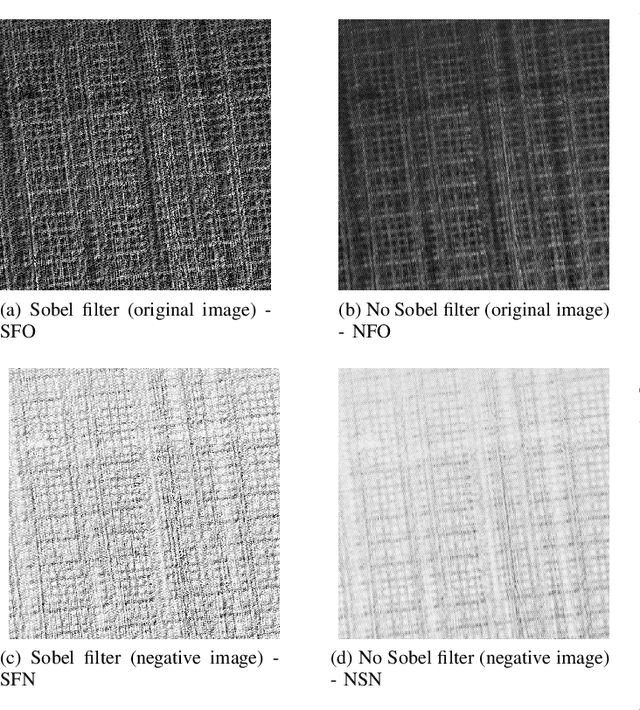
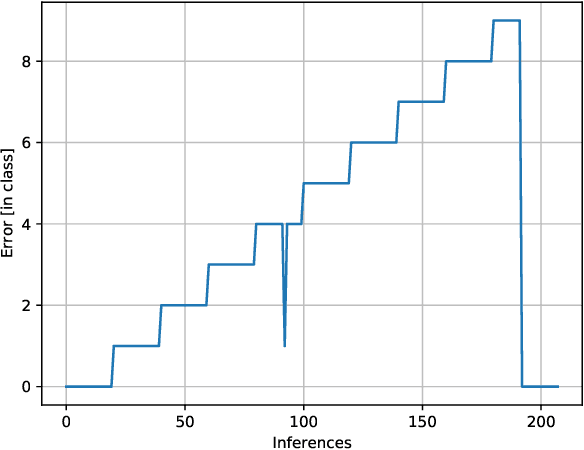
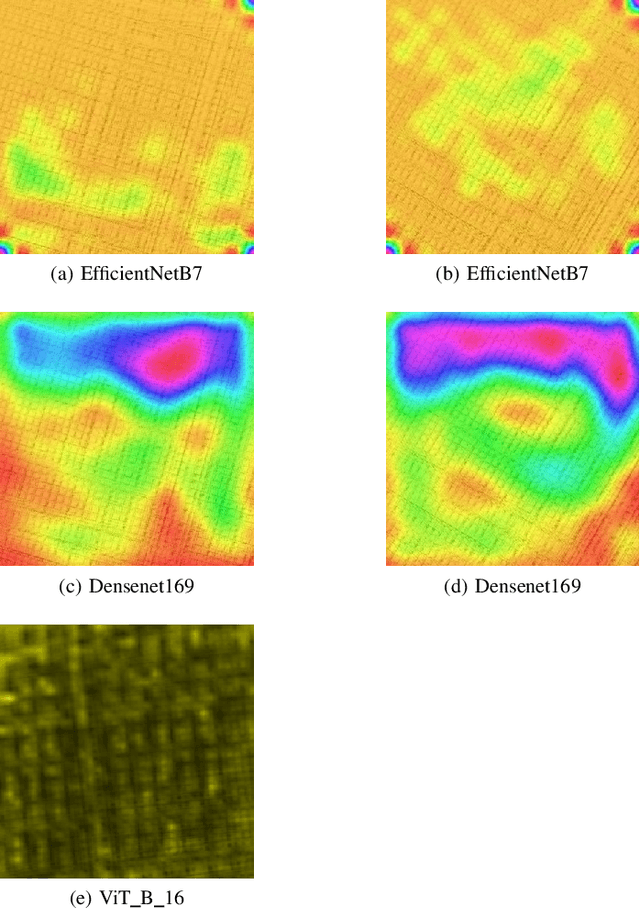

In Digital Holography (DH), it is crucial to extract the object distance from a hologram in order to reconstruct its amplitude and phase. This step is called auto-focusing and it is conventionally solved by first reconstructing a stack of images and then by sharpening each reconstructed image using a focus metric such as entropy or variance. The distance corresponding to the sharpest image is considered the focal position. This approach, while effective, is computationally demanding and time-consuming. In this paper, the determination of the distance is performed by Deep Learning (DL). Two deep learning (DL) architectures are compared: Convolutional Neural Network (CNN)and Visual transformer (ViT). ViT and CNN are used to cope with the problem of auto-focusing as a classification problem. Compared to a first attempt [11] in which the distance between two consecutive classes was 100{\mu}m, our proposal allows us to drastically reduce this distance to 1{\mu}m. Moreover, ViT reaches similar accuracy and is more robust than CNN.
DeeperLab: Single-Shot Image Parser
Feb 13, 2019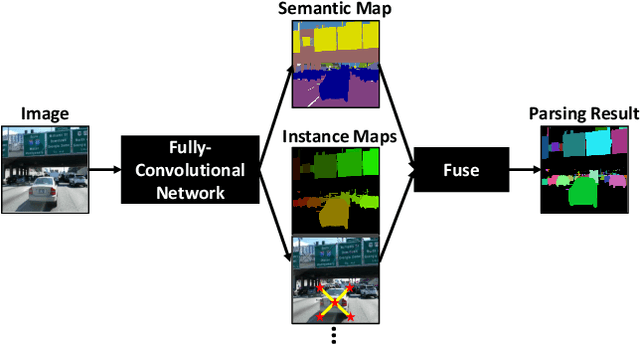
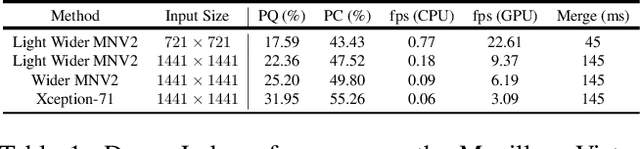
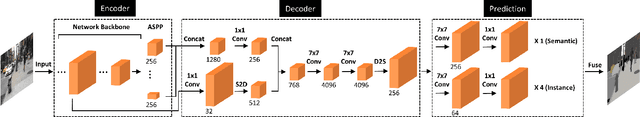
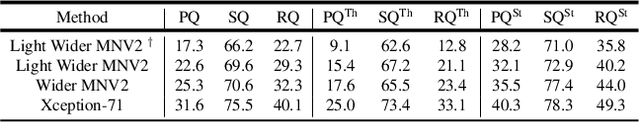
We present a single-shot, bottom-up approach for whole image parsing. Whole image parsing, also known as Panoptic Segmentation, generalizes the tasks of semantic segmentation for 'stuff' classes and instance segmentation for 'thing' classes, assigning both semantic and instance labels to every pixel in an image. Recent approaches to whole image parsing typically employ separate standalone modules for the constituent semantic and instance segmentation tasks and require multiple passes of inference. Instead, the proposed DeeperLab image parser performs whole image parsing with a significantly simpler, fully convolutional approach that jointly addresses the semantic and instance segmentation tasks in a single-shot manner, resulting in a streamlined system that better lends itself to fast processing. For quantitative evaluation, we use both the instance-based Panoptic Quality (PQ) metric and the proposed region-based Parsing Covering (PC) metric, which better captures the image parsing quality on 'stuff' classes and larger object instances. We report experimental results on the challenging Mapillary Vistas dataset, in which our single model achieves 31.95% (val) / 31.6% PQ (test) and 55.26% PC (val) with 3 frames per second (fps) on GPU or near real-time speed (22.6 fps on GPU) with reduced accuracy.
Deep Poisoning Functions: Towards Robust Privacy-safe Image Data Sharing
Dec 14, 2019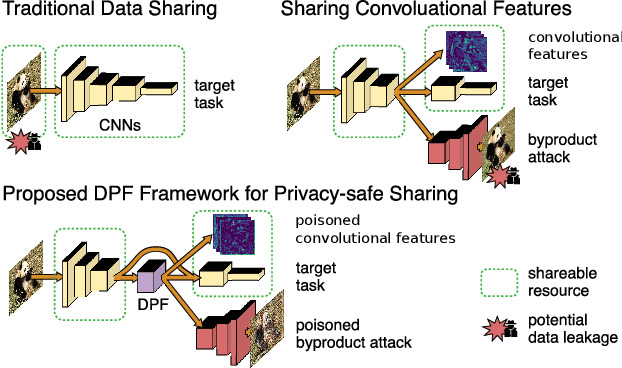
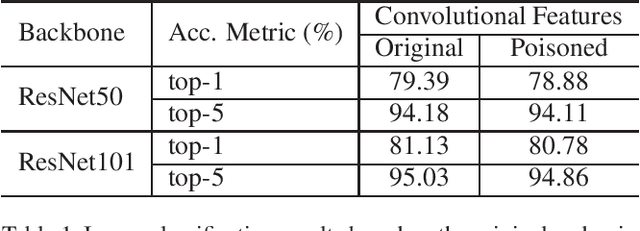
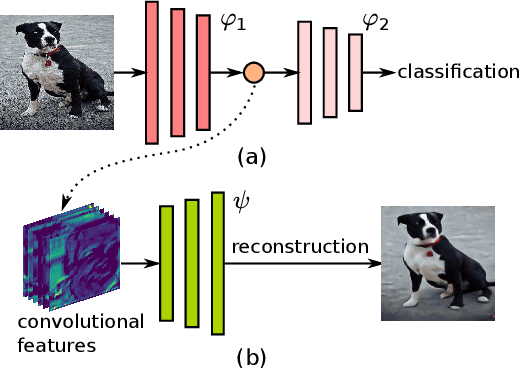
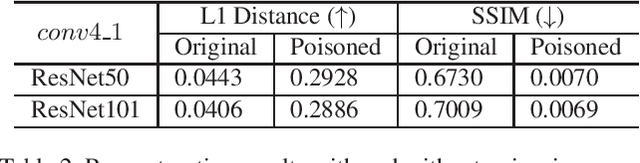
As deep networks are applied to an ever-expanding set of computer vision tasks, protecting general privacy in image data has become a critically important goal. This paper presents a new framework for privacy-preserving data sharing that is robust to adversarial attacks and overcomes the known issues existing in previous approaches. We introduce the concept of a Deep Poisoning Function (DPF), which is a module inserted into a pre-trained deep network designed to perform a specific vision task. The DPF is optimized to deliberately poison image data to prevent known adversarial attacks, while ensuring that the altered image data is functionally equivalent to the non-poisoned data for the original task. Given this equivalence, both poisoned and non-poisoned data can be used for further retraining or fine-tuning. Experimental results on image classification and face recognition tasks prove the efficacy of the proposed method.
Metric Learning for Image Registration
Apr 21, 2019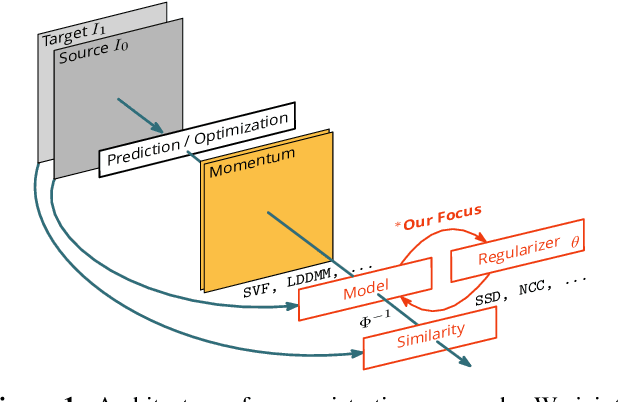
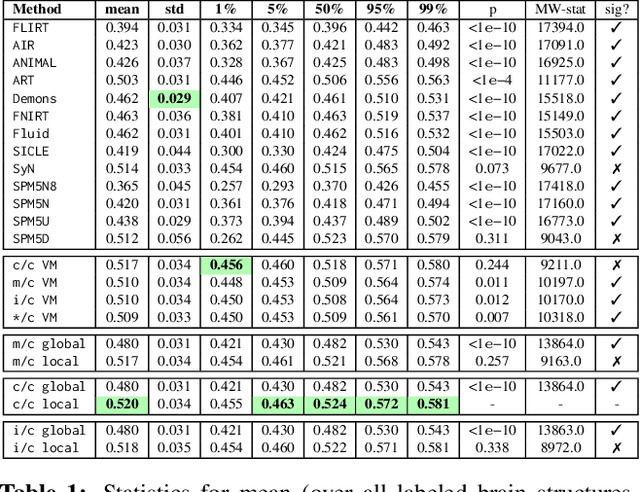
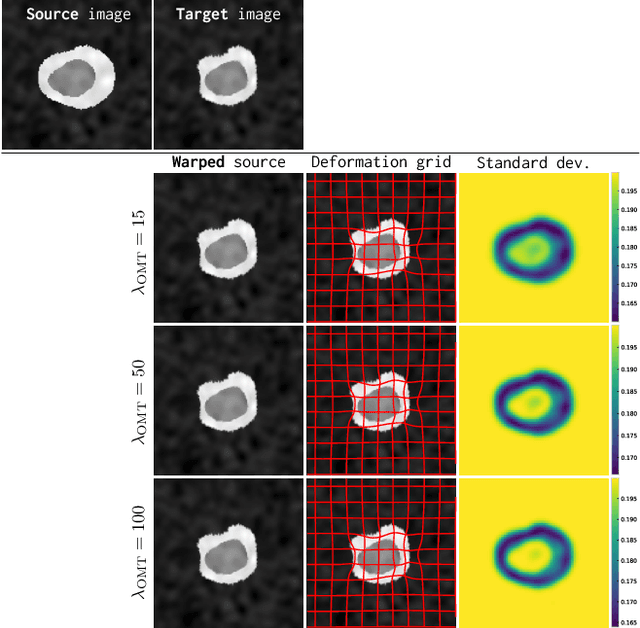
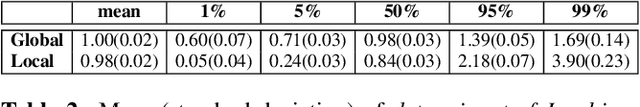
Image registration is a key technique in medical image analysis to estimate deformations between image pairs. A good deformation model is important for high-quality estimates. However, most existing approaches use ad-hoc deformation models chosen for mathematical convenience rather than to capture observed data variation. Recent deep learning approaches learn deformation models directly from data. However, they provide limited control over the spatial regularity of transformations. Instead of learning the entire registration approach, we learn a spatially-adaptive regularizer within a registration model. This allows controlling the desired level of regularity and preserving structural properties of a registration model. For example, diffeomorphic transformations can be attained. Our approach is a radical departure from existing deep learning approaches to image registration by embedding a deep learning model in an optimization-based registration algorithm to parameterize and data-adapt the registration model itself.
BIAS: Transparent reporting of biomedical image analysis challenges
Oct 23, 2019The number of biomedical image analysis challenges organized per year is steadily increasing. These international competitions have the purpose of benchmarking algorithms on common data sets, typically to identify the best method for a given problem. Recent research, however, revealed that common practice related to challenge reporting does not allow for adequate interpretation and reproducibility of results. To address the discrepancy between the impact of challenges and the quality (control), the Biomedical I mage Analysis ChallengeS (BIAS) initiative developed a set of recommendations for the reporting of challenges. The BIAS statement aims to improve the transparency of the reporting of a biomedical image analysis challenge regardless of field of application, image modality or task category assessed. This article describes how the BIAS statement was developed and presents a checklist which authors of biomedical image analysis challenges are encouraged to include in their submission when giving a paper on a challenge into review. The purpose of the checklist is to standardize and facilitate the review process and raise interpretability and reproducibility of challenge results by making relevant information explicit.
Single Independent Component Recovery and Applications
Oct 12, 2021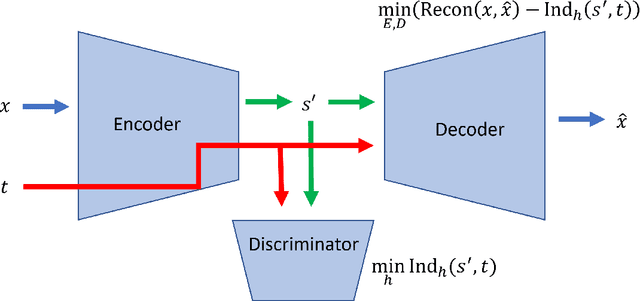
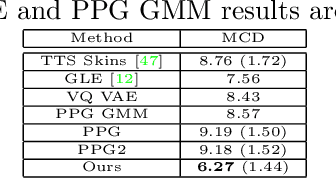
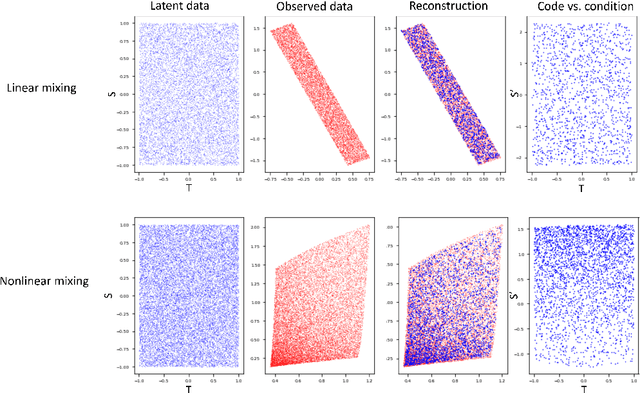
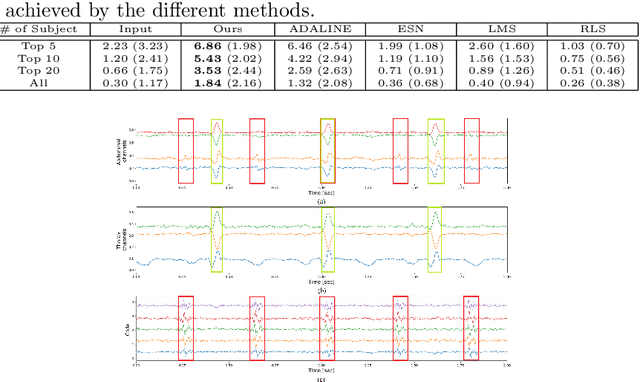
Latent variable discovery is a central problem in data analysis with a broad range of applications in applied science. In this work, we consider data given as an invertible mixture of two statistically independent components, and assume that one of the components is observed while the other is hidden. Our goal is to recover the hidden component. For this purpose, we propose an autoencoder equipped with a discriminator. Unlike the standard nonlinear ICA problem, which was shown to be non-identifiable, in the special case of ICA we consider here, we show that our approach can recover the component of interest up to entropy-preserving transformation. We demonstrate the performance of the proposed approach on several datasets, including image synthesis, voice cloning, and fetal ECG extraction.
Dynamic Fusion Network for RGBT Tracking
Sep 16, 2021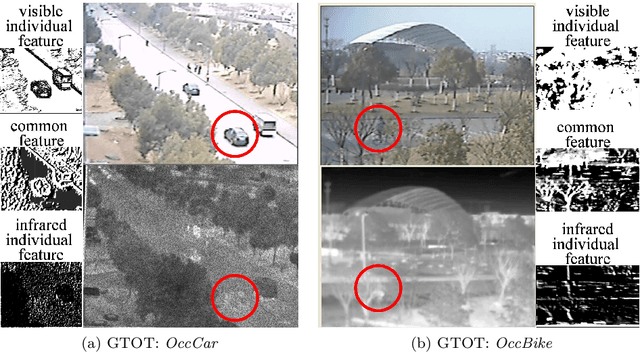
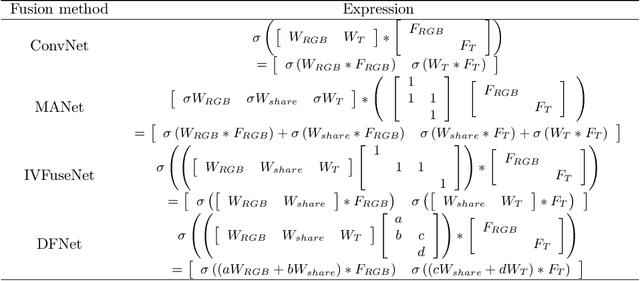
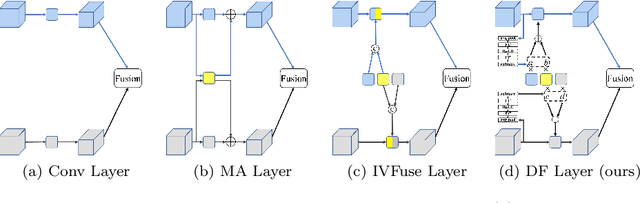
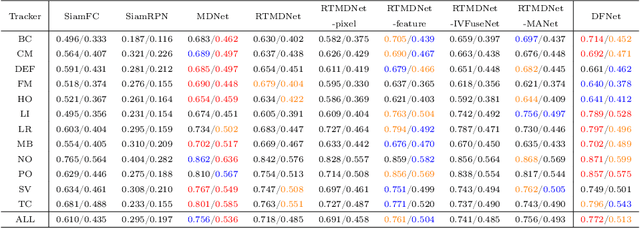
For both visible and infrared images have their own advantages and disadvantages, RGBT tracking has attracted more and more attention. The key points of RGBT tracking lie in feature extraction and feature fusion of visible and infrared images. Current RGBT tracking methods mostly pay attention to both individual features (features extracted from images of a single camera) and common features (features extracted and fused from an RGB camera and a thermal camera), while pay less attention to the different and dynamic contributions of individual features and common features for different sequences of registered image pairs. This paper proposes a novel RGBT tracking method, called Dynamic Fusion Network (DFNet), which adopts a two-stream structure, in which two non-shared convolution kernels are employed in each layer to extract individual features. Besides, DFNet has shared convolution kernels for each layer to extract common features. Non-shared convolution kernels and shared convolution kernels are adaptively weighted and summed according to different image pairs, so that DFNet can deal with different contributions for different sequences. DFNet has a fast speed, which is 28.658 FPS. The experimental results show that when DFNet only increases the Mult-Adds of 0.02% than the non-shared-convolution-kernel-based fusion method, Precision Rate (PR) and Success Rate (SR) reach 88.1% and 71.9% respectively.
Neural Image Captioning
Jul 02, 2019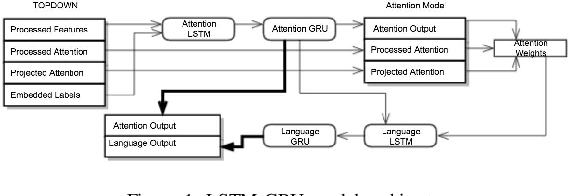



In recent years, the biggest advances in major Computer Vision tasks, such as object recognition, handwritten-digit identification, facial recognition, and many others., have all come through the use of Convolutional Neural Networks (CNNs). Similarly, in the domain of Natural Language Processing, Recurrent Neural Networks (RNNs), and Long Short Term Memory networks (LSTMs) in particular, have been crucial to some of the biggest breakthroughs in performance for tasks such as machine translation, part-of-speech tagging, sentiment analysis, and many others. These individual advances have greatly benefited tasks even at the intersection of NLP and Computer Vision, and inspired by this success, we studied some existing neural image captioning models that have proven to work well. In this work, we study some existing captioning models that provide near state-of-the-art performances, and try to enhance one such model. We also present a simple image captioning model that makes use of a CNN, an LSTM, and the beam search1 algorithm, and study its performance based on various qualitative and quantitative metrics.
Authentication Attacks on Projection-based Cancelable Biometric Schemes
Oct 28, 2021
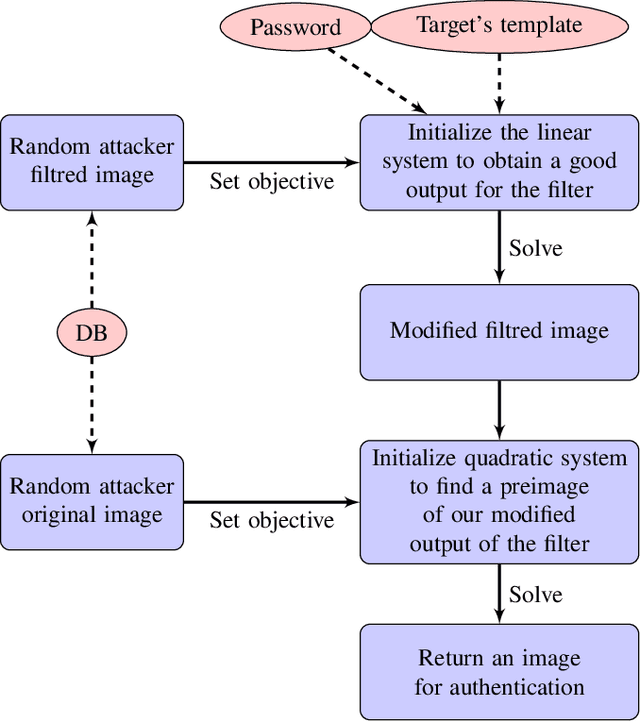
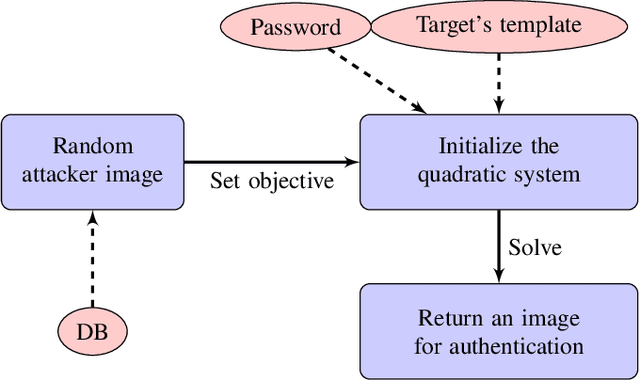
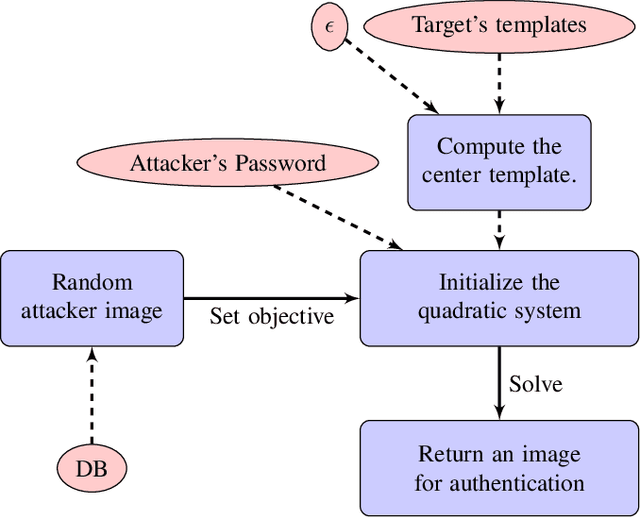
Cancelable biometric schemes aim at generating secure biometric templates by combining user specific tokens, such as password, stored secret or salt, along with biometric data. This type of transformation is constructed as a composition of a biometric transformation with a feature extraction algorithm. The security requirements of cancelable biometric schemes concern the irreversibility, unlinkability and revocability of templates, without losing in accuracy of comparison. While several schemes were recently attacked regarding these requirements, full reversibility of such a composition in order to produce colliding biometric characteristics, and specifically presentation attacks, were never demonstrated to the best of our knowledge. In this paper, we formalize these attacks for a traditional cancelable scheme with the help of integer linear programming (ILP) and quadratically constrained quadratic programming (QCQP). Solving these optimization problems allows an adversary to slightly alter its fingerprint image in order to impersonate any individual. Moreover, in an even more severe scenario, it is possible to simultaneously impersonate several individuals.
Supervised Segmentation with Domain Adaptation for Small Sampled Orbital CT Images
Jul 01, 2021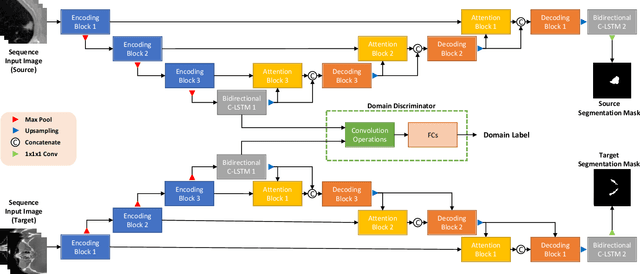
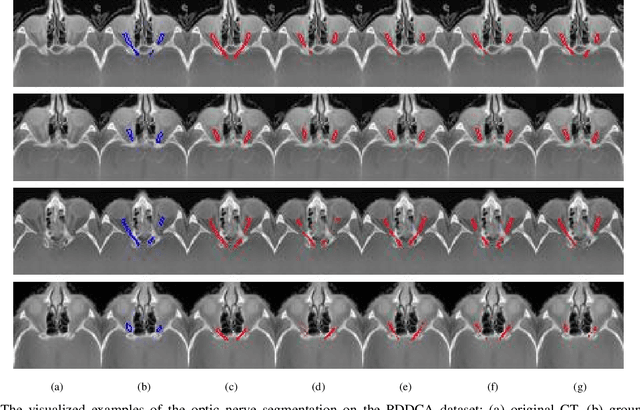
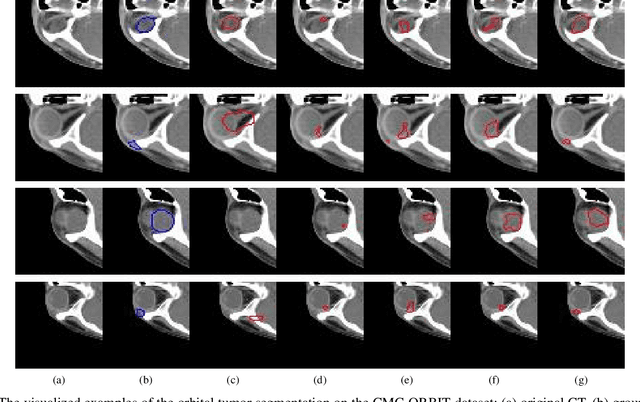
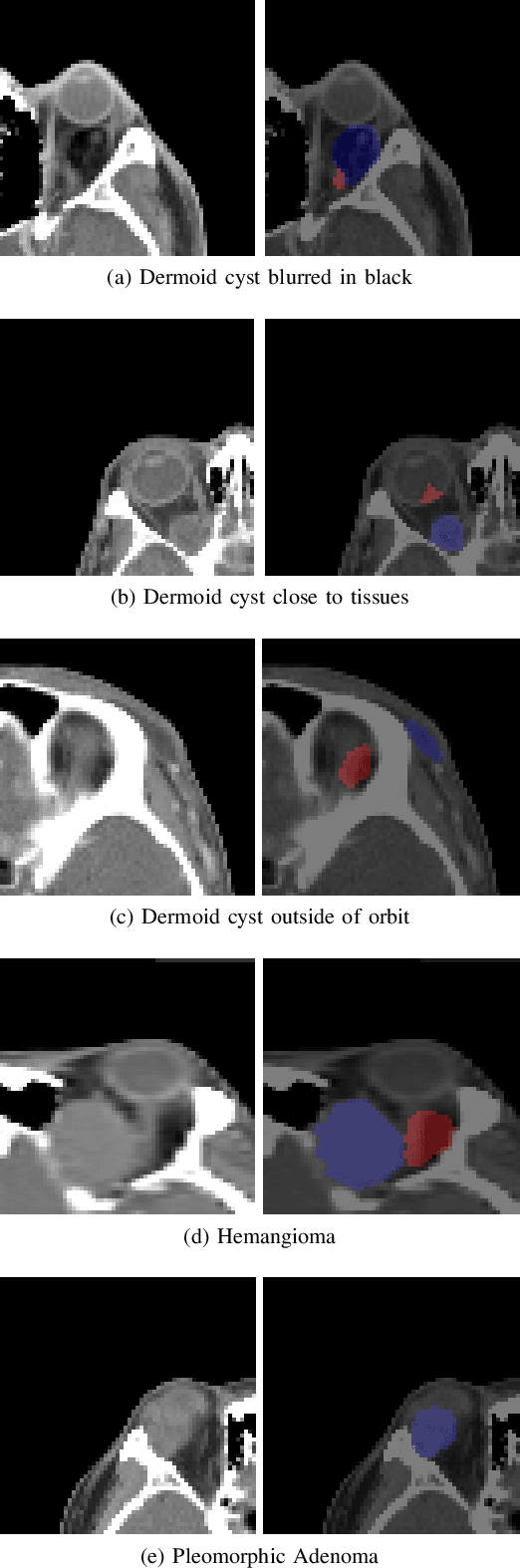
Deep neural networks (DNNs) have been widely used for medical image analysis. However, the lack of access a to large-scale annotated dataset poses a great challenge, especially in the case of rare diseases, or new domains for the research society. Transfer of pre-trained features, from the relatively large dataset is a considerable solution. In this paper, we have explored supervised segmentation using domain adaptation for optic nerve and orbital tumor, when only small sampled CT images are given. Even the lung image database consortium image collection (LIDC-IDRI) is a cross-domain to orbital CT, but the proposed domain adaptation method improved the performance of attention U-Net for the segmentation in public optic nerve dataset and our clinical orbital tumor dataset. The code and dataset are available at https://github.com/cmcbigdata.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge