"Image": models, code, and papers
Introduction to Medical Image Registration with DeepReg, Between Old and New
Sep 07, 2020
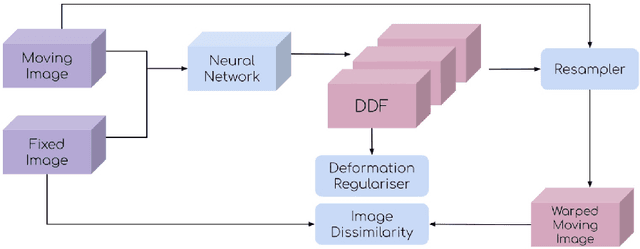
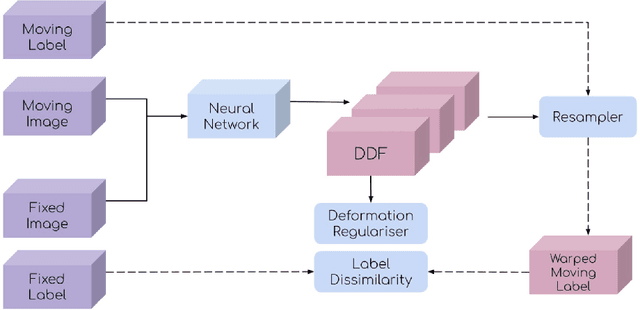
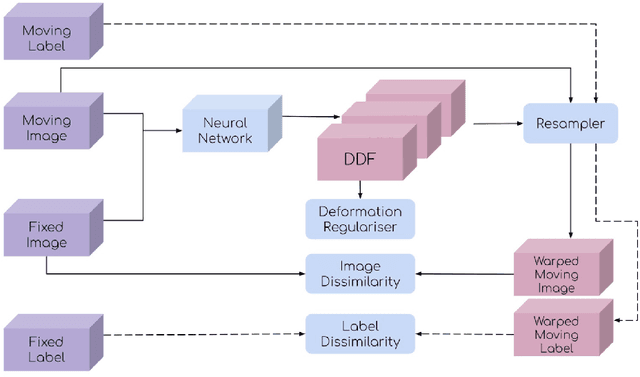
This document outlines a tutorial to get started with medical image registration using the open-source package DeepReg. The basic concepts of medical image registration are discussed, linking classical methods to newer methods using deep learning. Two iterative, classical algorithms using optimisation and one learning-based algorithm using deep learning are coded step-by-step using DeepReg utilities, all with real, open-accessible, medical data.
Algorithm for recognizing the contour of a honeycomb block
Dec 27, 2021The article discusses an algorithm for recognizing the contour of fragments of a honeycomb block. The inapplicability of ready-made functions of the OpenCV library is shown. Two proposed algorithms are considered. The direct scanning algorithm finds the extreme white pixels in the binarized image, it works adequately on convex shapes of products, but does not find a contour on concave areas and in cavities of products. To solve this problem, a scanning algorithm using a sliding matrix is proposed, which works correctly on products of any shape.
Efficient and Robust Classification for Sparse Attacks
Jan 23, 2022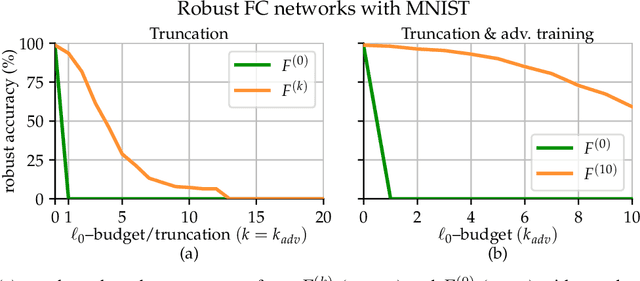
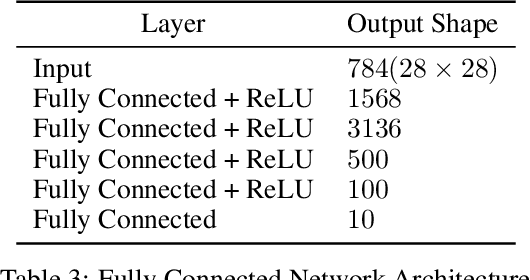
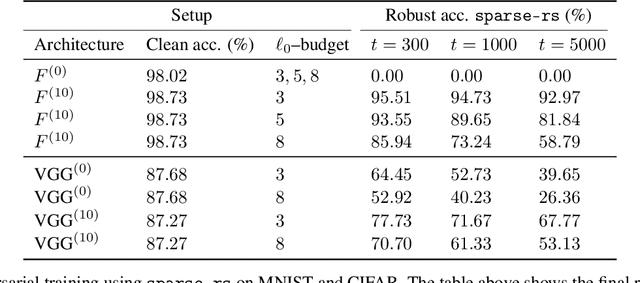
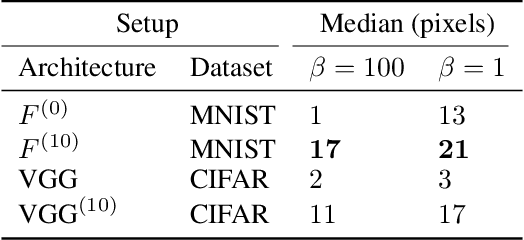

In the past two decades we have seen the popularity of neural networks increase in conjunction with their classification accuracy. Parallel to this, we have also witnessed how fragile the very same prediction models are: tiny perturbations to the inputs can cause misclassification errors throughout entire datasets. In this paper, we consider perturbations bounded by the $\ell_0$--norm, which have been shown as effective attacks in the domains of image-recognition, natural language processing, and malware-detection. To this end, we propose a novel defense method that consists of "truncation" and "adversarial training". We then theoretically study the Gaussian mixture setting and prove the asymptotic optimality of our proposed classifier. Motivated by the insights we obtain, we extend these components to neural network classifiers. We conduct numerical experiments in the domain of computer vision using the MNIST and CIFAR datasets, demonstrating significant improvement for the robust classification error of neural networks.
Swin UNETR: Swin Transformers for Semantic Segmentation of Brain Tumors in MRI Images
Jan 04, 2022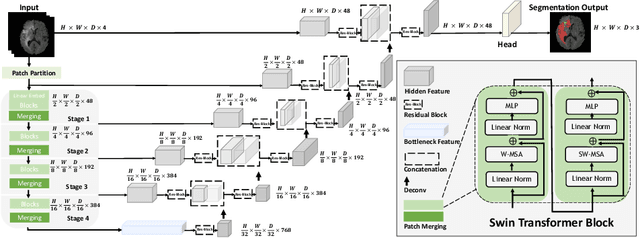

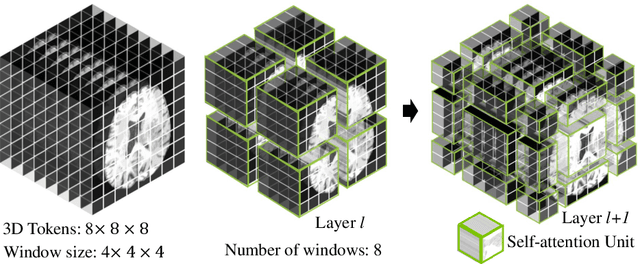
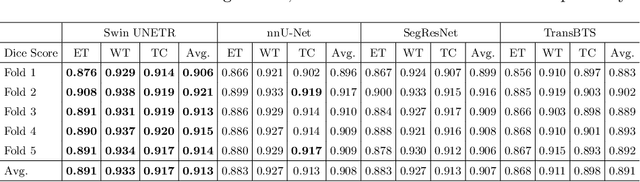
Semantic segmentation of brain tumors is a fundamental medical image analysis task involving multiple MRI imaging modalities that can assist clinicians in diagnosing the patient and successively studying the progression of the malignant entity. In recent years, Fully Convolutional Neural Networks (FCNNs) approaches have become the de facto standard for 3D medical image segmentation. The popular "U-shaped" network architecture has achieved state-of-the-art performance benchmarks on different 2D and 3D semantic segmentation tasks and across various imaging modalities. However, due to the limited kernel size of convolution layers in FCNNs, their performance of modeling long-range information is sub-optimal, and this can lead to deficiencies in the segmentation of tumors with variable sizes. On the other hand, transformer models have demonstrated excellent capabilities in capturing such long-range information in multiple domains, including natural language processing and computer vision. Inspired by the success of vision transformers and their variants, we propose a novel segmentation model termed Swin UNEt TRansformers (Swin UNETR). Specifically, the task of 3D brain tumor semantic segmentation is reformulated as a sequence to sequence prediction problem wherein multi-modal input data is projected into a 1D sequence of embedding and used as an input to a hierarchical Swin transformer as the encoder. The swin transformer encoder extracts features at five different resolutions by utilizing shifted windows for computing self-attention and is connected to an FCNN-based decoder at each resolution via skip connections. We have participated in BraTS 2021 segmentation challenge, and our proposed model ranks among the top-performing approaches in the validation phase. Code: https://monai.io/research/swin-unetr
Deep Image Compression using Decoder Side Information
Jan 14, 2020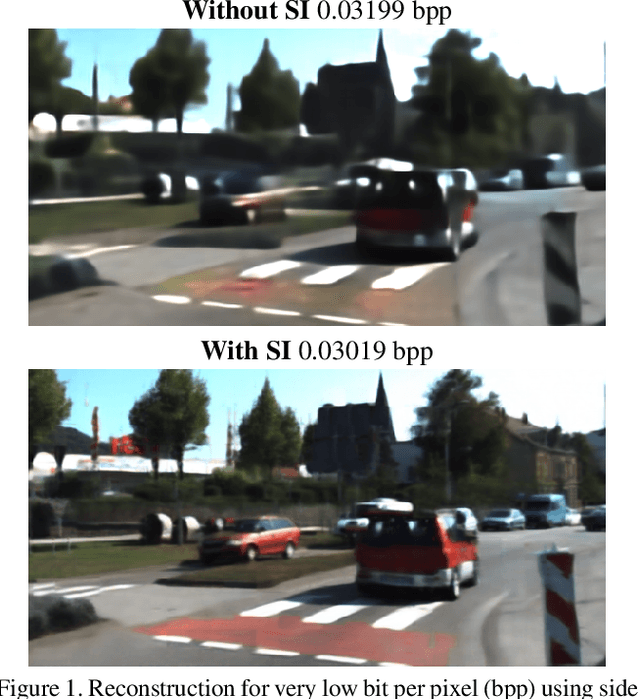
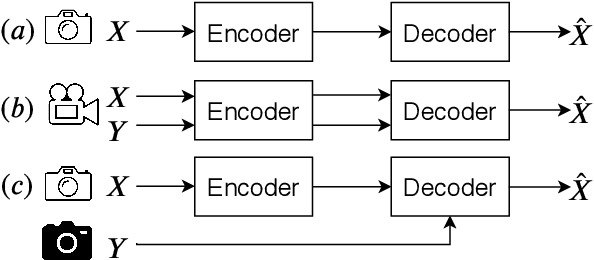
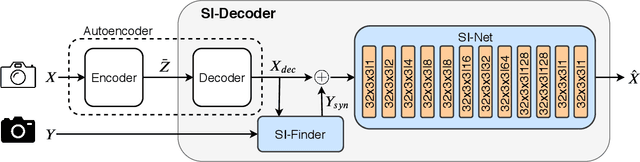
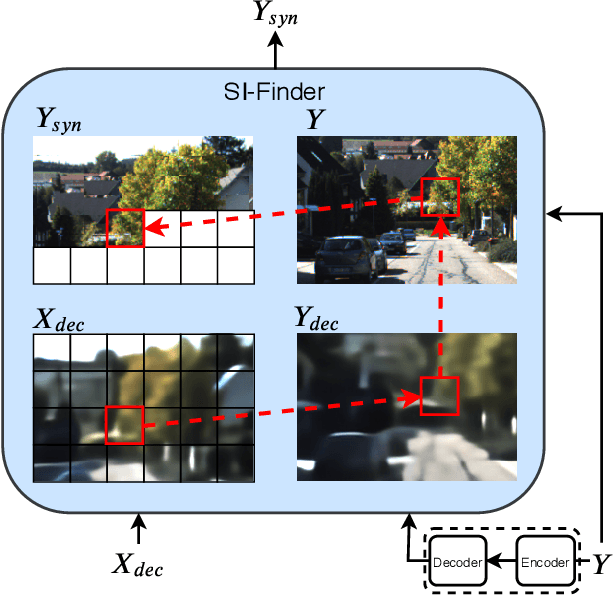
We present a Deep Image Compression neural network that relies on side information, which is only available to the decoder. We base our algorithm on the assumption that the image available to the encoder and the image available to the decoder are correlated, and we let the network learn these correlations in the training phase. Then, at run time, the encoder side encodes the input image without knowing anything about the decoder side image and sends it to the decoder. The decoder then uses the encoded input image and the side information image to reconstruct the original image. This problem is known as Distributed Source Coding in Information Theory, and we discuss several use cases for this technology. We compare our algorithm to several image compression algorithms and show that adding decoder-only side information does indeed improve results. Our code is publicly available at https://github.com/ayziksha/DSIN.
Bubble identification from images with machine learning methods
Feb 07, 2022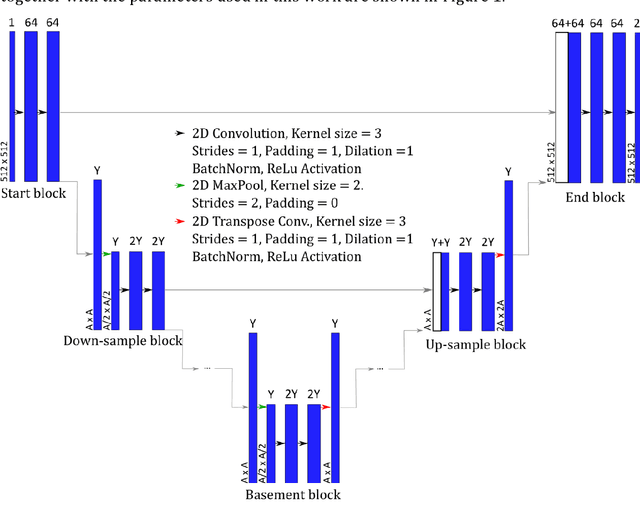
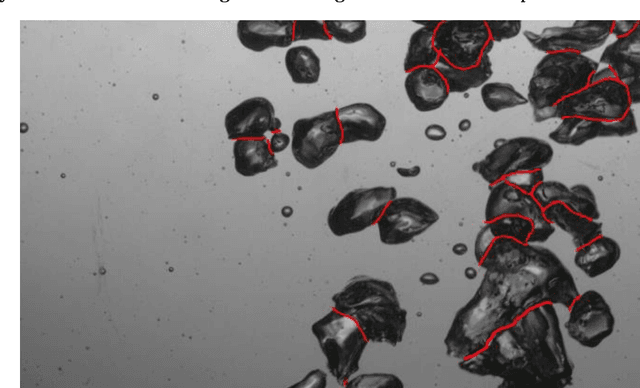
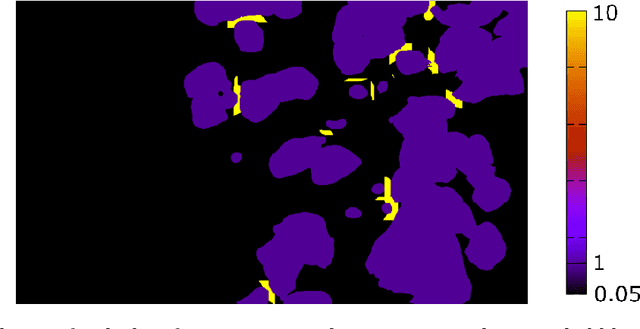
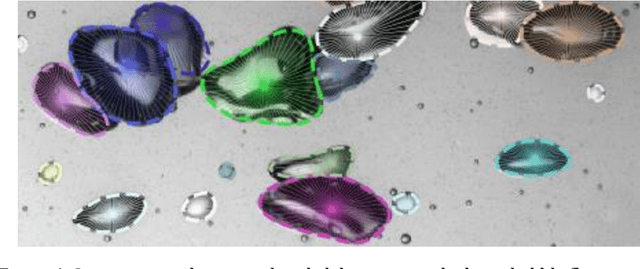
An automated and reliable processing of bubbly flow images is highly needed to analyse large data sets of comprehensive experimental series. A particular difficulty arises due to overlapping bubble projections in recorded images, which highly complicates the identification of individual bubbles. Recent approaches focus on the use of deep learning algorithms for this task and have already proven the high potential of such techniques. The main difficulties are the capability to handle different image conditions, higher gas volume fractions and a proper reconstruction of the hidden segment of a partly occluded bubble. In the present work, we try to tackle these points by testing three different methods based on Convolutional Neural Networks (CNNs) for the two former and two individual approaches that can be used subsequently to address the latter. To validate our methodology, we created test data sets with synthetic images that further demonstrate the capabilities as well as limitations of our combined approach. The generated data, code and trained models are made accessible to facilitate the use as well as further developments in the research field of bubble recognition in experimental images.
AI-Based Detection, Classification and Prediction/Prognosis in Medical Imaging: Towards Radiophenomics
Nov 01, 2021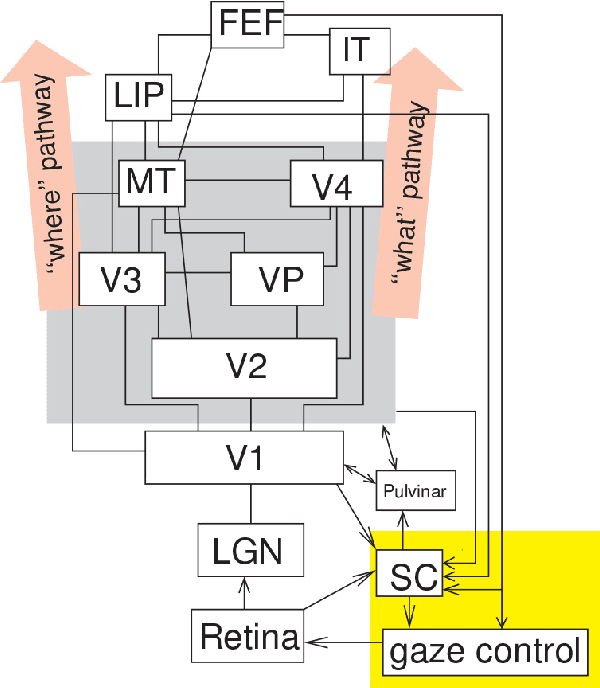
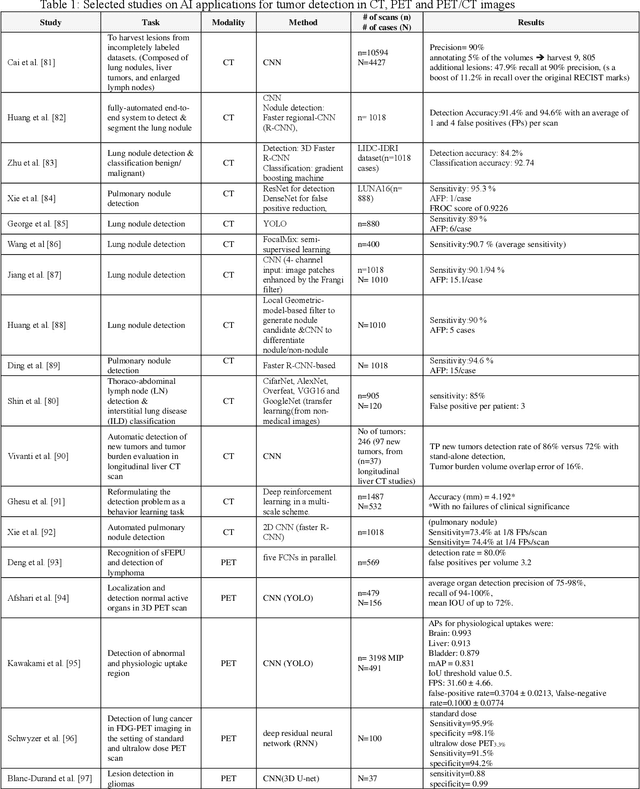

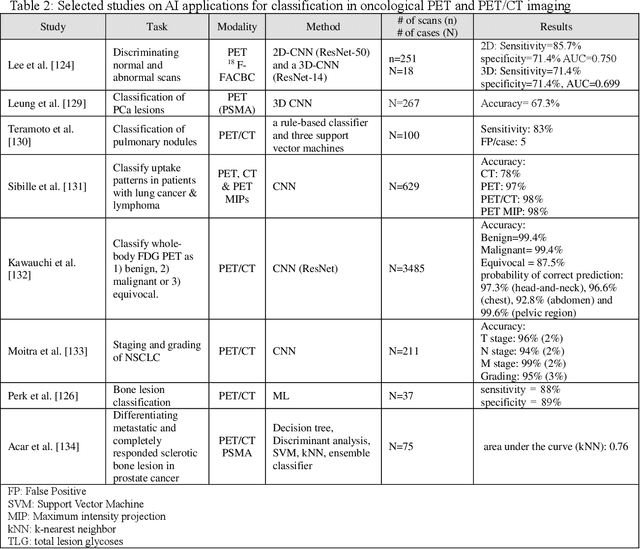
Artificial intelligence (AI) techniques have significant potential to enable effective, robust and automated image phenotyping including identification of subtle patterns. AI-based detection searches the image space to find the regions of interest based on patterns and features. There is a spectrum of tumor histologies from benign to malignant that can be identified by AI-based classification approaches using image features. The extraction of minable information from images gives way to the field of radiomics and can be explored via explicit (handcrafted/engineered) and deep radiomics frameworks. Radiomics analysis has the potential to be utilized as a noninvasive technique for the accurate characterization of tumors to improve diagnosis and treatment monitoring. This work reviews AI-based techniques, with a special focus on oncological PET and PET/CT imaging, for different detection, classification, and prediction/prognosis tasks. We also discuss needed efforts to enable the translation of AI techniques to routine clinical workflows, and potential improvements and complementary techniques such as the use of natural language processing on electronic health records and neuro-symbolic AI techniques.
RADA: Robust Adversarial Data Augmentation for Camera Localization in Challenging Weather
Dec 05, 2021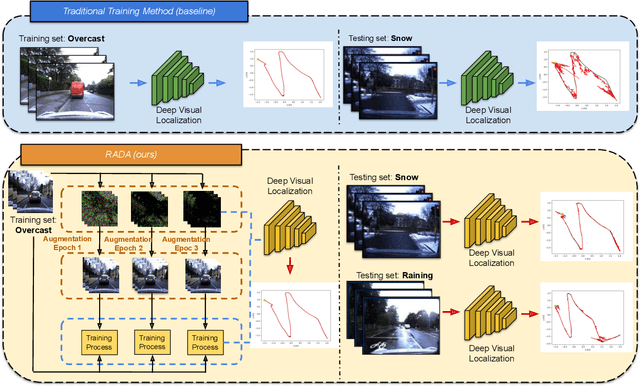
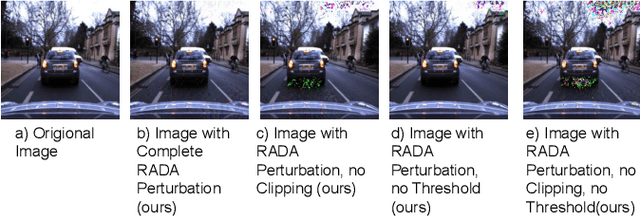
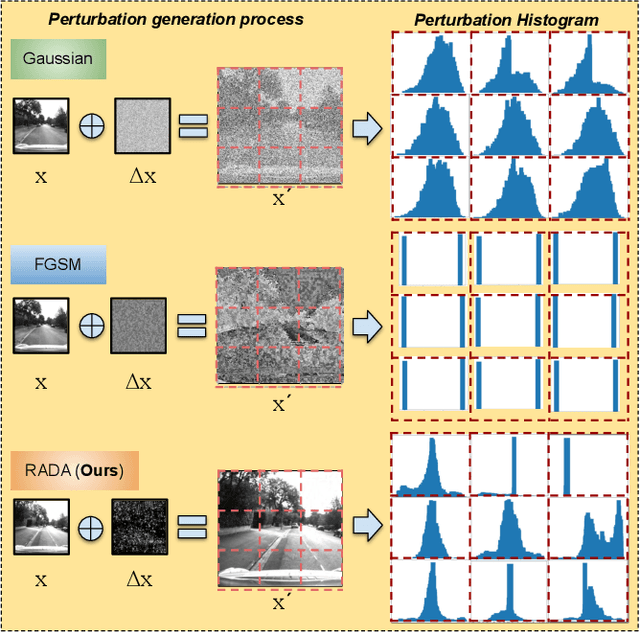
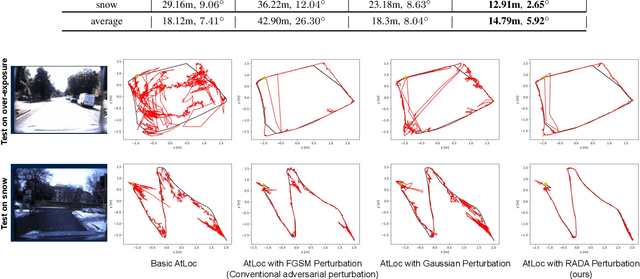

Camera localization is a fundamental and crucial problem for many robotic applications. In recent years, using deep-learning for camera-based localization has become a popular research direction. However, they lack robustness to large domain shifts, which can be caused by seasonal or illumination changes between training and testing data sets. Data augmentation is an attractive approach to tackle this problem, as it does not require additional data to be provided. However, existing augmentation methods blindly perturb all pixels and therefore cannot achieve satisfactory performance. To overcome this issue, we proposed RADA, a system whose aim is to concentrate on perturbing the geometrically informative parts of the image. As a result, it learns to generate minimal image perturbations that are still capable of perplexing the network. We show that when these examples are utilized as augmentation, it greatly improves robustness. We show that our method outperforms previous augmentation techniques and achieves up to two times higher accuracy than the SOTA localization models (e.g., AtLoc and MapNet) when tested on `unseen' challenging weather conditions.
PT-VTON: an Image-Based Virtual Try-On Network with Progressive Pose Attention Transfer
Nov 23, 2021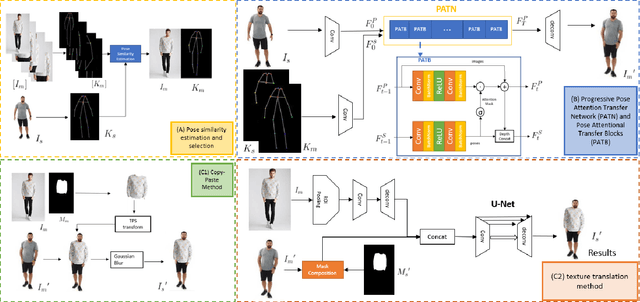
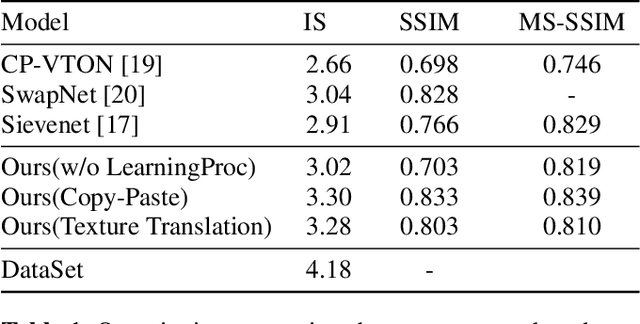
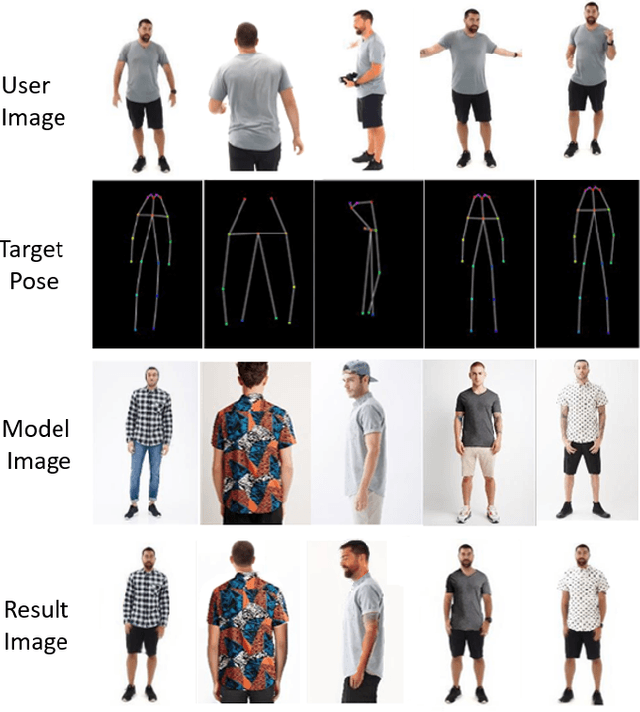

The virtual try-on system has gained great attention due to its potential to give customers a realistic, personalized product presentation in virtualized settings. In this paper, we present PT-VTON, a novel pose-transfer-based framework for cloth transfer that enables virtual try-on with arbitrary poses. PT-VTON can be applied to the fashion industry within minimal modification of existing systems while satisfying the overall visual fashionability and detailed fabric appearance requirements. It enables efficient clothes transferring between model and user images with arbitrary pose and body shape. We implement a prototype of PT-VTON and demonstrate that our system can match or surpass many other approaches when facing a drastic variation of poses by preserving detailed human and fabric characteristic appearances. PT-VTON is shown to outperform alternative approaches both on machine-based quantitative metrics and qualitative results.
An Ultra Fast Low Power Convolutional Neural Network Image Sensor with Pixel-level Computing
Jan 09, 2021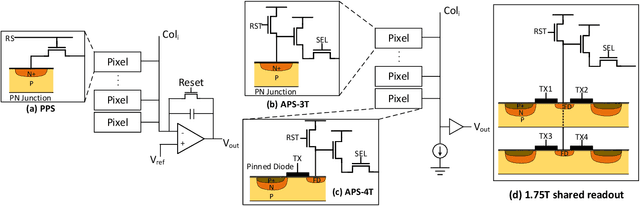
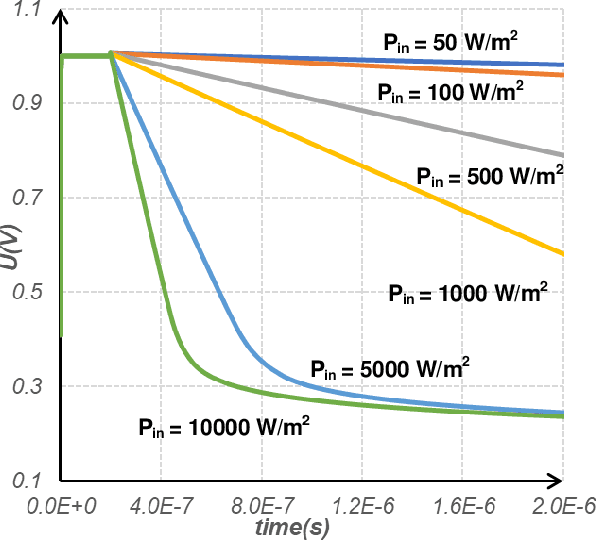
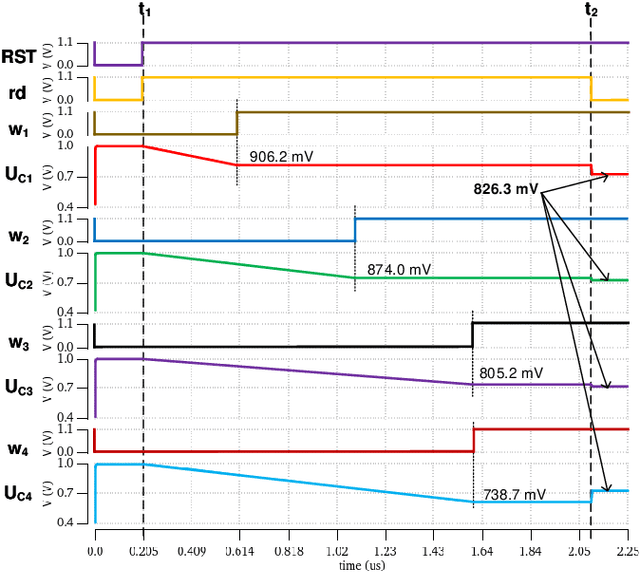
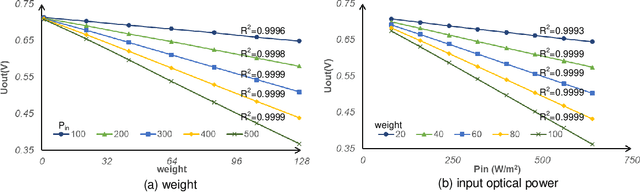
The separation of the data capture and analysis in modern vision systems has led to a massive amount of data transfer between the end devices and cloud computers, resulting in long latency, slow response, and high power consumption. Efficient hardware architectures are under focused development to enable Artificial Intelligence (AI) at the resource-limited end sensing devices. This paper proposes a Processing-In-Pixel (PIP) CMOS sensor architecture, which allows convolution operation before the column readout circuit to significantly improve the image reading speed with much lower power consumption. The simulation results show that the proposed architecture enables convolution operation (kernel size=3*3, stride=2, input channel=3, output channel=64) in a 1080P image sensor array with only 22.62 mW power consumption. In other words, the computational efficiency is 4.75 TOPS/w, which is about 3.6 times as higher as the state-of-the-art.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge