"Image": models, code, and papers
SSCAN: A Spatial-spectral Cross Attention Network for Hyperspectral Image Denoising
May 23, 2021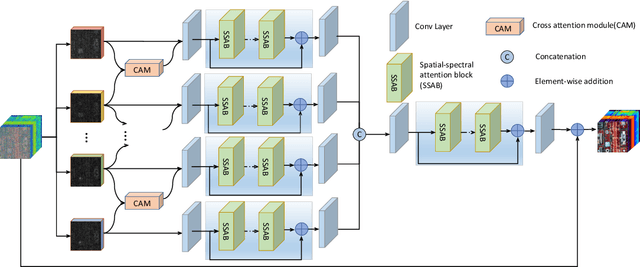
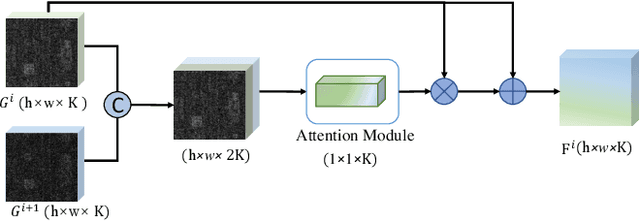
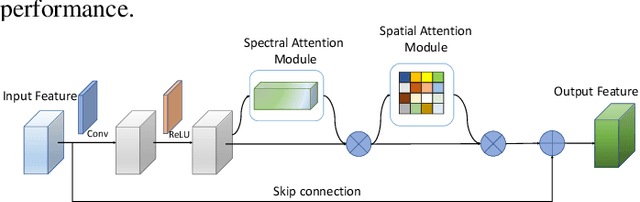
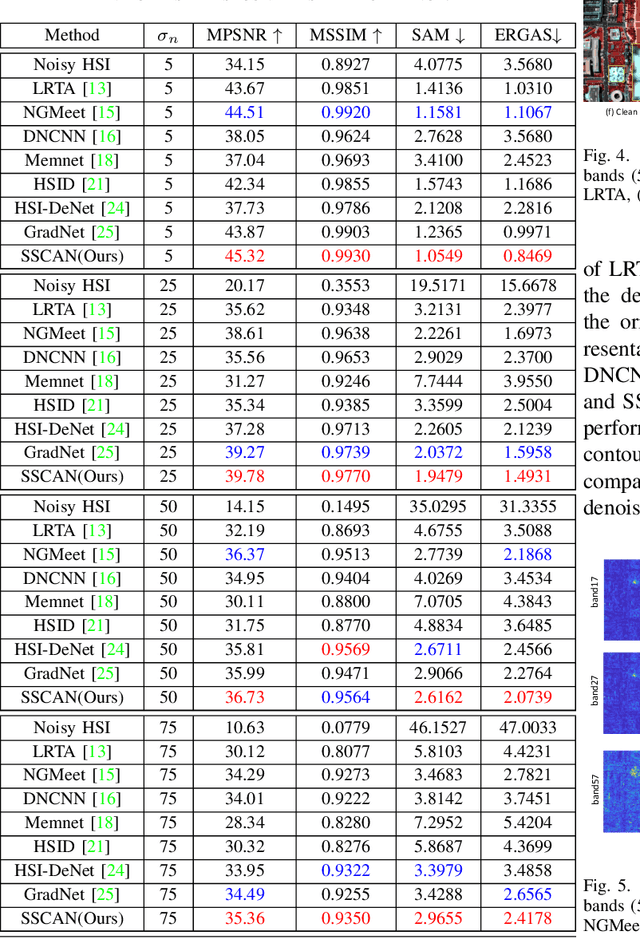
Hyperspectral images (HSIs) have been widely used in a variety of applications thanks to the rich spectral information they are able to provide. Among all HSI processing tasks, HSI denoising is a crucial step. Recently, deep learning-based image denoising methods have made great progress and achieved great performance. However, existing methods tend to ignore the correlations between adjacent spectral bands, leading to problems such as spectral distortion and blurred edges in denoised results. In this study, we propose a novel HSI denoising network, termed SSCAN, that combines group convolutions and attention modules. Specifically, we use a group convolution with a spatial attention module to facilitate feature extraction by directing models' attention to band-wise important features. We propose a spectral-spatial attention block (SSAB) to exploit the spatial and spectral information in hyperspectral images in an effective manner. In addition, we adopt residual learning operations with skip connections to ensure training stability. The experimental results indicate that the proposed SSCAN outperforms several state-of-the-art HSI denoising algorithms.
Improving Feature Extraction from Histopathological Images Through A Fine-tuning ImageNet Model
Jan 03, 2022Due to lack of annotated pathological images, transfer learning has been the predominant approach in the field of digital pathology.Pre-trained neural networks based on ImageNet database are often used to extract "off the shelf" features, achieving great success in predicting tissue types, molecular features, and clinical outcomes, etc. We hypothesize that fine-tuning the pre-trained models using histopathological images could further improve feature extraction, and downstream prediction performance.We used 100,000 annotated HE image patches for colorectal cancer (CRC) to finetune a pretrained Xception model via a twostep approach.The features extracted from finetuned Xception (FTX2048) model and Imagepretrained (IMGNET2048) model were compared through: (1) tissue classification for HE images from CRC, same image type that was used for finetuning; (2) prediction of immunerelated gene expression and (3) gene mutations for lung adenocarcinoma (LUAD).Fivefold cross validation was used for model performance evaluation. The extracted features from the finetuned FTX2048 exhibited significantly higher accuracy for predicting tisue types of CRC compared to the off the shelf feature directly from Xception based on ImageNet database. Particularly, FTX2048 markedly improved the accuracy for stroma from 87% to 94%. Similarly, features from FTX2048 boosted the prediction of transcriptomic expression of immunerelated genesin LUAD. For the genes that had signigicant relationships with image fetures, the features fgrom the finetuned model imprroved the prediction for the majority of the genes. Inaddition, fetures from FTX2048 improved prediction of mutation for 5 out of 9 most frequently mutated genes in LUAD.
Generative Adversarial Networks and Adversarial Autoencoders: Tutorial and Survey
Nov 26, 2021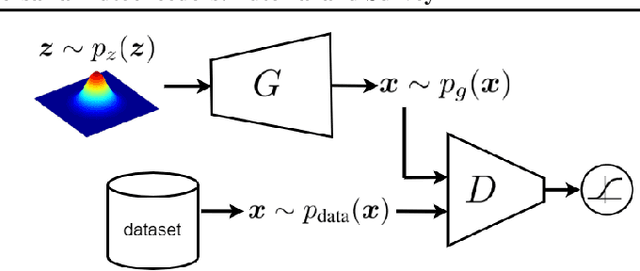
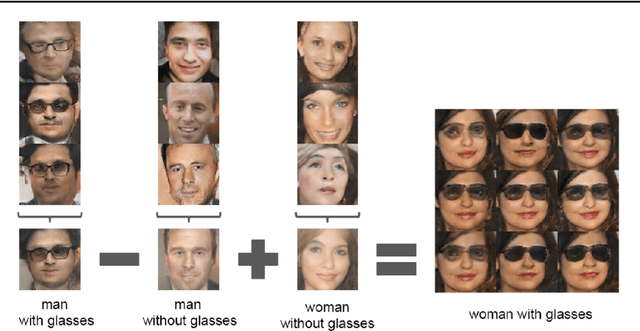

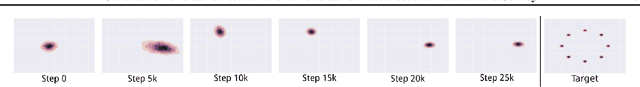
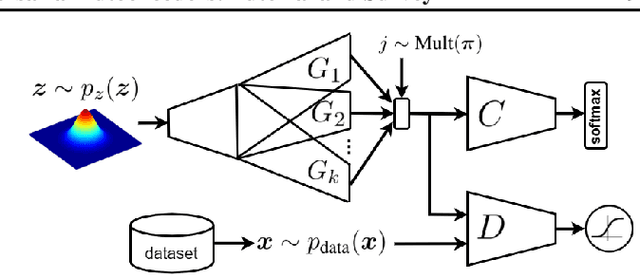
This is a tutorial and survey paper on Generative Adversarial Network (GAN), adversarial autoencoders, and their variants. We start with explaining adversarial learning and the vanilla GAN. Then, we explain the conditional GAN and DCGAN. The mode collapse problem is introduced and various methods, including minibatch GAN, unrolled GAN, BourGAN, mixture GAN, D2GAN, and Wasserstein GAN, are introduced for resolving this problem. Then, maximum likelihood estimation in GAN are explained along with f-GAN, adversarial variational Bayes, and Bayesian GAN. Then, we cover feature matching in GAN, InfoGAN, GRAN, LSGAN, energy-based GAN, CatGAN, MMD GAN, LapGAN, progressive GAN, triple GAN, LAG, GMAN, AdaGAN, CoGAN, inverse GAN, BiGAN, ALI, SAGAN, Few-shot GAN, SinGAN, and interpolation and evaluation of GAN. Then, we introduce some applications of GAN such as image-to-image translation (including PatchGAN, CycleGAN, DeepFaceDrawing, simulated GAN, interactive GAN), text-to-image translation (including StackGAN), and mixing image characteristics (including FineGAN and MixNMatch). Finally, we explain the autoencoders based on adversarial learning including adversarial autoencoder, PixelGAN, and implicit autoencoder.
Effect of Prior-based Losses on Segmentation Performance: A Benchmark
Jan 11, 2022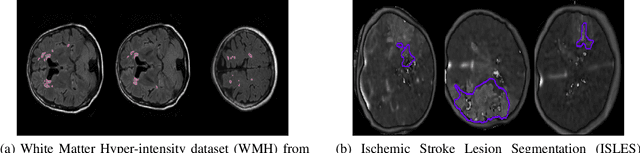
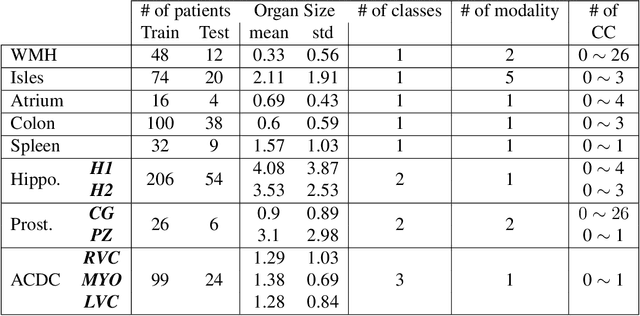
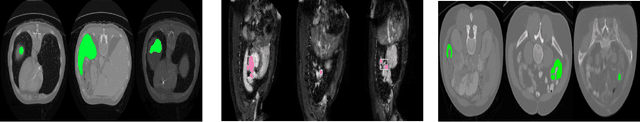
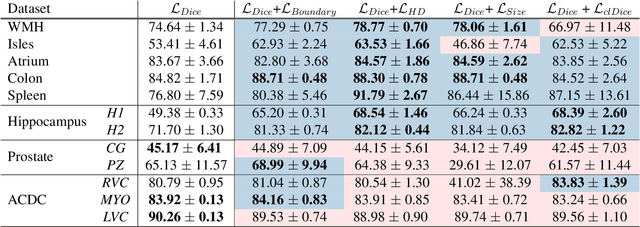
Today, deep convolutional neural networks (CNNs) have demonstrated state-of-the-art performance for medical image segmentation, on various imaging modalities and tasks. Despite early success, segmentation networks may still generate anatomically aberrant segmentations, with holes or inaccuracies near the object boundaries. To enforce anatomical plausibility, recent research studies have focused on incorporating prior knowledge such as object shape or boundary, as constraints in the loss function. Prior integrated could be low-level referring to reformulated representations extracted from the ground-truth segmentations, or high-level representing external medical information such as the organ's shape or size. Over the past few years, prior-based losses exhibited a rising interest in the research field since they allow integration of expert knowledge while still being architecture-agnostic. However, given the diversity of prior-based losses on different medical imaging challenges and tasks, it has become hard to identify what loss works best for which dataset. In this paper, we establish a benchmark of recent prior-based losses for medical image segmentation. The main objective is to provide intuition onto which losses to choose given a particular task or dataset. To this end, four low-level and high-level prior-based losses are selected. The considered losses are validated on 8 different datasets from a variety of medical image segmentation challenges including the Decathlon, the ISLES and the WMH challenge. Results show that whereas low-level prior-based losses can guarantee an increase in performance over the Dice loss baseline regardless of the dataset characteristics, high-level prior-based losses can increase anatomical plausibility as per data characteristics.
Image Differential Invariants
Nov 13, 2019



Inspired by the methods of systematic derivation of image moment invariants, we design two fundamental differential operators to generate image differential invariants for the action of 2D Euclidean, similarity and affine transformation groups. Each differential invariant obtained by using the new method can be expressed as a homogeneous polynomial of image partial derivatives. When setting the degree of the polynomial and the order of image partial derivatives are less than or equal to 4, we generate all Euclidean differential invariants and discuss the independence of them in detail. In the experimental part, we find the relation between Euclidean differential invariants and Gaussian-Hermite moment invariants when using the derivatives of Gaussian to estimate image partial derivatives. Texture classification and image patch verification are carried out on some synthetic and popular real databases. We mainly evaluate the stability and discriminability of Euclidean differential invariants and analyse the effects of various factors on performance of them. The experimental results validate image Euclidean differential invariants have better performance than some commonly used local image features in most cases.
Medical Image Segmentation Using Deep Learning: A Survey
Sep 28, 2020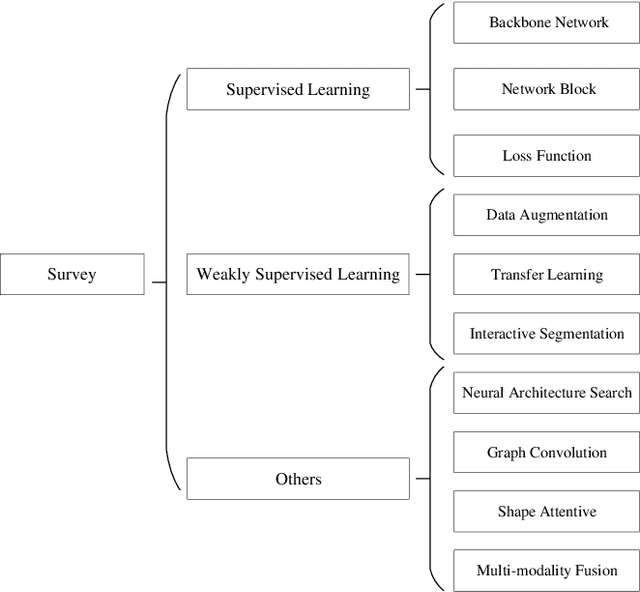
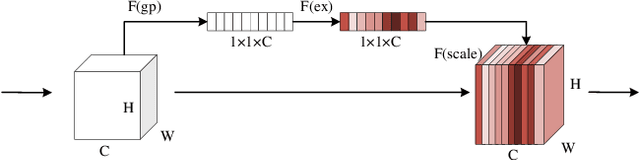
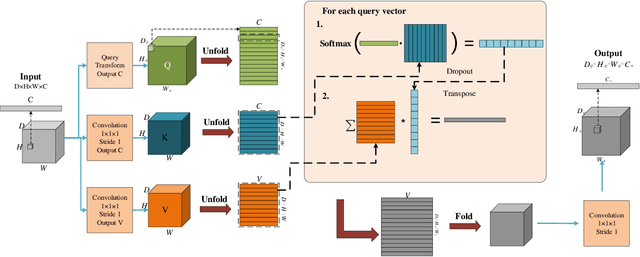
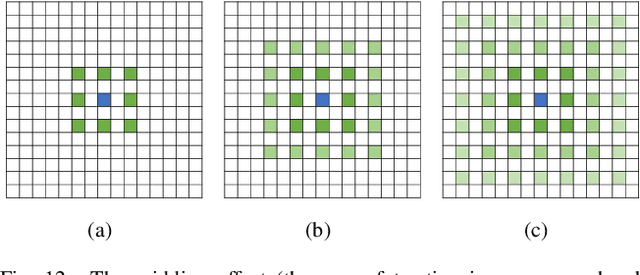
Deep learning has been widely used for medical image segmentation and a large number of papers has been presented recording the success of deep learning in the field. In this paper, we present a comprehensive thematic survey on medical image segmentation using deep learning techniques. This paper makes two original contributions. Firstly, compared to traditional surveys that directly divide literatures of deep learning on medical image segmentation into many groups and introduce literatures in detail for each group, we classify currently popular literatures according to a multi-level structure from coarse to fine. Secondly, this paper focuses on supervised and weakly supervised learning approaches, without including unsupervised approaches since they have been introduced in many old surveys and they are not popular currently. For supervised learning approaches, we analyze literatures in three aspects: the selection of backbone networks, the design of network blocks, and the improvement of loss functions. For weakly supervised learning approaches, we investigate literature according to data augmentation, transfer learning, and interactive segmentation, separately. Compared to existing surveys, this survey classifies the literatures very differently from before and is more convenient for readers to understand the relevant rationale and will guide them to think of appropriate improvements in medical image segmentation based on deep learning approaches.
PrivPAS: A real time Privacy-Preserving AI System and applied ethics
Feb 08, 2022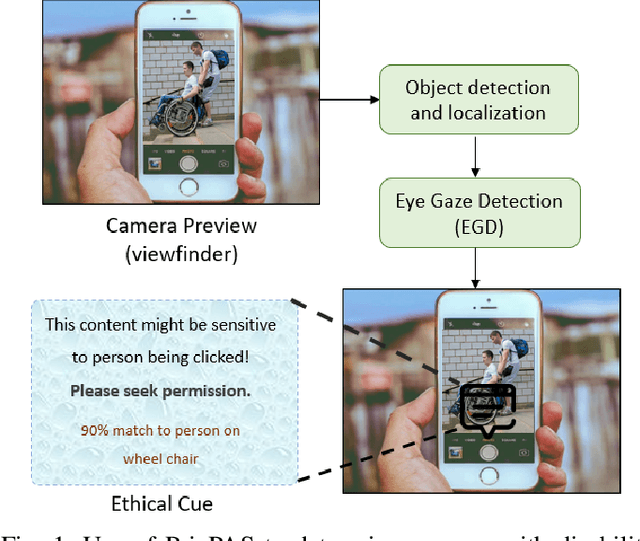
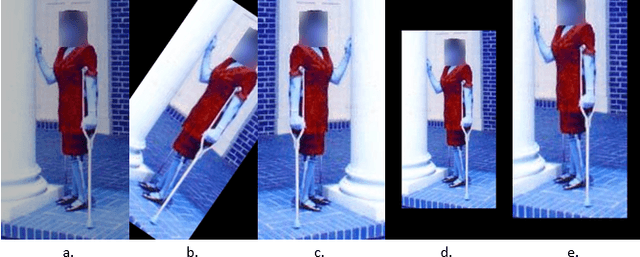

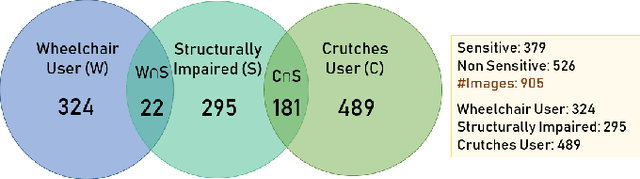

With 3.78 billion social media users worldwide in 2021 (48% of the human population), almost 3 billion images are shared daily. At the same time, a consistent evolution of smartphone cameras has led to a photography explosion with 85% of all new pictures being captured using smartphones. However, lately, there has been an increased discussion of privacy concerns when a person being photographed is unaware of the picture being taken or has reservations about the same being shared. These privacy violations are amplified for people with disabilities, who may find it challenging to raise dissent even if they are aware. Such unauthorized image captures may also be misused to gain sympathy by third-party organizations, leading to a privacy breach. Privacy for people with disabilities has so far received comparatively less attention from the AI community. This motivates us to work towards a solution to generate privacy-conscious cues for raising awareness in smartphone users of any sensitivity in their viewfinder content. To this end, we introduce PrivPAS (A real time Privacy-Preserving AI System) a novel framework to identify sensitive content. Additionally, we curate and annotate a dataset to identify and localize accessibility markers and classify whether an image is sensitive to a featured subject with a disability. We demonstrate that the proposed lightweight architecture, with a memory footprint of a mere 8.49MB, achieves a high mAP of 89.52% on resource-constrained devices. Furthermore, our pipeline, trained on face anonymized data, achieves an F1-score of 73.1%.
Rethinking Image Deraining via Rain Streaks and Vapors
Aug 03, 2020
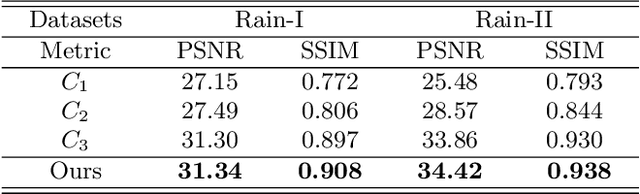
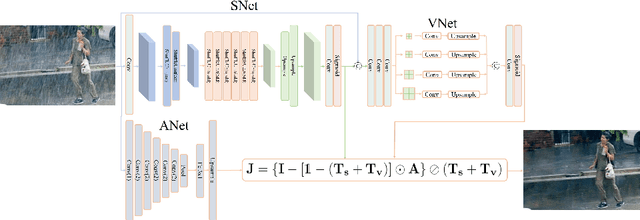
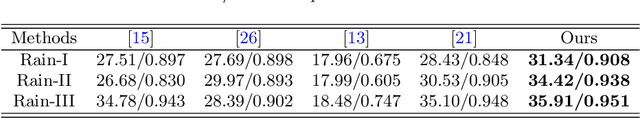
Single image deraining regards an input image as a fusion of a background image, a transmission map, rain streaks, and atmosphere light. While advanced models are proposed for image restoration (i.e., background image generation), they regard rain streaks with the same properties as background rather than transmission medium. As vapors (i.e., rain streaks accumulation or fog-like rain) are conveyed in the transmission map to model the veiling effect, the fusion of rain streaks and vapors do not naturally reflect the rain image formation. In this work, we reformulate rain streaks as transmission medium together with vapors to model rain imaging. We propose an encoder-decoder CNN named as SNet to learn the transmission map of rain streaks. As rain streaks appear with various shapes and directions, we use ShuffleNet units within SNet to capture their anisotropic representations. As vapors are brought by rain streaks, we propose a VNet containing spatial pyramid pooling (SSP) to predict the transmission map of vapors in multi-scales based on that of rain streaks. Meanwhile, we use an encoder CNN named ANet to estimate atmosphere light. The SNet, VNet, and ANet are jointly trained to predict transmission maps and atmosphere light for rain image restoration. Extensive experiments on the benchmark datasets demonstrate the effectiveness of the proposed visual model to predict rain streaks and vapors. The proposed deraining method performs favorably against state-of-the-art deraining approaches.
Problem-dependent attention and effort in neural networks with an application to image resolution
Jan 05, 2022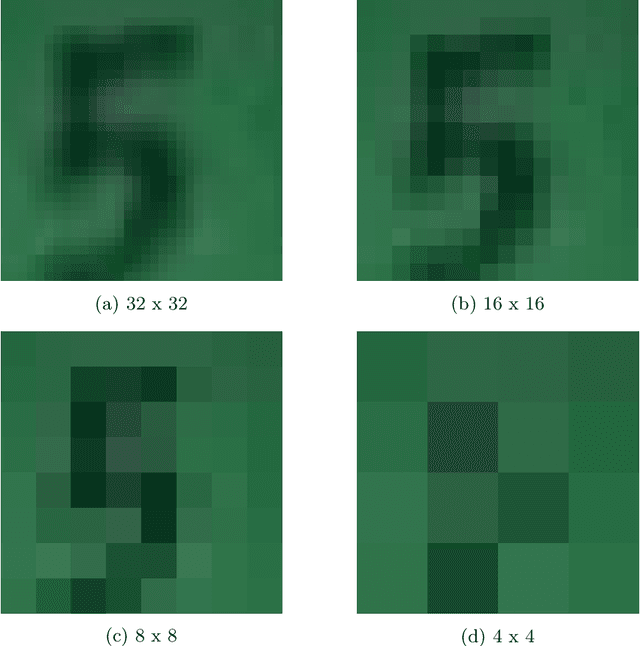
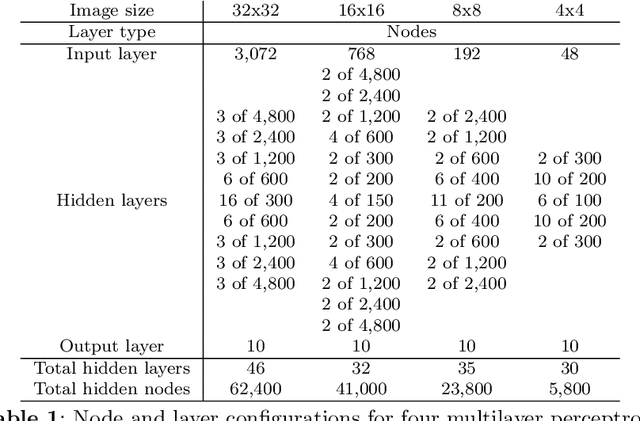
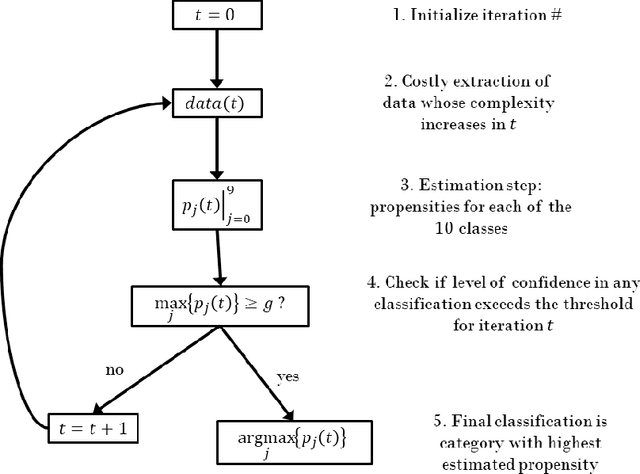
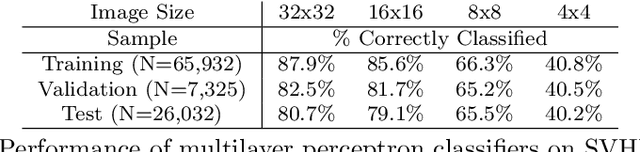
This paper introduces a new neural network-based estimation approach that is inspired by the biological phenomenon whereby humans and animals vary the levels of attention and effort that they dedicate to a problem depending upon its difficulty. The proposed approach leverages alternate models' internal levels of confidence in their own projections. If the least costly model is confident in its classification, then that is the classification used; if not, the model with the next lowest cost of implementation is run, and so on. This use of successively more complex models -- together with the models' internal propensity scores to evaluate their likelihood of being correct -- makes it possible to substantially reduce resource use while maintaining high standards for classification accuracy. The approach is applied to the digit recognition problem from Google's Street View House Numbers dataset, using Multilayer Perceptron (MLP) neural networks trained on high- and low-resolution versions of the digit images. The algorithm examines the low-resolution images first, only moving to higher resolution images if the classification from the initial low-resolution pass does not have a high degree of confidence. For the MLPs considered here, this sequential approach enables a reduction in resource usage of more than 50\% without any sacrifice in classification accuracy.
Unified Visual Transformer Compression
Mar 15, 2022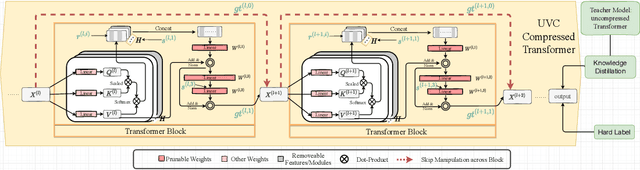
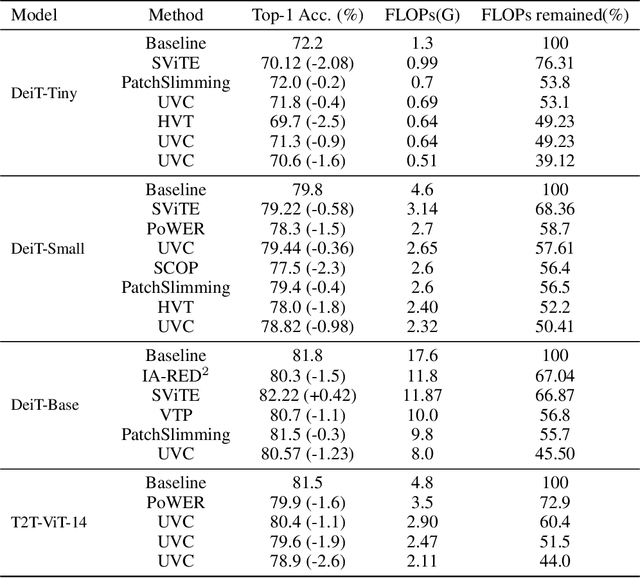
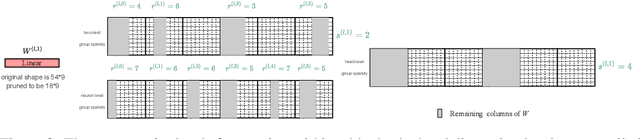
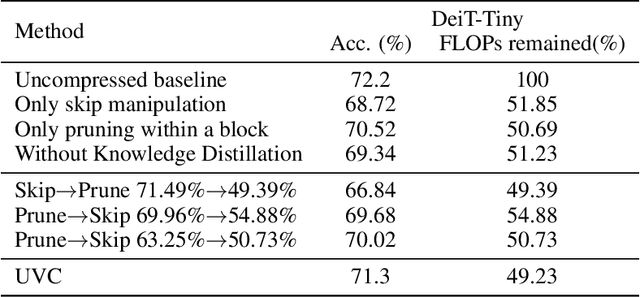
Vision transformers (ViTs) have gained popularity recently. Even without customized image operators such as convolutions, ViTs can yield competitive performance when properly trained on massive data. However, the computational overhead of ViTs remains prohibitive, due to stacking multi-head self-attention modules and else. Compared to the vast literature and prevailing success in compressing convolutional neural networks, the study of Vision Transformer compression has also just emerged, and existing works focused on one or two aspects of compression. This paper proposes a unified ViT compression framework that seamlessly assembles three effective techniques: pruning, layer skipping, and knowledge distillation. We formulate a budget-constrained, end-to-end optimization framework, targeting jointly learning model weights, layer-wise pruning ratios/masks, and skip configurations, under a distillation loss. The optimization problem is then solved using the primal-dual algorithm. Experiments are conducted with several ViT variants, e.g. DeiT and T2T-ViT backbones on the ImageNet dataset, and our approach consistently outperforms recent competitors. For example, DeiT-Tiny can be trimmed down to 50\% of the original FLOPs almost without losing accuracy. Codes are available online:~\url{https://github.com/VITA-Group/UVC}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge