"Image": models, code, and papers
Learning Fixed Points in Generative Adversarial Networks: From Image-to-Image Translation to Disease Detection and Localization
Aug 29, 2019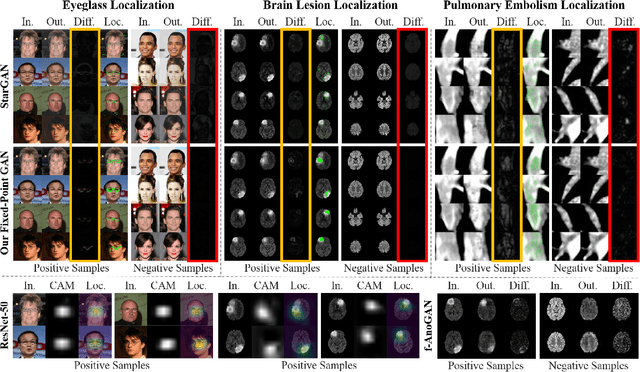
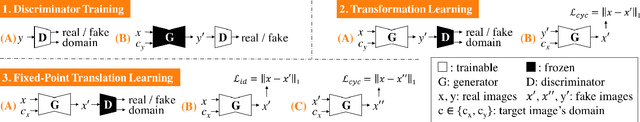
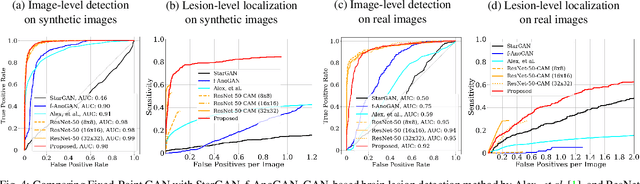
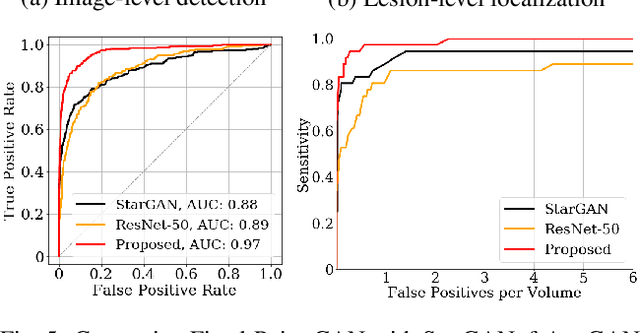
Generative adversarial networks (GANs) have ushered in a revolution in image-to-image translation. The development and proliferation of GANs raises an interesting question: can we train a GAN to remove an object, if present, from an image while otherwise preserving the image? Specifically, can a GAN "virtually heal" anyone by turning his medical image, with an unknown health status (diseased or healthy), into a healthy one, so that diseased regions could be revealed by subtracting those two images? Such a task requires a GAN to identify a minimal subset of target pixels for domain translation, an ability that we call fixed-point translation, which no GAN is equipped with yet. Therefore, we propose a new GAN, called Fixed-Point GAN, trained by (1) supervising same-domain translation through a conditional identity loss, and (2) regularizing cross-domain translation through revised adversarial, domain classification, and cycle consistency loss. Based on fixed-point translation, we further derive a novel framework for disease detection and localization using only image-level annotation. Qualitative and quantitative evaluations demonstrate that the proposed method outperforms the state of the art in multi-domain image-to-image translation and that it surpasses predominant weakly-supervised localization methods in both disease detection and localization. Implementation is available at https://github.com/jlianglab/Fixed-Point-GAN.
Learning to Manipulate Individual Objects in an Image
Apr 11, 2020
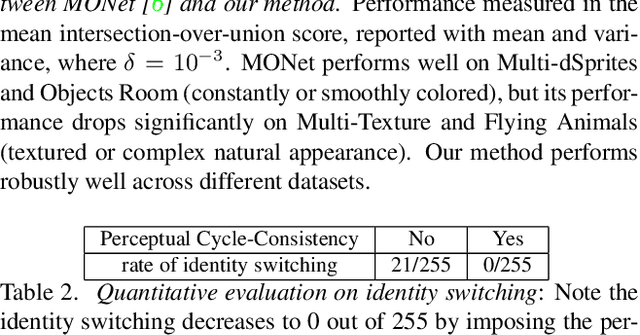

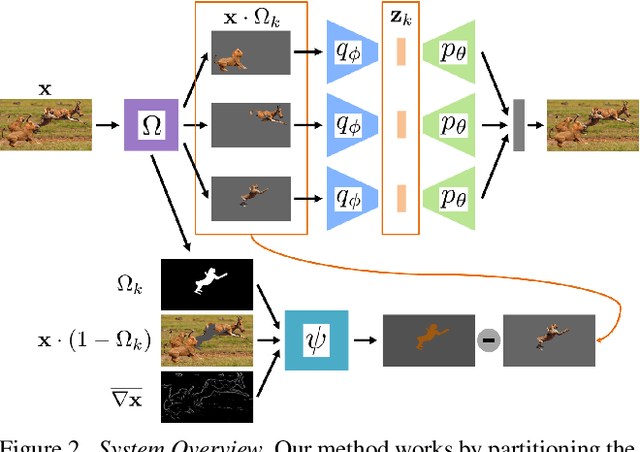
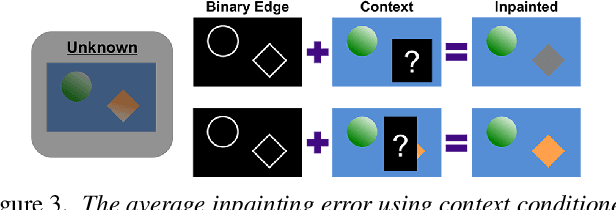
We describe a method to train a generative model with latent factors that are (approximately) independent and localized. This means that perturbing the latent variables affects only local regions of the synthesized image, corresponding to objects. Unlike other unsupervised generative models, ours enables object-centric manipulation, without requiring object-level annotations, or any form of annotation for that matter. The key to our method is the combination of spatial disentanglement, enforced by a Contextual Information Separation loss, and perceptual cycle-consistency, enforced by a loss that penalizes changes in the image partition in response to perturbations of the latent factors. We test our method's ability to allow independent control of spatial and semantic factors of variability on existing datasets and also introduce two new ones that highlight the limitations of current methods.
Deep Image Deraining Via Intrinsic Rainy Image Priors and Multi-scale Auxiliary Decoding
Nov 25, 2019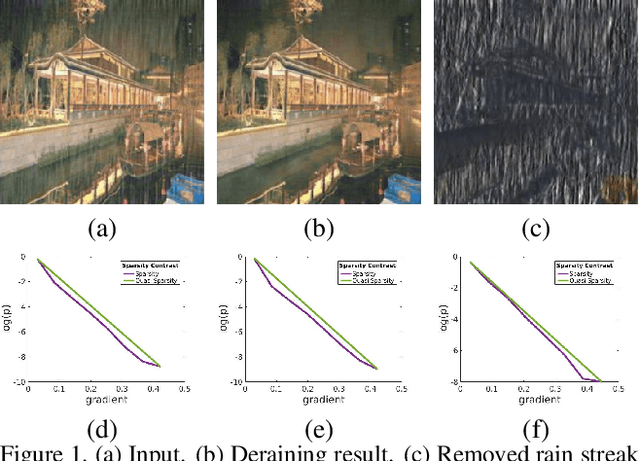
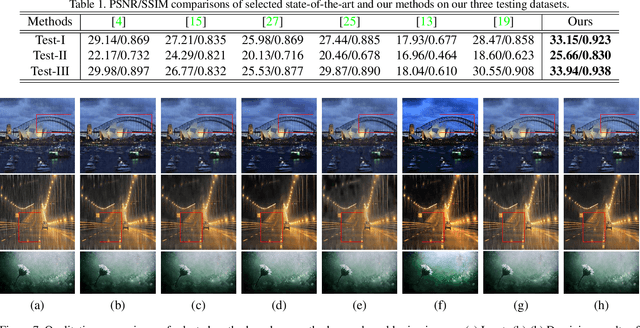
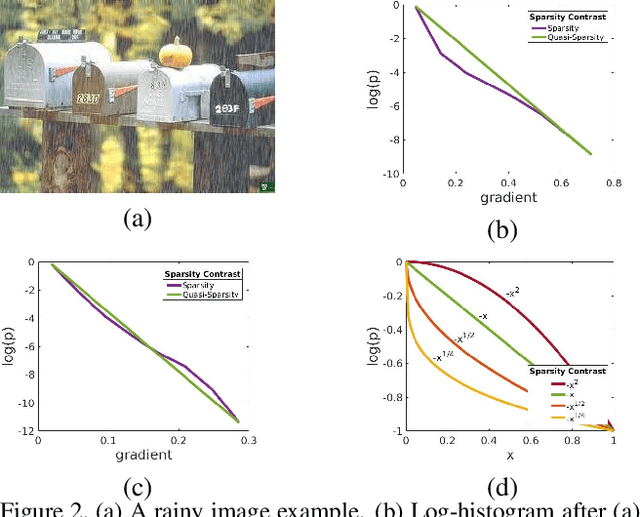
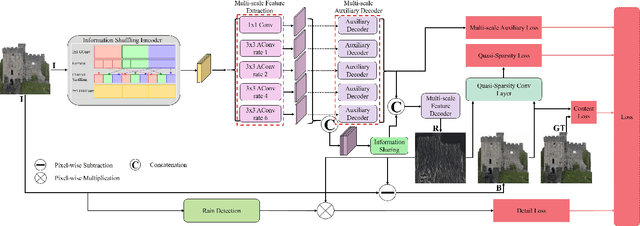
Different rain models and novel network structures have been proposed to remove rain streaks from single rainy images. In this work, we bring attention to the intrinsic priors and multi-scale features of the rainy images, and develop several intrinsic loss functions to train a CNN deraining network. We first study the sparse priors of rainy images, which have been verified to preserve unbroken edges in image decomposition. However, its mathematical formulation usually leads to an intractable solution, we propose quasi-sparsity priors to decrease complexity, so that our network can be trained under the supervision of sparse properties of rainy images. Quasi-sparsity supervises network training in different gradient domain which is still ill-posed to decompose a rainy image into rain layer and background layer. We develop another $L_1$ loss based on the intrinsic low-value property of rain layer to restore image contents together with the commonly-used $L_1$ similarity loss. Multi-scale features are further explored via a multi-scale auxiliary decoding structure to show which kinds of features contribute the most to the deraining task, and the corresponding multi-scale auxiliary loss improves the deraining performance further. In our network, more efficient group convolution and feature sharing are utilized to obtain an one order of magnitude improvement in network running speed. The proposed deraining method performs favorably against state-of-the-art deraining approaches.
In-painting Radiography Images for Unsupervised Anomaly Detection
Nov 26, 2021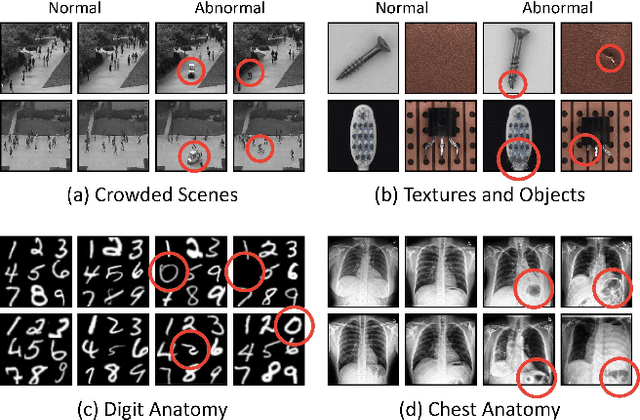
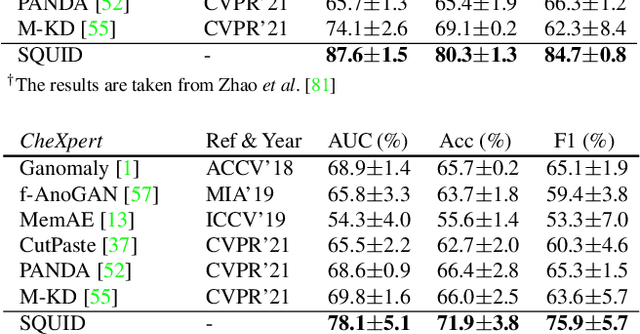
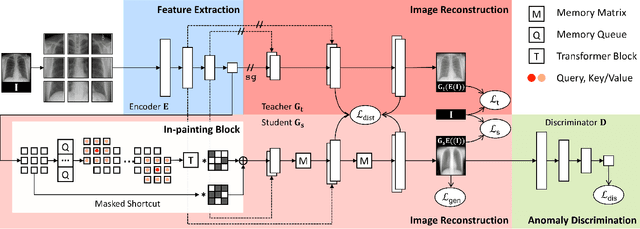
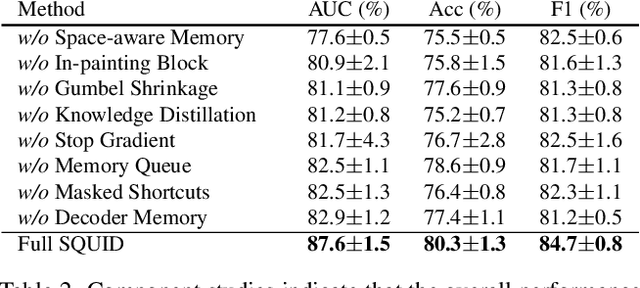
We propose space-aware memory queues for in-painting and detecting anomalies from radiography images (abbreviated as SQUID). Radiography imaging protocols focus on particular body regions, therefore producing images of great similarity and yielding recurrent anatomical structures across patients. To exploit this structured information, our SQUID consists of a new Memory Queue and a novel in-painting block in the feature space. We show that SQUID can taxonomize the ingrained anatomical structures into recurrent patterns; and in the inference, SQUID can identify anomalies (unseen/modified patterns) in the image. SQUID surpasses the state of the art in unsupervised anomaly detection by over 5 points on two chest X-ray benchmark datasets. Additionally, we have created a new dataset (DigitAnatomy), which synthesizes the spatial correlation and consistent shape in chest anatomy. We hope DigitAnatomy can prompt the development, evaluation, and interpretability of anomaly detection methods, particularly for radiography imaging.
Asymmetric Modality Translation For Face Presentation Attack Detection
Oct 20, 2021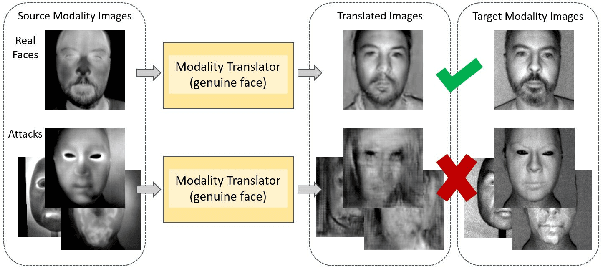
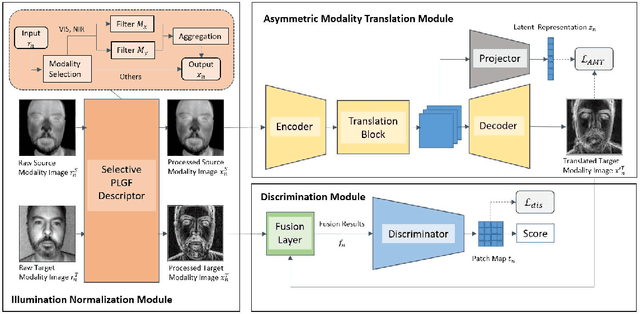
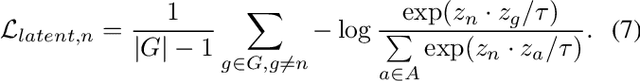
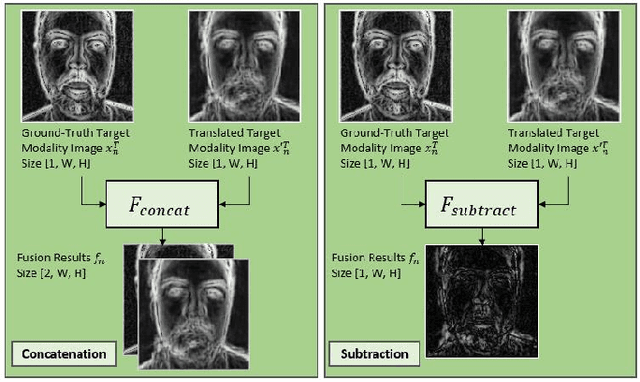
Face presentation attack detection (PAD) is an essential measure to protect face recognition systems from being spoofed by malicious users and has attracted great attention from both academia and industry. Although most of the existing methods can achieve desired performance to some extent, the generalization issue of face presentation attack detection under cross-domain settings (e.g., the setting of unseen attacks and varying illumination) remains to be solved. In this paper, we propose a novel framework based on asymmetric modality translation for face presentation attack detection in bi-modality scenarios. Under the framework, we establish connections between two modality images of genuine faces. Specifically, a novel modality fusion scheme is presented that the image of one modality is translated to the other one through an asymmetric modality translator, then fused with its corresponding paired image. The fusion result is fed as the input to a discriminator for inference. The training of the translator is supervised by an asymmetric modality translation loss. Besides, an illumination normalization module based on Pattern of Local Gravitational Force (PLGF) representation is used to reduce the impact of illumination variation. We conduct extensive experiments on three public datasets, which validate that our method is effective in detecting various types of attacks and achieves state-of-the-art performance under different evaluation protocols.
Test-Time Detection of Backdoor Triggers for Poisoned Deep Neural Networks
Dec 06, 2021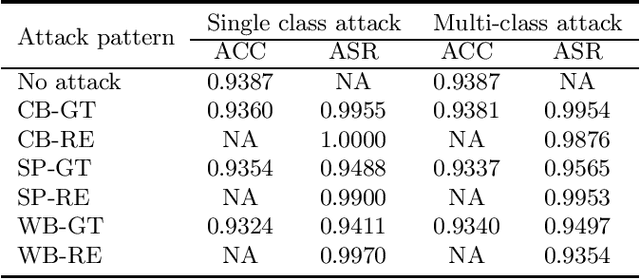
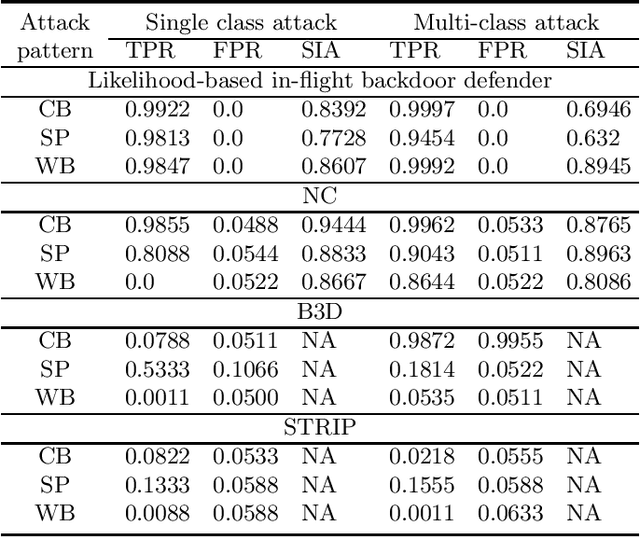

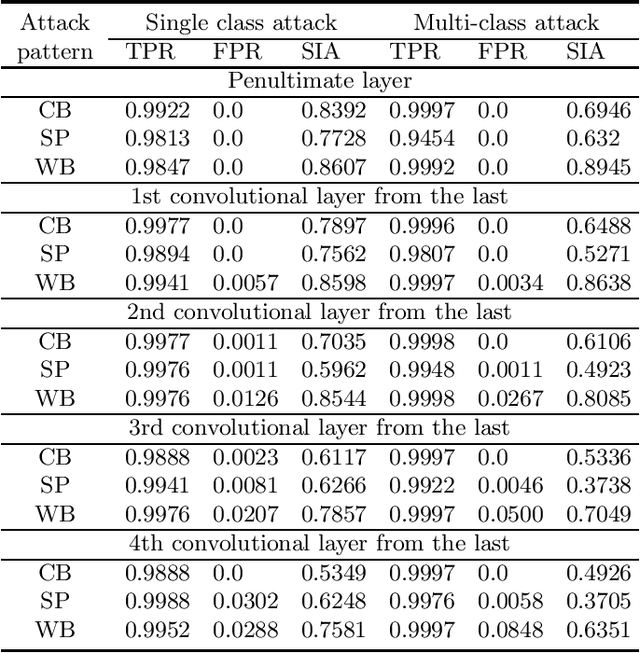
Backdoor (Trojan) attacks are emerging threats against deep neural networks (DNN). A DNN being attacked will predict to an attacker-desired target class whenever a test sample from any source class is embedded with a backdoor pattern; while correctly classifying clean (attack-free) test samples. Existing backdoor defenses have shown success in detecting whether a DNN is attacked and in reverse-engineering the backdoor pattern in a "post-training" regime: the defender has access to the DNN to be inspected and a small, clean dataset collected independently, but has no access to the (possibly poisoned) training set of the DNN. However, these defenses neither catch culprits in the act of triggering the backdoor mapping, nor mitigate the backdoor attack at test-time. In this paper, we propose an "in-flight" defense against backdoor attacks on image classification that 1) detects use of a backdoor trigger at test-time; and 2) infers the class of origin (source class) for a detected trigger example. The effectiveness of our defense is demonstrated experimentally against different strong backdoor attacks.
Optimized 2D CA-CFAR for Drone-Mounted Radar Signal Processing Using Integral Image Algorithm
Dec 21, 2020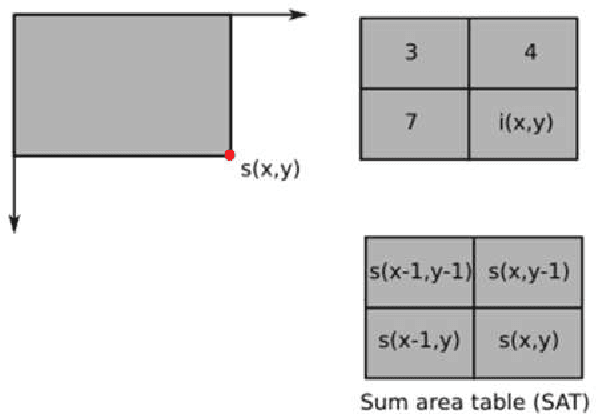
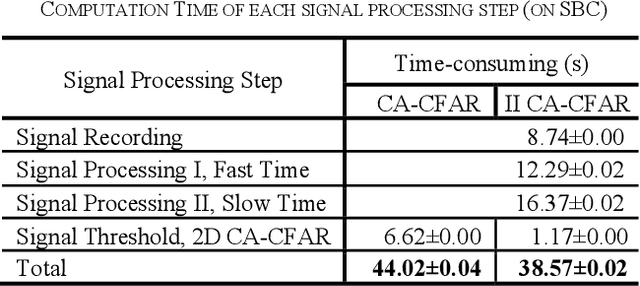
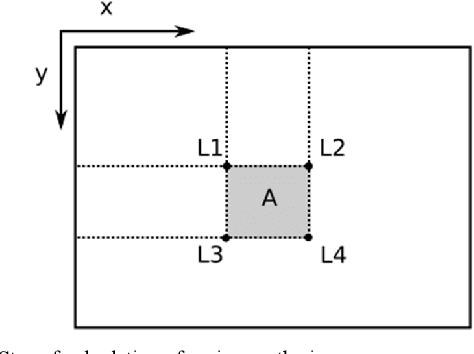
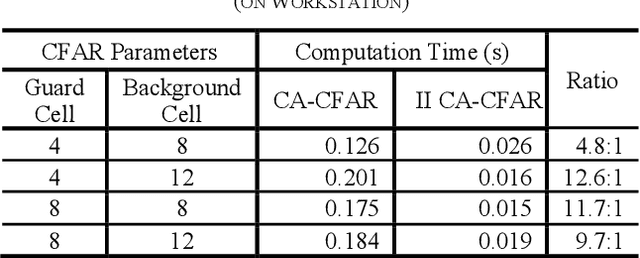

Buried survivor detection in the post-disaster environment by employing radar as sensor is an appealing approach. However, the implementation in the real field is challenging especially for large observation missions. Mounting the radar on the flying drone is the most promising solution. In this case, since the limitations of drones such as low computer specification and limited power resources, an efficient radar data processing is crucially required. Hence, this paper study about the implementation of the integral image technique to optimize the computation of the signal processing step of ultra-wideband impulse radar signatures. The evaluation was held on the single board computer mounted on the developed multisensory drone. The results confirm that the developed method can relatively reduce the data processing time.
Adversarial Attacks in Cooperative AI
Dec 06, 2021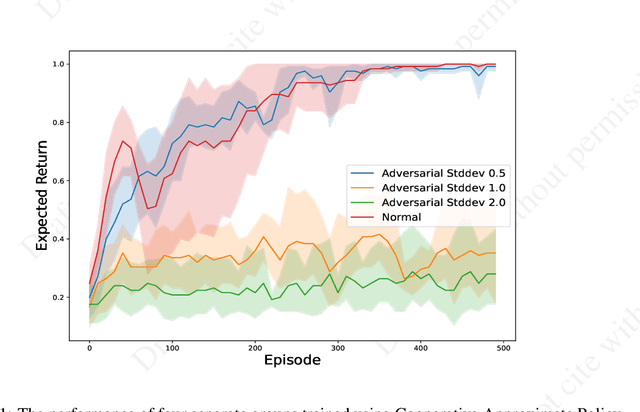
Single-agent reinforcement learning algorithms in a multi-agent environment are inadequate for fostering cooperation. If intelligent agents are to interact and work together to solve complex problems, methods that counter non-cooperative behavior are needed to facilitate the training of multiple agents. This is the goal of cooperative AI. Recent work in adversarial machine learning, however, shows that models (e.g., image classifiers) can be easily deceived into making incorrect decisions. In addition, some past research in cooperative AI has relied on new notions of representations, like public beliefs, to accelerate the learning of optimally cooperative behavior. Hence, cooperative AI might introduce new weaknesses not investigated in previous machine learning research. In this paper, our contributions include: (1) arguing that three algorithms inspired by human-like social intelligence introduce new vulnerabilities, unique to cooperative AI, that adversaries can exploit, and (2) an experiment showing that simple, adversarial perturbations on the agents' beliefs can negatively impact performance. This evidence points to the possibility that formal representations of social behavior are vulnerable to adversarial attacks.
A review: Deep learning for medical image segmentation using multi-modality fusion
Apr 22, 2020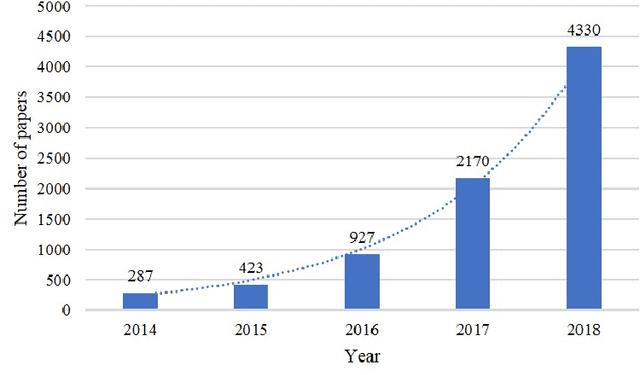
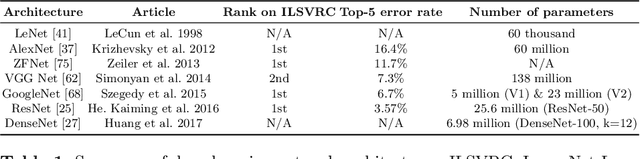
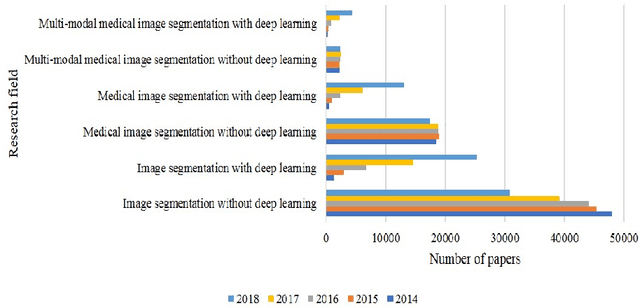
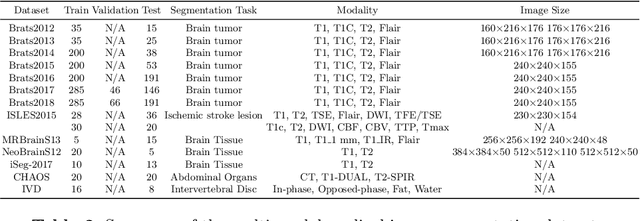
Multi-modality is widely used in medical imaging, because it can provide multiinformation about a target (tumor, organ or tissue). Segmentation using multimodality consists of fusing multi-information to improve the segmentation. Recently, deep learning-based approaches have presented the state-of-the-art performance in image classification, segmentation, object detection and tracking tasks. Due to their self-learning and generalization ability over large amounts of data, deep learning recently has also gained great interest in multi-modal medical image segmentation. In this paper, we give an overview of deep learning-based approaches for multi-modal medical image segmentation task. Firstly, we introduce the general principle of deep learning and multi-modal medical image segmentation. Secondly, we present different deep learning network architectures, then analyze their fusion strategies and compare their results. The earlier fusion is commonly used, since it's simple and it focuses on the subsequent segmentation network architecture. However, the later fusion gives more attention on fusion strategy to learn the complex relationship between different modalities. In general, compared to the earlier fusion, the later fusion can give more accurate result if the fusion method is effective enough. We also discuss some common problems in medical image segmentation. Finally, we summarize and provide some perspectives on the future research.
* 26 pages, 8 figures
Learning to Rearrange Voxels in Binary Segmentation Masks for Smooth Manifold Triangulation
Aug 11, 2021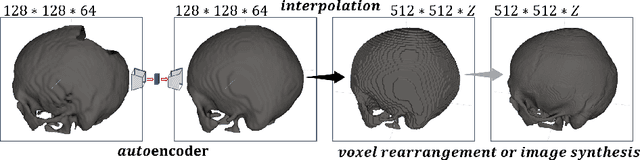
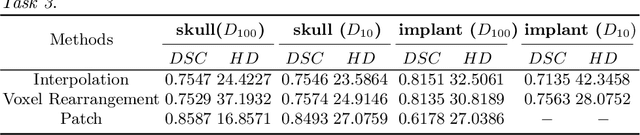
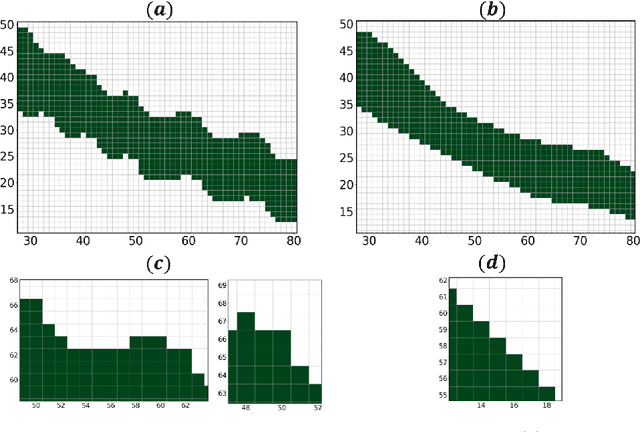
Medical images, especially volumetric images, are of high resolution and often exceed the capacity of standard desktop GPUs. As a result, most deep learning-based medical image analysis tasks require the input images to be downsampled, often substantially, before these can be fed to a neural network. However, downsampling can lead to a loss of image quality, which is undesirable especially in reconstruction tasks, where the fine geometric details need to be preserved. In this paper, we propose that high-resolution images can be reconstructed in a coarse-to-fine fashion, where a deep learning algorithm is only responsible for generating a coarse representation of the image, which consumes moderate GPU memory. For producing the high-resolution outcome, we propose two novel methods: learned voxel rearrangement of the coarse output and hierarchical image synthesis. Compared to the coarse output, the high-resolution counterpart allows for smooth surface triangulation, which can be 3D-printed in the highest possible quality. Experiments of this paper are carried out on the dataset of AutoImplant 2021 (https://autoimplant2021.grand-challenge.org/), a MICCAI challenge on cranial implant design. The dataset contains high-resolution skulls that can be viewed as 2D manifolds embedded in a 3D space. Codes associated with this study can be accessed at https://github.com/Jianningli/voxel_rearrangement.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge