"Image": models, code, and papers
Stain Normalized Breast Histopathology Image Recognition using Convolutional Neural Networks for Cancer Detection
Jan 04, 2022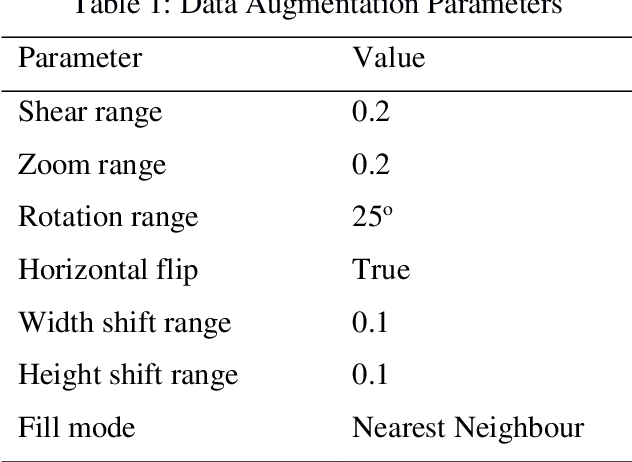
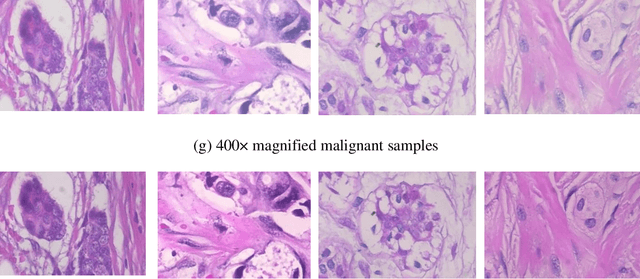
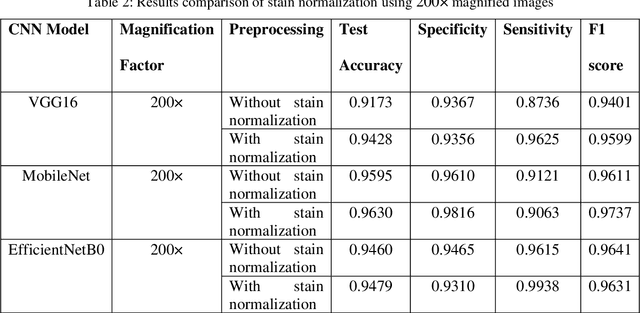
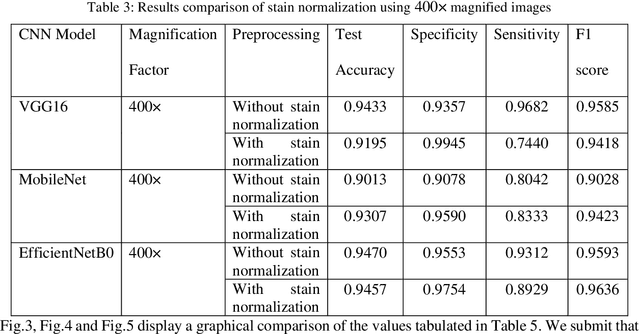
Computer assisted diagnosis in digital pathology is becoming ubiquitous as it can provide more efficient and objective healthcare diagnostics. Recent advances have shown that the convolutional Neural Network (CNN) architectures, a well-established deep learning paradigm, can be used to design a Computer Aided Diagnostic (CAD) System for breast cancer detection. However, the challenges due to stain variability and the effect of stain normalization with such deep learning frameworks are yet to be well explored. Moreover, performance analysis with arguably more efficient network models, which may be important for high throughput screening, is also not well explored.To address this challenge, we consider some contemporary CNN models for binary classification of breast histopathology images that involves (1) the data preprocessing with stain normalized images using an adaptive colour deconvolution (ACD) based color normalization algorithm to handle the stain variabilities; and (2) applying transfer learning based training of some arguably more efficient CNN models, namely Visual Geometry Group Network (VGG16), MobileNet and EfficientNet. We have validated the trained CNN networks on a publicly available BreaKHis dataset, for 200x and 400x magnified histopathology images. The experimental analysis shows that pretrained networks in most cases yield better quality results on data augmented breast histopathology images with stain normalization, than the case without stain normalization. Further, we evaluated the performance and efficiency of popular lightweight networks using stain normalized images and found that EfficientNet outperforms VGG16 and MobileNet in terms of test accuracy and F1 Score. We observed that efficiency in terms of test time is better in EfficientNet than other networks; VGG Net, MobileNet, without much drop in the classification accuracy.
Semi-supervised Learning for COVID-19 Image Classification via ResNet
Feb 27, 2021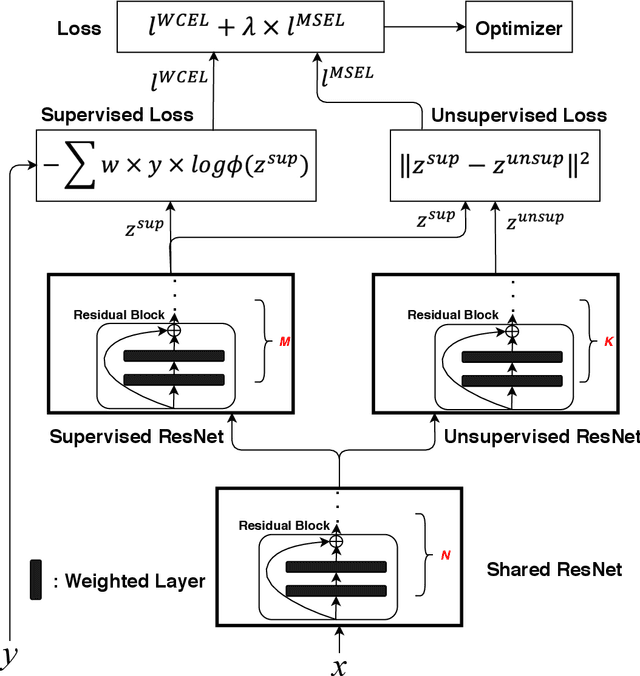
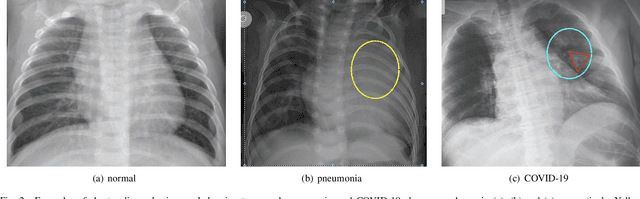
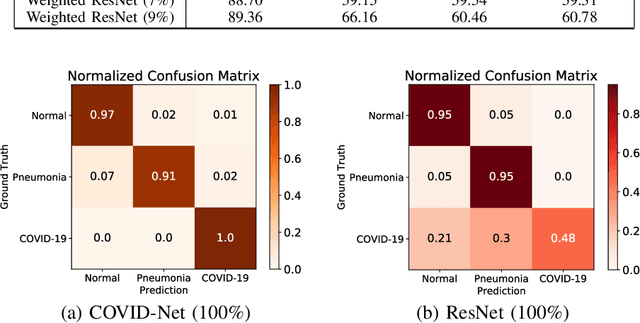
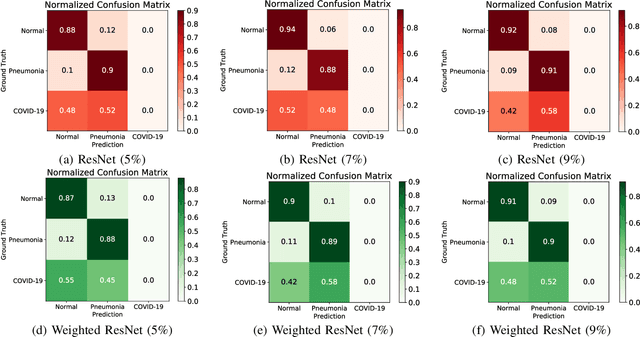
Coronavirus disease 2019 (COVID-19) is an ongoing global pandemic in over 200 countries and territories, which has resulted in a great public health concern across the international community. Analysis of X-ray imaging data can play a critical role in timely and accurate screening and fighting against COVID-19. Supervised deep learning has been successfully applied to recognize COVID-19 pathology from X-ray imaging datasets. However, it requires a substantial amount of annotated X-ray images to train models, which is often not applicable to data analysis for emerging events such as COVID-19 outbreak, especially in the early stage of the outbreak. To address this challenge, this paper proposes a two-path semi-supervised deep learning model, ssResNet, based on Residual Neural Network (ResNet) for COVID-19 image classification, where two paths refer to a supervised path and an unsupervised path, respectively. Moreover, we design a weighted supervised loss that assigns higher weight for the minority classes in the training process to resolve the data imbalance. Experimental results on a large-scale of X-ray image dataset COVIDx demonstrate that the proposed model can achieve promising performance even when trained on very few labeled training images.
ComBiNet: Compact Convolutional Bayesian Neural Network for Image Segmentation
Apr 14, 2021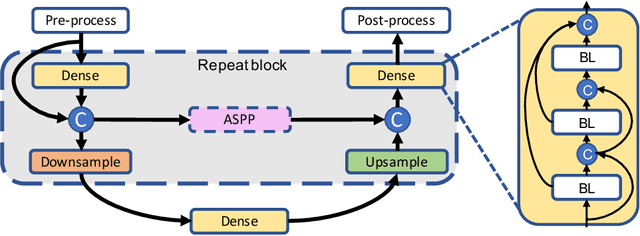
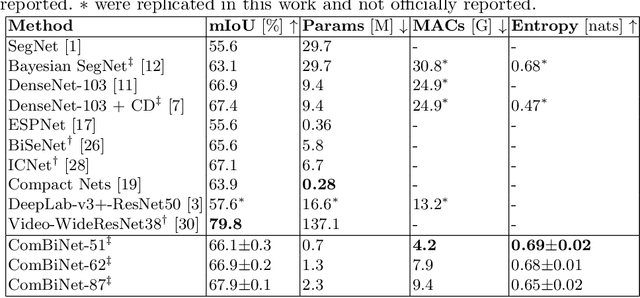
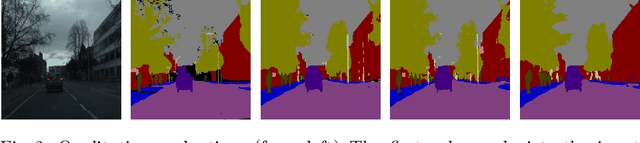
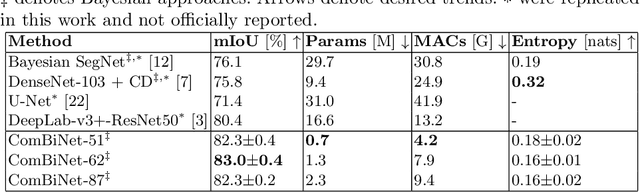
Fully convolutional U-shaped neural networks have largely been the dominant approach for pixel-wise image segmentation. In this work, we tackle two defects that hinder their deployment in real-world applications: 1) Predictions lack uncertainty quantification that may be crucial to many decision making systems; 2) Large memory storage and computational consumption demanding extensive hardware resources. To address these issues and improve their practicality we demonstrate a few-parameter compact Bayesian convolutional architecture, that achieves a marginal improvement in accuracy in comparison to related work using significantly fewer parameters and compute operations. The architecture combines parameter-efficient operations such as separable convolutions, bi-linear interpolation, multi-scale feature propagation and Bayesian inference for per-pixel uncertainty quantification through Monte Carlo Dropout. The best performing configurations required fewer than 2.5 million parameters on diverse challenging datasets with few observations.
Background-aware Classification Activation Map for Weakly Supervised Object Localization
Dec 29, 2021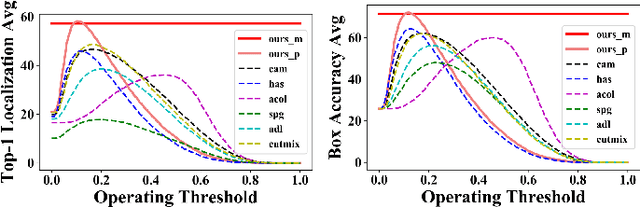
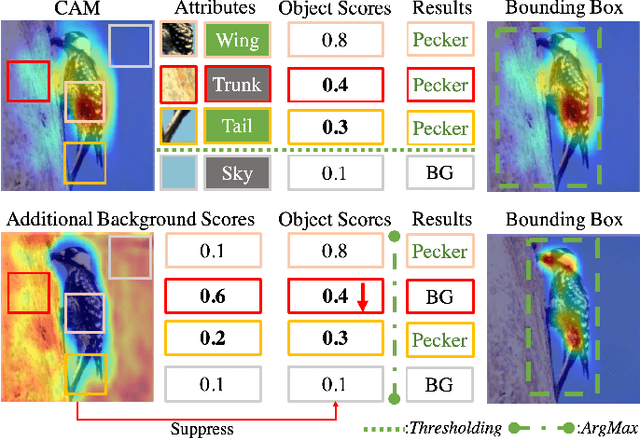
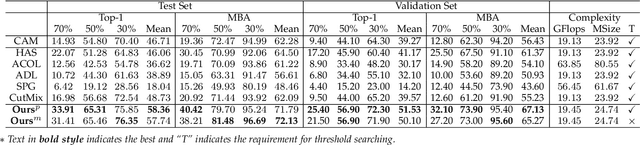

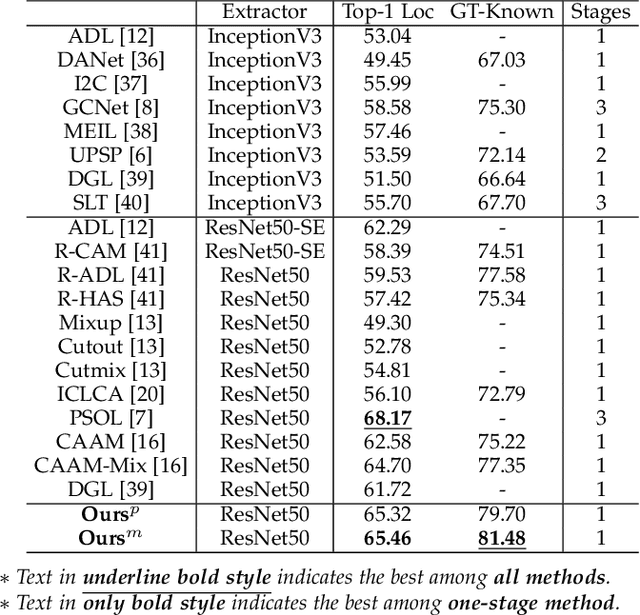
Weakly supervised object localization (WSOL) relaxes the requirement of dense annotations for object localization by using image-level classification masks to supervise its learning process. However, current WSOL methods suffer from excessive activation of background locations and need post-processing to obtain the localization mask. This paper attributes these issues to the unawareness of background cues, and propose the background-aware classification activation map (B-CAM) to simultaneously learn localization scores of both object and background with only image-level labels. In our B-CAM, two image-level features, aggregated by pixel-level features of potential background and object locations, are used to purify the object feature from the object-related background and to represent the feature of the pure-background sample, respectively. Then based on these two features, both the object classifier and the background classifier are learned to determine the binary object localization mask. Our B-CAM can be trained in end-to-end manner based on a proposed stagger classification loss, which not only improves the objects localization but also suppresses the background activation. Experiments show that our B-CAM outperforms one-stage WSOL methods on the CUB-200, OpenImages and VOC2012 datasets.
ViKiNG: Vision-Based Kilometer-Scale Navigation with Geographic Hints
Feb 23, 2022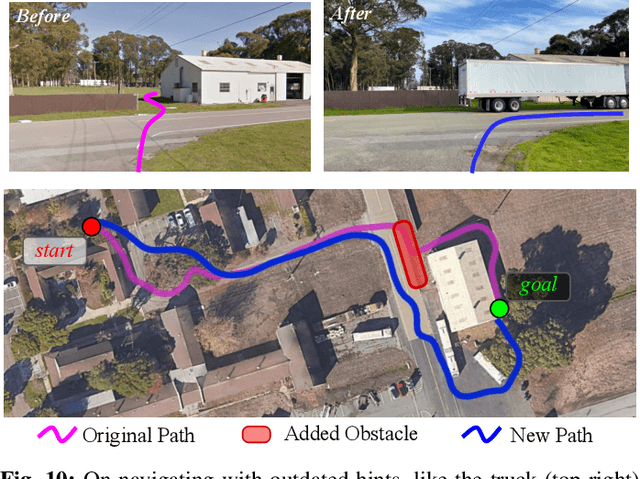
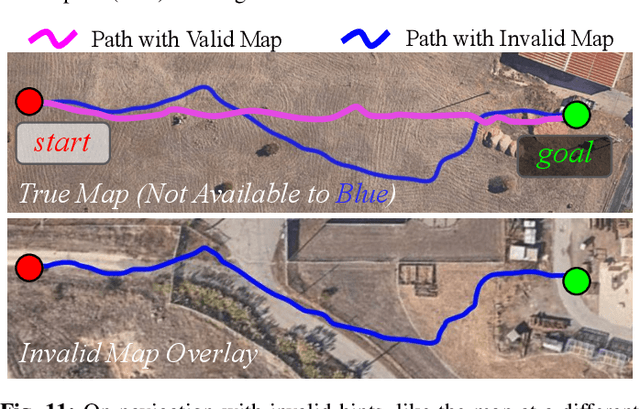
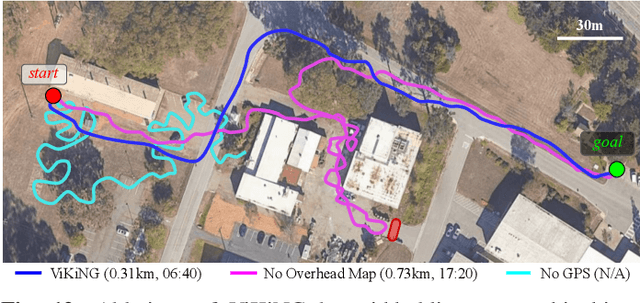
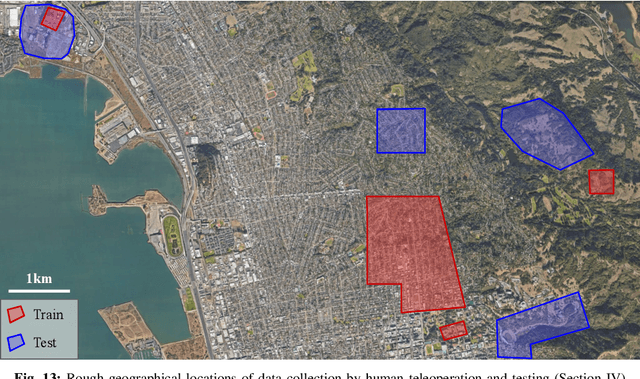
Robotic navigation has been approached as a problem of 3D reconstruction and planning, as well as an end-to-end learning problem. However, long-range navigation requires both planning and reasoning about local traversability, as well as being able to utilize information about global geography, in the form of a roadmap, GPS, or other side information, which provides important navigational hints but may be low-fidelity or unreliable. In this work, we propose a learning-based approach that integrates learning and planning, and can utilize side information such as schematic roadmaps, satellite maps and GPS coordinates as a planning heuristic, without relying on them being accurate. Our method, ViKiNG, incorporates a local traversability model, which looks at the robot's current camera observation and a potential subgoal to infer how easily that subgoal can be reached, as well as a heuristic model, which looks at overhead maps and attempts to estimate the distance to the destination for various subgoals. These models are used by a heuristic planner to decide the best next subgoal in order to reach the final destination. Our method performs no explicit geometric reconstruction, utilizing only a topological representation of the environment. Despite having never seen trajectories longer than 80 meters in its training dataset, ViKiNG can leverage its image-based learned controller and goal-directed heuristic to navigate to goals up to 3 kilometers away in previously unseen environments, and exhibit complex behaviors such as probing potential paths and doubling back when they are found to be non-viable. ViKiNG is also robust to unreliable maps and GPS, since the low-level controller ultimately makes decisions based on egocentric image observations, using maps only as planning heuristics. For videos of our experiments, please check out https://sites.google.com/view/viking-release.
Semantic Segmentation by Early Region Proxy
Mar 26, 2022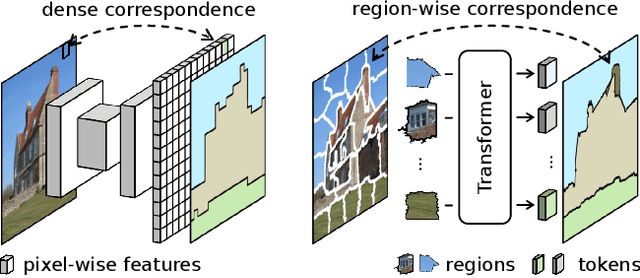
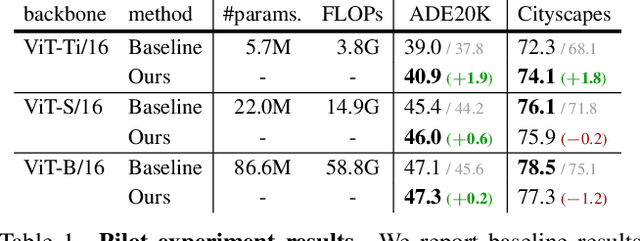
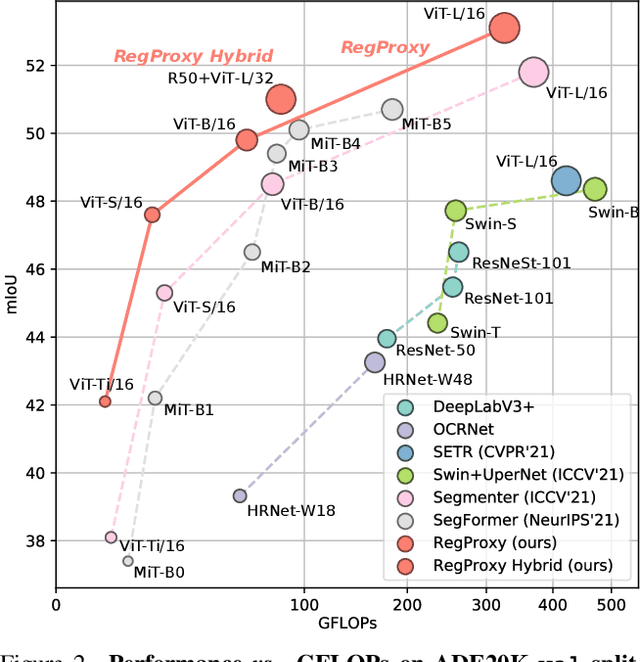
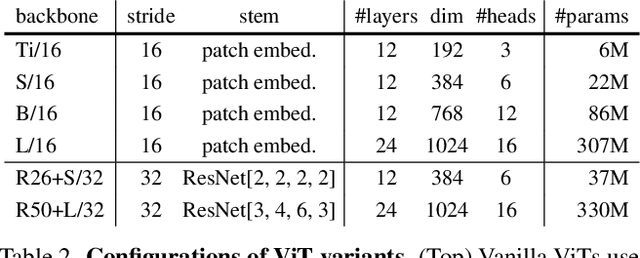
Typical vision backbones manipulate structured features. As a compromise, semantic segmentation has long been modeled as per-point prediction on dense regular grids. In this work, we present a novel and efficient modeling that starts from interpreting the image as a tessellation of learnable regions, each of which has flexible geometrics and carries homogeneous semantics. To model region-wise context, we exploit Transformer to encode regions in a sequence-to-sequence manner by applying multi-layer self-attention on the region embeddings, which serve as proxies of specific regions. Semantic segmentation is now carried out as per-region prediction on top of the encoded region embeddings using a single linear classifier, where a decoder is no longer needed. The proposed RegProxy model discards the common Cartesian feature layout and operates purely at region level. Hence, it exhibits the most competitive performance-efficiency trade-off compared with the conventional dense prediction methods. For example, on ADE20K, the small-sized RegProxy-S/16 outperforms the best CNN model using 25% parameters and 4% computation, while the largest RegProxy-L/16 achieves 52.9mIoU which outperforms the state-of-the-art by 2.1% with fewer resources. Codes and models are available at https://github.com/YiF-Zhang/RegionProxy.
Incorporating Gradient Similarity for Robust Time Delay Estimation in Ultrasound Elastography
Mar 30, 2022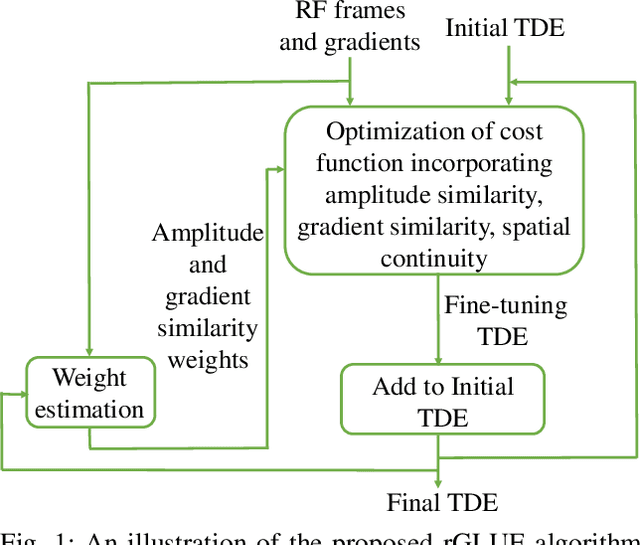
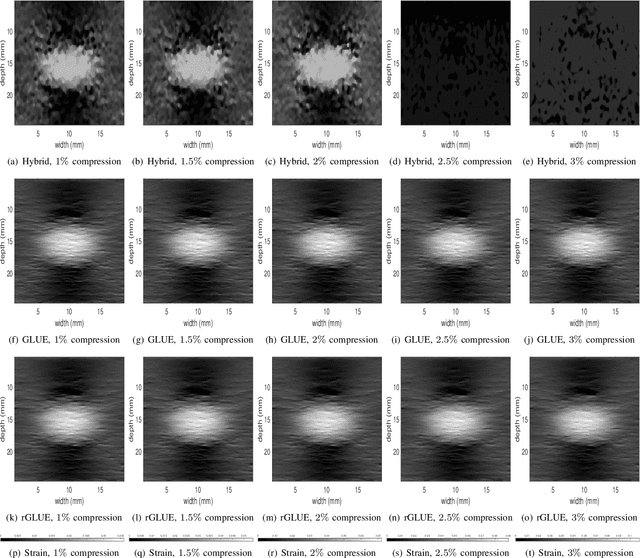

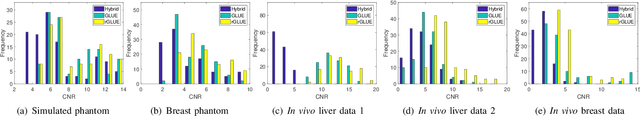
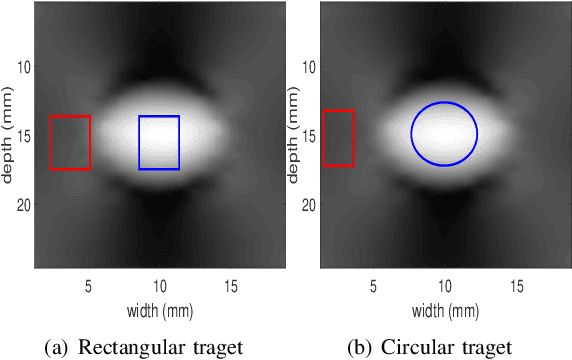
Energy-based ultrasound elastography techniques minimize a regularized cost function consisting of data and continuity terms to obtain local displacement estimates based on the local time-delay estimation (TDE) between radio-frequency (RF) frames. The data term associated with the existing techniques takes only the amplitude similarity into account and hence is not sufficiently robust to the outlier samples present in the RF frames under consideration. This drawback creates noticeable artifacts in the strain image. To resolve this issue, we propose to formulate the data function as a linear combination of the amplitude and gradient similarity constraints. We estimate the adaptive weight concerning each similarity term following an iterative scheme. Finally, we optimize the non-linear cost function in an efficient manner to convert the problem to a sparse system of linear equations which are solved for millions of variables. We call our technique rGLUE: robust data term in GLobal Ultrasound Elastography. rGLUE has been validated using simulation, phantom, in vivo liver, and breast datasets. In all of our experiments, rGLUE substantially outperforms the recent elastography methods both visually and quantitatively. For simulated, phantom, and in vivo datasets, respectively, rGLUE achieves 107%, 18%, and 23% improvements of signal-to-noise ratio (SNR) and 61%, 19%, and 25% improvements of contrast-to-noise ratio (CNR) over GLUE, a recently-published elastography algorithm.
Improving Point Cloud Based Place Recognition with Ranking-based Loss and Large Batch Training
Mar 02, 2022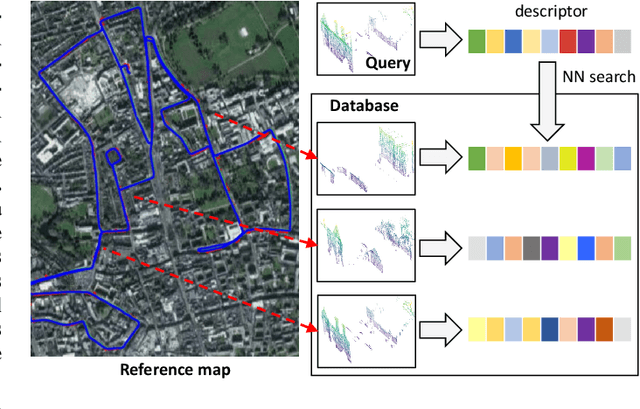
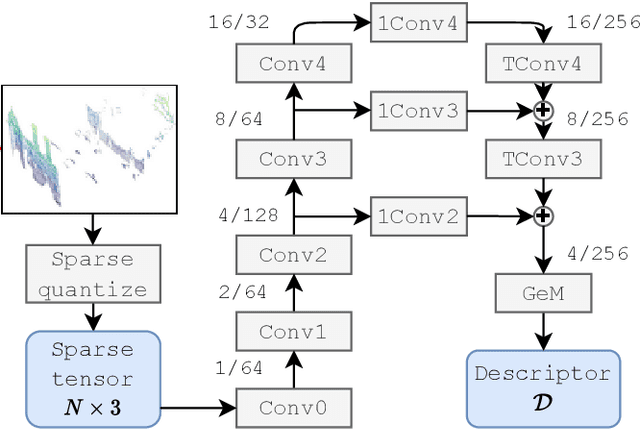
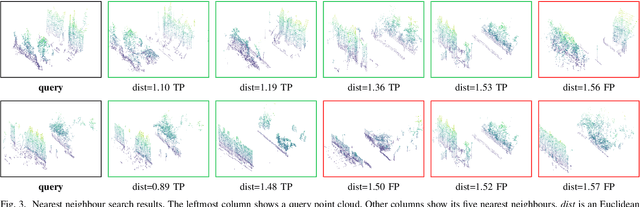
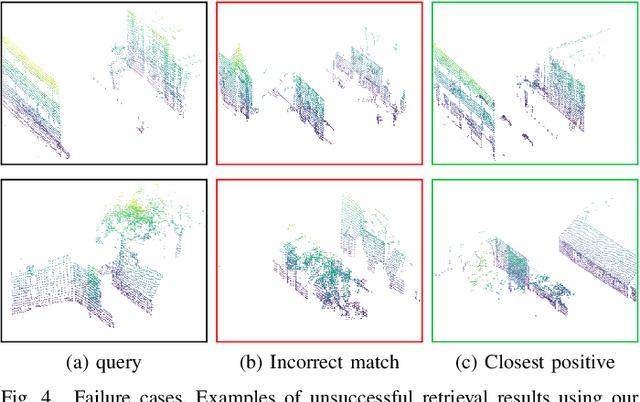
The paper presents a simple and effective learning-based method for computing a discriminative 3D point cloud descriptor for place recognition purposes. Recent state-of-the-art methods have relatively complex architectures such as multi-scale oyramid of point Transformers combined with a pyramid of feature aggregation modules. Our method uses a simple and efficient 3D convolutional feature extraction, based on a sparse voxelized representation, enhanced with channel attention blocks. We employ recent advances in image retrieval and propose a modified version of a loss function based on a differentiable average precision approximation. Such loss function requires training with very large batches for the best results. This is enabled by using multistaged backpropagation. Experimental evaluation on the popular benchmarks proves the effectiveness of our approach, with a consistent improvement over the state of the art
WSSS4LUAD: Grand Challenge on Weakly-supervised Tissue Semantic Segmentation for Lung Adenocarcinoma
Apr 13, 2022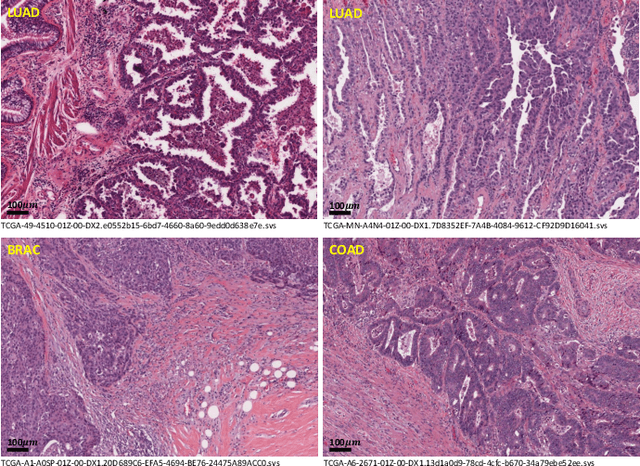
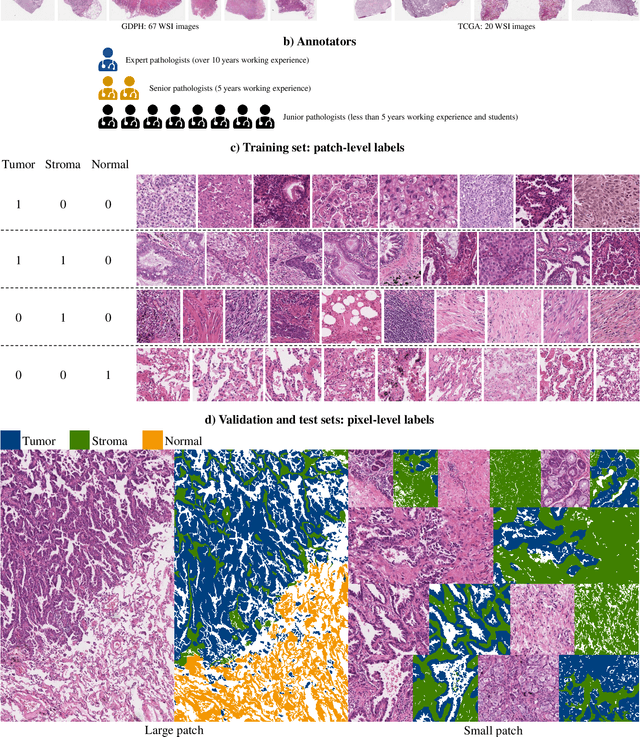
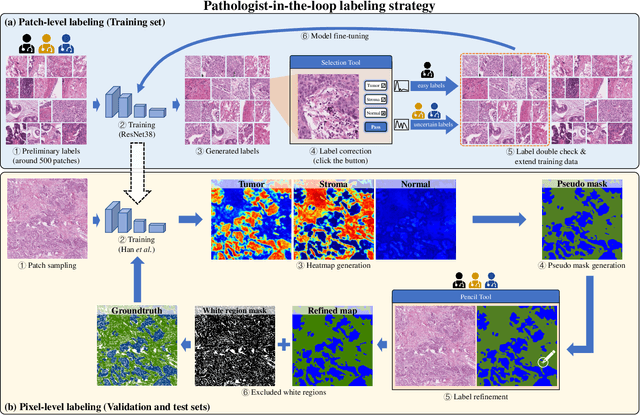
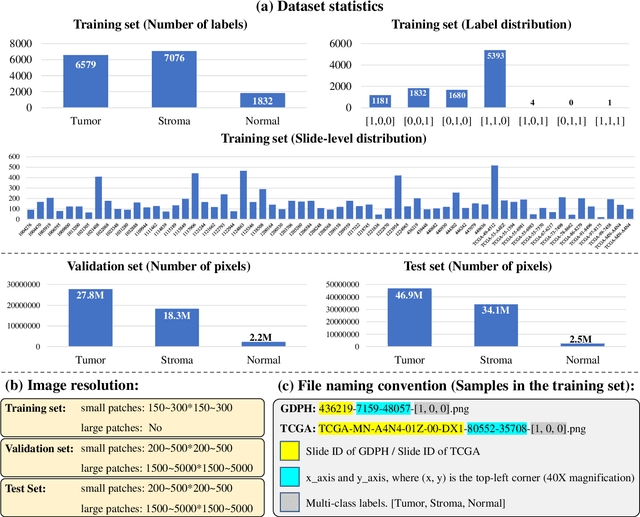
Lung cancer is the leading cause of cancer death worldwide, and adenocarcinoma (LUAD) is the most common subtype. Exploiting the potential value of the histopathology images can promote precision medicine in oncology. Tissue segmentation is the basic upstream task of histopathology image analysis. Existing deep learning models have achieved superior segmentation performance but require sufficient pixel-level annotations, which is time-consuming and expensive. To enrich the label resources of LUAD and to alleviate the annotation efforts, we organize this challenge WSSS4LUAD to call for the outstanding weakly-supervised semantic segmentation techniques for histopathology images of LUAD. Participants have to design the algorithm to segment tumor epithelial, tumor-associated stroma and normal tissue with only patch-level labels. This challenge includes 10,091 patch-level annotations (the training set) and over 130 million labeled pixels (the validation and test sets), from 67 WSIs (47 from GDPH, 20 from TCGA). All the labels were generated by a pathologist-in-the-loop pipeline with the help of AI models and checked by the label review board. Among 532 registrations, 28 teams submitted the results in the test phase with over 1,000 submissions. Finally, the first place team achieved mIoU of 0.8413 (tumor: 0.8389, stroma: 0.7931, normal: 0.8919). According to the technical reports of the top-tier teams, CAM is still the most popular approach in WSSS. Cutmix data augmentation has been widely adopted to generate more reliable samples. With the success of this challenge, we believe that WSSS approaches with patch-level annotations can replace the traditional pixel annotations while reducing the annotation efforts. The entire dataset has been released to encourage more researches on computational pathology in LUAD and more novel WSSS techniques.
MedSelect: Selective Labeling for Medical Image Classification Combining Meta-Learning with Deep Reinforcement Learning
Mar 26, 2021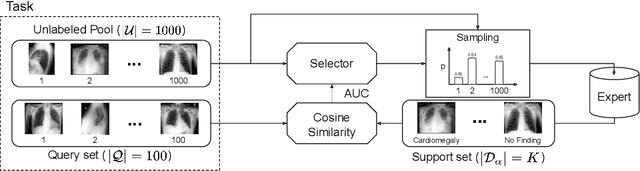
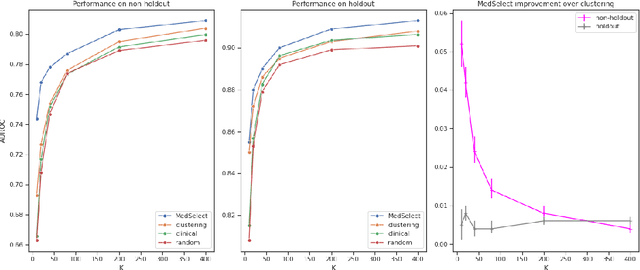
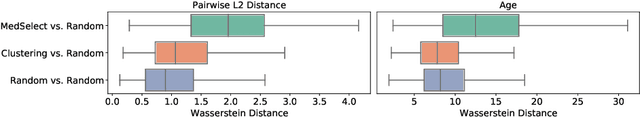
We propose a selective learning method using meta-learning and deep reinforcement learning for medical image interpretation in the setting of limited labeling resources. Our method, MedSelect, consists of a trainable deep learning selector that uses image embeddings obtained from contrastive pretraining for determining which images to label, and a non-parametric selector that uses cosine similarity to classify unseen images. We demonstrate that MedSelect learns an effective selection strategy outperforming baseline selection strategies across seen and unseen medical conditions for chest X-ray interpretation. We also perform an analysis of the selections performed by MedSelect comparing the distribution of latent embeddings and clinical features, and find significant differences compared to the strongest performing baseline. We believe that our method may be broadly applicable across medical imaging settings where labels are expensive to acquire.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge