"Image": models, code, and papers
Towards Privacy-preserving Explanations in Medical Image Analysis
Jul 20, 2021
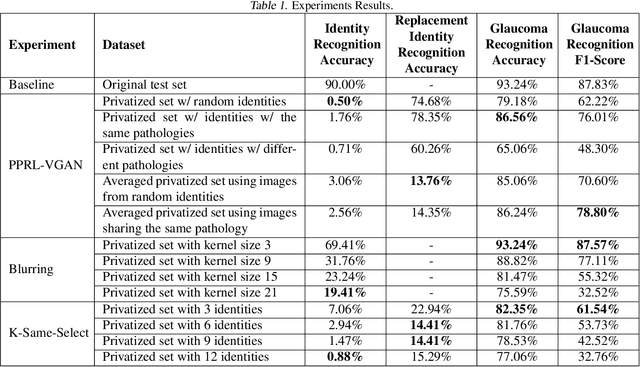
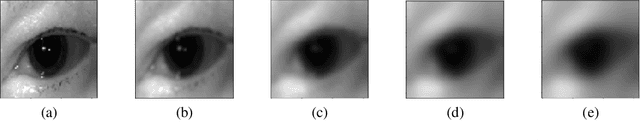
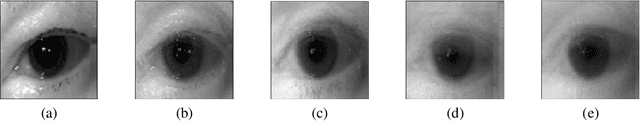
The use of Deep Learning in the medical field is hindered by the lack of interpretability. Case-based interpretability strategies can provide intuitive explanations for deep learning models' decisions, thus, enhancing trust. However, the resulting explanations threaten patient privacy, motivating the development of privacy-preserving methods compatible with the specifics of medical data. In this work, we analyze existing privacy-preserving methods and their respective capacity to anonymize medical data while preserving disease-related semantic features. We find that the PPRL-VGAN deep learning method was the best at preserving the disease-related semantic features while guaranteeing a high level of privacy among the compared state-of-the-art methods. Nevertheless, we emphasize the need to improve privacy-preserving methods for medical imaging, as we identified relevant drawbacks in all existing privacy-preserving approaches.
Image Outpainting and Harmonization using Generative Adversarial Networks
Feb 15, 2020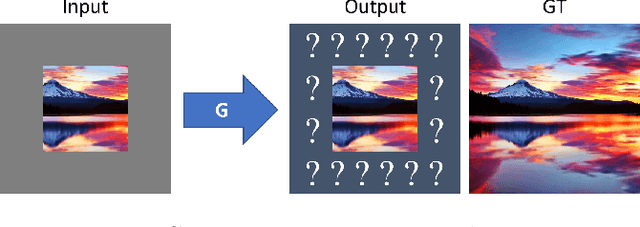



Although the inherently ambiguous task of predicting what resides beyond all four edges of an image has rarely been explored before, we demonstrate that GANs hold powerful potential in producing reasonable extrapolations. Two outpainting methods are proposed that aim to instigate this line of research: the first approach uses a context encoder inspired by common inpainting architectures and paradigms, while the second approach adds an extra post-processing step using a single-image generative model. This way, the hallucinated details are integrated with the style of the original image, in an attempt to further boost the quality of the result and possibly allow for arbitrary output resolutions to be supported.
CarveMix: A Simple Data Augmentation Method for Brain Lesion Segmentation
Aug 16, 2021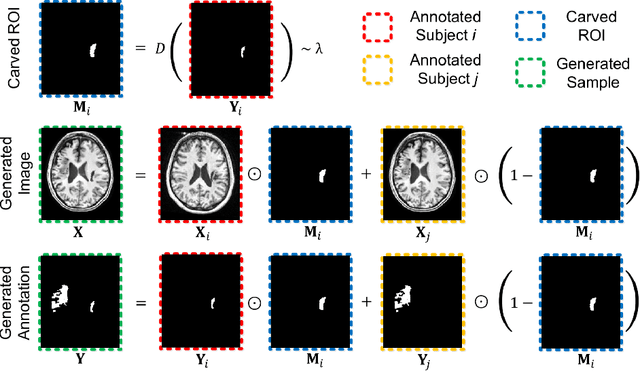
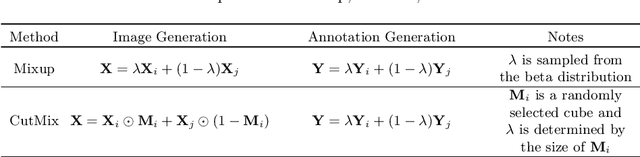

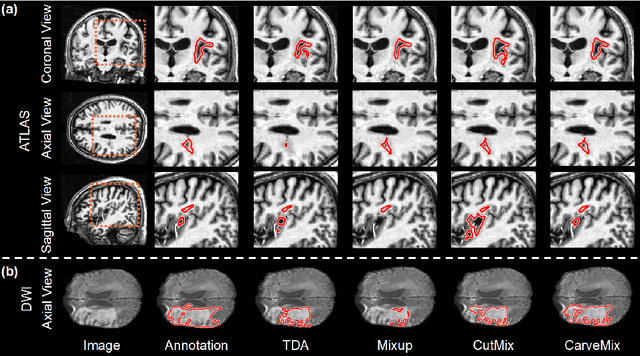
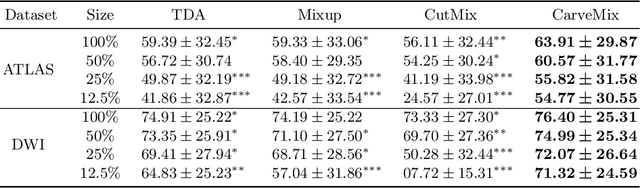
Brain lesion segmentation provides a valuable tool for clinical diagnosis, and convolutional neural networks (CNNs) have achieved unprecedented success in the task. Data augmentation is a widely used strategy that improves the training of CNNs, and the design of the augmentation method for brain lesion segmentation is still an open problem. In this work, we propose a simple data augmentation approach, dubbed as CarveMix, for CNN-based brain lesion segmentation. Like other "mix"-based methods, such as Mixup and CutMix, CarveMix stochastically combines two existing labeled images to generate new labeled samples. Yet, unlike these augmentation strategies based on image combination, CarveMix is lesion-aware, where the combination is performed with an attention on the lesions and a proper annotation is created for the generated image. Specifically, from one labeled image we carve a region of interest (ROI) according to the lesion location and geometry, and the size of the ROI is sampled from a probability distribution. The carved ROI then replaces the corresponding voxels in a second labeled image, and the annotation of the second image is replaced accordingly as well. In this way, we generate new labeled images for network training and the lesion information is preserved. To evaluate the proposed method, experiments were performed on two brain lesion datasets. The results show that our method improves the segmentation accuracy compared with other simple data augmentation approaches.
Fast Strain Estimation and Frame Selection in Ultrasound Elastography using Machine Learning
Oct 16, 2021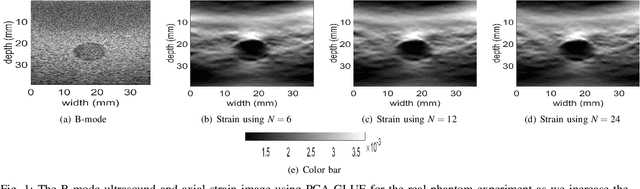
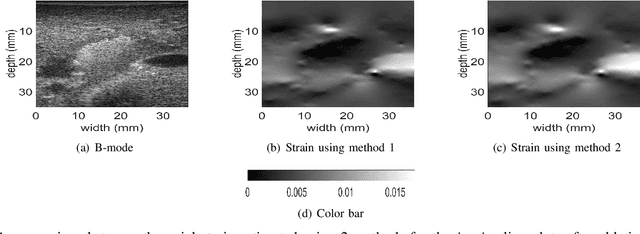
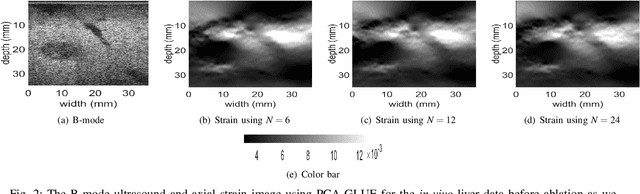
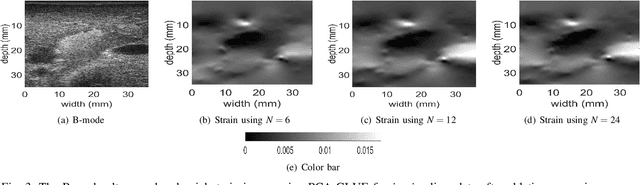
Ultrasound Elastography aims to determine the mechanical properties of the tissue by monitoring tissue deformation due to internal or external forces. Tissue deformations are estimated from ultrasound radio frequency (RF) signals and are often referred to as time delay estimation (TDE). Given two RF frames I1 and I2, we can compute a displacement image which shows the change in the position of each sample in I1 to a new position in I2. Two important challenges in TDE include high computational complexity and the difficulty in choosing suitable RF frames. Selecting suitable frames is of high importance because many pairs of RF frames either do not have acceptable deformation for extracting informative strain images or are decorrelated and deformation cannot be reliably estimated. Herein, we introduce a method that learns 12 displacement modes in quasi-static elastography by performing Principal Component Analysis (PCA) on displacement fields of a large training database. In the inference stage, we use dynamic programming (DP) to compute an initial displacement estimate of around 1% of the samples, and then decompose this sparse displacement into a linear combination of the 12 displacement modes. Our method assumes that the displacement of the whole image could also be described by this linear combination of principal components. We then use the GLobal Ultrasound Elastography (GLUE) method to fine-tune the result yielding the exact displacement image. Our method, which we call PCA-GLUE, is more than 10 times faster than DP in calculating the initial displacement map while giving the same result. Our second contribution in this paper is determining the suitability of the frame pair I1 and I2 for strain estimation, which we achieve by using the weight vector that we calculated for PCA-GLUE as an input to a multi-layer perceptron (MLP) classifier.
ViT-Inception-GAN for Image Colourising
Jun 11, 2021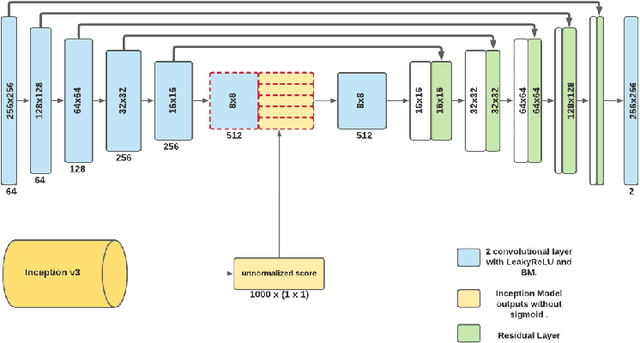
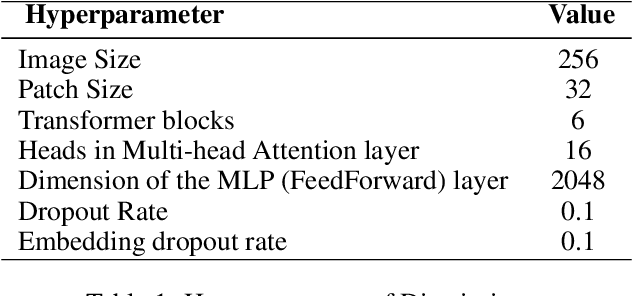
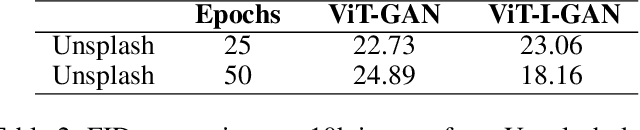

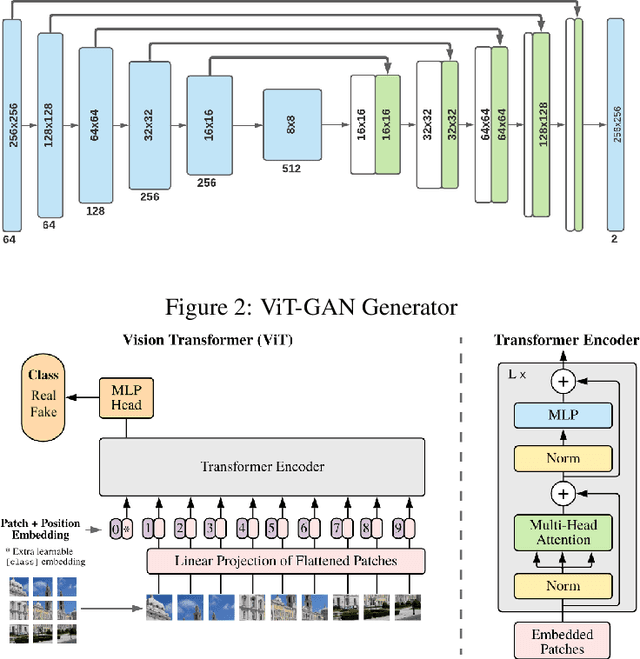

Studies involving colourising images has been garnering researchers' keen attention over time, assisted by significant advances in various Machine Learning techniques and compute power availability. Traditionally, colourising images have been an intricate task that gave a substantial degree of freedom during the assignment of chromatic information. In our proposed method, we attempt to colourise images using Vision Transformer - Inception - Generative Adversarial Network (ViT-I-GAN), which has an Inception-v3 fusion embedding in the generator. For a stable and robust network, we have used Vision Transformer (ViT) as the discriminator. We trained the model on the Unsplash and the COCO dataset for demonstrating the improvement made by the Inception-v3 embedding. We have compared the results between ViT-GANs with and without Inception-v3 embedding.
Adversarial normalization for multi domain image segmentation
Feb 01, 2020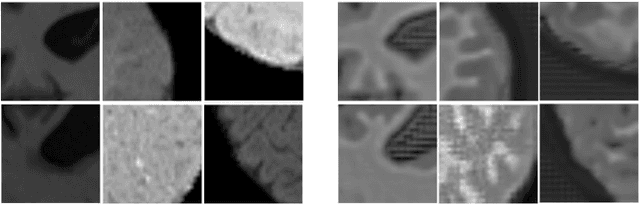
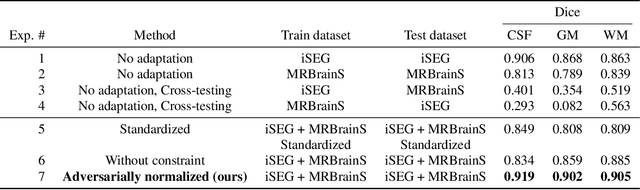
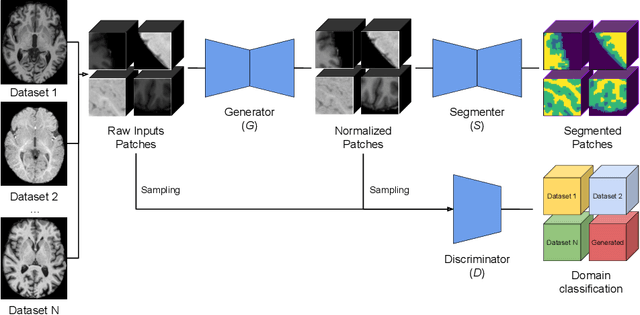

Image normalization is a critical step in medical imaging. This step is often done on a per-dataset basis, preventing current segmentation algorithms from the full potential of exploiting jointly normalized information across multiple datasets. To solve this problem, we propose an adversarial normalization approach for image segmentation which learns common normalizing functions across multiple datasets while retaining image realism. The adversarial training provides an optimal normalizer that improves both the segmentation accuracy and the discrimination of unrealistic normalizing functions. Our contribution therefore leverages common imaging information from multiple domains. The optimality of our common normalizer is evaluated by combining brain images from both infants and adults. Results on the challenging iSEG and MRBrainS datasets reveal the potential of our adversarial normalization approach for segmentation, with Dice improvements of up to 59.6% over the baseline.
3D-OOCS: Learning Prostate Segmentation with Inductive Bias
Oct 29, 2021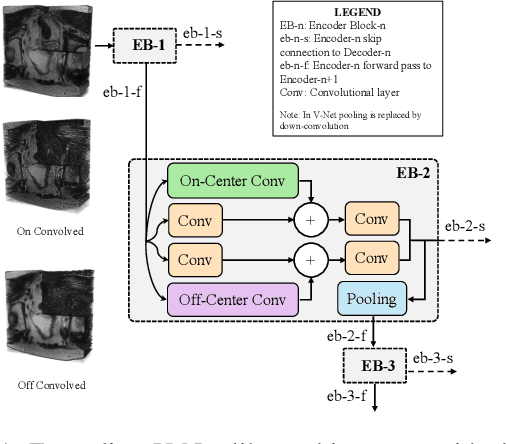
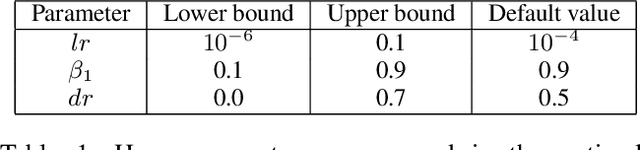
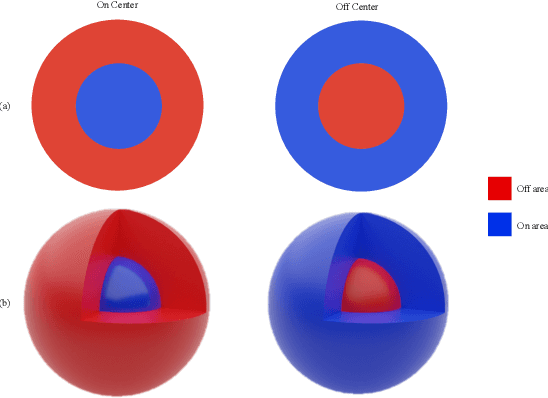
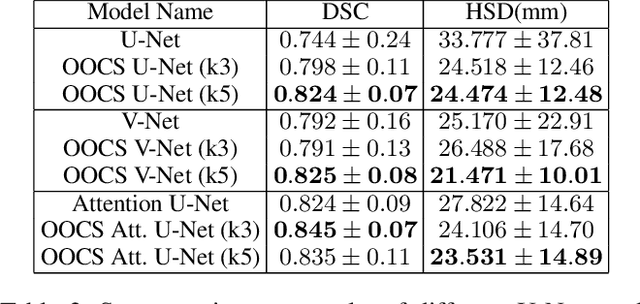
Despite the great success of convolutional neural networks (CNN) in 3D medical image segmentation tasks, the methods currently in use are still not robust enough to the different protocols utilized by different scanners, and to the variety of image properties or artefacts they produce. To this end, we introduce OOCS-enhanced networks, a novel architecture inspired by the innate nature of visual processing in the vertebrates. With different 3D U-Net variants as the base, we add two 3D residual components to the second encoder blocks: on and off center-surround (OOCS). They generalise the ganglion pathways in the retina to a 3D setting. The use of 2D-OOCS in any standard CNN network complements the feedforward framework with sharp edge-detection inductive biases. The use of 3D-OOCS also helps 3D U-Nets to scrutinise and delineate anatomical structures present in 3D images with increased accuracy.We compared the state-of-the-art 3D U-Nets with their 3D-OOCS extensions and showed the superior accuracy and robustness of the latter in automatic prostate segmentation from 3D Magnetic Resonance Images (MRIs). For a fair comparison, we trained and tested all the investigated 3D U-Nets with the same pipeline, including automatic hyperparameter optimisation and data augmentation.
A Novel BiLevel Paradigm for Image-to-Image Translation
Apr 18, 2019
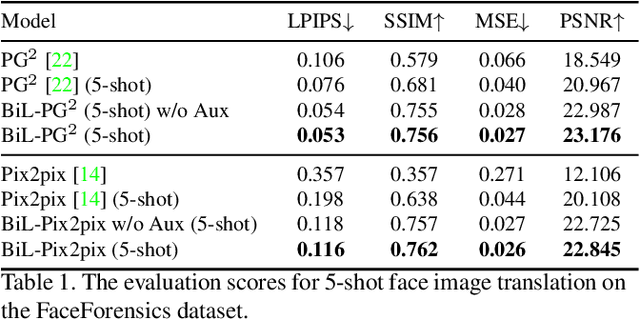
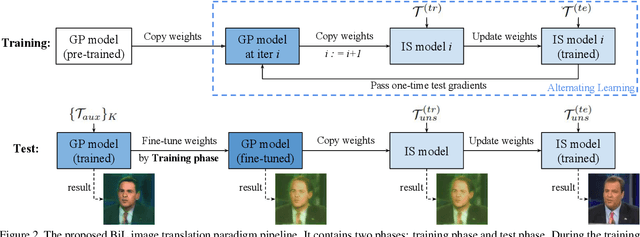
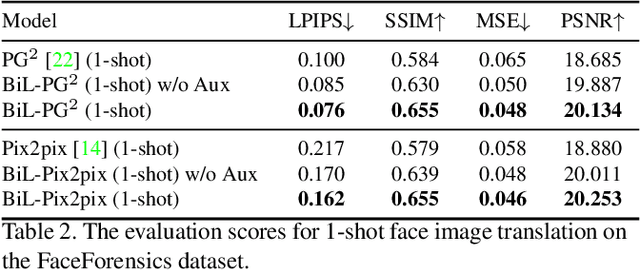
Image-to-image (I2I) translation is a pixel-level mapping that requires a large number of paired training data and often suffers from the problems of high diversity and strong category bias in image scenes. In order to tackle these problems, we propose a novel BiLevel (BiL) learning paradigm that alternates the learning of two models, respectively at an instance-specific (IS) and a general-purpose (GP) level. In each scene, the IS model learns to maintain the specific scene attributes. It is initialized by the GP model that learns from all the scenes to obtain the generalizable translation knowledge. This GP initialization gives the IS model an efficient starting point, thus enabling its fast adaptation to the new scene with scarce training data. We conduct extensive I2I translation experiments on human face and street view datasets. Quantitative results validate that our approach can significantly boost the performance of classical I2I translation models, such as PG2 and Pix2Pix. Our visualization results show both higher image quality and more appropriate instance-specific details, e.g., the translated image of a person looks more like that person in terms of identity.
Quality-aware Cine Cardiac MRI Reconstruction and Analysis from Undersampled k-space Data
Sep 16, 2021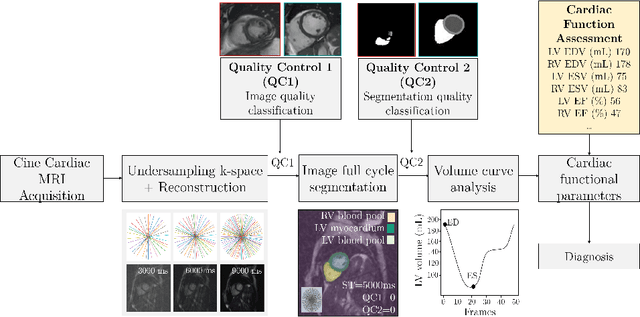
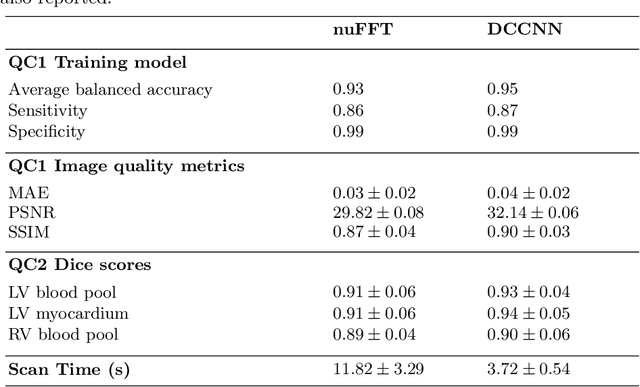
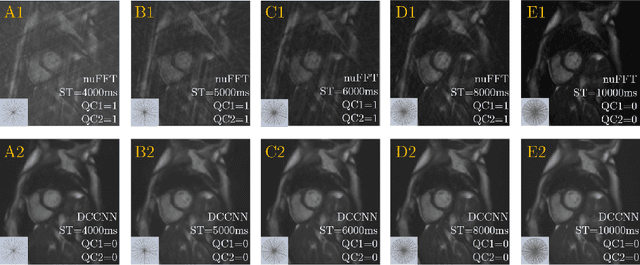
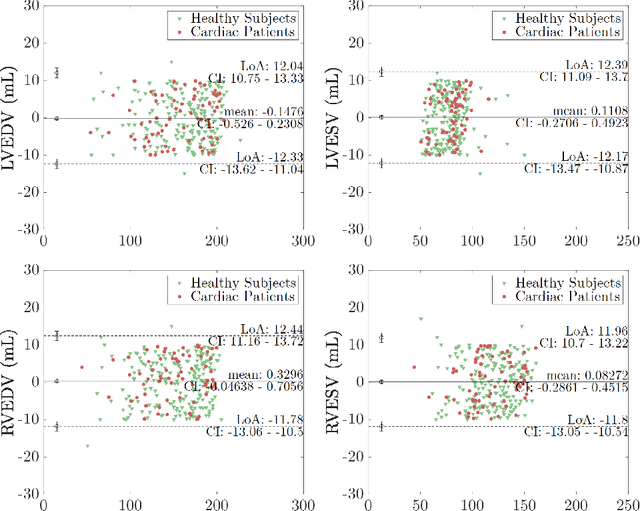
Cine cardiac MRI is routinely acquired for the assessment of cardiac health, but the imaging process is slow and typically requires several breath-holds to acquire sufficient k-space profiles to ensure good image quality. Several undersampling-based reconstruction techniques have been proposed during the last decades to speed up cine cardiac MRI acquisition. However, the undersampling factor is commonly fixed to conservative values before acquisition to ensure diagnostic image quality, potentially leading to unnecessarily long scan times. In this paper, we propose an end-to-end quality-aware cine short-axis cardiac MRI framework that combines image acquisition and reconstruction with downstream tasks such as segmentation, volume curve analysis and estimation of cardiac functional parameters. The goal is to reduce scan time by acquiring only a fraction of k-space data to enable the reconstruction of images that can pass quality control checks and produce reliable estimates of cardiac functional parameters. The framework consists of a deep learning model for the reconstruction of 2D+t cardiac cine MRI images from undersampled data, an image quality-control step to detect good quality reconstructions, followed by a deep learning model for bi-ventricular segmentation, a quality-control step to detect good quality segmentations and automated calculation of cardiac functional parameters. To demonstrate the feasibility of the proposed approach, we perform simulations using a cohort of selected participants from the UK Biobank (n=270), 200 healthy subjects and 70 patients with cardiomyopathies. Our results show that we can produce quality-controlled images in a scan time reduced from 12 to 4 seconds per slice, enabling reliable estimates of cardiac functional parameters such as ejection fraction within 5% mean absolute error.
3D-StyleGAN: A Style-Based Generative Adversarial Network for Generative Modeling of Three-Dimensional Medical Images
Jul 20, 2021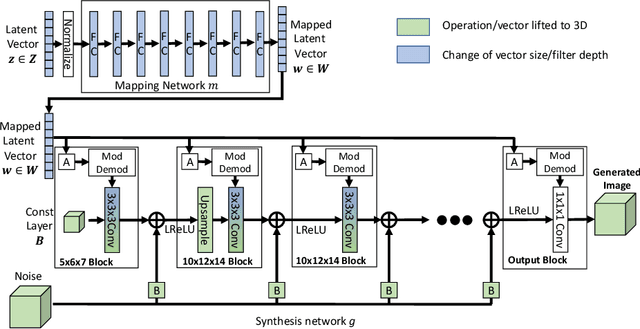
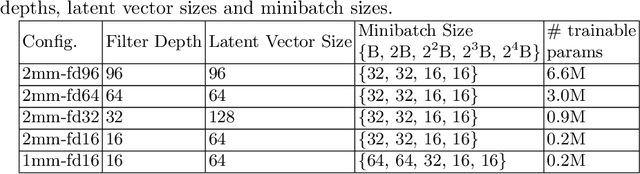
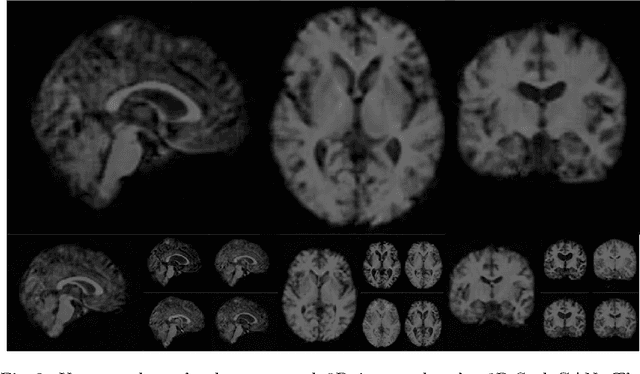
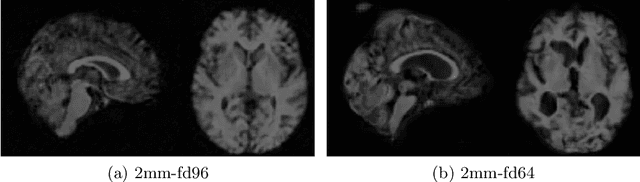
Image synthesis via Generative Adversarial Networks (GANs) of three-dimensional (3D) medical images has great potential that can be extended to many medical applications, such as, image enhancement and disease progression modeling. However, current GAN technologies for 3D medical image synthesis need to be significantly improved to be readily adapted to real-world medical problems. In this paper, we extend the state-of-the-art StyleGAN2 model, which natively works with two-dimensional images, to enable 3D image synthesis. In addition to the image synthesis, we investigate the controllability and interpretability of the 3D-StyleGAN via style vectors inherited form the original StyleGAN2 that are highly suitable for medical applications: (i) the latent space projection and reconstruction of unseen real images, and (ii) style mixing. We demonstrate the 3D-StyleGAN's performance and feasibility with ~12,000 three-dimensional full brain MR T1 images, although it can be applied to any 3D volumetric images. Furthermore, we explore different configurations of hyperparameters to investigate potential improvement of the image synthesis with larger networks. The codes and pre-trained networks are available online: https://github.com/sh4174/3DStyleGAN.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge