"Image": models, code, and papers
Predicting 4D Liver MRI for MR-guided Interventions
Feb 25, 2022

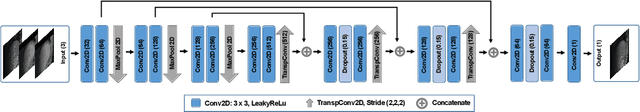
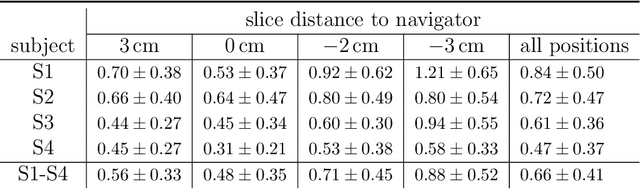
Organ motion poses an unresolved challenge in image-guided interventions. In the pursuit of solving this problem, the research field of time-resolved volumetric magnetic resonance imaging (4D MRI) has evolved. However, current techniques are unsuitable for most interventional settings because they lack sufficient temporal and/or spatial resolution or have long acquisition times. In this work, we propose a novel approach for real-time, high-resolution 4D MRI with large fields of view for MR-guided interventions. To this end, we trained a convolutional neural network (CNN) end-to-end to predict a 3D liver MRI that correctly predicts the liver's respiratory state from a live 2D navigator MRI of a subject. Our method can be used in two ways: First, it can reconstruct near real-time 4D MRI with high quality and high resolution (209x128x128 matrix size with isotropic 1.8mm voxel size and 0.6s/volume) given a dynamic interventional 2D navigator slice for guidance during an intervention. Second, it can be used for retrospective 4D reconstruction with a temporal resolution of below 0.2s/volume for motion analysis and use in radiation therapy. We report a mean target registration error (TRE) of 1.19 $\pm$0.74mm, which is below voxel size. We compare our results with a state-of-the-art retrospective 4D MRI reconstruction. Visual evaluation shows comparable quality. We show that small training sizes with short acquisition times down to 2min can already achieve promising results and 24min are sufficient for high quality results. Because our method can be readily combined with earlier methods, acquisition time can be further decreased while also limiting quality loss. We show that an end-to-end, deep learning formulation is highly promising for 4D MRI reconstruction.
Deep-CNN based Robotic Multi-Class Under-Canopy Weed Control in Precision Farming
Dec 28, 2021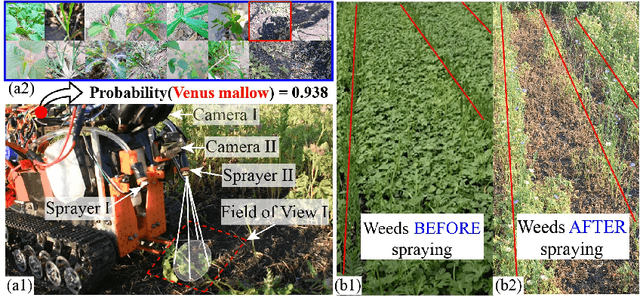
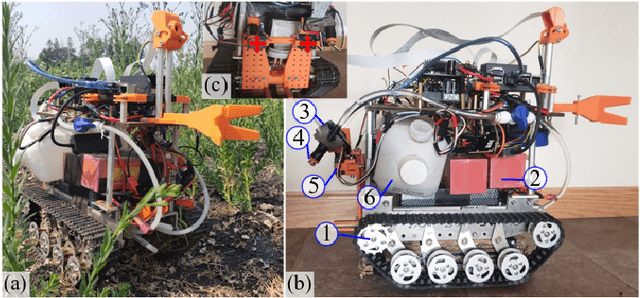


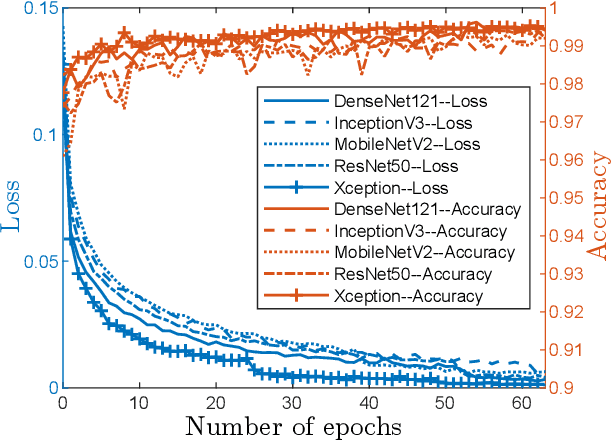
Smart weeding systems to perform plant-specific operations can contribute to the sustainability of agriculture and the environment. Despite monumental advances in autonomous robotic technologies for precision weed management in recent years, work on under-canopy weeding in fields is yet to be realized. A prerequisite of such systems is reliable detection and classification of weeds to avoid mistakenly spraying and, thus, damaging the surrounding plants. Real-time multi-class weed identification enables species-specific treatment of weeds and significantly reduces the amount of herbicide use. Here, our first contribution is the first adequately large realistic image dataset \textit{AIWeeds} (one/multiple kinds of weeds in one image), a library of about 10,000 annotated images of flax, and the 14 most common weeds in fields and gardens taken from 20 different locations in North Dakota, California, and Central China. Second, we provide a full pipeline from model training with maximum efficiency to deploying the TensorRT-optimized model onto a single board computer. Based on \textit{AIWeeds} and the pipeline, we present a baseline for classification performance using five benchmark CNN models. Among them, MobileNetV2, with both the shortest inference time and lowest memory consumption, is the qualified candidate for real-time applications. Finally, we deploy MobileNetV2 onto our own compact autonomous robot \textit{SAMBot} for real-time weed detection. The 90\% test accuracy realized in previously unseen scenes in flax fields (with a row spacing of 0.2-0.3 m), with crops and weeds, distortion, blur, and shadows, is a milestone towards precision weed control in the real world. We have publicly released the dataset and code to generate the results at \url{https://github.com/StructuresComp/Multi-class-Weed-Classification}.
Unpaired Image Super-Resolution using Pseudo-Supervision
Feb 26, 2020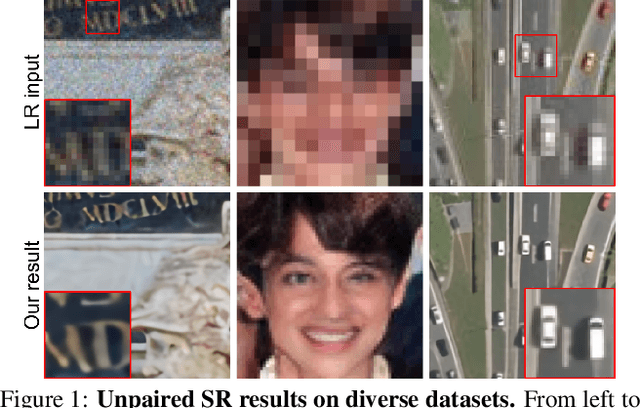
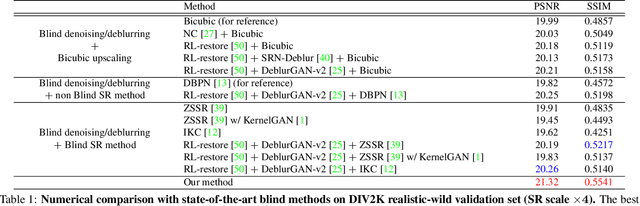

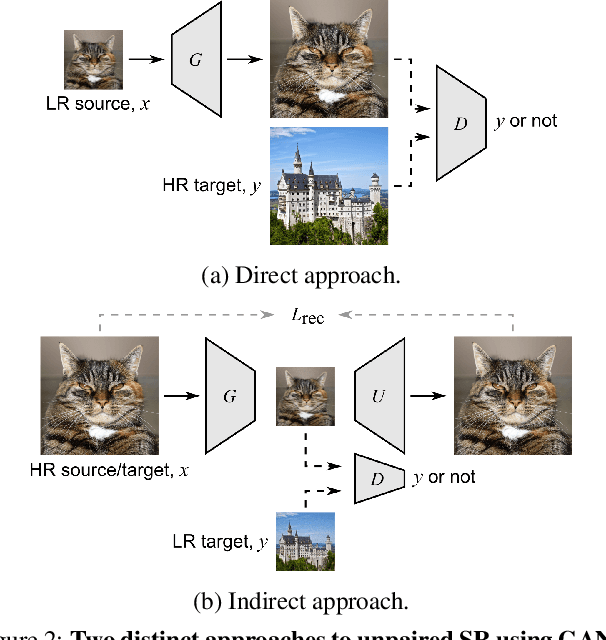
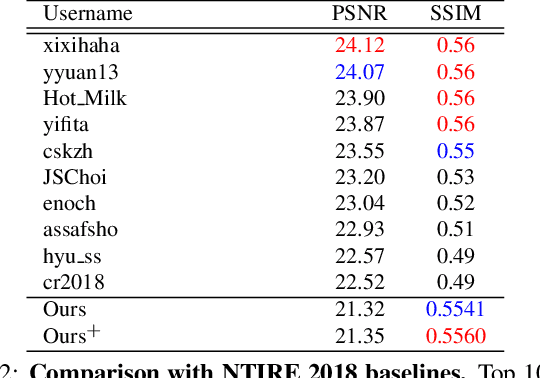
In most studies on learning-based image super-resolution (SR), the paired training dataset is created by downscaling high-resolution (HR) images with a predetermined operation (e.g., bicubic). However, these methods fail to super-resolve real-world low-resolution (LR) images, for which the degradation process is much more complicated and unknown. In this paper, we propose an unpaired SR method using a generative adversarial network that does not require a paired/aligned training dataset. Our network consists of an unpaired kernel/noise correction network and a pseudo-paired SR network. The correction network removes noise and adjusts the kernel of the inputted LR image; then, the corrected clean LR image is upscaled by the SR network. In the training phase, the correction network also produces a pseudo-clean LR image from the inputted HR image, and then a mapping from the pseudo-clean LR image to the inputted HR image is learned by the SR network in a paired manner. Because our SR network is independent of the correction network, well-studied existing network architectures and pixel-wise loss functions can be integrated with the proposed framework. Experiments on diverse datasets show that the proposed method is superior to existing solutions to the unpaired SR problem.
Assisted Probe Positioning for Ultrasound Guided Radiotherapy Using Image Sequence Classification
Oct 06, 2020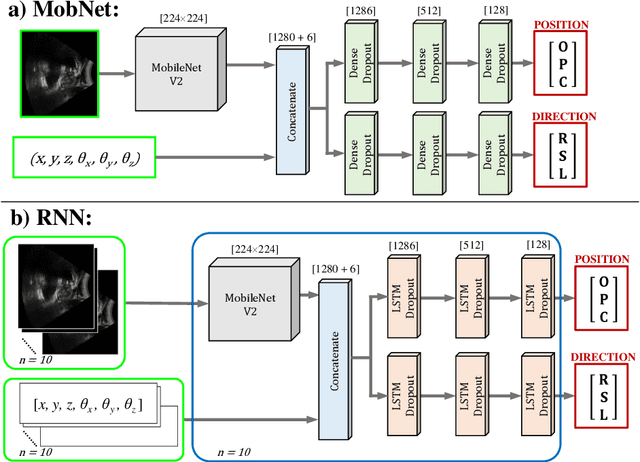
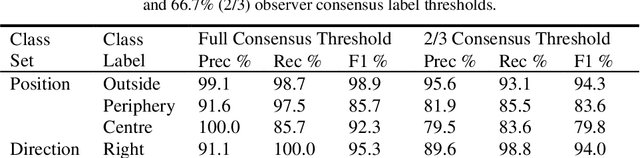
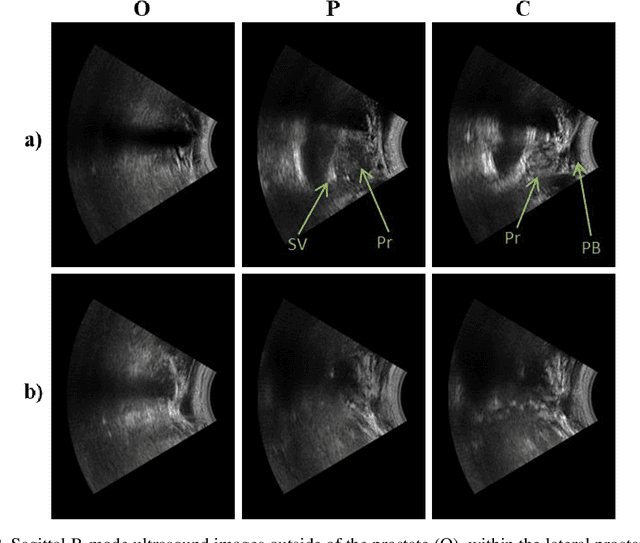
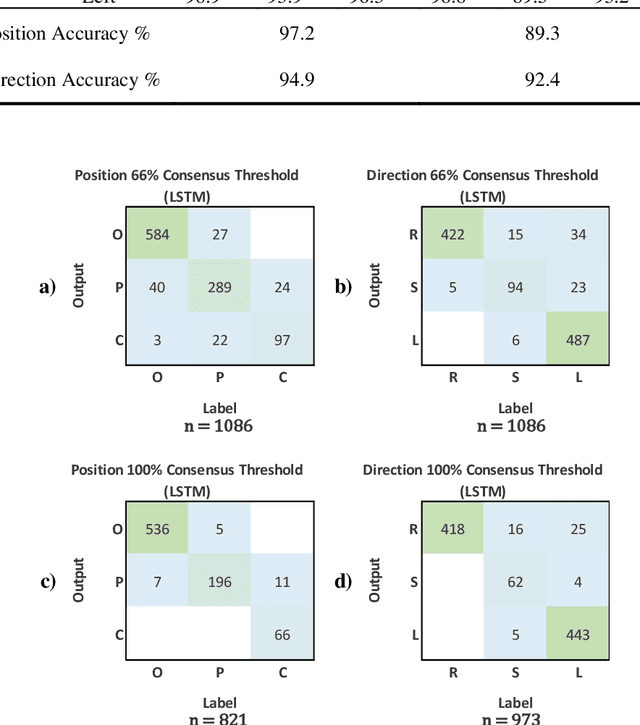
Effective transperineal ultrasound image guidance in prostate external beam radiotherapy requires consistent alignment between probe and prostate at each session during patient set-up. Probe placement and ultrasound image inter-pretation are manual tasks contingent upon operator skill, leading to interoperator uncertainties that degrade radiotherapy precision. We demonstrate a method for ensuring accurate probe placement through joint classification of images and probe position data. Using a multi-input multi-task algorithm, spatial coordinate data from an optically tracked ultrasound probe is combined with an image clas-sifier using a recurrent neural network to generate two sets of predictions in real-time. The first set identifies relevant prostate anatomy visible in the field of view using the classes: outside prostate, prostate periphery, prostate centre. The second set recommends a probe angular adjustment to achieve alignment between the probe and prostate centre with the classes: move left, move right, stop. The algo-rithm was trained and tested on 9,743 clinical images from 61 treatment sessions across 32 patients. We evaluated classification accuracy against class labels de-rived from three experienced observers at 2/3 and 3/3 agreement thresholds. For images with unanimous consensus between observers, anatomical classification accuracy was 97.2% and probe adjustment accuracy was 94.9%. The algorithm identified optimal probe alignment within a mean (standard deviation) range of 3.7$^{\circ}$ (1.2$^{\circ}$) from angle labels with full observer consensus, comparable to the 2.8$^{\circ}$ (2.6$^{\circ}$) mean interobserver range. We propose such an algorithm could assist ra-diotherapy practitioners with limited experience of ultrasound image interpreta-tion by providing effective real-time feedback during patient set-up.
Learning Deep Image Priors for Blind Image Denoising
Jun 04, 2019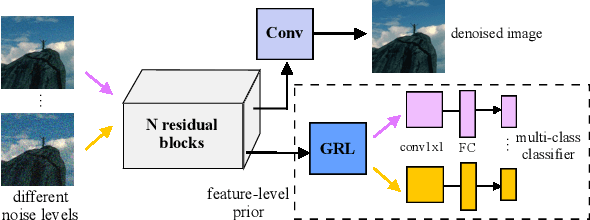
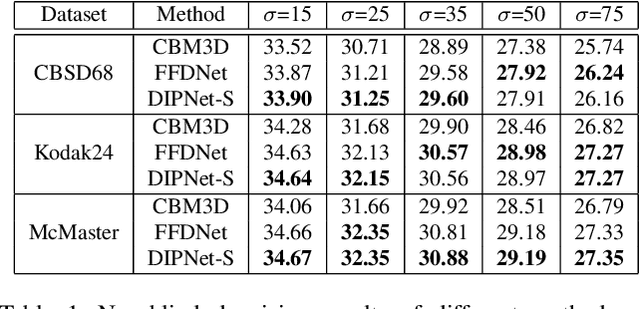
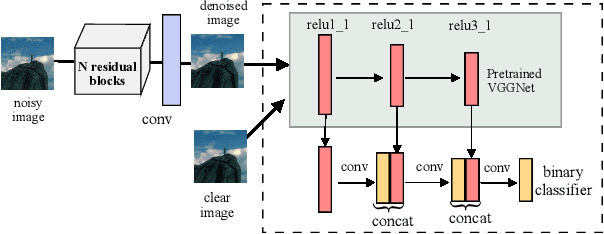
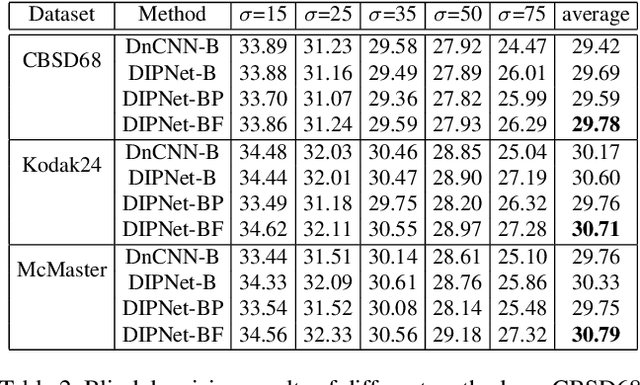
Image denoising is the process of removing noise from noisy images, which is an image domain transferring task, i.e., from a single or several noise level domains to a photo-realistic domain. In this paper, we propose an effective image denoising method by learning two image priors from the perspective of domain alignment. We tackle the domain alignment on two levels. 1) the feature-level prior is to learn domain-invariant features for corrupted images with different level noise; 2) the pixel-level prior is used to push the denoised images to the natural image manifold. The two image priors are based on $\mathcal{H}$-divergence theory and implemented by learning classifiers in adversarial training manners. We evaluate our approach on multiple datasets. The results demonstrate the effectiveness of our approach for robust image denoising on both synthetic and real-world noisy images. Furthermore, we show that the feature-level prior is capable of alleviating the discrepancy between different level noise. It can be used to improve the blind denoising performance in terms of distortion measures (PSNR and SSIM), while pixel-level prior can effectively improve the perceptual quality to ensure the realistic outputs, which is further validated by subjective evaluation.
Semantic Similarity Computing Model Based on Multi Model Fine-Grained Nonlinear Fusion
Feb 05, 2022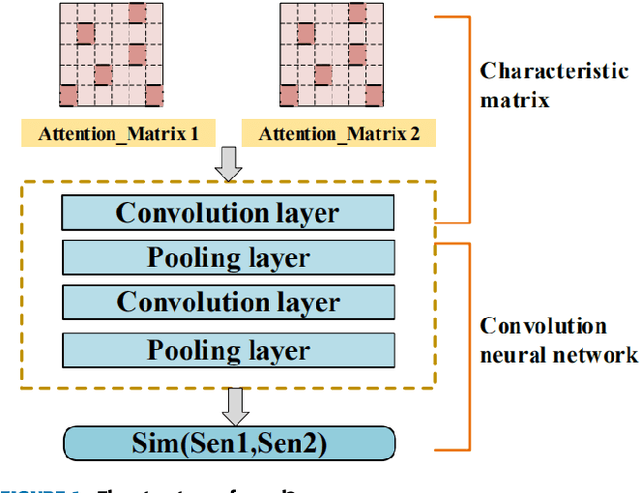
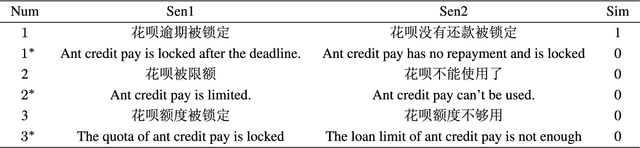

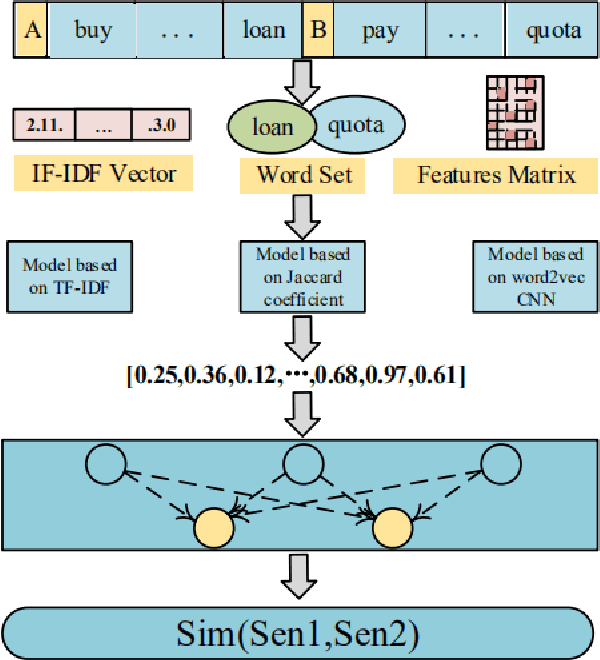
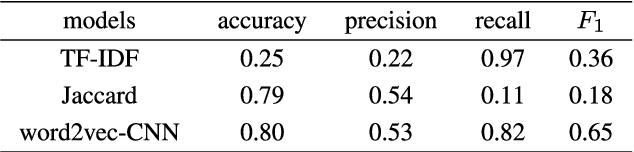
Natural language processing (NLP) task has achieved excellent performance in many fields, including semantic understanding, automatic summarization, image recognition and so on. However, most of the neural network models for NLP extract the text in a fine-grained way, which is not conducive to grasp the meaning of the text from a global perspective. To alleviate the problem, the combination of the traditional statistical method and deep learning model as well as a novel model based on multi model nonlinear fusion are proposed in this paper. The model uses the Jaccard coefficient based on part of speech, Term Frequency-Inverse Document Frequency (TF-IDF) and word2vec-CNN algorithm to measure the similarity of sentences respectively. According to the calculation accuracy of each model, the normalized weight coefficient is obtained and the calculation results are compared. The weighted vector is input into the fully connected neural network to give the final classification results. As a result, the statistical sentence similarity evaluation algorithm reduces the granularity of feature extraction, so it can grasp the sentence features globally. Experimental results show that the matching of sentence similarity calculation method based on multi model nonlinear fusion is 84%, and the F1 value of the model is 75%.
MT-TransUNet: Mediating Multi-Task Tokens in Transformers for Skin Lesion Segmentation and Classification
Dec 03, 2021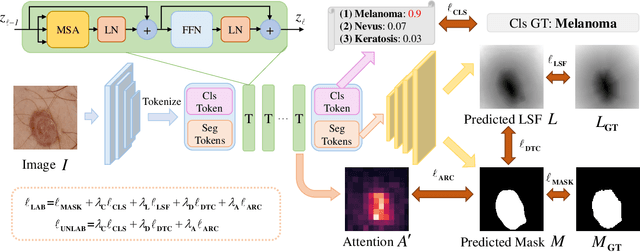
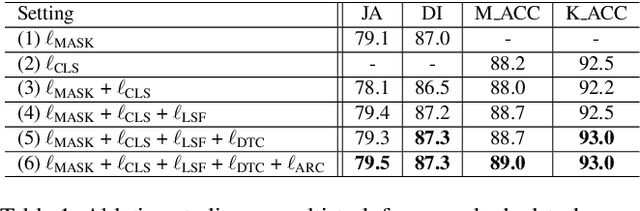
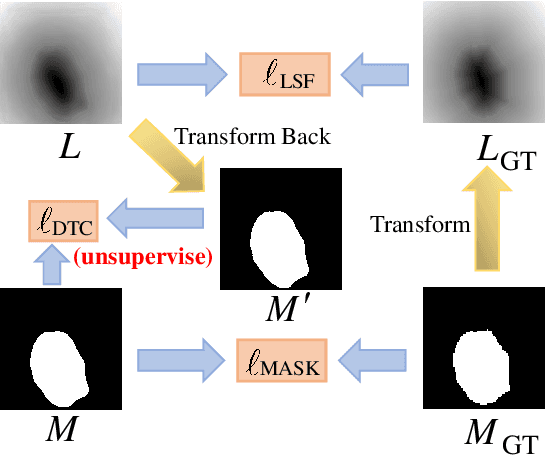
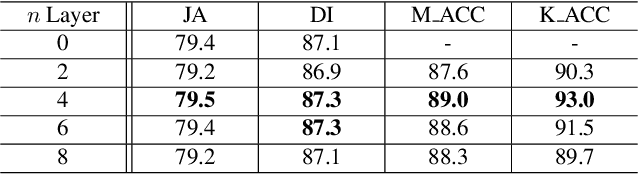
Recent advances in automated skin cancer diagnosis have yielded performance on par with board-certified dermatologists. However, these approaches formulated skin cancer diagnosis as a simple classification task, dismissing the potential benefit from lesion segmentation. We argue that an accurate lesion segmentation can supplement the classification task with additive lesion information, such as asymmetry, border, intensity, and physical size; in turn, a faithful lesion classification can support the segmentation task with discriminant lesion features. To this end, this paper proposes a new multi-task framework, named MT-TransUNet, which is capable of segmenting and classifying skin lesions collaboratively by mediating multi-task tokens in Transformers. Furthermore, we have introduced dual-task and attended region consistency losses to take advantage of those images without pixel-level annotation, ensuring the model's robustness when it encounters the same image with an account of augmentation. Our MT-TransUNet exceeds the previous state of the art for lesion segmentation and classification tasks in ISIC-2017 and PH2; more importantly, it preserves compelling computational efficiency regarding model parameters (48M~vs.~130M) and inference speed (0.17s~vs.~2.02s per image). Code will be available at https://github.com/JingyeChen/MT-TransUNet.
SAGAN: Adversarial Spatial-asymmetric Attention for Noisy Nona-Bayer Reconstruction
Oct 19, 2021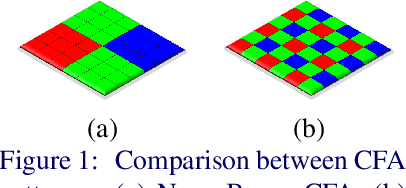
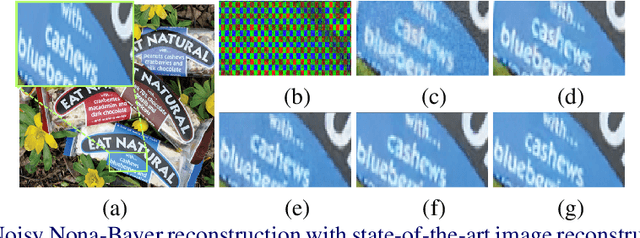
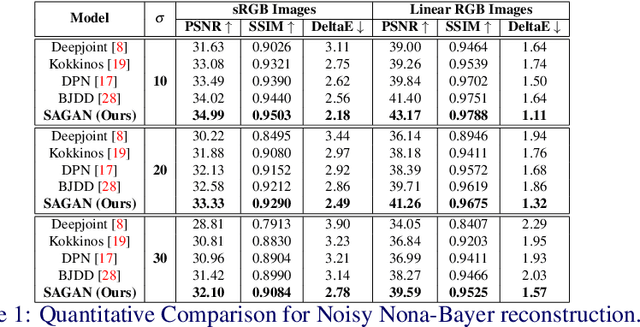

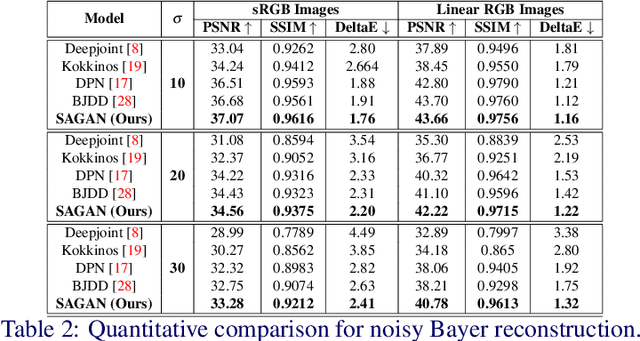
Nona-Bayer colour filter array (CFA) pattern is considered one of the most viable alternatives to traditional Bayer patterns. Despite the substantial advantages, such non-Bayer CFA patterns are susceptible to produce visual artefacts while reconstructing RGB images from noisy sensor data. This study addresses the challenges of learning RGB image reconstruction from noisy Nona-Bayer CFA comprehensively. We propose a novel spatial-asymmetric attention module to jointly learn bi-direction transformation and large-kernel global attention to reduce the visual artefacts. We combine our proposed module with adversarial learning to produce plausible images from Nona-Bayer CFA. The feasibility of the proposed method has been verified and compared with the state-of-the-art image reconstruction method. The experiments reveal that the proposed method can reconstruct RGB images from noisy Nona-Bayer CFA without producing any visually disturbing artefacts. Also, it can outperform the state-of-the-art image reconstruction method in both qualitative and quantitative comparison. Code available: https://github.com/sharif-apu/SAGAN_BMVC21.
Heuristic Hyperparameter Optimization for Convolutional Neural Networks using Genetic Algorithm
Dec 14, 2021
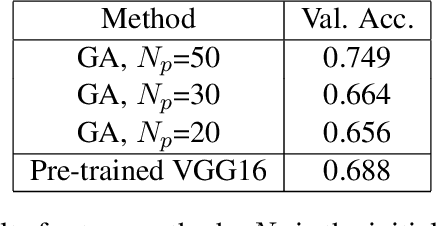
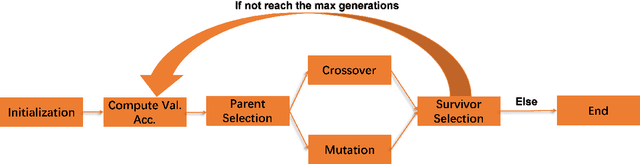

In recent years, people from all over the world are suffering from one of the most severe diseases in history, known as Coronavirus disease 2019, COVID-19 for short. When the virus reaches the lungs, it has a higher probability to cause lung pneumonia and sepsis. X-ray image is a powerful tool in identifying the typical features of the infection for COVID-19 patients. The radiologists and pathologists observe that ground-glass opacity appears in the chest X-ray for infected patient \cite{cozzi2021ground}, and it could be used as one of the criteria during the diagnosis process. In the past few years, deep learning has proven to be one of the most powerful methods in the field of image classification. Due to significant differences in Chest X-Ray between normal and infected people \cite{rousan2020chest}, deep models could be used to identify the presence of the disease given a patient's Chest X-Ray. Many deep models are complex, and it evolves with lots of input parameters. Designers sometimes struggle with the tuning process for deep models, especially when they build up the model from scratch. Genetic Algorithm, inspired by the biological evolution process, plays a key role in solving such complex problems. In this paper, I proposed a genetic-based approach to optimize the Convolutional Neural Network(CNN) for the Chest X-Ray classification task.
Predicting 3D shapes, masks, and properties of materials, liquids, and objects inside transparent containers, using the TransProteus CGI dataset
Sep 15, 2021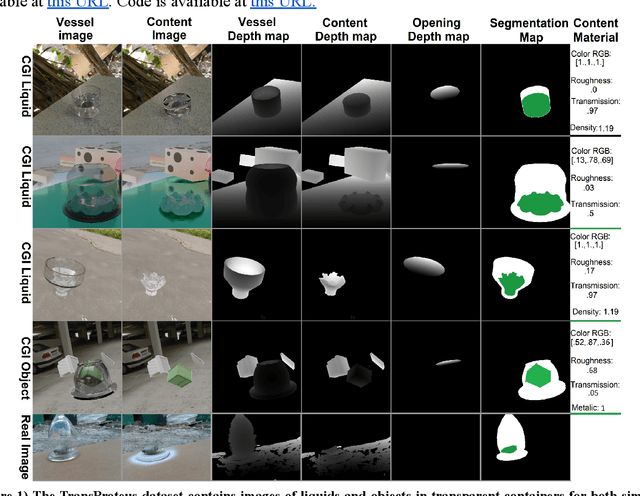
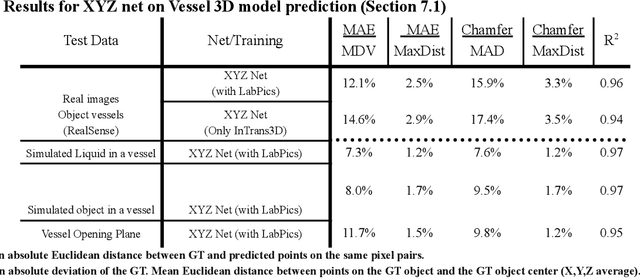
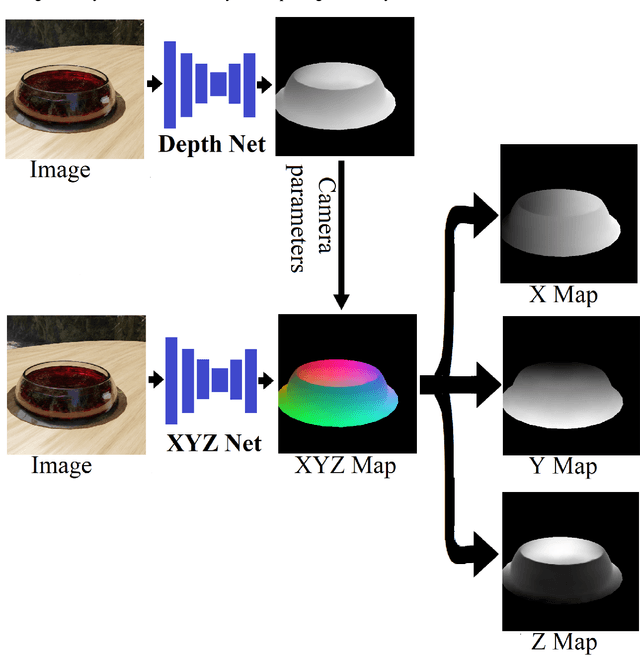
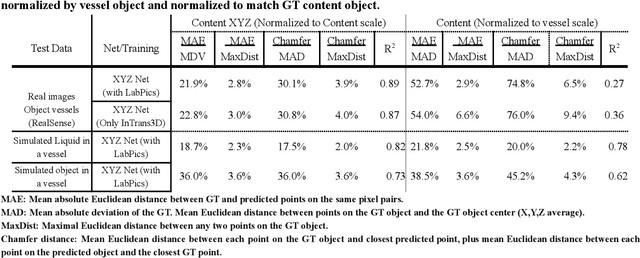
We present TransProteus, a dataset, and methods for predicting the 3D structure, masks, and properties of materials, liquids, and objects inside transparent vessels from a single image without prior knowledge of the image source and camera parameters. Manipulating materials in transparent containers is essential in many fields and depends heavily on vision. This work supplies a new procedurally generated dataset consisting of 50k images of liquids and solid objects inside transparent containers. The image annotations include 3D models, material properties (color/transparency/roughness...), and segmentation masks for the vessel and its content. The synthetic (CGI) part of the dataset was procedurally generated using 13k different objects, 500 different environments (HDRI), and 1450 material textures (PBR) combined with simulated liquids and procedurally generated vessels. In addition, we supply 104 real-world images of objects inside transparent vessels with depth maps of both the vessel and its content. We propose a camera agnostic method that predicts 3D models from an image as an XYZ map. This allows the trained net to predict the 3D model as a map with XYZ coordinates per pixel without prior knowledge of the image source. To calculate the training loss, we use the distance between pairs of points inside the 3D model instead of the absolute XYZ coordinates. This makes the loss function translation invariant. We use this to predict 3D models of vessels and their content from a single image. Finally, we demonstrate a net that uses a single image to predict the material properties of the vessel content and surface.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge