"Image": models, code, and papers
DeeperLab: Single-Shot Image Parser
Feb 13, 2019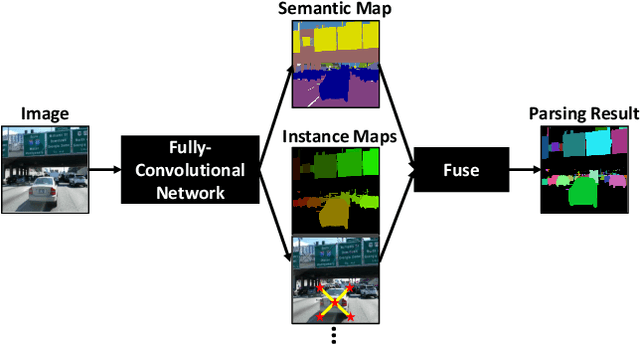
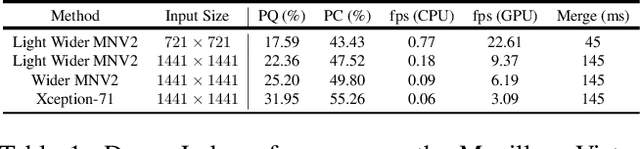
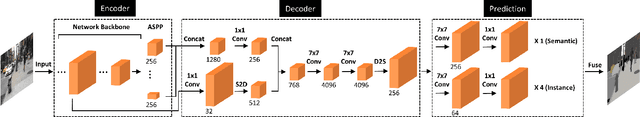
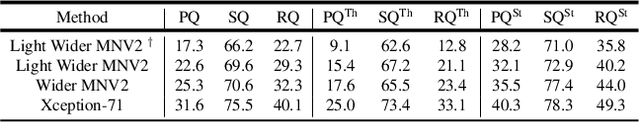
We present a single-shot, bottom-up approach for whole image parsing. Whole image parsing, also known as Panoptic Segmentation, generalizes the tasks of semantic segmentation for 'stuff' classes and instance segmentation for 'thing' classes, assigning both semantic and instance labels to every pixel in an image. Recent approaches to whole image parsing typically employ separate standalone modules for the constituent semantic and instance segmentation tasks and require multiple passes of inference. Instead, the proposed DeeperLab image parser performs whole image parsing with a significantly simpler, fully convolutional approach that jointly addresses the semantic and instance segmentation tasks in a single-shot manner, resulting in a streamlined system that better lends itself to fast processing. For quantitative evaluation, we use both the instance-based Panoptic Quality (PQ) metric and the proposed region-based Parsing Covering (PC) metric, which better captures the image parsing quality on 'stuff' classes and larger object instances. We report experimental results on the challenging Mapillary Vistas dataset, in which our single model achieves 31.95% (val) / 31.6% PQ (test) and 55.26% PC (val) with 3 frames per second (fps) on GPU or near real-time speed (22.6 fps on GPU) with reduced accuracy.
QDCNN: Quantum Dilated Convolutional Neural Network
Oct 29, 2021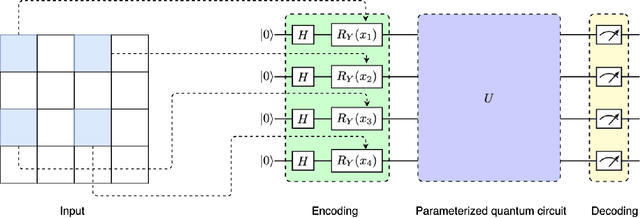
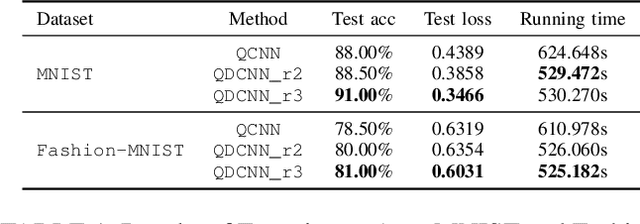
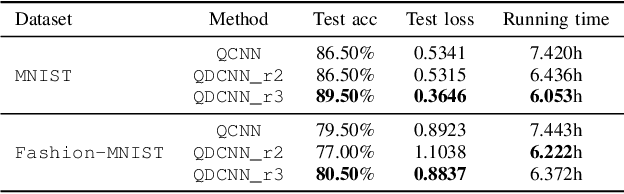
In recent years, with rapid progress in the development of quantum technologies, quantum machine learning has attracted a lot of interest. In particular, a family of hybrid quantum-classical neural networks, consisting of classical and quantum elements, has been massively explored for the purpose of improving the performance of classical neural networks. In this paper, we propose a novel hybrid quantum-classical algorithm called quantum dilated convolutional neural networks (QDCNNs). Our method extends the concept of dilated convolution, which has been widely applied in modern deep learning algorithms, to the context of hybrid neural networks. The proposed QDCNNs are able to capture larger context during the quantum convolution process while reducing the computational cost. We perform empirical experiments on MNIST and Fashion-MNIST datasets for the task of image recognition and demonstrate that QDCNN models generally enjoy better performances in terms of both accuracy and computation efficiency compared to existing quantum convolutional neural networks (QCNNs).
Growing an architecture for a neural network
Aug 04, 2021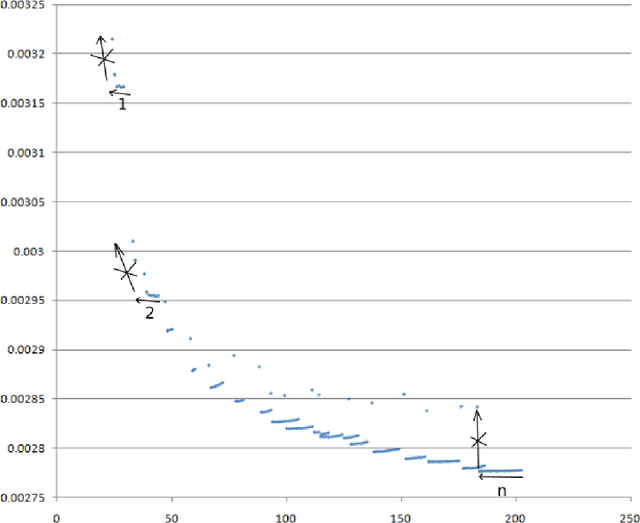
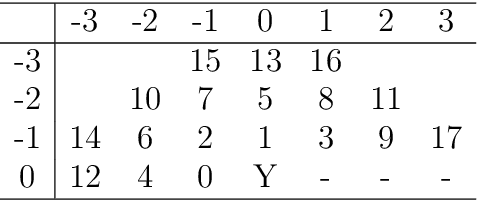
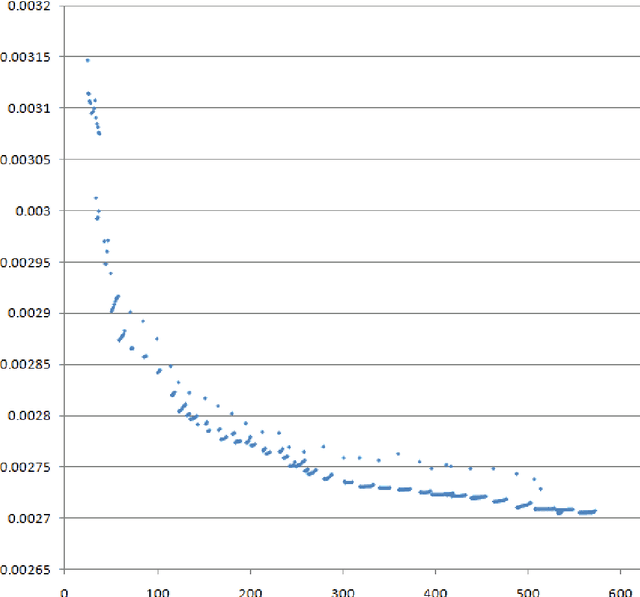

We propose a new kind of automatic architecture search algorithm. The algorithm alternates pruning connections and adding neurons, and it is not restricted to layered architectures only. Here architecture is an arbitrary oriented graph with some weights (along with some biases and an activation function), so there may be no layered structure in such a network. The algorithm minimizes the complexity of staying within a given error. We demonstrate our algorithm on the brightness prediction problem of the next point through the previous points on an image. Our second test problem is the approximation of the bivariate function defining the brightness of a black and white image. Our optimized networks significantly outperform the standard solution for neural network architectures in both cases.
Metric Learning for Image Registration
Apr 21, 2019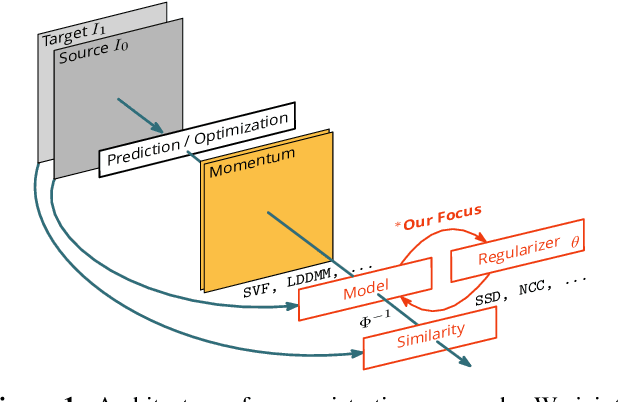
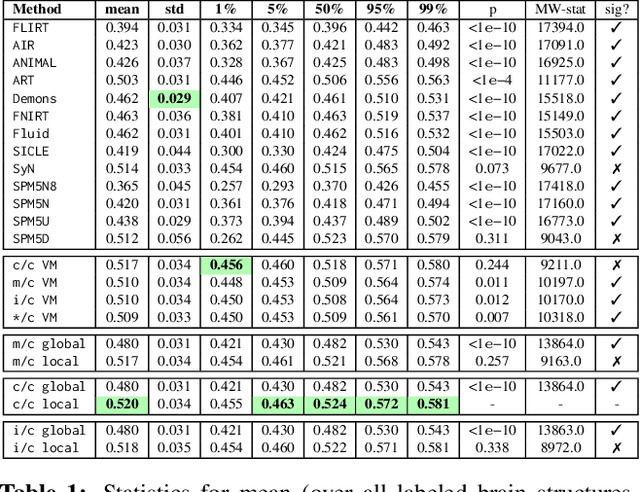
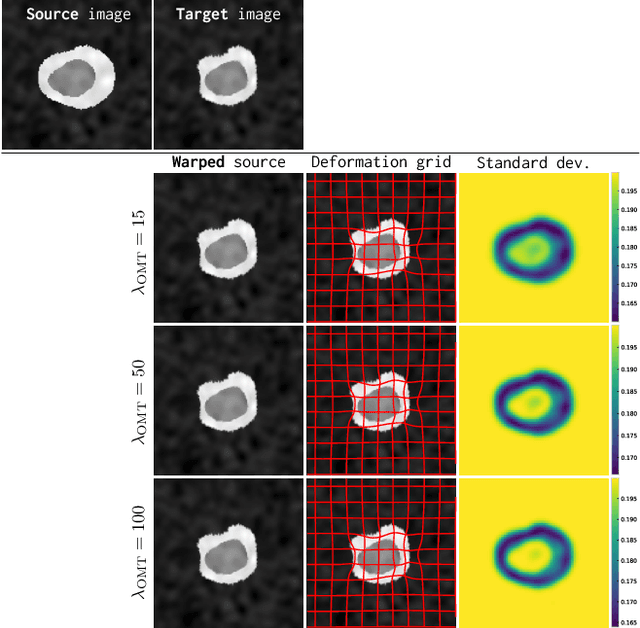
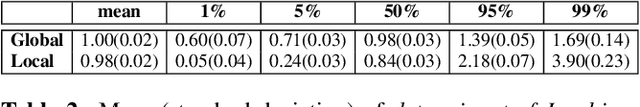
Image registration is a key technique in medical image analysis to estimate deformations between image pairs. A good deformation model is important for high-quality estimates. However, most existing approaches use ad-hoc deformation models chosen for mathematical convenience rather than to capture observed data variation. Recent deep learning approaches learn deformation models directly from data. However, they provide limited control over the spatial regularity of transformations. Instead of learning the entire registration approach, we learn a spatially-adaptive regularizer within a registration model. This allows controlling the desired level of regularity and preserving structural properties of a registration model. For example, diffeomorphic transformations can be attained. Our approach is a radical departure from existing deep learning approaches to image registration by embedding a deep learning model in an optimization-based registration algorithm to parameterize and data-adapt the registration model itself.
View Blind-spot as Inpainting: Self-Supervised Denoising with Mask Guided Residual Convolution
Sep 10, 2021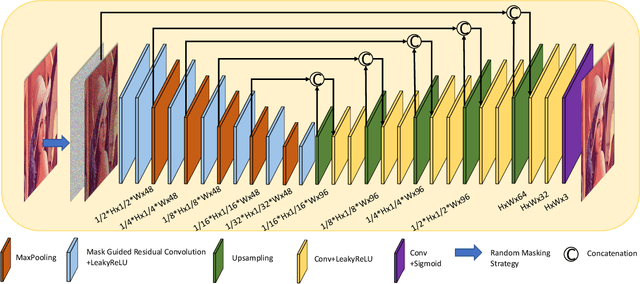
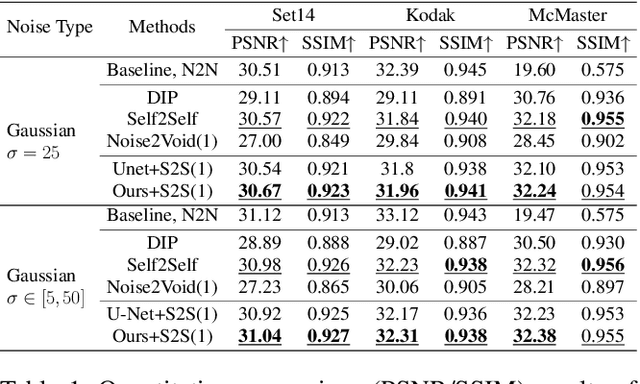
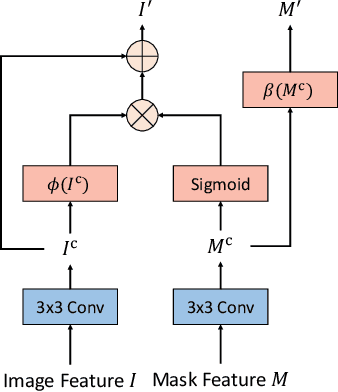
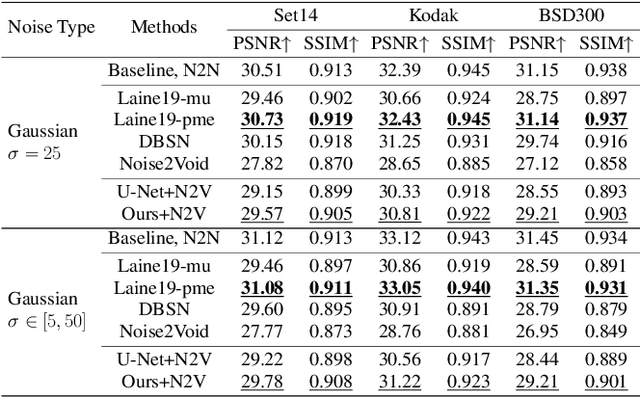
In recent years, self-supervised denoising methods have shown impressive performance, which circumvent painstaking collection procedure of noisy-clean image pairs in supervised denoising methods and boost denoising applicability in real world. One of well-known self-supervised denoising strategies is the blind-spot training scheme. However, a few works attempt to improve blind-spot based self-denoiser in the aspect of network architecture. In this paper, we take an intuitive view of blind-spot strategy and consider its process of using neighbor pixels to predict manipulated pixels as an inpainting process. Therefore, we propose a novel Mask Guided Residual Convolution (MGRConv) into common convolutional neural networks, e.g. U-Net, to promote blind-spot based denoising. Our MGRConv can be regarded as soft partial convolution and find a trade-off among partial convolution, learnable attention maps, and gated convolution. It enables dynamic mask learning with appropriate mask constrain. Different from partial convolution and gated convolution, it provides moderate freedom for network learning. It also avoids leveraging external learnable parameters for mask activation, unlike learnable attention maps. The experiments show that our proposed plug-and-play MGRConv can assist blind-spot based denoising network to reach promising results on both existing single-image based and dataset-based methods.
Billion-Scale Pretraining with Vision Transformers for Multi-Task Visual Representations
Aug 12, 2021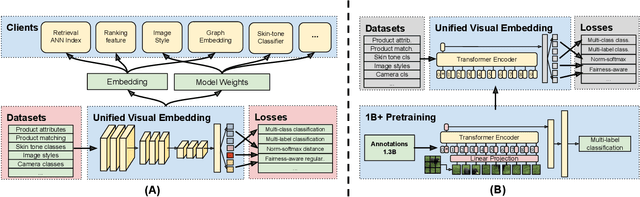
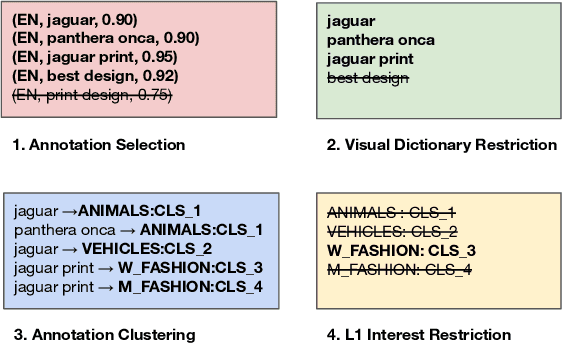
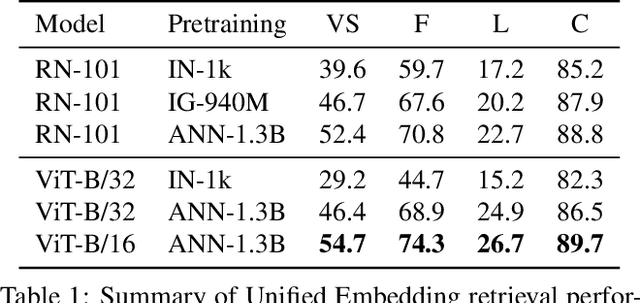

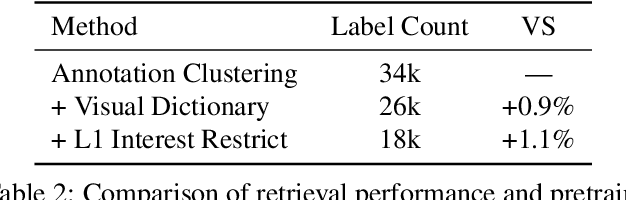
Large-scale pretraining of visual representations has led to state-of-the-art performance on a range of benchmark computer vision tasks, yet the benefits of these techniques at extreme scale in complex production systems has been relatively unexplored. We consider the case of a popular visual discovery product, where these representations are trained with multi-task learning, from use-case specific visual understanding (e.g. skin tone classification) to general representation learning for all visual content (e.g. embeddings for retrieval). In this work, we describe how we (1) generate a dataset with over a billion images via large weakly-supervised pretraining to improve the performance of these visual representations, and (2) leverage Transformers to replace the traditional convolutional backbone, with insights into both system and performance improvements, especially at 1B+ image scale. To support this backbone model, we detail a systematic approach to deriving weakly-supervised image annotations from heterogenous text signals, demonstrating the benefits of clustering techniques to handle the long-tail distribution of image labels. Through a comprehensive study of offline and online evaluation, we show that large-scale Transformer-based pretraining provides significant benefits to industry computer vision applications. The model is deployed in a production visual shopping system, with 36% improvement in top-1 relevance and 23% improvement in click-through volume. We conduct extensive experiments to better understand the empirical relationships between Transformer-based architectures, dataset scale, and the performance of production vision systems.
Combining chest X-rays and EHR data using machine learning to diagnose acute respiratory failure
Aug 27, 2021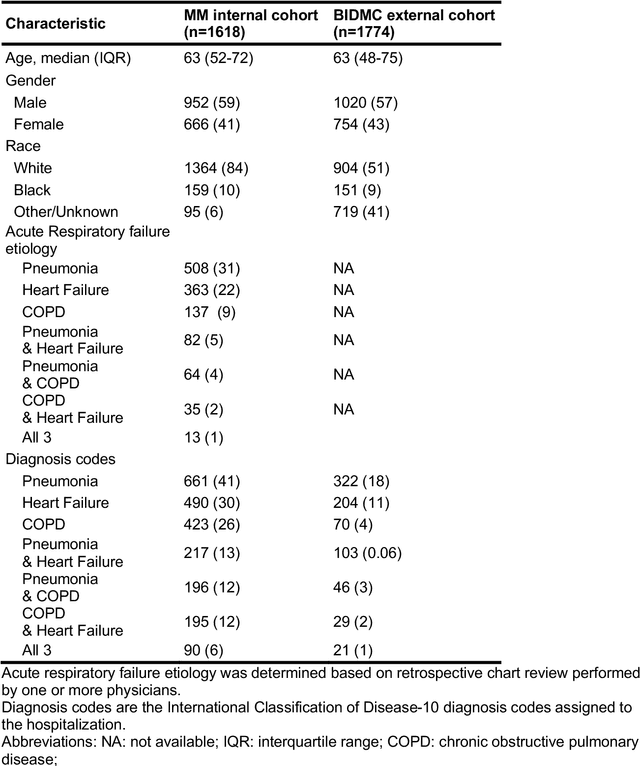
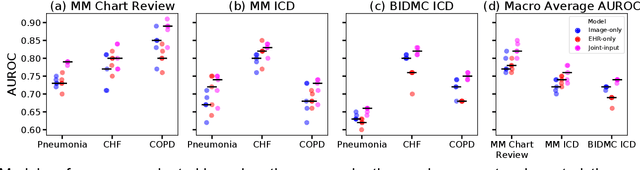
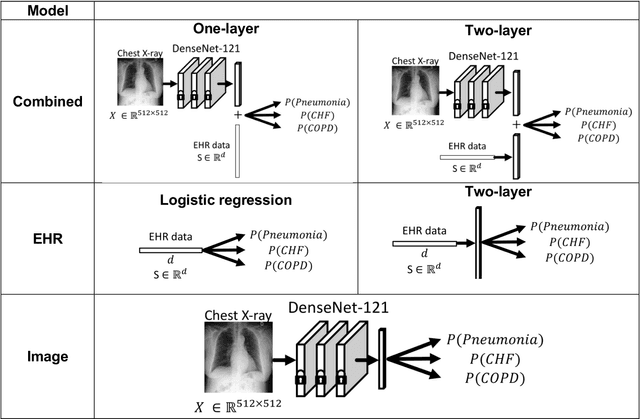
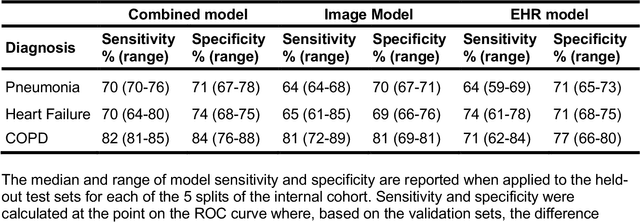
When patients develop acute respiratory failure, accurately identifying the underlying etiology is essential for determining the best treatment, but it can be challenging to differentiate between common diagnoses in clinical practice. Machine learning models could improve medical diagnosis by augmenting clinical decision making and play a role in the diagnostic evaluation of patients with acute respiratory failure. While machine learning models have been developed to identify common findings on chest radiographs (e.g. pneumonia), augmenting these approaches by also analyzing clinically relevant data from the electronic health record (EHR) could aid in the diagnosis of acute respiratory failure. Machine learning models were trained to predict the cause of acute respiratory failure (pneumonia, heart failure, and/or COPD) using chest radiographs and EHR data from patients within an internal cohort using diagnoses based on physician chart review. Models were also tested on patients in an external cohort using discharge diagnosis codes. A model combining chest radiographs and EHR data outperformed models based on each modality alone for pneumonia and COPD. For pneumonia, the combined model AUROC was 0.79 (0.78-0.79), image model AUROC was 0.73 (0.72-0.75), and EHR model AUROC was 0.73 (0.70-0.76); for COPD, combined: 0.89 (0.83-0.91), image: 0.85 (0.77-0.89), and EHR: 0.80 (0.76-0.84); for heart failure, combined: 0.80 (0.77-0.84), image: 0.77 (0.71-0.81), and EHR: 0.80 (0.75-0.82). In the external cohort, performance was consistent for heart failure and COPD, but declined slightly for pneumonia. Overall, machine learning models combing chest radiographs and EHR data can accurately differentiate between common causes of acute respiratory failure. Further work is needed to determine whether these models could aid clinicians in the diagnosis of acute respiratory failure in clinical settings.
WarpedGANSpace: Finding non-linear RBF paths in GAN latent space
Sep 27, 2021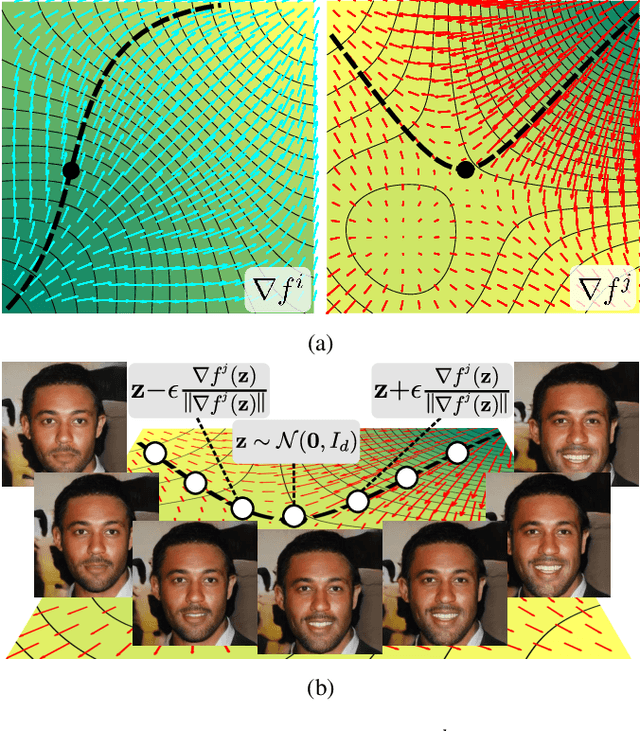
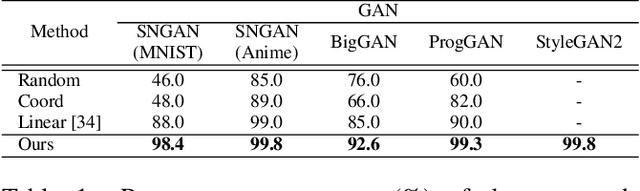
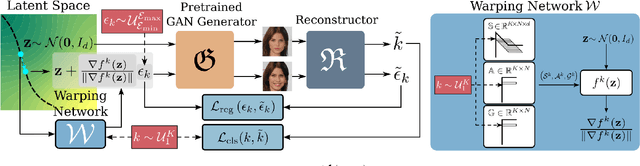
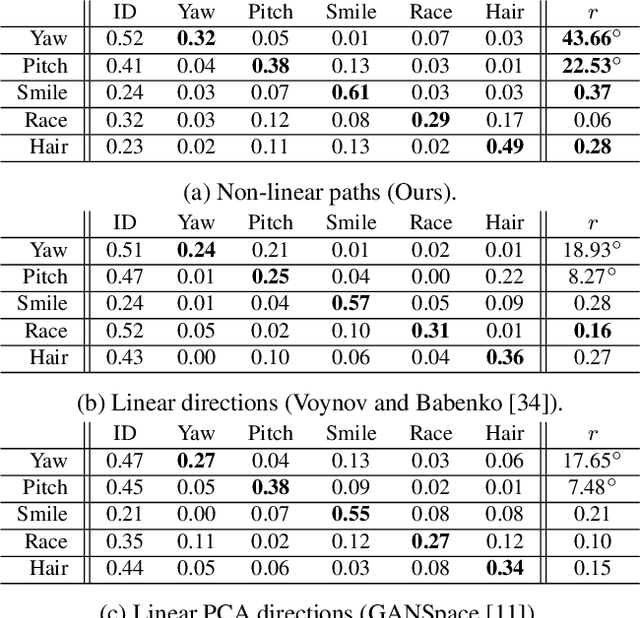
This work addresses the problem of discovering, in an unsupervised manner, interpretable paths in the latent space of pretrained GANs, so as to provide an intuitive and easy way of controlling the underlying generative factors. In doing so, it addresses some of the limitations of the state-of-the-art works, namely, a) that they discover directions that are independent of the latent code, i.e., paths that are linear, and b) that their evaluation relies either on visual inspection or on laborious human labeling. More specifically, we propose to learn non-linear warpings on the latent space, each one parametrized by a set of RBF-based latent space warping functions, and where each warping gives rise to a family of non-linear paths via the gradient of the function. Building on the work of Voynov and Babenko, that discovers linear paths, we optimize the trainable parameters of the set of RBFs, so as that images that are generated by codes along different paths, are easily distinguishable by a discriminator network. This leads to easily distinguishable image transformations, such as pose and facial expressions in facial images. We show that linear paths can be derived as a special case of our method, and show experimentally that non-linear paths in the latent space lead to steeper, more disentangled and interpretable changes in the image space than in state-of-the art methods, both qualitatively and quantitatively. We make the code and the pretrained models publicly available at: https://github.com/chi0tzp/WarpedGANSpace.
Deep Poisoning Functions: Towards Robust Privacy-safe Image Data Sharing
Dec 14, 2019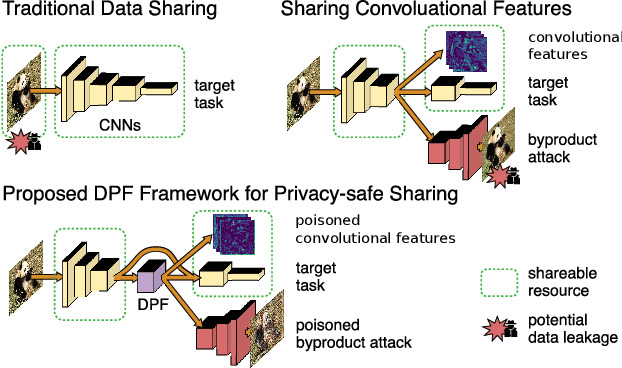
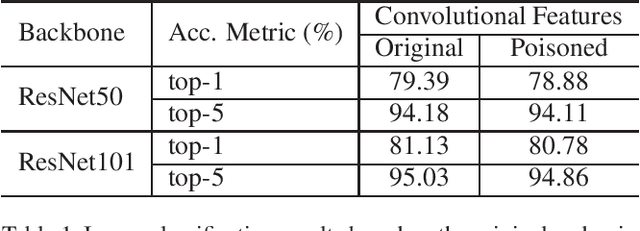
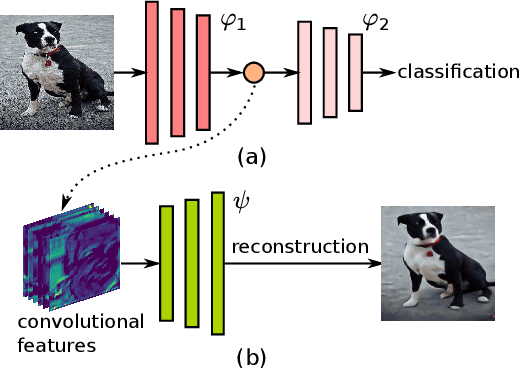
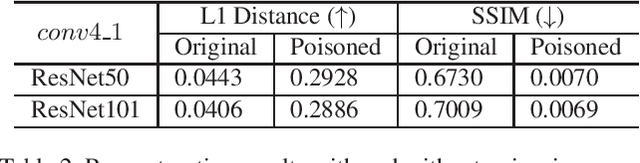
As deep networks are applied to an ever-expanding set of computer vision tasks, protecting general privacy in image data has become a critically important goal. This paper presents a new framework for privacy-preserving data sharing that is robust to adversarial attacks and overcomes the known issues existing in previous approaches. We introduce the concept of a Deep Poisoning Function (DPF), which is a module inserted into a pre-trained deep network designed to perform a specific vision task. The DPF is optimized to deliberately poison image data to prevent known adversarial attacks, while ensuring that the altered image data is functionally equivalent to the non-poisoned data for the original task. Given this equivalence, both poisoned and non-poisoned data can be used for further retraining or fine-tuning. Experimental results on image classification and face recognition tasks prove the efficacy of the proposed method.
BIAS: Transparent reporting of biomedical image analysis challenges
Oct 23, 2019The number of biomedical image analysis challenges organized per year is steadily increasing. These international competitions have the purpose of benchmarking algorithms on common data sets, typically to identify the best method for a given problem. Recent research, however, revealed that common practice related to challenge reporting does not allow for adequate interpretation and reproducibility of results. To address the discrepancy between the impact of challenges and the quality (control), the Biomedical I mage Analysis ChallengeS (BIAS) initiative developed a set of recommendations for the reporting of challenges. The BIAS statement aims to improve the transparency of the reporting of a biomedical image analysis challenge regardless of field of application, image modality or task category assessed. This article describes how the BIAS statement was developed and presents a checklist which authors of biomedical image analysis challenges are encouraged to include in their submission when giving a paper on a challenge into review. The purpose of the checklist is to standardize and facilitate the review process and raise interpretability and reproducibility of challenge results by making relevant information explicit.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge