"Image": models, code, and papers
Image Harmonization Datasets: HCOCO, HAdobe5k, HFlickr, and Hday2night
Aug 28, 2019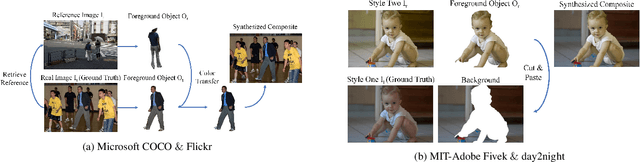
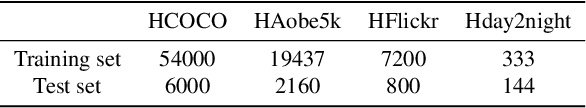
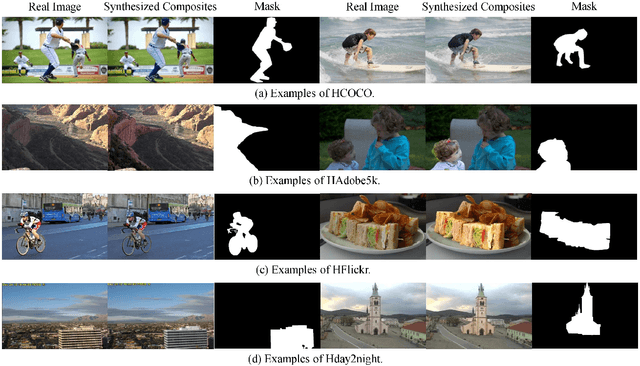
Image composition is an important operation in image processing, but the inconsistency between foreground and background significantly degrades the quality of composite image. Image harmonization, which aims to make the foreground compatible with the background, is a promising yet challenging task. However, the lack of high-quality public dataset for image harmonization, which significantly hinders the development of image harmonization techniques. Therefore, we create synthesized composite images based on existing COCO (resp., Adobe5k, day2night) dataset, leading to our HCOCO (resp., HAdobe5k, Hday2night) dataset. To enrich the diversity of our datasets, we also generate synthesized composite images based on our collected Flick images, leading to our HFlickr dataset. All four datasets are released in https://github.com/bcmi/Image_Harmonization_Datasets.
Unrolling of Deep Graph Total Variation for Image Denoising
Oct 21, 2020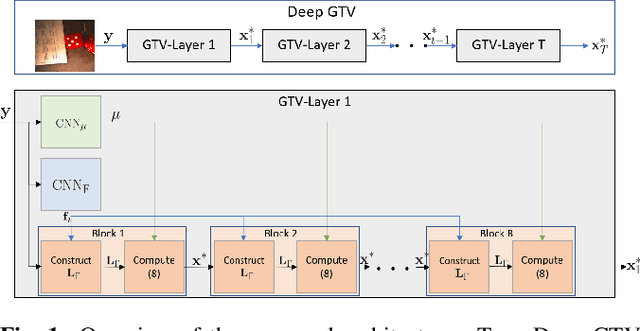
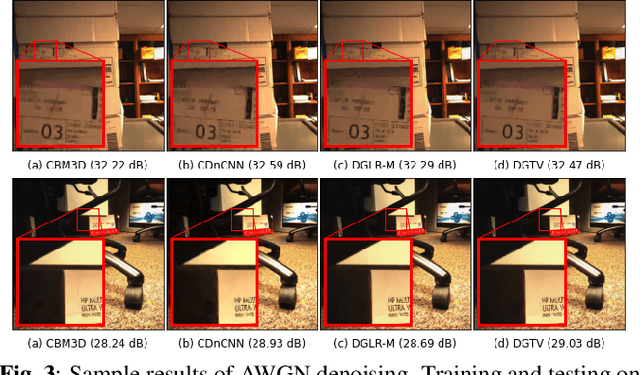
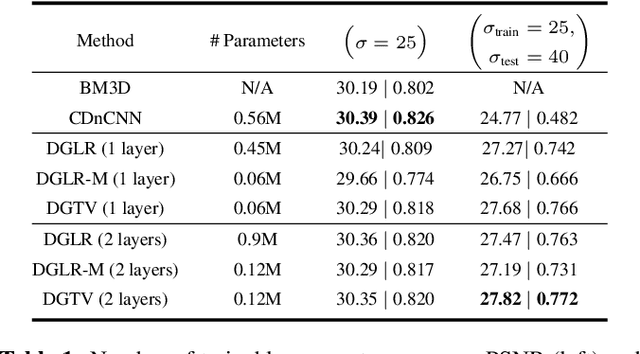
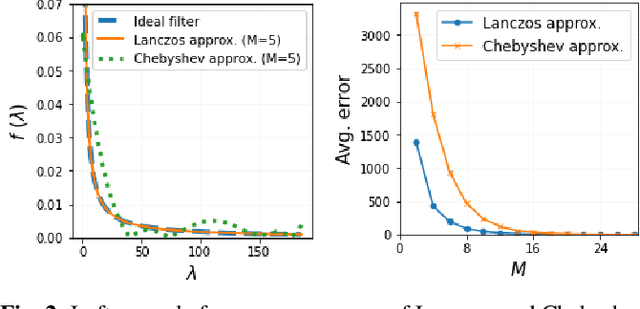

While deep learning (DL) architectures like convolutional neural networks (CNNs) have enabled effective solutions in image denoising, in general their implementations overly rely on training data, lack interpretability, and require tuning of a large parameter set. In this paper, we combine classical graph signal filtering with deep feature learning into a competitive hybrid design---one that utilizes interpretable analytical low-pass graph filters and employs 80% fewer network parameters than state-of-the-art DL denoising scheme DnCNN. Specifically, to construct a suitable similarity graph for graph spectral filtering, we first adopt a CNN to learn feature representations per pixel, and then compute feature distances to establish edge weights. Given a constructed graph, we next formulate a convex optimization problem for denoising using a graph total variation (GTV) prior. Via a $l_1$ graph Laplacian reformulation, we interpret its solution in an iterative procedure as a graph low-pass filter and derive its frequency response. For fast filter implementation, we realize this response using a Lanczos approximation. Experimental results show that in the case of statistical mistmatch, our algorithm outperformed DnCNN by up to 3dB in PSNR.
RadFusion: Benchmarking Performance and Fairness for Multimodal Pulmonary Embolism Detection from CT and EHR
Nov 27, 2021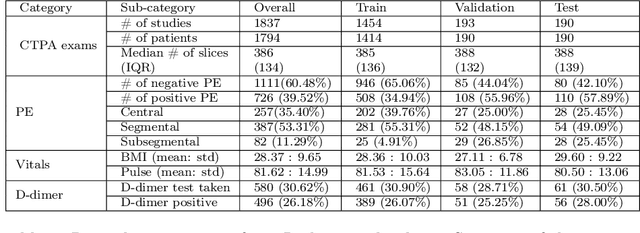
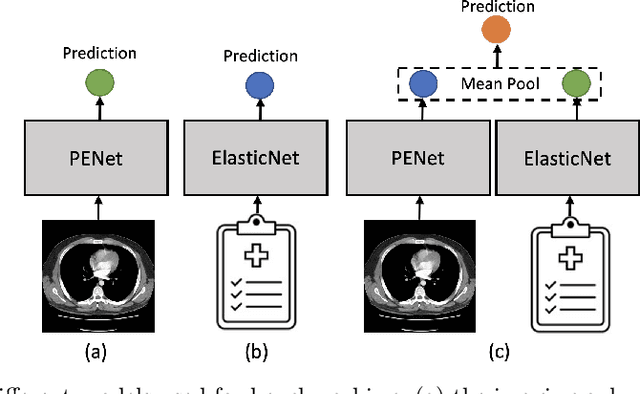
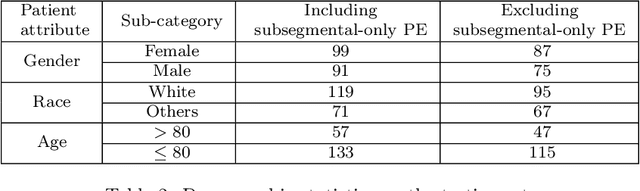
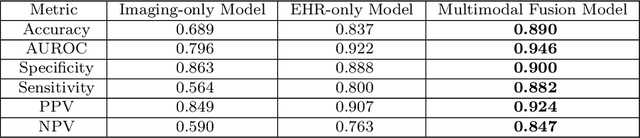
Despite the routine use of electronic health record (EHR) data by radiologists to contextualize clinical history and inform image interpretation, the majority of deep learning architectures for medical imaging are unimodal, i.e., they only learn features from pixel-level information. Recent research revealing how race can be recovered from pixel data alone highlights the potential for serious biases in models which fail to account for demographics and other key patient attributes. Yet the lack of imaging datasets which capture clinical context, inclusive of demographics and longitudinal medical history, has left multimodal medical imaging underexplored. To better assess these challenges, we present RadFusion, a multimodal, benchmark dataset of 1794 patients with corresponding EHR data and high-resolution computed tomography (CT) scans labeled for pulmonary embolism. We evaluate several representative multimodal fusion models and benchmark their fairness properties across protected subgroups, e.g., gender, race/ethnicity, age. Our results suggest that integrating imaging and EHR data can improve classification performance and robustness without introducing large disparities in the true positive rate between population groups.
An Auto-Encoder Strategy for Adaptive Image Segmentation
Apr 29, 2020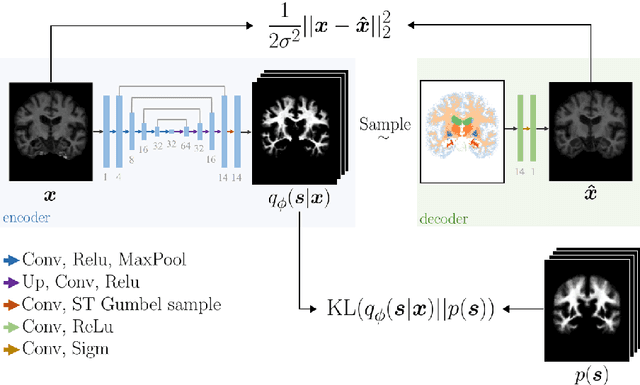
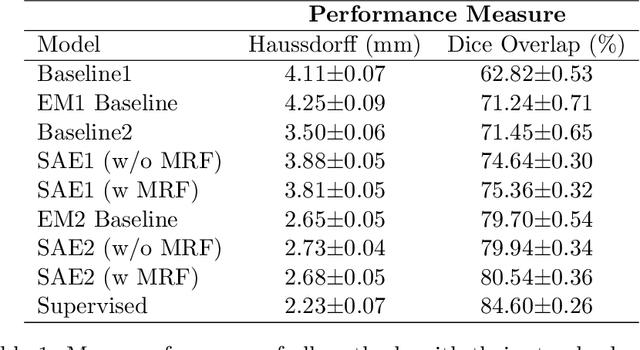
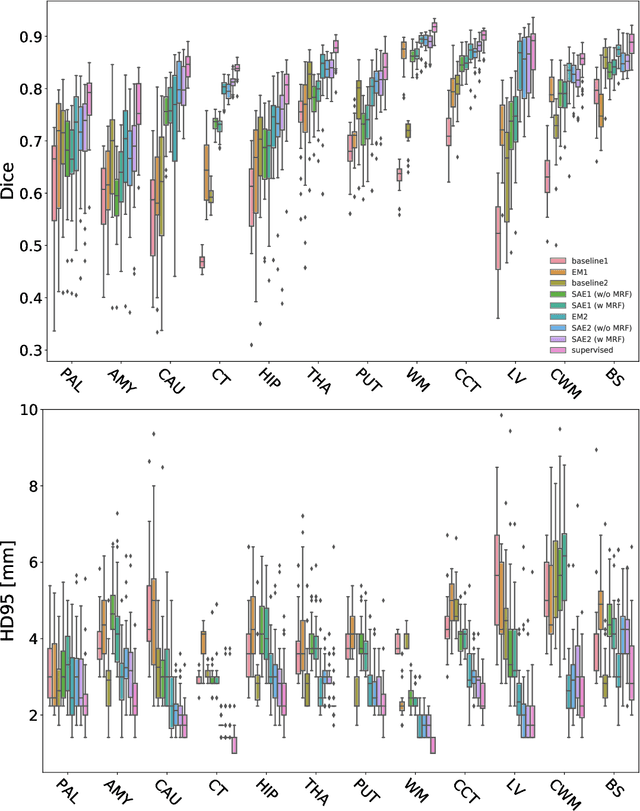
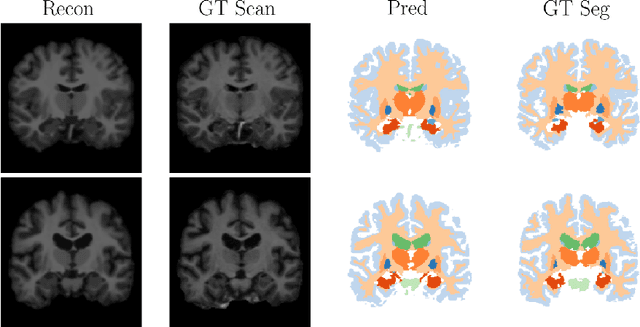
Deep neural networks are powerful tools for biomedical image segmentation. These models are often trained with heavy supervision, relying on pairs of images and corresponding voxel-level labels. However, obtaining segmentations of anatomical regions on a large number of cases can be prohibitively expensive. Thus there is a strong need for deep learning-based segmentation tools that do not require heavy supervision and can continuously adapt. In this paper, we propose a novel perspective of segmentation as a discrete representation learning problem, and present a variational autoencoder segmentation strategy that is flexible and adaptive. Our method, called Segmentation Auto-Encoder (SAE), leverages all available unlabeled scans and merely requires a segmentation prior, which can be a single unpaired segmentation image. In experiments, we apply SAE to brain MRI scans. Our results show that SAE can produce good quality segmentations, particularly when the prior is good. We demonstrate that a Markov Random Field prior can yield significantly better results than a spatially independent prior. Our code is freely available at https://github.com/evanmy/sae.
Semi-supervised Domain Adaptation for Semantic Segmentation
Oct 20, 2021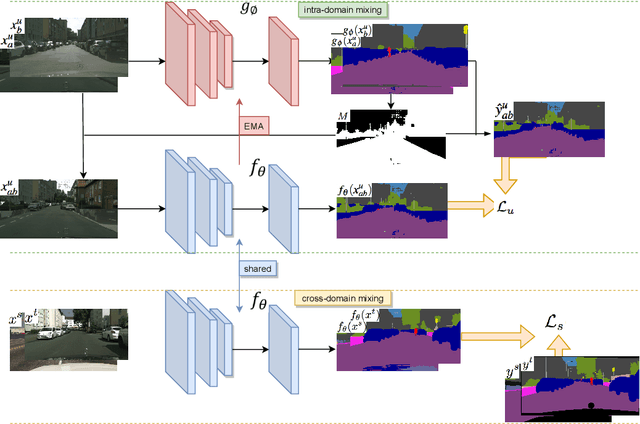
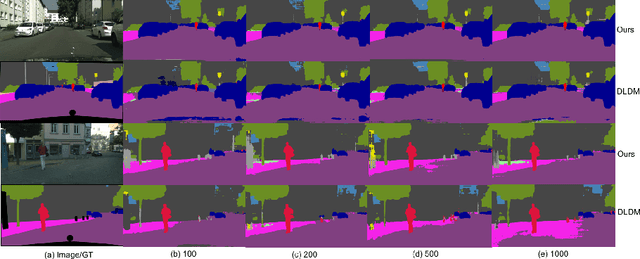
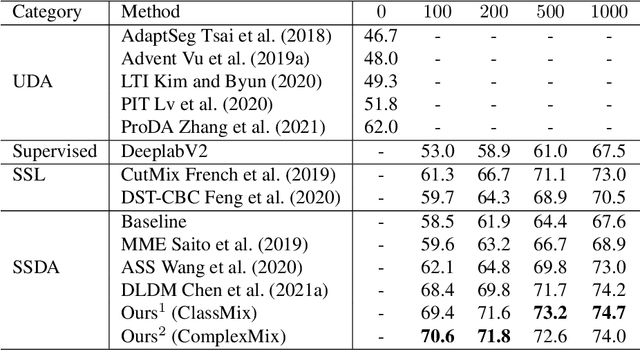
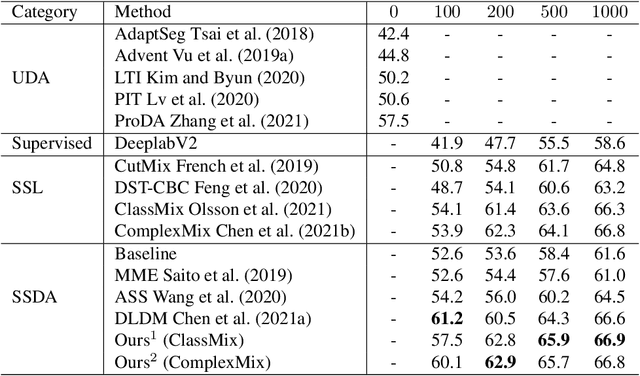

Deep learning approaches for semantic segmentation rely primarily on supervised learning approaches and require substantial efforts in producing pixel-level annotations. Further, such approaches may perform poorly when applied to unseen image domains. To cope with these limitations, both unsupervised domain adaptation (UDA) with full source supervision but without target supervision and semi-supervised learning (SSL) with partial supervision have been proposed. While such methods are effective at aligning different feature distributions, there is still a need to efficiently exploit unlabeled data to address the performance gap with respect to fully-supervised methods. In this paper we address semi-supervised domain adaptation (SSDA) for semantic segmentation, where a large amount of labeled source data as well as a small amount of labeled target data are available. We propose a novel and effective two-step semi-supervised dual-domain adaptation (SSDDA) approach to address both cross- and intra-domain gaps in semantic segmentation. The proposed framework is comprised of two mixing modules. First, we conduct a cross-domain adaptation via an image-level mixing strategy, which learns to align the distribution shift of features between the source data and target data. Second, intra-domain adaptation is achieved using a separate student-teacher network which is built to generate category-level data augmentation by mixing unlabeled target data in a way that respects predicted object boundaries. We demonstrate that the proposed approach outperforms state-of-the-art methods on two common synthetic-to-real semantic segmentation benchmarks. An extensive ablation study is provided to further validate the effectiveness of our approach.
Color Image Segmentation using Adaptive Particle Swarm Optimization and Fuzzy C-means
Apr 18, 2020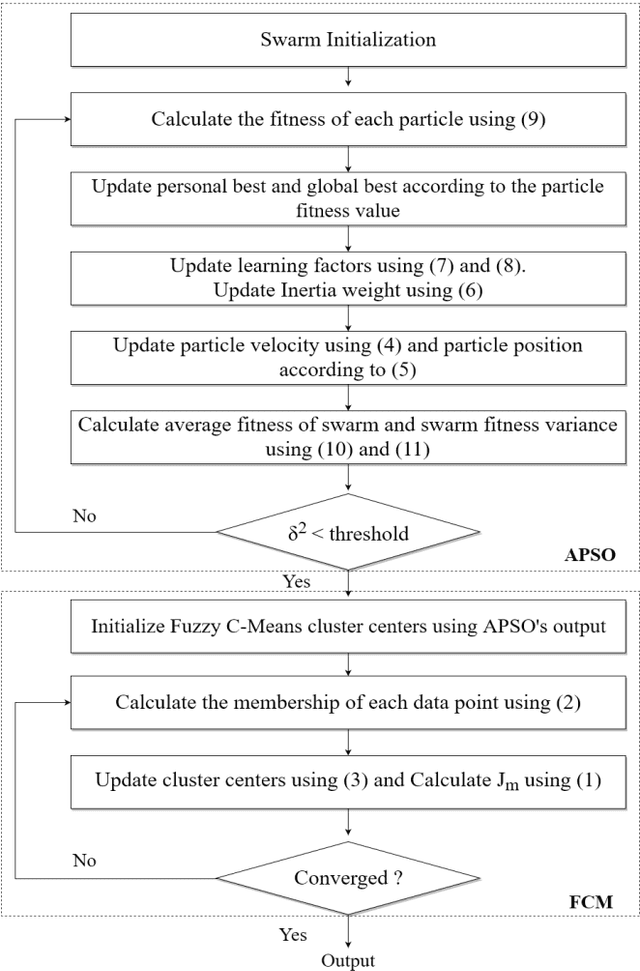
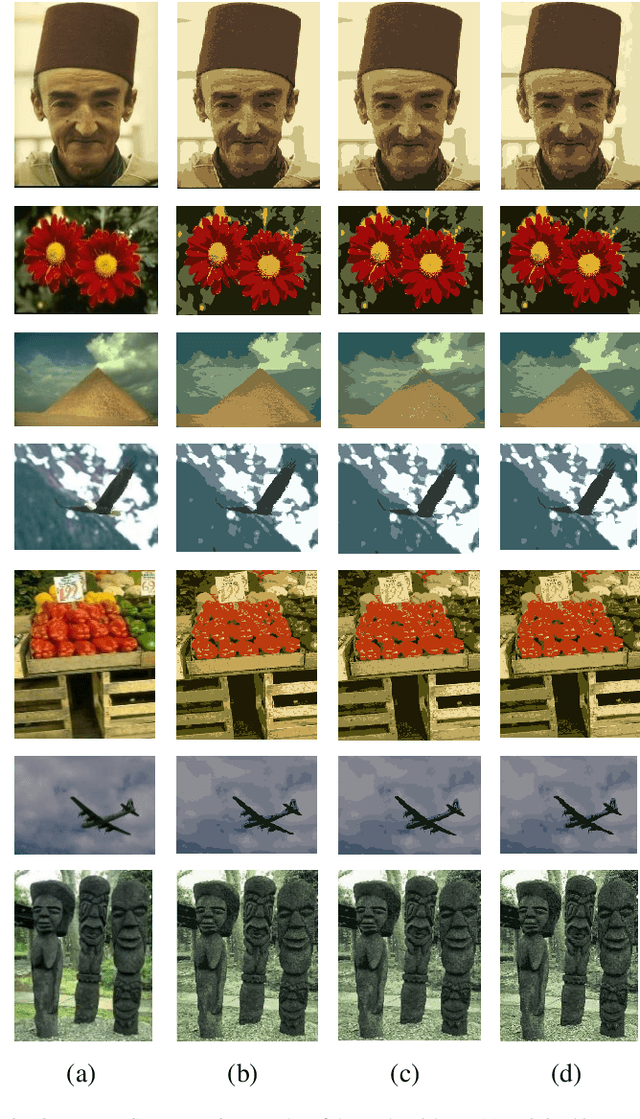

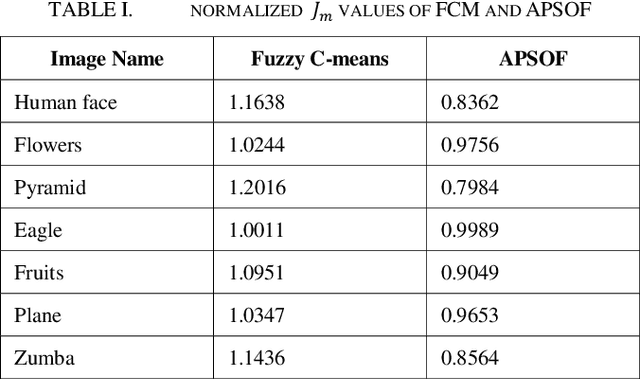
Segmentation partitions an image into different regions containing pixels with similar attributes. A standard non-contextual variant of Fuzzy C-means clustering algorithm (FCM), considering its simplicity is generally used in image segmentation. Using FCM has its disadvantages like it is dependent on the initial guess of the number of clusters and highly sensitive to noise. Satisfactory visual segments cannot be obtained using FCM. Particle Swarm Optimization (PSO) belongs to the class of evolutionary algorithms and has good convergence speed and fewer parameters compared to Genetic Algorithms (GAs). An optimized version of PSO can be combined with FCM to act as a proper initializer for the algorithm thereby reducing its sensitivity to initial guess. A hybrid PSO algorithm named Adaptive Particle Swarm Optimization (APSO) which improves in the calculation of various hyper parameters like inertia weight, learning factors over standard PSO, using insights from swarm behaviour, leading to improvement in cluster quality can be used. This paper presents a new image segmentation algorithm called Adaptive Particle Swarm Optimization and Fuzzy C-means Clustering Algorithm (APSOF), which is based on Adaptive Particle Swarm Optimization (APSO) and Fuzzy C-means clustering. Experimental results show that APSOF algorithm has edge over FCM in correctly identifying the optimum cluster centers, there by leading to accurate classification of the image pixels. Hence, APSOF algorithm has superior performance in comparison with classic Particle Swarm Optimization (PSO) and Fuzzy C-means clustering algorithm (FCM) for image segmentation.
A color temperature-based high-speed decolorization: an empirical approach for tone mapping applications
Aug 31, 2021Grayscale images are fundamental to many image processing applications like data compression, feature extraction, printing and tone mapping. However, some image information is lost when converting from color to grayscale. In this paper, we propose a light-weight and high-speed image decolorization method based on human perception of color temperatures. Chromatic aberration results from differential refraction of light depending on its wavelength. It causes some rays corresponding to cooler colors (like blue, green) to converge before the warmer colors (like red, orange). This phenomena creates a perception of warm colors "advancing" toward the eye, while the cool colors to be "receding" away. In this proposed color to gray conversion model, we implement a weighted blending function to combine red (perceived warm) and blue (perceived cool) channel. Our main contribution is threefold: First, we implement a high-speed color processing method using exact pixel by pixel processing, and we report a $5.7\times$ speed up when compared to other new algorithms. Second, our optimal color conversion method produces luminance in images that are comparable to other state of the art methods which we quantified using the objective metrics (E-score and C2G-SSIM) and a subjective user study. Third, we demonstrate that an effective luminance distribution can be achieved using our algorithm by using global and local tone mapping applications.
Discrete Representations Strengthen Vision Transformer Robustness
Nov 20, 2021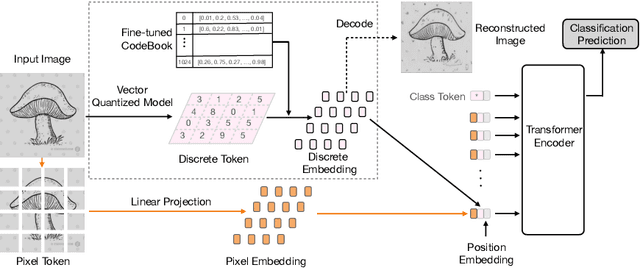
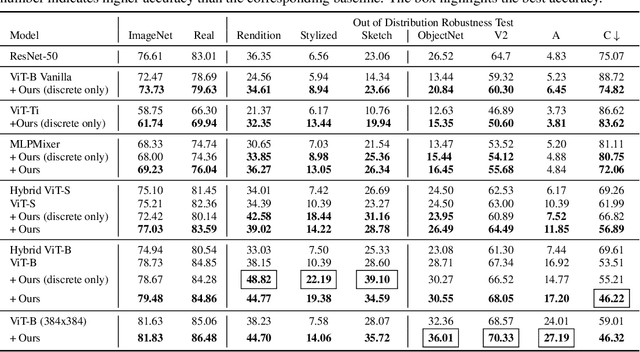

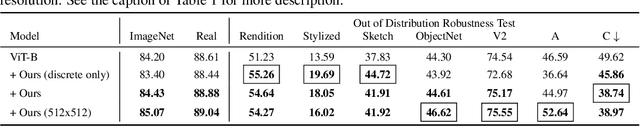
Vision Transformer (ViT) is emerging as the state-of-the-art architecture for image recognition. While recent studies suggest that ViTs are more robust than their convolutional counterparts, our experiments find that ViTs are overly reliant on local features (e.g., nuisances and texture) and fail to make adequate use of global context (e.g., shape and structure). As a result, ViTs fail to generalize to out-of-distribution, real-world data. To address this deficiency, we present a simple and effective architecture modification to ViT's input layer by adding discrete tokens produced by a vector-quantized encoder. Different from the standard continuous pixel tokens, discrete tokens are invariant under small perturbations and contain less information individually, which promote ViTs to learn global information that is invariant. Experimental results demonstrate that adding discrete representation on four architecture variants strengthens ViT robustness by up to 12% across seven ImageNet robustness benchmarks while maintaining the performance on ImageNet.
SMDS-Net: Model Guided Spectral-Spatial Network for Hyperspectral Image Denoising
Dec 03, 2020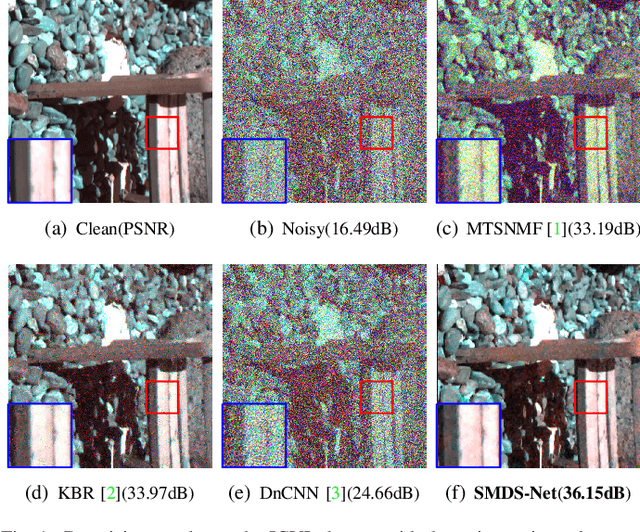
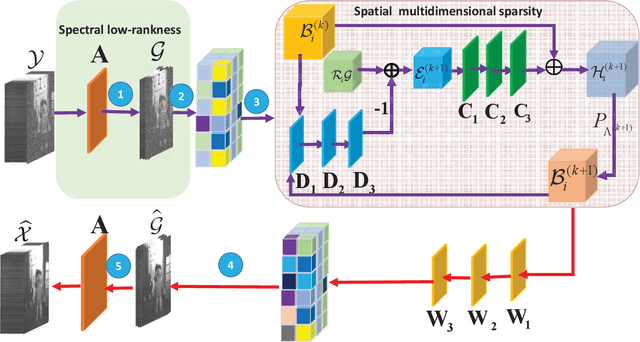


Deep learning (DL) based hyperspectral images (HSIs) denoising approaches directly learn the nonlinear mapping between observed noisy images and underlying clean images. They normally do not consider the physical characteristics of HSIs, therefore making them lack of interpretability that is key to understand their denoising mechanism.. In order to tackle this problem, we introduce a novel model guided interpretable network for HSI denoising. Specifically, fully considering the spatial redundancy, spectral low-rankness and spectral-spatial properties of HSIs, we first establish a subspace based multi-dimensional sparse model. This model first projects the observed HSIs into a low-dimensional orthogonal subspace, and then represents the projected image with a multidimensional dictionary. After that, the model is unfolded into an end-to-end network named SMDS-Net whose fundamental modules are seamlessly connected with the denoising procedure and optimization of the model. This makes SMDS-Net convey clear physical meanings, i.e., learning the low-rankness and sparsity of HSIs. Finally, all key variables including dictionaries and thresholding parameters are obtained by the end-to-end training. Extensive experiments and comprehensive analysis confirm the denoising ability and interpretability of our method against the state-of-the-art HSI denoising methods.
Predição da Idade Cerebral a partir de Imagens de Ressonância Magnética utilizando Redes Neurais Convolucionais
Dec 23, 2021In this work, deep learning techniques for brain age prediction from magnetic resonance images are investigated, aiming to assist in the identification of biomarkers of the natural aging process. The identification of biomarkers is useful for detecting an early-stage neurodegenerative process, as well as for predicting age-related or non-age-related cognitive decline. Two techniques are implemented and compared in this work: a 3D Convolutional Neural Network applied to the volumetric image and a 2D Convolutional Neural Network applied to slices from the axial plane, with subsequent fusion of individual predictions. The best result was obtained by the 2D model, which achieved a mean absolute error of 3.83 years. -- Neste trabalho s\~ao investigadas t\'ecnicas de aprendizado profundo para a predi\c{c}\~ao da idade cerebral a partir de imagens de resson\^ancia magn\'etica, visando auxiliar na identifica\c{c}\~ao de biomarcadores do processo natural de envelhecimento. A identifica\c{c}\~ao de biomarcadores \'e \'util para a detec\c{c}\~ao de um processo neurodegenerativo em est\'agio inicial, al\'em de possibilitar prever um decl\'inio cognitivo relacionado ou n\~ao \`a idade. Duas t\'ecnicas s\~ao implementadas e comparadas neste trabalho: uma Rede Neural Convolucional 3D aplicada na imagem volum\'etrica e uma Rede Neural Convolucional 2D aplicada a fatias do plano axial, com posterior fus\~ao das predi\c{c}\~oes individuais. O melhor resultado foi obtido pelo modelo 2D, que alcan\c{c}ou um erro m\'edio absoluto de 3.83 anos.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge