"Image": models, code, and papers
Uncertainty Quantification using Variational Inference for Biomedical Image Segmentation
Aug 12, 2020
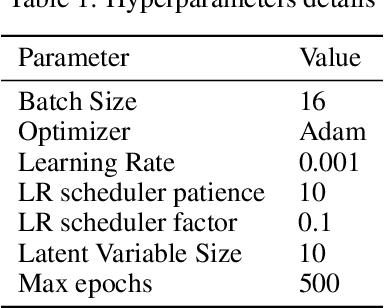
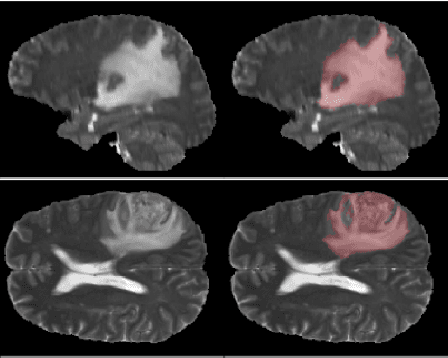
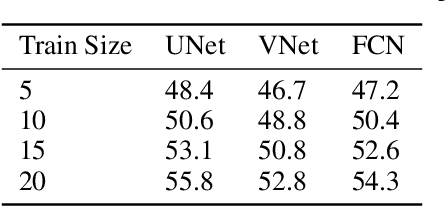
Deep learning motivated by convolutional neural networks has been highly successful in a range of medical imaging problems like image classification, image segmentation, image synthesis etc. However for validation and interpretability, not only do we need the predictions made by the model but also how confident it is while making those predictions. This is important in safety critical applications for the people to accept it. In this work, we used an encoder decoder architecture based on variational inference techniques for segmenting brain tumour images. We compare different backbones architectures like U-Net, V-Net and FCN as sampling data from the conditional distribution for the encoder. We evaluate our work on the publicly available BRATS dataset using Dice Similarity Coefficient (DSC) and Intersection Over Union (IOU) as the evaluation metrics. Our model outperforms previous state of the art results while making use of uncertainty quantification in a principled bayesian manner.
GMM-UNIT: Unsupervised Multi-Domain and Multi-Modal Image-to-Image Translation via Attribute Gaussian Mixture Modeling
Mar 15, 2020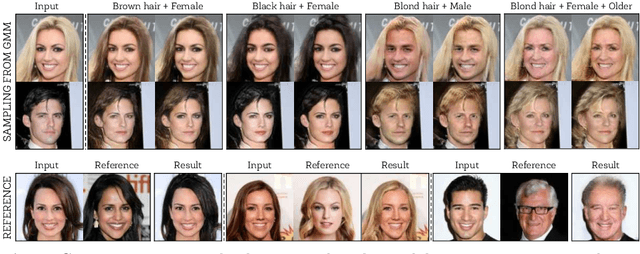
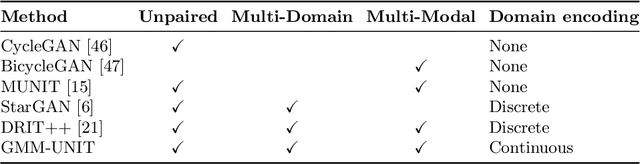

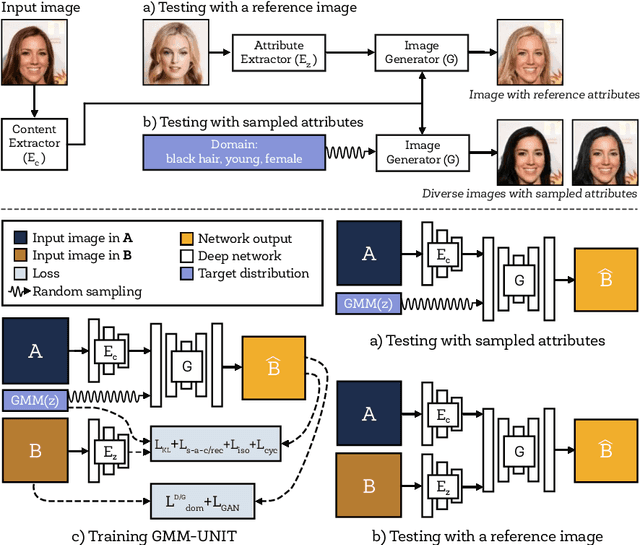

Unsupervised image-to-image translation (UNIT) aims at learning a mapping between several visual domains by using unpaired training images. Recent studies have shown remarkable success for multiple domains but they suffer from two main limitations: they are either built from several two-domain mappings that are required to be learned independently, or they generate low-diversity results, a problem known as model collapse. To overcome these limitations, we propose a method named GMM-UNIT, which is based on a content-attribute disentangled representation where the attribute space is fitted with a GMM. Each GMM component represents a domain, and this simple assumption has two prominent advantages. First, it can be easily extended to most multi-domain and multi-modal image-to-image translation tasks. Second, the continuous domain encoding allows for interpolation between domains and for extrapolation to unseen domains and translations. Additionally, we show how GMM-UNIT can be constrained down to different methods in the literature, meaning that GMM-UNIT is a unifying framework for unsupervised image-to-image translation.
Image fusion using symmetric skip autoencodervia an Adversarial Regulariser
Jun 04, 2020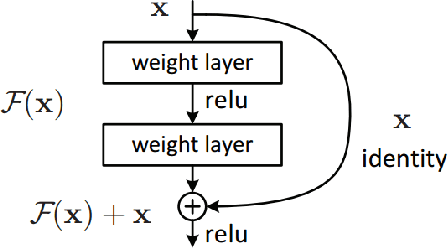
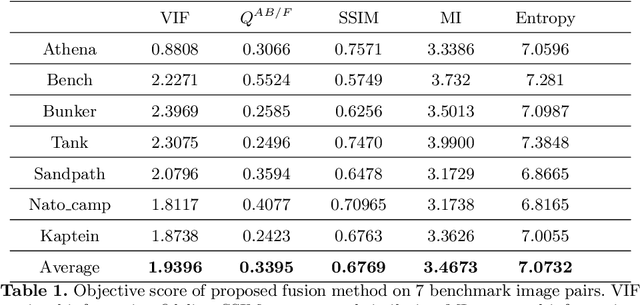
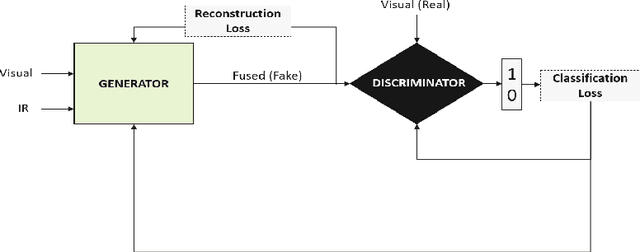
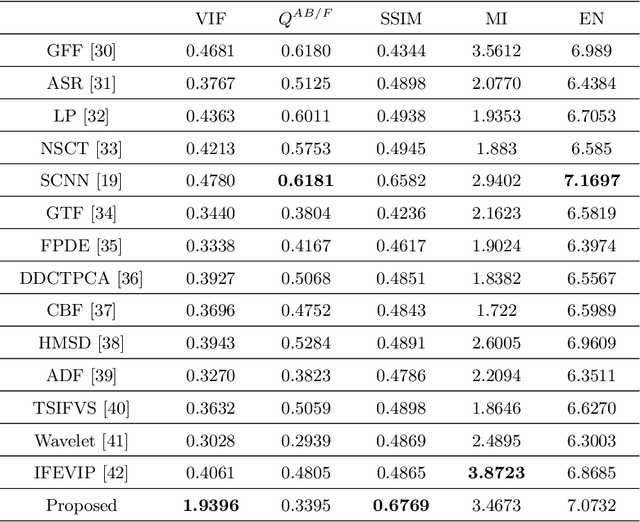
It is a challenging task to extract the best of both worlds by combining the spatial characteristics of a visible image and the spectral content of an infrared image. In this work, we propose a spatially constrained adversarial autoencoder that extracts deep features from the infrared and visible images to obtain a more exhaustive and global representation. In this paper, we propose a residual autoencoder architecture, regularised by a residual adversarial network, to generate a more realistic fused image. The residual module serves as primary building for the encoder, decoder and adversarial network, as an add on the symmetric skip connections perform the functionality of embedding the spatial characteristics directly from the initial layers of encoder structure to the decoder part of the network. The spectral information in the infrared image is incorporated by adding the feature maps over several layers in the encoder part of the fusion structure, which makes inference on both the visual and infrared images separately. In order to efficiently optimize the parameters of the network, we propose an adversarial regulariser network which would perform supervised learning on the fused image and the original visual image.
An Efficient Cervical Whole Slide Image Analysis Framework Based on Multi-scale Semantic and Spatial Deep Features
Jul 03, 2021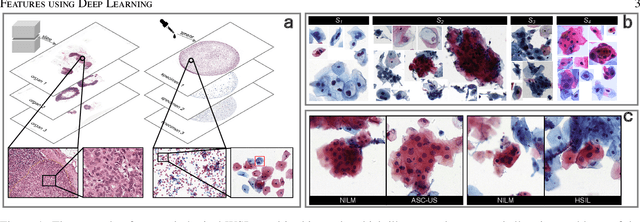
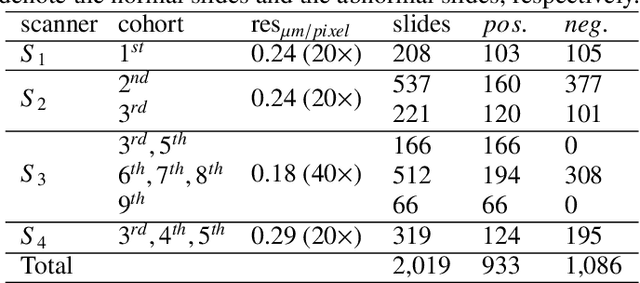
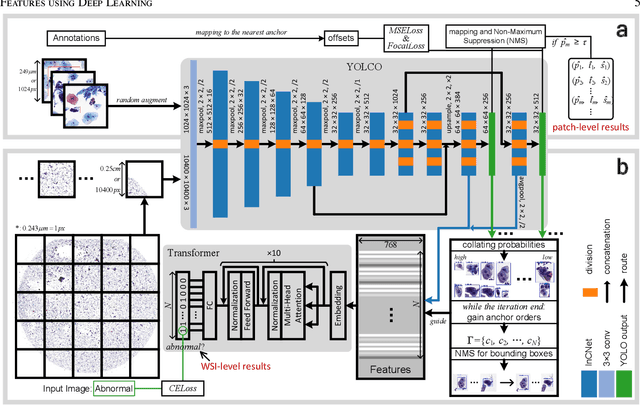
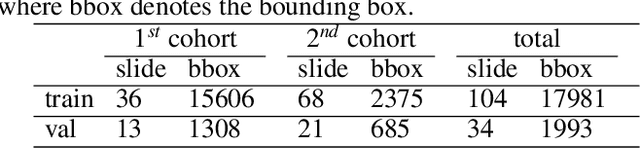
Digital gigapixel whole slide image (WSI) is widely used in clinical diagnosis, and automated WSI analysis is key for computer-aided diagnosis. Currently, analyzing the integrated descriptor of probabilities or feature maps from massive local patches encoded by ResNet classifier is the main manner for WSI-level prediction. Feature representations of the sparse and tiny lesion cells in cervical slides, however, are still challengeable for the under-promoted upstream encoders, while the unused spatial representations of cervical cells are the available features to supply the semantics analysis. As well as patches sampling with overlap and repetitive processing incur the inefficiency and the unpredictable side effect. This study designs a novel inline connection network (InCNet) by enriching the multi-scale connectivity to build the lightweight model named You Only Look Cytopathology Once (YOLCO) with the additional supervision of spatial information. The proposed model allows the input size enlarged to megapixel that can stitch the WSI without any overlap by the average repeats decreased from $10^3\sim10^4$ to $10^1\sim10^2$ for collecting features and predictions at two scales. Based on Transformer for classifying the integrated multi-scale multi-task features, the experimental results appear $0.872$ AUC score better and $2.51\times$ faster than the best conventional method in WSI classification on multicohort datasets of 2,019 slides from four scanning devices.
Dense Relational Image Captioning via Multi-task Triple-Stream Networks
Oct 12, 2020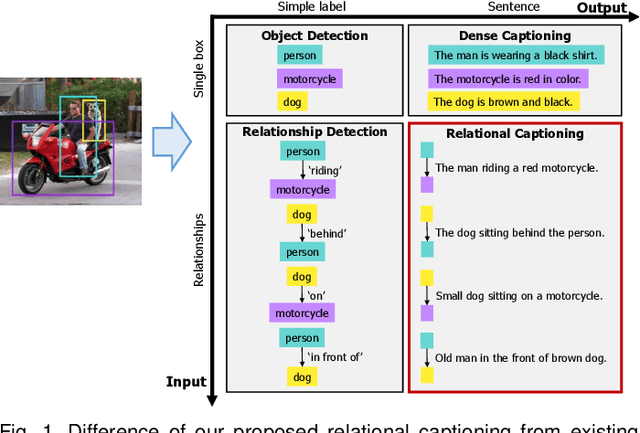
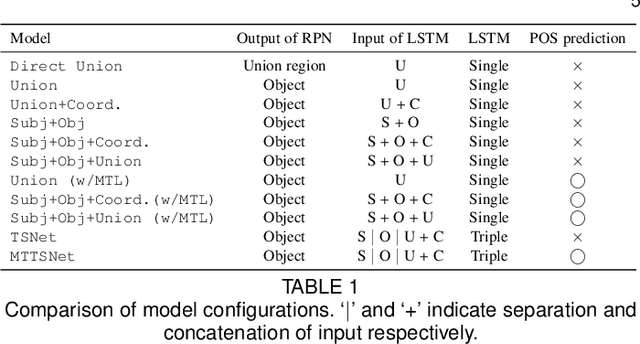
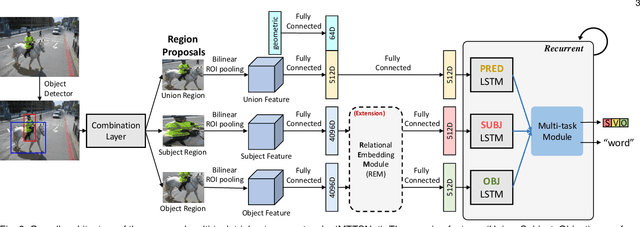
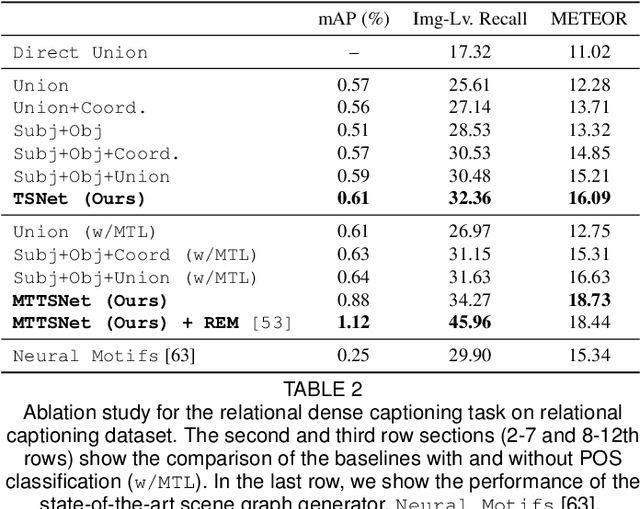
We introduce dense relational captioning, a novel image captioning task which aims to generate multiple captions with respect to relational information between objects in a visual scene. Relational captioning provides explicit descriptions of each relationship between object combinations. This framework is advantageous in both diversity and amount of information, leading to a comprehensive image understanding based on relationships, e.g., relational proposal generation. For relational understanding between objects, the part-of-speech (POS, i.e., subject-object-predicate categories) can be a valuable prior information to guide the causal sequence of words in a caption. We enforce our framework to not only learn to generate captions but also predict the POS of each word. To this end, we propose the multi-task triple-stream network (MTTSNet) which consists of three recurrent units responsible for each POS which is trained by jointly predicting the correct captions and POS for each word. In addition, we found that the performance of MTTSNet can be improved by modulating the object embeddings with an explicit relational module. We demonstrate that our proposed model can generate more diverse and richer captions, via extensive experimental analysis on large scale datasets and several metrics. We additionally extend analysis to an ablation study, applications on holistic image captioning, scene graph generation, and retrieval tasks.
HYPERLOCK: In-Memory Hyperdimensional Encryption in Memristor Crossbar Array
Jan 27, 2022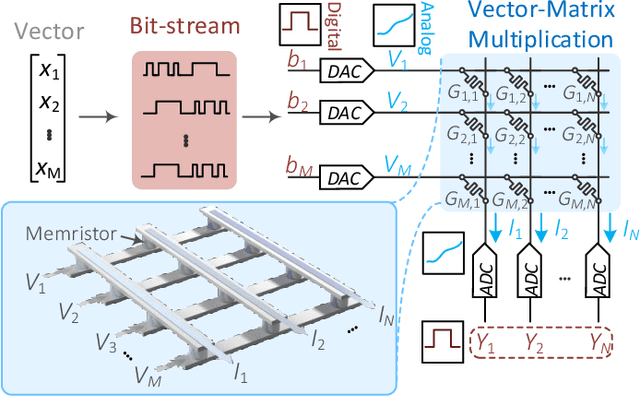
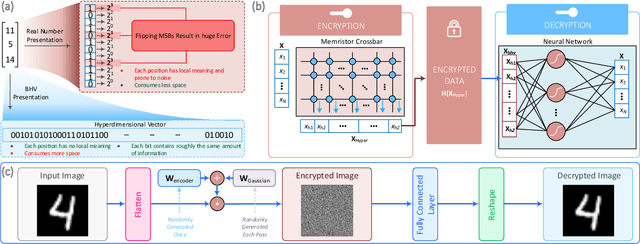
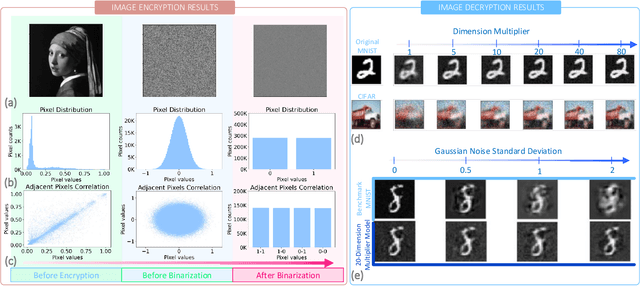
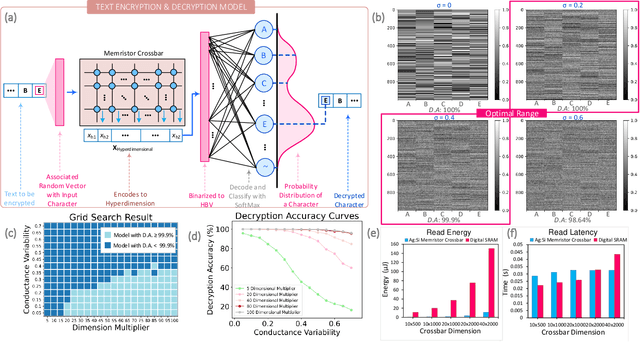
We present a novel cryptography architecture based on memristor crossbar array, binary hypervectors, and neural network. Utilizing the stochastic and unclonable nature of memristor crossbar and error tolerance of binary hypervectors and neural network, implementation of the algorithm on memristor crossbar simulation is made possible. We demonstrate that with an increasing dimension of the binary hypervectors, the non-idealities in the memristor circuit can be effectively controlled. At the fine level of controlled crossbar non-ideality, noise from memristor circuit can be used to encrypt data while being sufficiently interpretable by neural network for decryption. We applied our algorithm on image cryptography for proof of concept, and to text en/decryption with 100% decryption accuracy despite crossbar noises. Our work shows the potential and feasibility of using memristor crossbars as an unclonable stochastic encoder unit of cryptography on top of their existing functionality as a vector-matrix multiplication acceleration device.
Free-Form Image Inpainting via Contrastive Attention Network
Oct 29, 2020
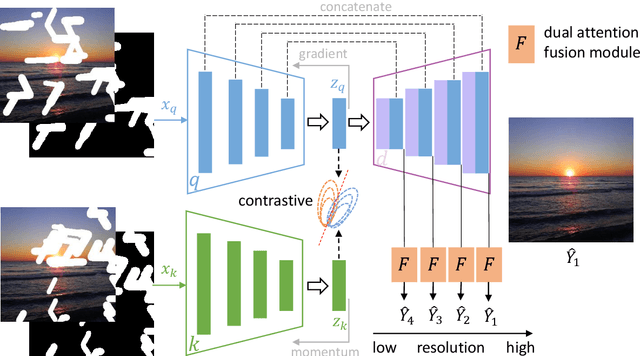
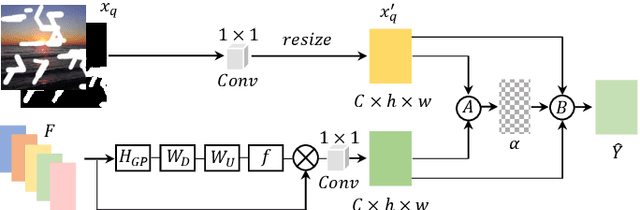

Most deep learning based image inpainting approaches adopt autoencoder or its variants to fill missing regions in images. Encoders are usually utilized to learn powerful representational spaces, which are important for dealing with sophisticated learning tasks. Specifically, in image inpainting tasks, masks with any shapes can appear anywhere in images (i.e., free-form masks) which form complex patterns. It is difficult for encoders to capture such powerful representations under this complex situation. To tackle this problem, we propose a self-supervised Siamese inference network to improve the robustness and generalization. It can encode contextual semantics from full resolution images and obtain more discriminative representations. we further propose a multi-scale decoder with a novel dual attention fusion module (DAF), which can combine both the restored and known regions in a smooth way. This multi-scale architecture is beneficial for decoding discriminative representations learned by encoders into images layer by layer. In this way, unknown regions will be filled naturally from outside to inside. Qualitative and quantitative experiments on multiple datasets, including facial and natural datasets (i.e., Celeb-HQ, Pairs Street View, Places2 and ImageNet), demonstrate that our proposed method outperforms state-of-the-art methods in generating high-quality inpainting results.
Panoptic Segmentation Meets Remote Sensing
Nov 30, 2021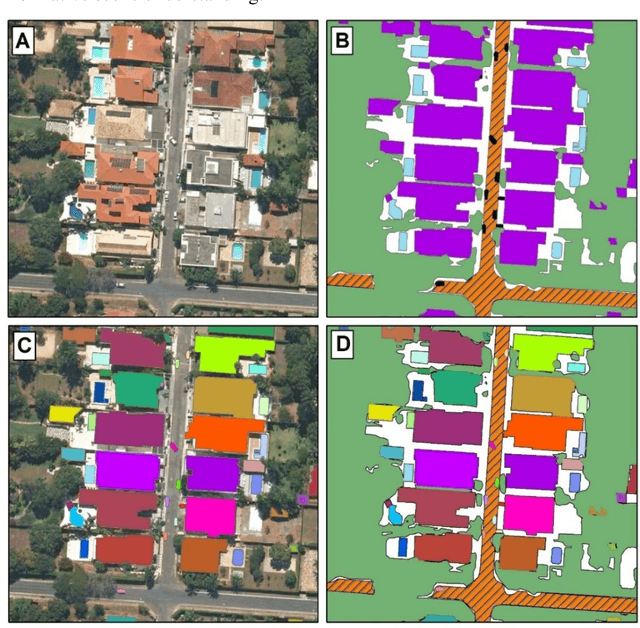
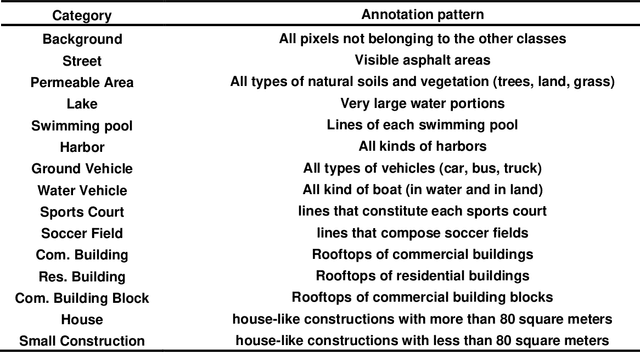
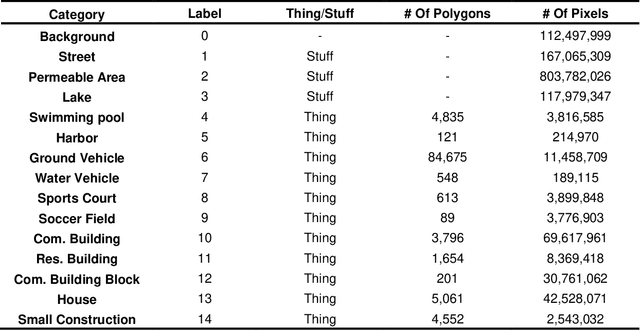
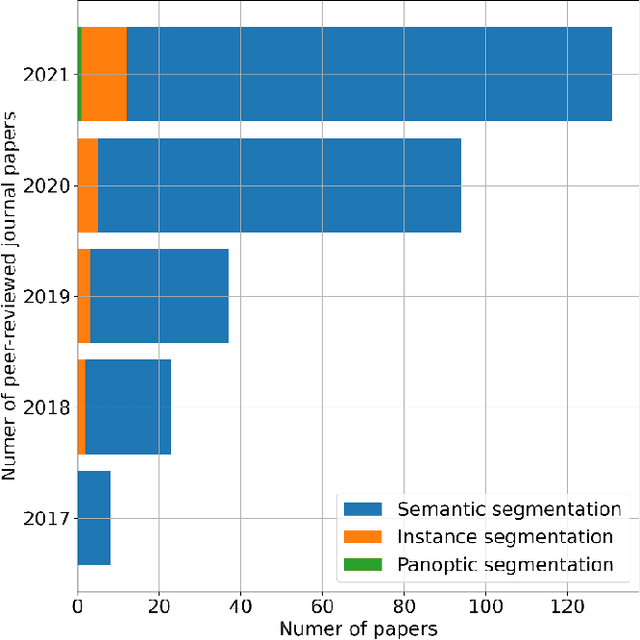

Panoptic segmentation combines instance and semantic predictions, allowing the detection of "things" and "stuff" simultaneously. Effectively approaching panoptic segmentation in remotely sensed data can be auspicious in many challenging problems since it allows continuous mapping and specific target counting. Several difficulties have prevented the growth of this task in remote sensing: (a) most algorithms are designed for traditional images, (b) image labelling must encompass "things" and "stuff" classes, and (c) the annotation format is complex. Thus, aiming to solve and increase the operability of panoptic segmentation in remote sensing, this study has five objectives: (1) create a novel data preparation pipeline for panoptic segmentation, (2) propose an annotation conversion software to generate panoptic annotations; (3) propose a novel dataset on urban areas, (4) modify the Detectron2 for the task, and (5) evaluate difficulties of this task in the urban setting. We used an aerial image with a 0,24-meter spatial resolution considering 14 classes. Our pipeline considers three image inputs, and the proposed software uses point shapefiles for creating samples in the COCO format. Our study generated 3,400 samples with 512x512 pixel dimensions. We used the Panoptic-FPN with two backbones (ResNet-50 and ResNet-101), and the model evaluation considered semantic instance and panoptic metrics. We obtained 93.9, 47.7, and 64.9 for the mean IoU, box AP, and PQ. Our study presents the first effective pipeline for panoptic segmentation and an extensive database for other researchers to use and deal with other data or related problems requiring a thorough scene understanding.
Training End-to-end Single Image Generators without GANs
Apr 07, 2020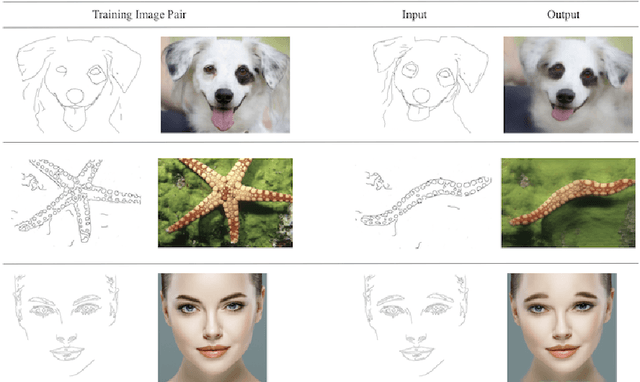
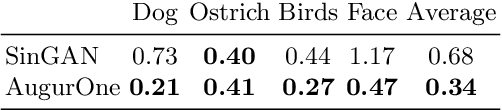


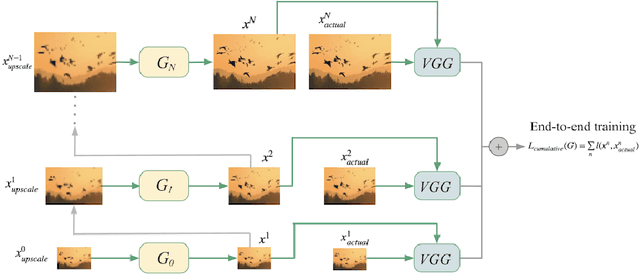
We present AugurOne, a novel approach for training single image generative models. Our approach trains an upscaling neural network using non-affine augmentations of the (single) input image, particularly including non-rigid thin plate spline image warps. The extensive augmentations significantly increase the in-sample distribution for the upsampling network enabling the upscaling of highly variable inputs. A compact latent space is jointly learned allowing for controlled image synthesis. Differently from Single Image GAN, our approach does not require GAN training and takes place in an end-to-end fashion allowing fast and stable training. We experimentally evaluate our method and show that it obtains compelling novel animations of single-image, as well as, state-of-the-art performance on conditional generation tasks e.g. paint-to-image and edges-to-image.
Deep Learning based Prediction of MSI in Colorectal Cancer via Prediction of the Status of MMR Markers
Feb 24, 2022

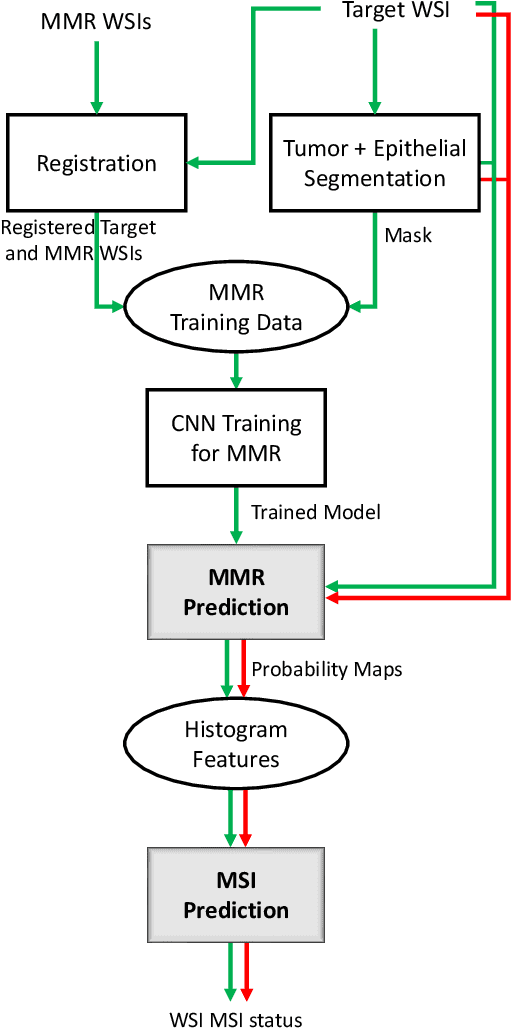
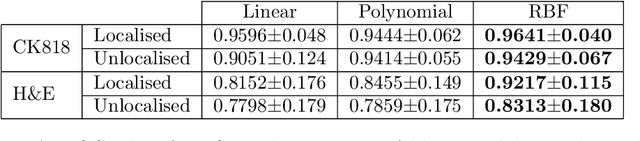
An accurate diagnosis and profiling of tumour are critical to the best treatment choices for cancer patients. In addition to the cancer type and its aggressiveness, molecular heterogeneity also plays a vital role in treatment selection. MSI or MMR deficiency is one of the well-studied aberrations in terms of molecular changes. Colorectal cancer patients with MMR deficiency respond well to immunotherapy, hence assessment of the relevant molecular markers can assist clinicians in making optimal treatment selections for patients. Immunohistochemistry is one of the ways for identifying these molecular changes which requires additional sections of tumour tissue. Introduction of automated methods that can predict MSI or MMR status from a target image without the need for additional sections can substantially reduce the cost associated with it. In this work, we present our work on predicting MSI status in a two-stage process using a single target slide either stained with CK818 or H\&E. First, we train a multi-headed convolutional neural network model where each head is responsible for predicting one of the MMR protein expressions. To this end, we perform registration of MMR slides to the target slide as a pre-processing step. In the second stage, statistical features computed from the MMR prediction maps are used for the final MSI prediction. Our results demonstrate that MSI classification can be improved on incorporating fine-grained MMR labels in comparison to the previous approaches in which coarse labels (MSI/MSS) are utilised.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge