"Image": models, code, and papers
A Comprehensive Comparison of Multi-Dimensional Image Denoising Methods
Nov 06, 2020
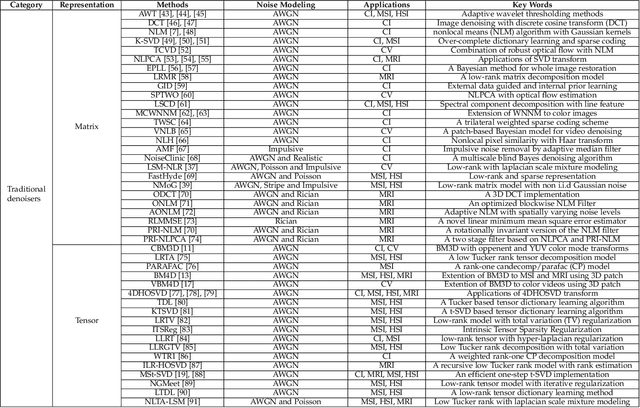
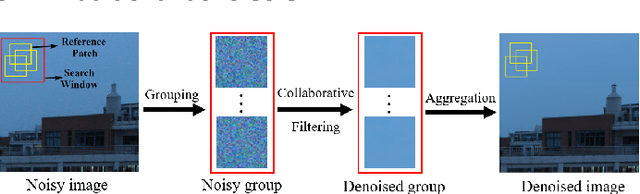
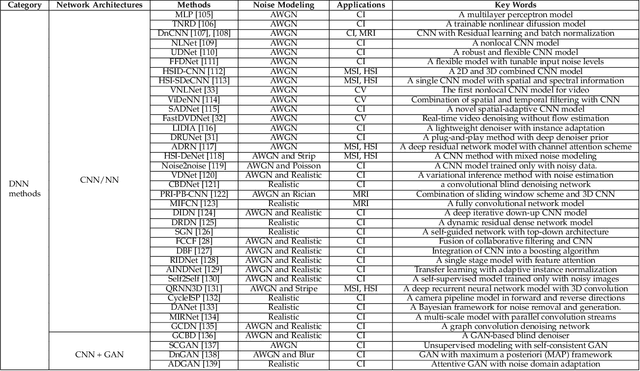
Filtering multi-dimensional images such as color images, color videos, multispectral images and magnetic resonance images is challenging in terms of both effectiveness and efficiency. Leveraging the nonlocal self-similarity (NLSS) characteristic of images and sparse representation in the transform domain, the block-matching and 3D filtering (BM3D) based methods show powerful denoising performance. Recently, numerous new approaches with different regularization terms, transforms and advanced deep neural network (DNN) architectures are proposed to improve denoising quality. In this paper, we extensively compare over 60 methods on both synthetic and real-world datasets. We also introduce a new color image and video dataset for benchmarking, and our evaluations are performed from four different perspectives including quantitative metrics, visual effects, human ratings and computational cost. Comprehensive experiments demonstrate: (i) the effectiveness and efficiency of the BM3D family for various denoising tasks, (ii) a simple matrix-based algorithm could produce similar results compared with its tensor counterparts, and (iii) several DNN models trained with synthetic Gaussian noise show state-of-the-art performance on real-world color image and video datasets. Despite the progress in recent years, we discuss shortcomings and possible extensions of existing techniques. Datasets and codes for evaluation are made publicly available at https://github.com/ZhaomingKong/Denoising-Comparison.
Using Self-Supervised Pretext Tasks for Active Learning
Jan 19, 2022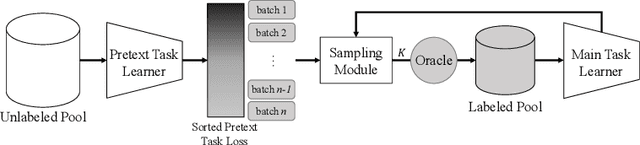
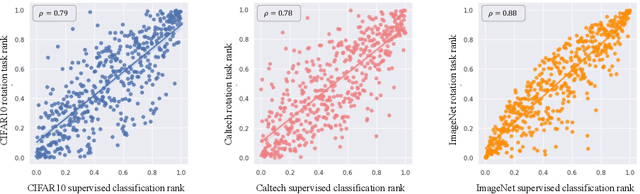
Labeling a large set of data is expensive. Active learning aims to tackle this problem by asking to annotate only the most informative data from the unlabeled set. We propose a novel active learning approach that utilizes self-supervised pretext tasks and a unique data sampler to select data that are both difficult and representative. We discover that the loss of a simple self-supervised pretext task, such as rotation prediction, is closely correlated to the downstream task loss. The pretext task learner is trained on the unlabeled set, and the unlabeled data are sorted and grouped into batches by their pretext task losses. In each iteration, the main task model is used to sample the most uncertain data in a batch to be annotated. We evaluate our method on various image classification and segmentation benchmarks and achieve compelling performances on CIFAR10, Caltech-101, ImageNet, and CityScapes.
PlantStereo: A Stereo Matching Benchmark for Plant Surface Dense Reconstruction
Nov 30, 2021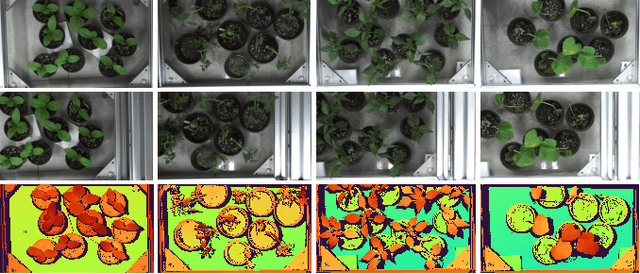


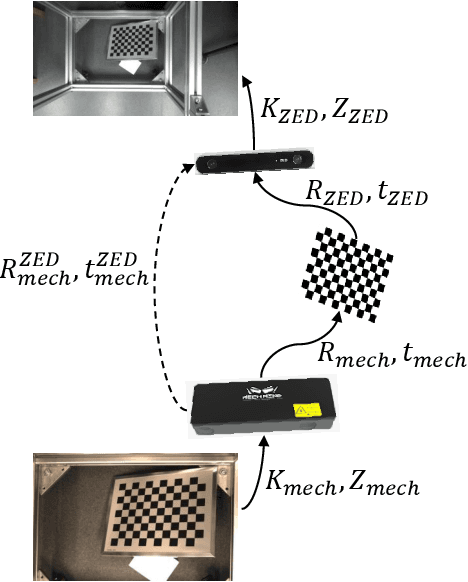
Stereo matching is an important task in computer vision which has drawn tremendous research attention for decades. While in terms of disparity accuracy, density and data size, public stereo datasets are difficult to meet the requirements of models. In this paper, we aim to address the issue between datasets and models and propose a large scale stereo dataset with high accuracy disparity ground truth named PlantStereo. We used a semi-automatic way to construct the dataset: after camera calibration and image registration, high accuracy disparity images can be obtained from the depth images. In total, PlantStereo contains 812 image pairs covering a diverse set of plants: spinach, tomato, pepper and pumpkin. We firstly evaluated our PlantStereo dataset on four different stereo matching methods. Extensive experiments on different models and plants show that compared with ground truth in integer accuracy, high accuracy disparity images provided by PlantStereo can remarkably improve the training effect of deep learning models. This paper provided a feasible and reliable method to realize plant surface dense reconstruction. The PlantStereo dataset and relative code are available at: https://www.github.com/wangqingyu985/PlantStereo
Deep Implicit Statistical Shape Models for 3D Medical Image Delineation
Apr 07, 2021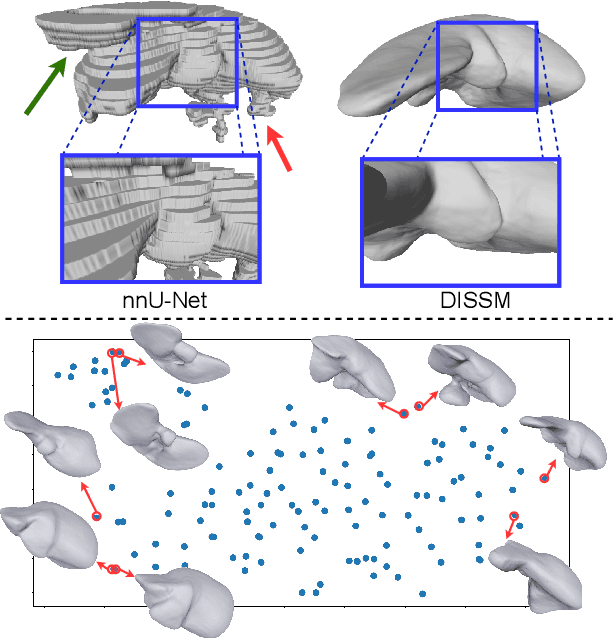
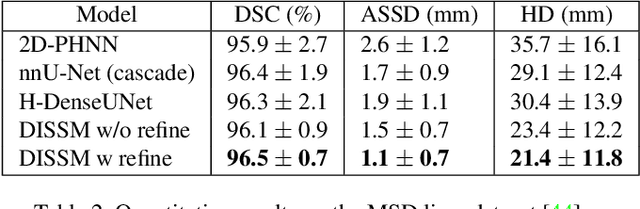
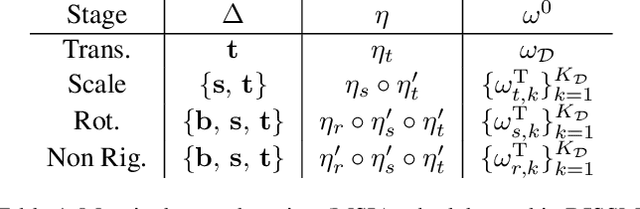
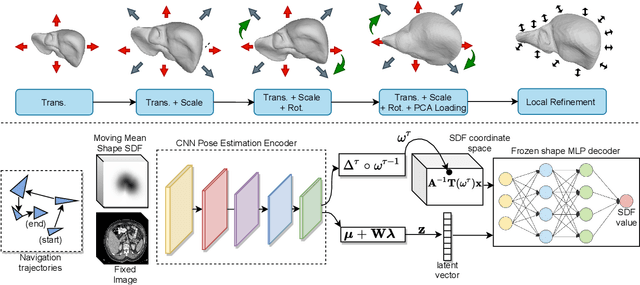

3D delineation of anatomical structures is a cardinal goal in medical imaging analysis. Prior to deep learning, statistical shape models that imposed anatomical constraints and produced high quality surfaces were a core technology. Prior to deep learning, statistical shape models that imposed anatomical constraints and produced high quality surfaces were a core technology. Today fully-convolutional networks (FCNs), while dominant, do not offer these capabilities. We present deep implicit statistical shape models (DISSMs), a new approach to delineation that marries the representation power of convolutional neural networks (CNNs) with the robustness of SSMs. DISSMs use a deep implicit surface representation to produce a compact and descriptive shape latent space that permits statistical models of anatomical variance. To reliably fit anatomically plausible shapes to an image, we introduce a novel rigid and non-rigid pose estimation pipeline that is modelled as a Markov decision process(MDP). We outline a training regime that includes inverted episodic training and a deep realization of marginal space learning (MSL). Intra-dataset experiments on the task of pathological liver segmentation demonstrate that DISSMs can perform more robustly than three leading FCN models, including nnU-Net: reducing the mean Hausdorff distance (HD) by 7.7-14.3mm and improving the worst case Dice-Sorensen coefficient (DSC) by 1.2-2.3%. More critically, cross-dataset experiments on a dataset directly reflecting clinical deployment scenarios demonstrate that DISSMs improve the mean DSC and HD by 3.5-5.9% and 12.3-24.5mm, respectively, and the worst-case DSC by 5.4-7.3%. These improvements are over and above any benefits from representing delineations with high-quality surface.
HEAT: Holistic Edge Attention Transformer for Structured Reconstruction
Nov 30, 2021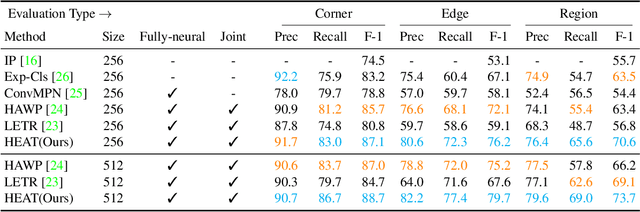
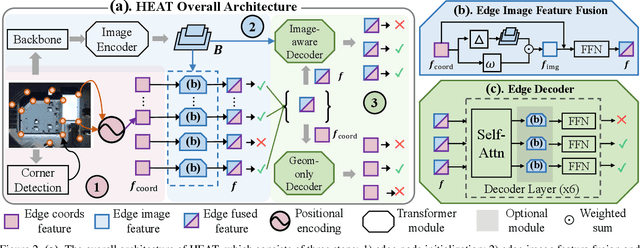
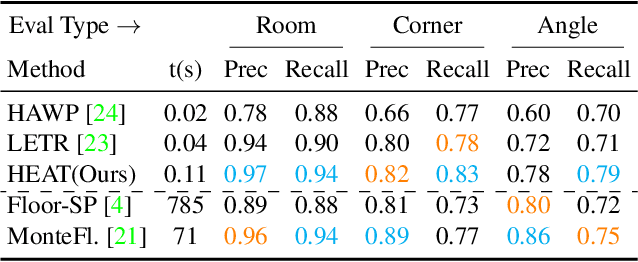
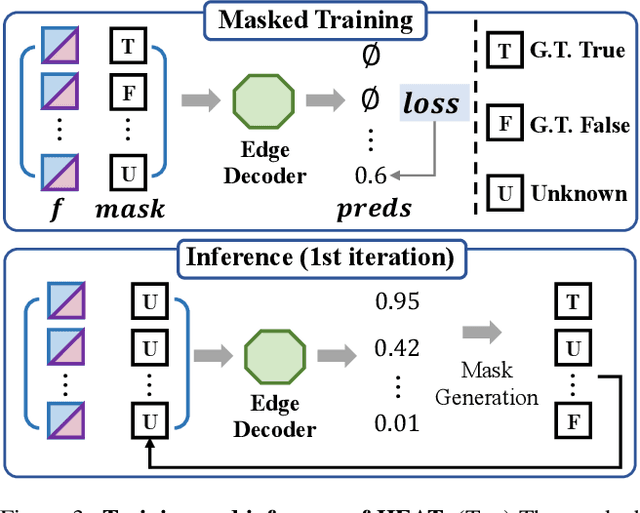
This paper presents a novel attention-based neural network for structured reconstruction, which takes a 2D raster image as an input and reconstructs a planar graph depicting an underlying geometric structure. The approach detects corners and classifies edge candidates between corners in an end-to-end manner. Our contribution is a holistic edge classification architecture, which 1) initializes the feature of an edge candidate by a trigonometric positional encoding of its end-points; 2) fuses image feature to each edge candidate by deformable attention; 3) employs two weight-sharing Transformer decoders to learn holistic structural patterns over the graph edge candidates; and 4) is trained with a masked learning strategy. The corner detector is a variant of the edge classification architecture, adapted to operate on pixels as corner candidates. We conduct experiments on two structured reconstruction tasks: outdoor building architecture and indoor floorplan planar graph reconstruction. Extensive qualitative and quantitative evaluations demonstrate the superiority of our approach over the state of the art. We will share code and models.
Computational analysis of pathological image enables interpretable prediction for microsatellite instability
Oct 07, 2020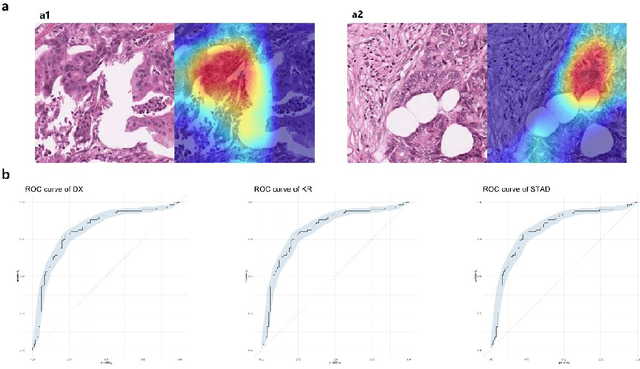
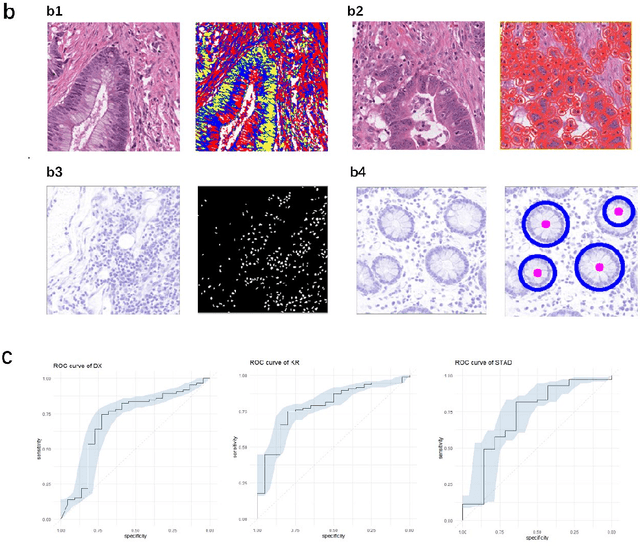
Microsatellite instability (MSI) is associated with several tumor types and its status has become increasingly vital in guiding patient treatment decisions. However, in clinical practice, distinguishing MSI from its counterpart is challenging since the diagnosis of MSI requires additional genetic or immunohistochemical tests. In this study, interpretable pathological image analysis strategies are established to help medical experts to automatically identify MSI. The strategies only require ubiquitous Haematoxylin and eosin-stained whole-slide images and can achieve decent performance in the three cohorts collected from The Cancer Genome Atlas. The strategies provide interpretability in two aspects. On the one hand, the image-level interpretability is achieved by generating localization heat maps of important regions based on the deep learning network; on the other hand, the feature-level interpretability is attained through feature importance and pathological feature interaction analysis. More interestingly, both from the image-level and feature-level interpretability, color features and texture characteristics are shown to contribute the most to the MSI predictions. Therefore, the classification models under the proposed strategies can not only serve as an efficient tool for predicting the MSI status of patients, but also provide more insights to pathologists with clinical understanding.
LC-GAN: Image-to-image Translation Based on Generative Adversarial Network for Endoscopic Images
Mar 10, 2020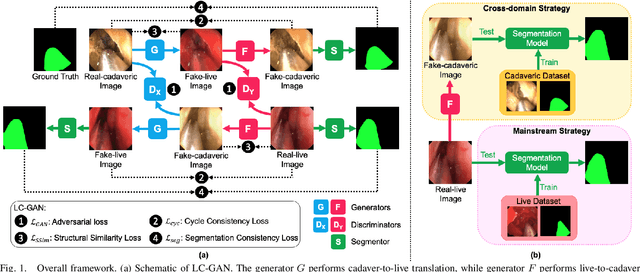
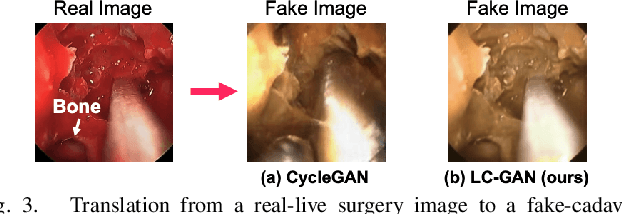
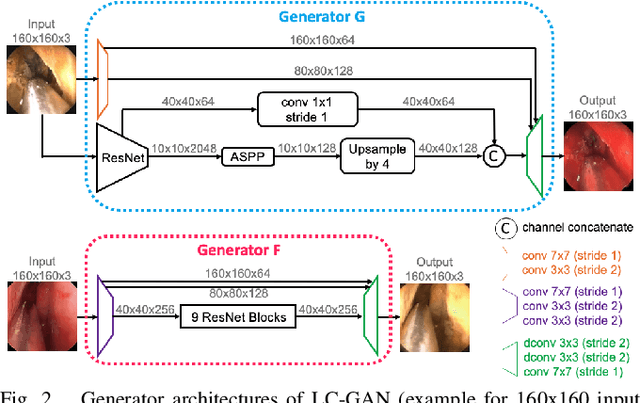

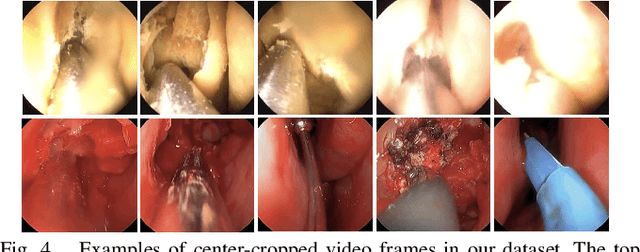
The intelligent perception of endoscopic vision is appealing in many computer-assisted and robotic surgeries. Achieving good vision-based analysis with deep learning techniques requires large labeled datasets, but manual data labeling is expensive and time-consuming in medical problems. When applying a trained model to a different but relevant dataset, a new labeled dataset may be required for training to avoid performance degradation. In this work, we investigate a novel cross-domain strategy to reduce the need for manual data labeling by proposing an image-to-image translation model called live-cadaver GAN (LC-GAN) based on generative adversarial networks (GANs). More specifically, we consider a situation when a labeled cadaveric surgery dataset is available while the task is instrument segmentation on a live surgery dataset. We train LC-GAN to learn the mappings between the cadaveric and live datasets. To achieve instrument segmentation on live images, we can first translate the live images to fake-cadaveric images with LC-GAN, and then perform segmentation on the fake-cadaveric images with models trained on the real cadaveric dataset. With this cross-domain strategy, we fully leverage the labeled cadaveric dataset for segmentation on live images without the need to label the live dataset again. Two generators with different architectures are designed for LC-GAN to make use of the deep feature representation learned from the cadaveric image based instrument segmentation task. Moreover, we propose structural similarity loss and segmentation consistency loss to improve the semantic consistency during translation. The results demonstrate that LC-GAN achieves better image-to-image translation results, and leads to improved segmentation performance in the proposed cross-domain segmentation task.
Correlation based Imaging for rotating satellites
Nov 01, 2021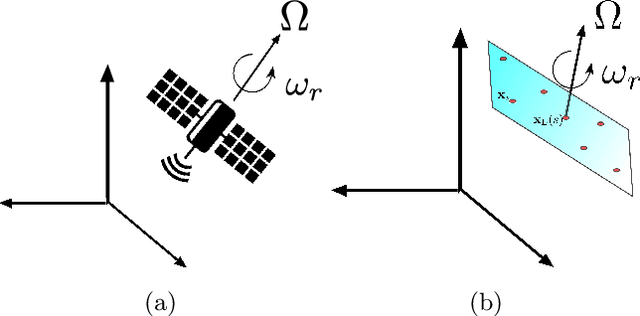
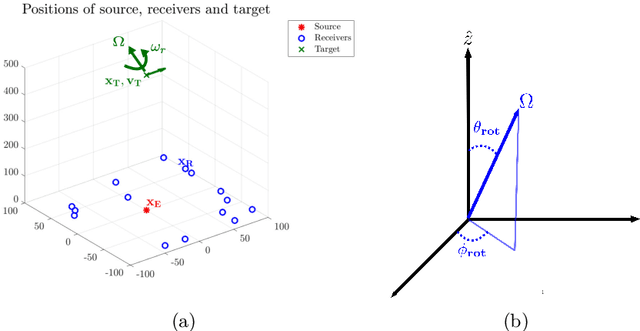
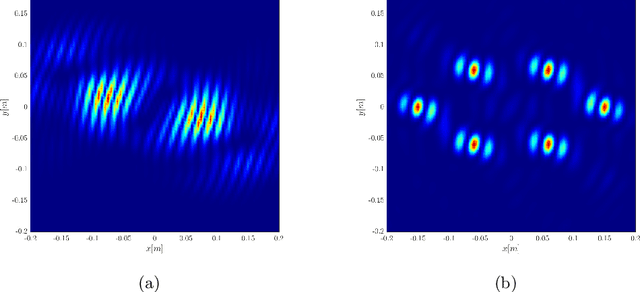
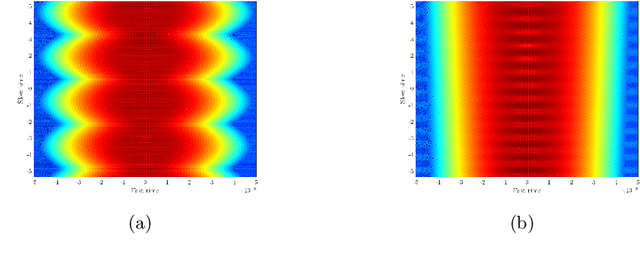
We consider imaging of fast moving small objects in space, such as low earth orbit satellites, which are also rotating around a fixed axis. The imaging system consists of ground based, asynchronous sources of radiation and several passive receivers above the dense atmosphere. We use the cross-correlation of the received signals to reduce distortions from ambient medium fluctuations. Imaging with correlations also has the advantage of not requiring any knowledge about the probing pulse and depends weakly on the emitter positions. We account for the target's orbital velocity by introducing the necessary Doppler compensation. To image a fast rotating object we also need to compensate for the rotation. We show that the rotation parameters can be extracted directly from the auto-correlation of the data before the formation of the image. We then investigate and analyze an imaging method that relies on backpropagating the cross-correlation data structure to two points rather than one, thus forming an interference matrix. The proposed imaging method consists of estimating the reflectivity as the top eigenvector of the migrated cross-correlation data interference matrix. We call this the rank-1 image and show that it provides superior image resolution compared to the usual single-point migration scheme for fast moving and rotating objects. Moreover, we observe a significant improvement in resolution due to the rotation leading to a diffraction limited resolution. We carry out a theoretical analysis that illustrates the role of the two point migration method as well as that of the inverse aperture and rotation in improving resolution. Extensive numerical simulations support the theoretical results.
* arXiv admin note: text overlap with arXiv:2003.00131
Hierarchical Kinematic Probability Distributions for 3D Human Shape and Pose Estimation from Images in the Wild
Oct 03, 2021
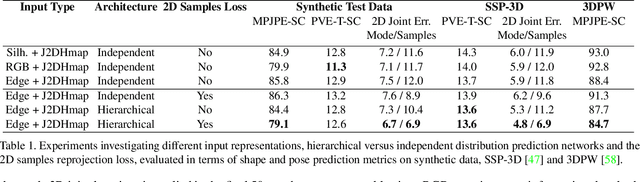
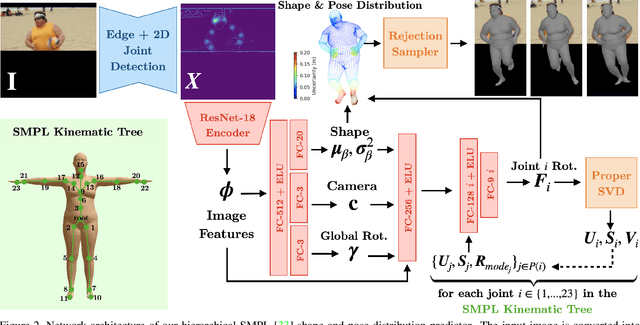
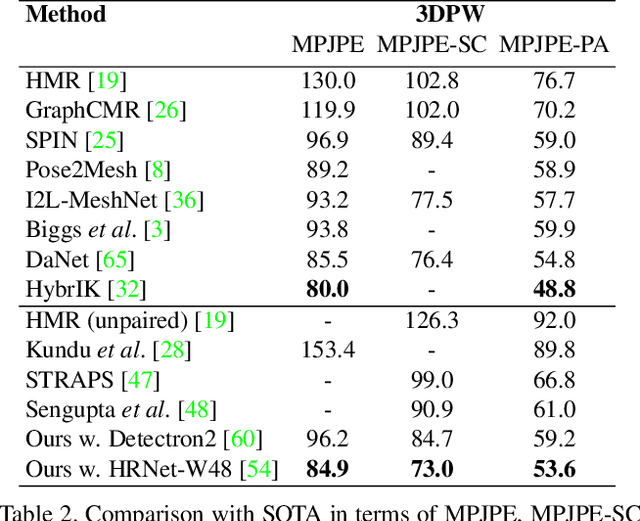
This paper addresses the problem of 3D human body shape and pose estimation from an RGB image. This is often an ill-posed problem, since multiple plausible 3D bodies may match the visual evidence present in the input - particularly when the subject is occluded. Thus, it is desirable to estimate a distribution over 3D body shape and pose conditioned on the input image instead of a single 3D reconstruction. We train a deep neural network to estimate a hierarchical matrix-Fisher distribution over relative 3D joint rotation matrices (i.e. body pose), which exploits the human body's kinematic tree structure, as well as a Gaussian distribution over SMPL body shape parameters. To further ensure that the predicted shape and pose distributions match the visual evidence in the input image, we implement a differentiable rejection sampler to impose a reprojection loss between ground-truth 2D joint coordinates and samples from the predicted distributions, projected onto the image plane. We show that our method is competitive with the state-of-the-art in terms of 3D shape and pose metrics on the SSP-3D and 3DPW datasets, while also yielding a structured probability distribution over 3D body shape and pose, with which we can meaningfully quantify prediction uncertainty and sample multiple plausible 3D reconstructions to explain a given input image. Code is available at https://github.com/akashsengupta1997/HierarchicalProbabilistic3DHuman .
Assisted Probe Positioning for Ultrasound Guided Radiotherapy Using Image Sequence Classification
Oct 06, 2020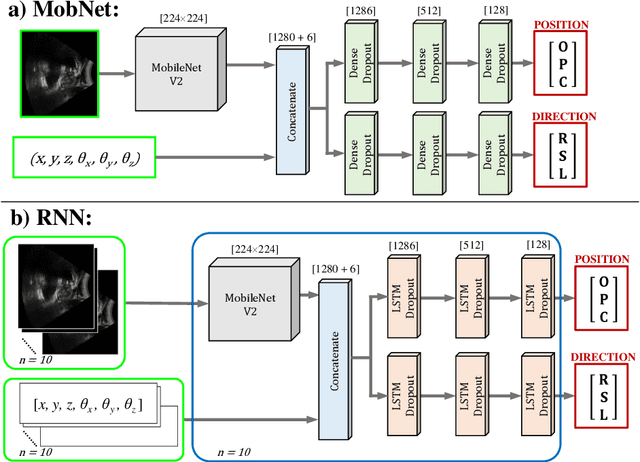
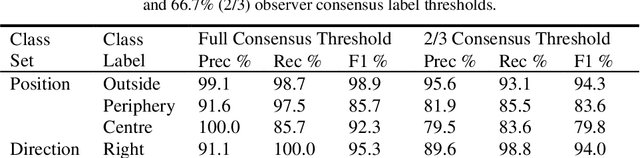
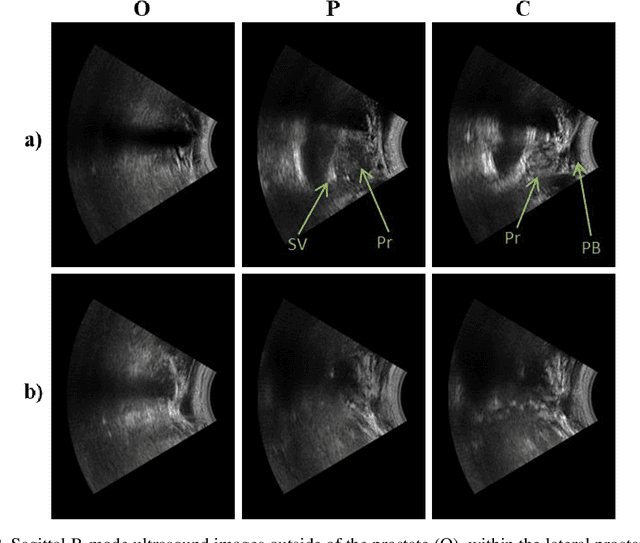
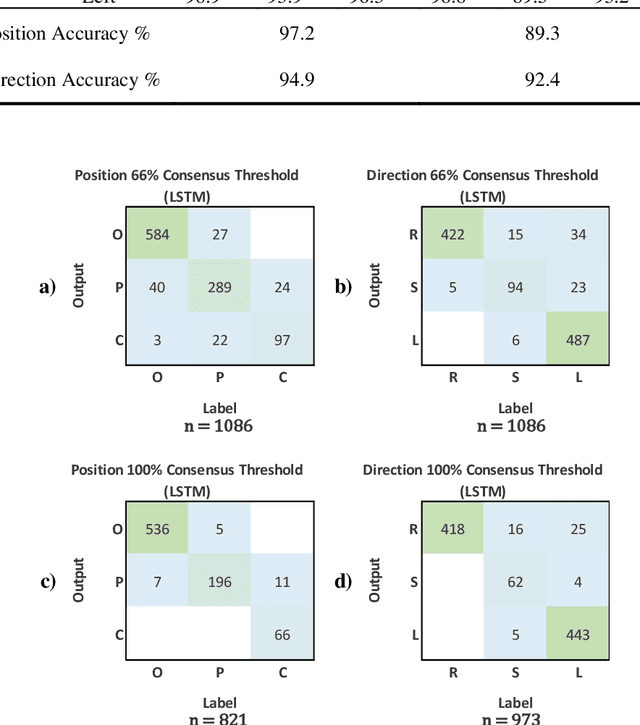
Effective transperineal ultrasound image guidance in prostate external beam radiotherapy requires consistent alignment between probe and prostate at each session during patient set-up. Probe placement and ultrasound image inter-pretation are manual tasks contingent upon operator skill, leading to interoperator uncertainties that degrade radiotherapy precision. We demonstrate a method for ensuring accurate probe placement through joint classification of images and probe position data. Using a multi-input multi-task algorithm, spatial coordinate data from an optically tracked ultrasound probe is combined with an image clas-sifier using a recurrent neural network to generate two sets of predictions in real-time. The first set identifies relevant prostate anatomy visible in the field of view using the classes: outside prostate, prostate periphery, prostate centre. The second set recommends a probe angular adjustment to achieve alignment between the probe and prostate centre with the classes: move left, move right, stop. The algo-rithm was trained and tested on 9,743 clinical images from 61 treatment sessions across 32 patients. We evaluated classification accuracy against class labels de-rived from three experienced observers at 2/3 and 3/3 agreement thresholds. For images with unanimous consensus between observers, anatomical classification accuracy was 97.2% and probe adjustment accuracy was 94.9%. The algorithm identified optimal probe alignment within a mean (standard deviation) range of 3.7$^{\circ}$ (1.2$^{\circ}$) from angle labels with full observer consensus, comparable to the 2.8$^{\circ}$ (2.6$^{\circ}$) mean interobserver range. We propose such an algorithm could assist ra-diotherapy practitioners with limited experience of ultrasound image interpreta-tion by providing effective real-time feedback during patient set-up.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge