"Image": models, code, and papers
Improving Augmentation and Evaluation Schemes for Semantic Image Synthesis
Dec 08, 2020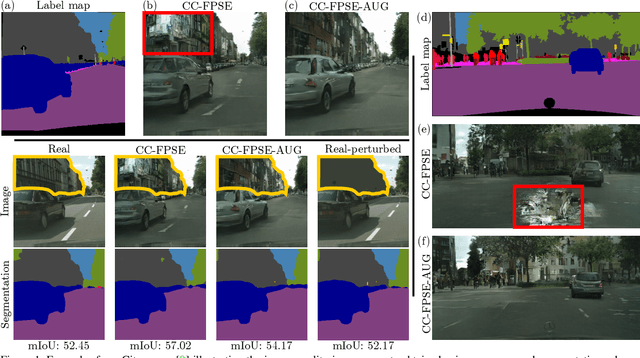
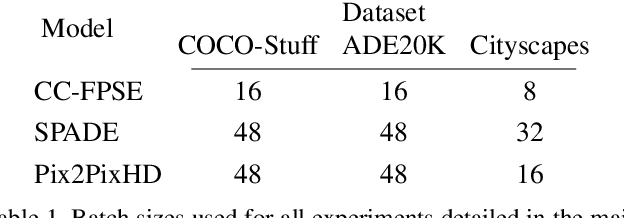

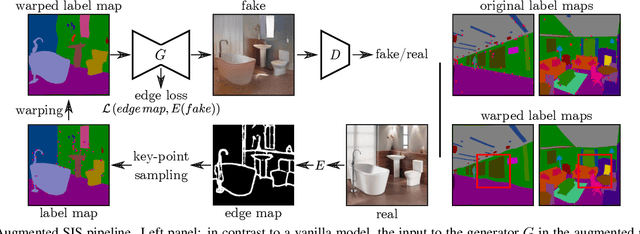
Despite data augmentation being a de facto technique for boosting the performance of deep neural networks, little attention has been paid to developing augmentation strategies for generative adversarial networks (GANs). To this end, we introduce a novel augmentation scheme designed specifically for GAN-based semantic image synthesis models. We propose to randomly warp object shapes in the semantic label maps used as an input to the generator. The local shape discrepancies between the warped and non-warped label maps and images enable the GAN to learn better the structural and geometric details of the scene and thus to improve the quality of generated images. While benchmarking the augmented GAN models against their vanilla counterparts, we discover that the quantification metrics reported in the previous semantic image synthesis studies are strongly biased towards specific semantic classes as they are derived via an external pre-trained segmentation network. We therefore propose to improve the established semantic image synthesis evaluation scheme by analyzing separately the performance of generated images on the biased and unbiased classes for the given segmentation network. Finally, we show strong quantitative and qualitative improvements obtained with our augmentation scheme, on both class splits, using state-of-the-art semantic image synthesis models across three different datasets. On average across COCO-Stuff, ADE20K and Cityscapes datasets, the augmented models outperform their vanilla counterparts by ~3 mIoU and ~10 FID points.
Auto-Encoding for Shared Cross Domain Feature Representation and Image-to-Image Translation
Jun 11, 2020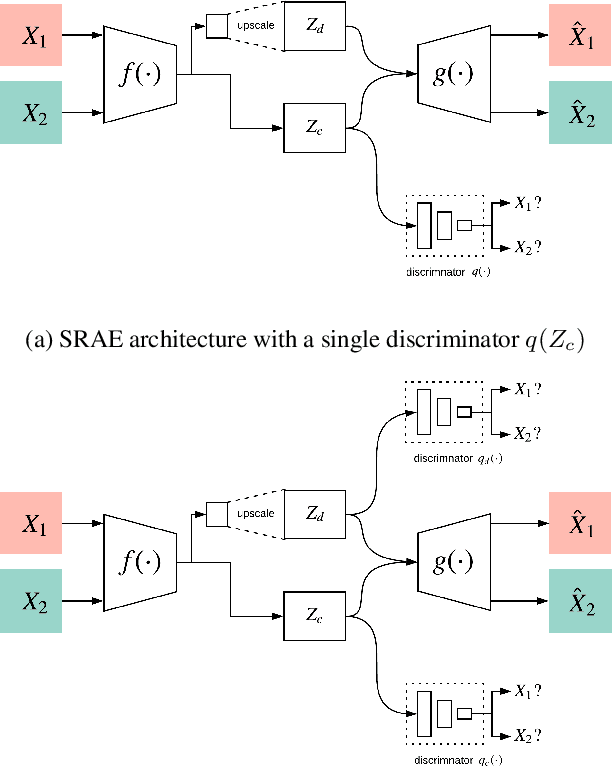
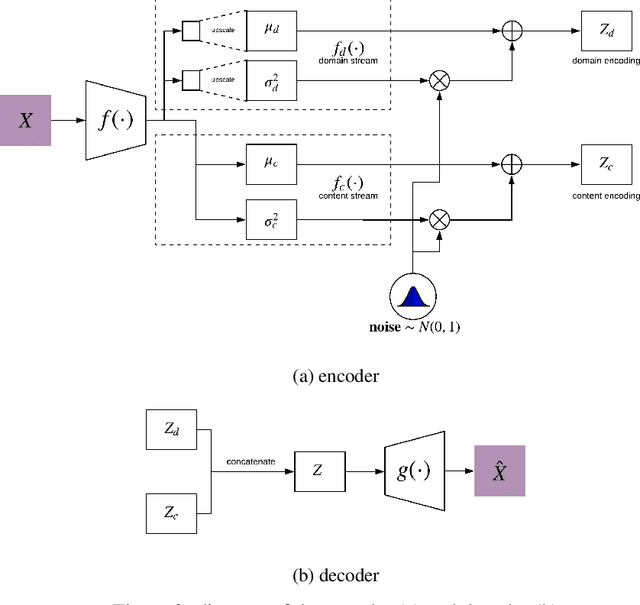
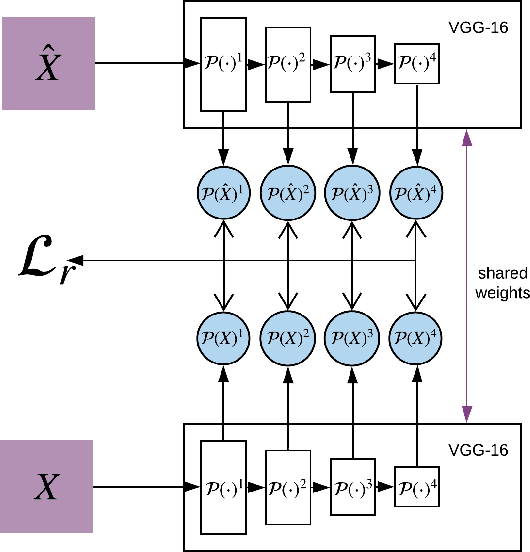

Image-to-image translation is a subset of computer vision and pattern recognition problems where our goal is to learn a mapping between input images of domain $\mathbf{X}_1$ and output images of domain $\mathbf{X}_2$. Current methods use neural networks with an encoder-decoder structure to learn a mapping $G:\mathbf{X}_1 \to\mathbf{X}_2$ such that the distribution of images from $\mathbf{X}_2$ and $G(\mathbf{X}_1)$ are identical, where $G(\mathbf{X}_1) = d_G (f_G (\mathbf{X}_1))$ and $f_G (\cdot)$ is referred as the encoder and $d_G(\cdot)$ is referred to as the decoder. Currently, such methods which also compute an inverse mapping $F:\mathbf{X}_2 \to \mathbf{X}_1$ use a separate encoder-decoder pair $d_F (f_F (\mathbf{X}_2))$ or at least a separate decoder $d_F (\cdot)$ to do so. Here we introduce a method to perform cross domain image-to-image translation across multiple domains using a single encoder-decoder architecture. We use an auto-encoder network which given an input image $\mathbf{X}_1$, first computes a latent domain encoding $Z_d = f_d (\mathbf{X}_1)$ and a latent content encoding $Z_c = f_c (\mathbf{X}_1)$, where the domain encoding $Z_d$ and content encoding $Z_c$ are independent. And then a decoder network $g(Z_d,Z_c)$ creates a reconstruction of the original image $\mathbf{\widehat{X}}_1=g(Z_d,Z_c )\approx \mathbf{X}_1$. Ideally, the domain encoding $Z_d$ contains no information regarding the content of the image and the content encoding $Z_c$ contains no information regarding the domain of the image. We use this property of the encodings to find the mapping across domains $G: X\to Y$ by simply changing the domain encoding $Z_d$ of the decoder's input. $G(\mathbf{X}_1 )=d(f_d (\mathbf{x}_2^i ),f_c (\mathbf{X}_1))$ where $\mathbf{x}_2^i$ is the $i^{th}$ observation of $\mathbf{X}_2$.
Fast Neural Architecture Search for Lightweight Dense Prediction Networks
Mar 09, 2022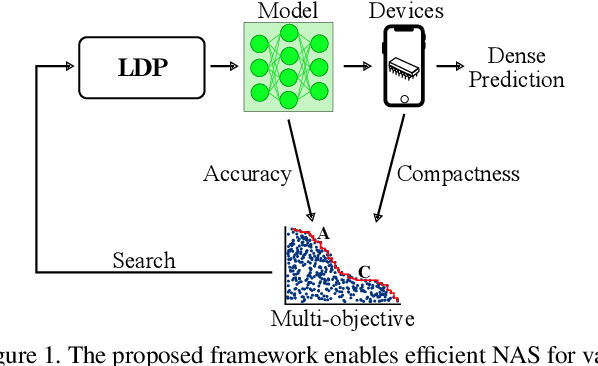
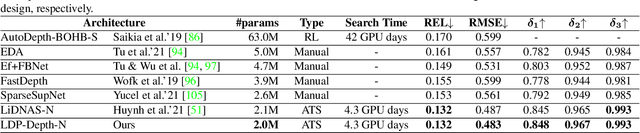
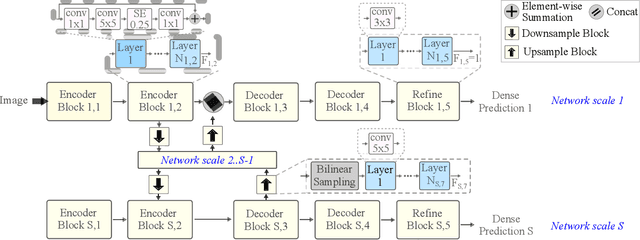
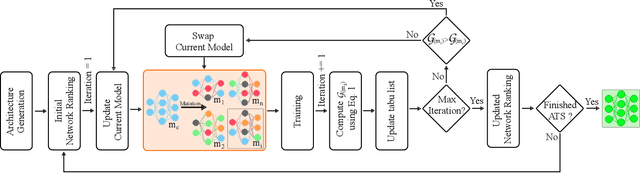
We present LDP, a lightweight dense prediction neural architecture search (NAS) framework. Starting from a pre-defined generic backbone, LDP applies the novel Assisted Tabu Search for efficient architecture exploration. LDP is fast and suitable for various dense estimation problems, unlike previous NAS methods that are either computational demanding or deployed only for a single subtask. The performance of LPD is evaluated on monocular depth estimation, semantic segmentation, and image super-resolution tasks on diverse datasets, including NYU-Depth-v2, KITTI, Cityscapes, COCO-stuff, DIV2K, Set5, Set14, BSD100, Urban100. Experiments show that the proposed framework yields consistent improvements on all tested dense prediction tasks, while being $5\%-315\%$ more compact in terms of the number of model parameters than prior arts.
Survival Analysis for Idiopathic Pulmonary Fibrosis using CT Images and Incomplete Clinical Data
Mar 21, 2022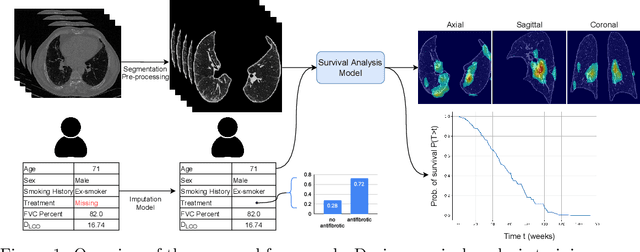
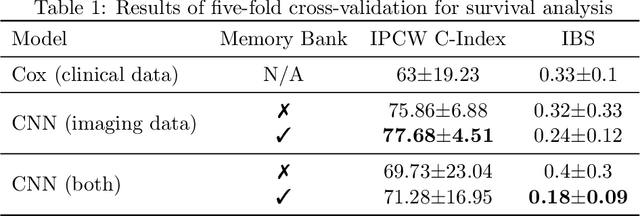
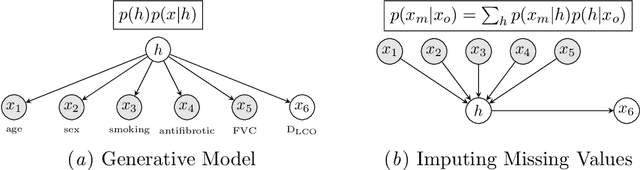

Idiopathic Pulmonary Fibrosis (IPF) is an inexorably progressive fibrotic lung disease with a variable and unpredictable rate of progression. CT scans of the lungs inform clinical assessment of IPF patients and contain pertinent information related to disease progression. In this work, we propose a multi-modal method that uses neural networks and memory banks to predict the survival of IPF patients using clinical and imaging data. The majority of clinical IPF patient records have missing data (e.g. missing lung function tests). To this end, we propose a probabilistic model that captures the dependencies between the observed clinical variables and imputes missing ones. This principled approach to missing data imputation can be naturally combined with a deep survival analysis model. We show that the proposed framework yields significantly better survival analysis results than baselines in terms of concordance index and integrated Brier score. Our work also provides insights into novel image-based biomarkers that are linked to mortality.
Practical Noise Simulation for RGB Images
Jan 30, 2022This document describes a noise generator that simulates realistic noise found in smartphone cameras. The generator simulates Poissonian-Gaussian noise whose parameters have been estimated on the Smartphone Image Denoising Dataset (SIDD). The generator is available online, and is currently being used in compressed-domain denoising exploration experiments in JPEG AI.
Image Translation for Medical Image Generation -- Ischemic Stroke Lesions
Oct 05, 2020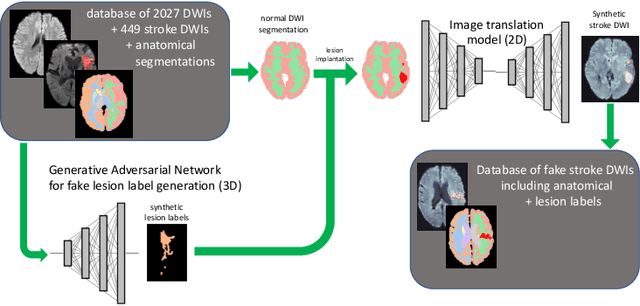
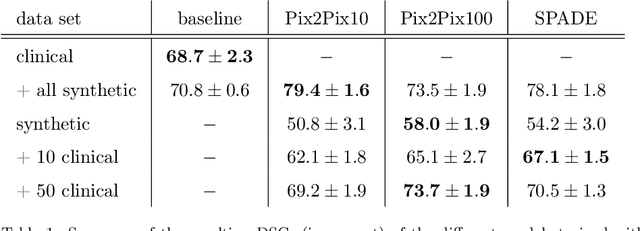
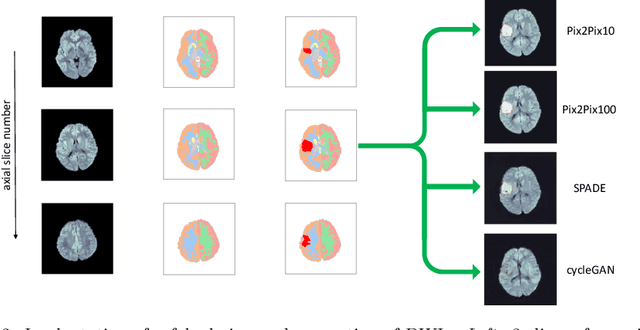
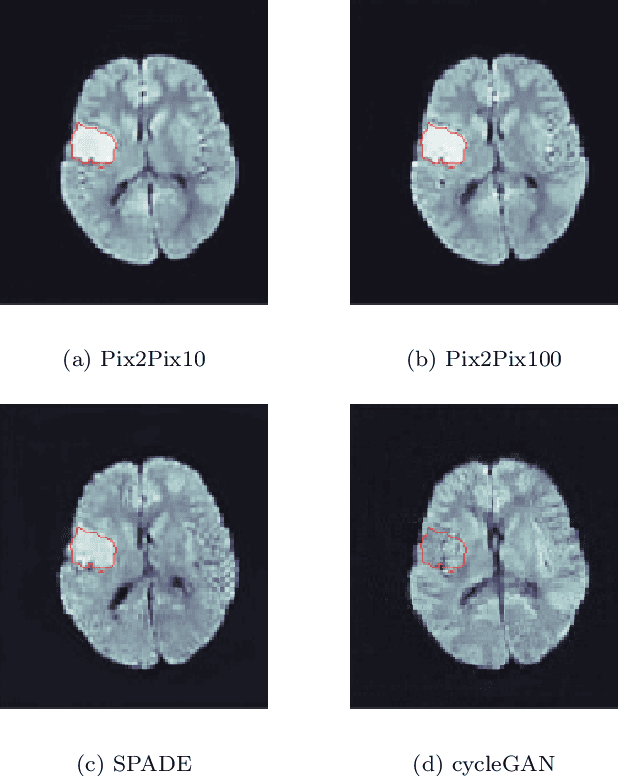
Deep learning-based automated disease detection and segmentation algorithms promise to accelerate and improve many clinical processes. However, such algorithms require vast amounts of annotated training data, which are typically not available in a medical context, e.g., due to data privacy concerns, legal obstructions, and non-uniform data formats. Synthetic databases of annotated pathologies could provide the required amounts of training data. Here, we demonstrate with the example of ischemic stroke that a significant improvement in lesion segmentation is feasible using deep learning-based data augmentation. To this end, we train different image-to-image translation models to synthesize diffusion-weighted magnetic resonance images (DWIs) of brain volumes with and without stroke lesions from semantic segmentation maps. In addition, we train a generative adversarial network to generate synthetic lesion masks. Subsequently, we combine these two components to build a large database of synthetic stroke DWIs. The performance of the various generative models is evaluated using a U-Net which is trained to segment stroke lesions on a clinical test set. We compare the results to human expert inter-reader scores. For the model with the best performance, we report a maximum Dice score of 82.6\%, which significantly outperforms the model trained on the clinical images alone (74.8\%), and also the inter-reader Dice score of two human readers of 76.9\%. Moreover, we show that for a very limited database of only 10 or 50 clinical cases, synthetic data can be used to pre-train the segmentation algorithms, which ultimately yields an improvement by a factor of as high as 8 compared to a setting where no synthetic data is used.
MOST-Net: A Memory Oriented Style Transfer Network for Face Sketch Synthesis
Feb 08, 2022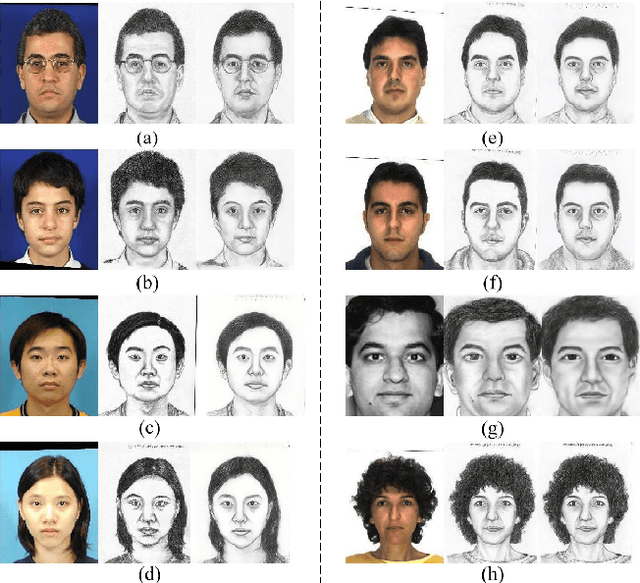
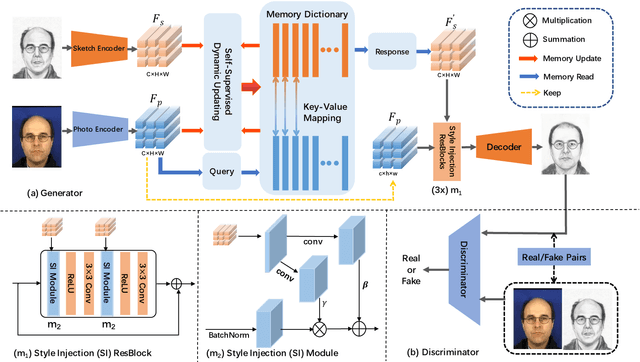
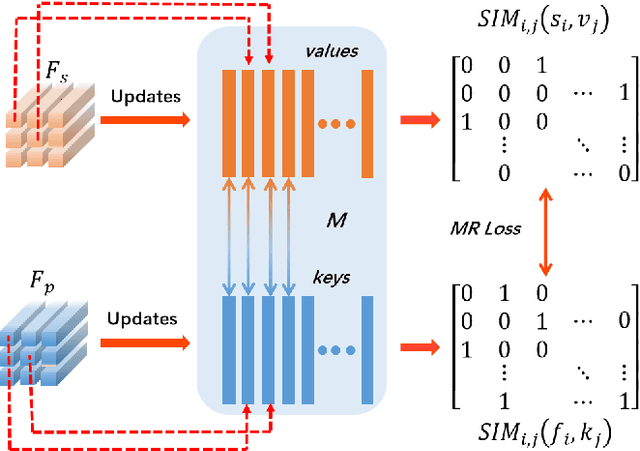
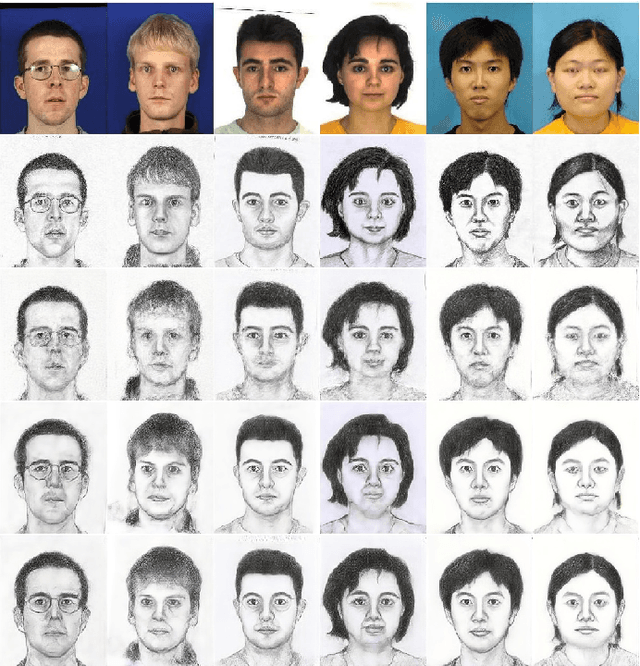
Face sketch synthesis has been widely used in multi-media entertainment and law enforcement. Despite the recent developments in deep neural networks, accurate and realistic face sketch synthesis is still a challenging task due to the diversity and complexity of human faces. Current image-to-image translation-based face sketch synthesis frequently encounters over-fitting problems when it comes to small-scale datasets. To tackle this problem, we present an end-to-end Memory Oriented Style Transfer Network (MOST-Net) for face sketch synthesis which can produce high-fidelity sketches with limited data. Specifically, an external self-supervised dynamic memory module is introduced to capture the domain alignment knowledge in the long term. In this way, our proposed model could obtain the domain-transfer ability by establishing the durable relationship between faces and corresponding sketches on the feature level. Furthermore, we design a novel Memory Refinement Loss (MR Loss) for feature alignment in the memory module, which enhances the accuracy of memory slots in an unsupervised manner. Extensive experiments on the CUFS and the CUFSF datasets show that our MOST-Net achieves state-of-the-art performance, especially in terms of the Structural Similarity Index(SSIM).
CapillaryX: A Software Design Pattern for Analyzing Medical Images in Real-time using Deep Learning
Apr 13, 2022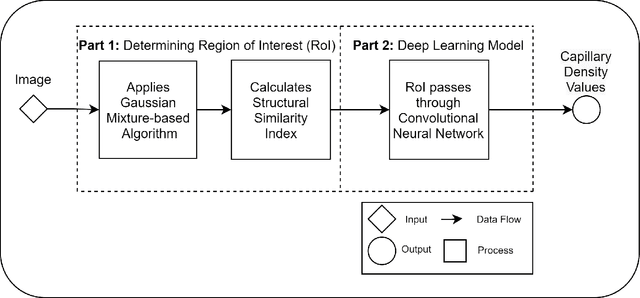
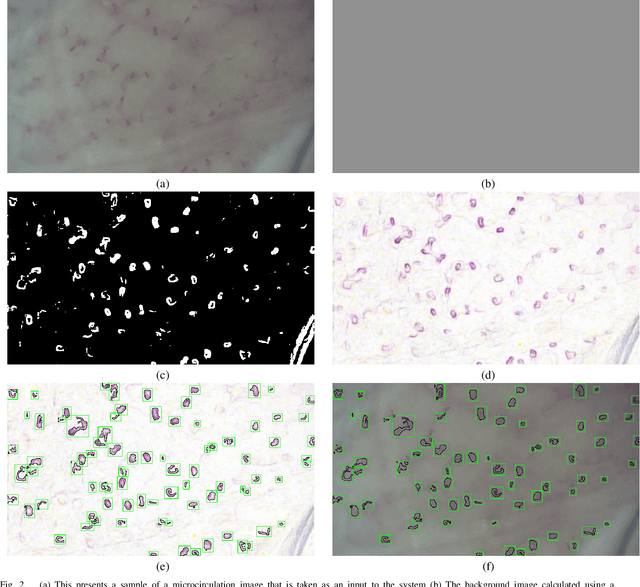
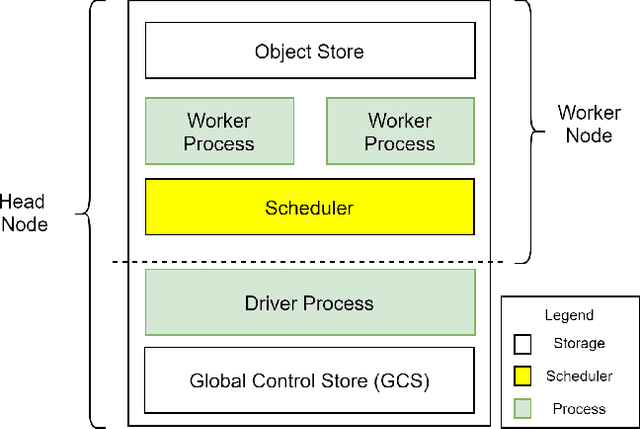
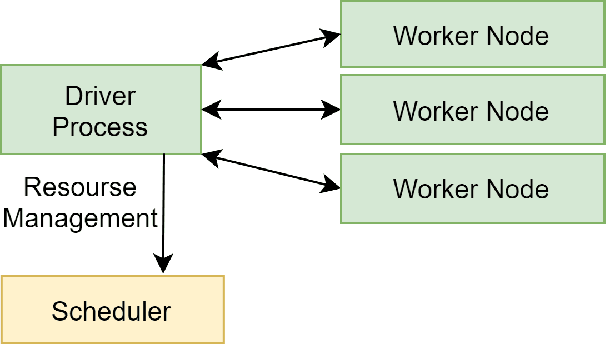
Recent advances in digital imaging, e.g., increased number of pixels captured, have meant that the volume of data to be processed and analyzed from these images has also increased. Deep learning algorithms are state-of-the-art for analyzing such images, given their high accuracy when trained with a large data volume of data. Nevertheless, such analysis requires considerable computational power, making such algorithms time- and resource-demanding. Such high demands can be met by using third-party cloud service providers. However, analyzing medical images using such services raises several legal and privacy challenges and does not necessarily provide real-time results. This paper provides a computing architecture that locally and in parallel can analyze medical images in real-time using deep learning thus avoiding the legal and privacy challenges stemming from uploading data to a third-party cloud provider. To make local image processing efficient on modern multi-core processors, we utilize parallel execution to offset the resource-intensive demands of deep neural networks. We focus on a specific medical-industrial case study, namely the quantifying of blood vessels in microcirculation images for which we have developed a working system. It is currently used in an industrial, clinical research setting as part of an e-health application. Our results show that our system is approximately 78% faster than its serial system counterpart and 12% faster than a master-slave parallel system architecture.
Fetal Brain Tissue Annotation and Segmentation Challenge Results
Apr 20, 2022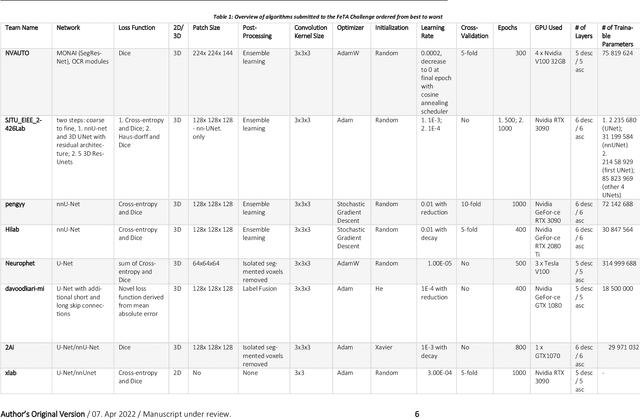
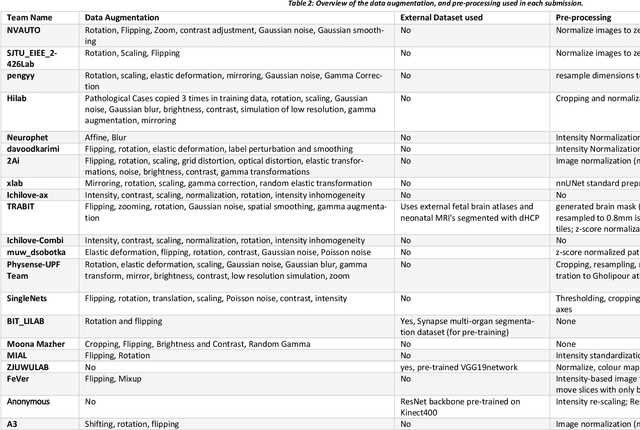
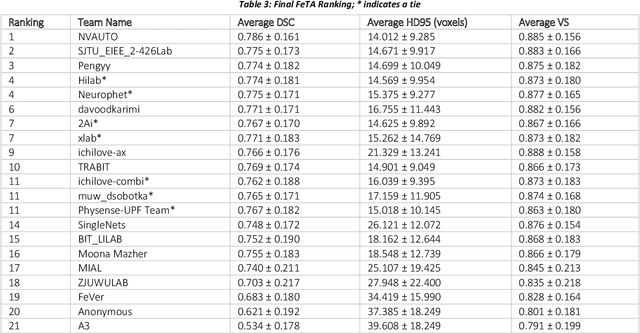
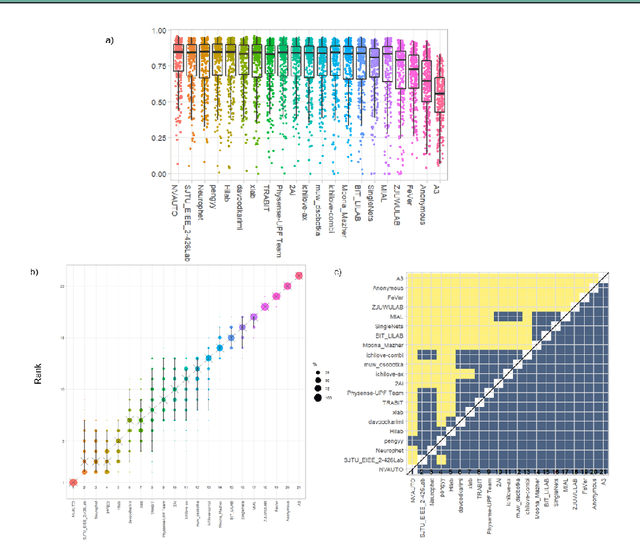
In-utero fetal MRI is emerging as an important tool in the diagnosis and analysis of the developing human brain. Automatic segmentation of the developing fetal brain is a vital step in the quantitative analysis of prenatal neurodevelopment both in the research and clinical context. However, manual segmentation of cerebral structures is time-consuming and prone to error and inter-observer variability. Therefore, we organized the Fetal Tissue Annotation (FeTA) Challenge in 2021 in order to encourage the development of automatic segmentation algorithms on an international level. The challenge utilized FeTA Dataset, an open dataset of fetal brain MRI reconstructions segmented into seven different tissues (external cerebrospinal fluid, grey matter, white matter, ventricles, cerebellum, brainstem, deep grey matter). 20 international teams participated in this challenge, submitting a total of 21 algorithms for evaluation. In this paper, we provide a detailed analysis of the results from both a technical and clinical perspective. All participants relied on deep learning methods, mainly U-Nets, with some variability present in the network architecture, optimization, and image pre- and post-processing. The majority of teams used existing medical imaging deep learning frameworks. The main differences between the submissions were the fine tuning done during training, and the specific pre- and post-processing steps performed. The challenge results showed that almost all submissions performed similarly. Four of the top five teams used ensemble learning methods. However, one team's algorithm performed significantly superior to the other submissions, and consisted of an asymmetrical U-Net network architecture. This paper provides a first of its kind benchmark for future automatic multi-tissue segmentation algorithms for the developing human brain in utero.
CNN Filter DB: An Empirical Investigation of Trained Convolutional Filters
Apr 09, 2022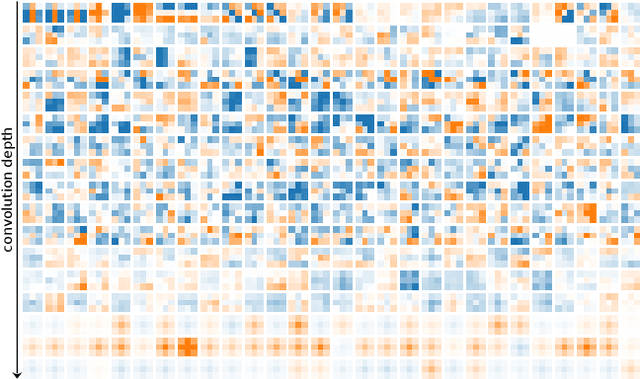

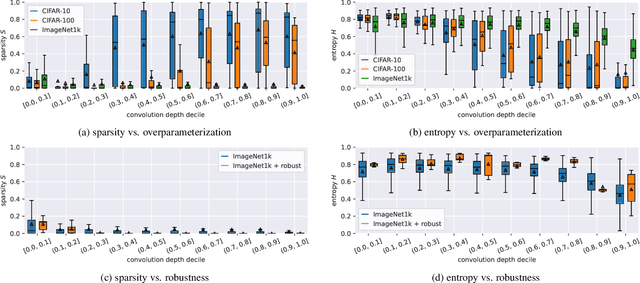
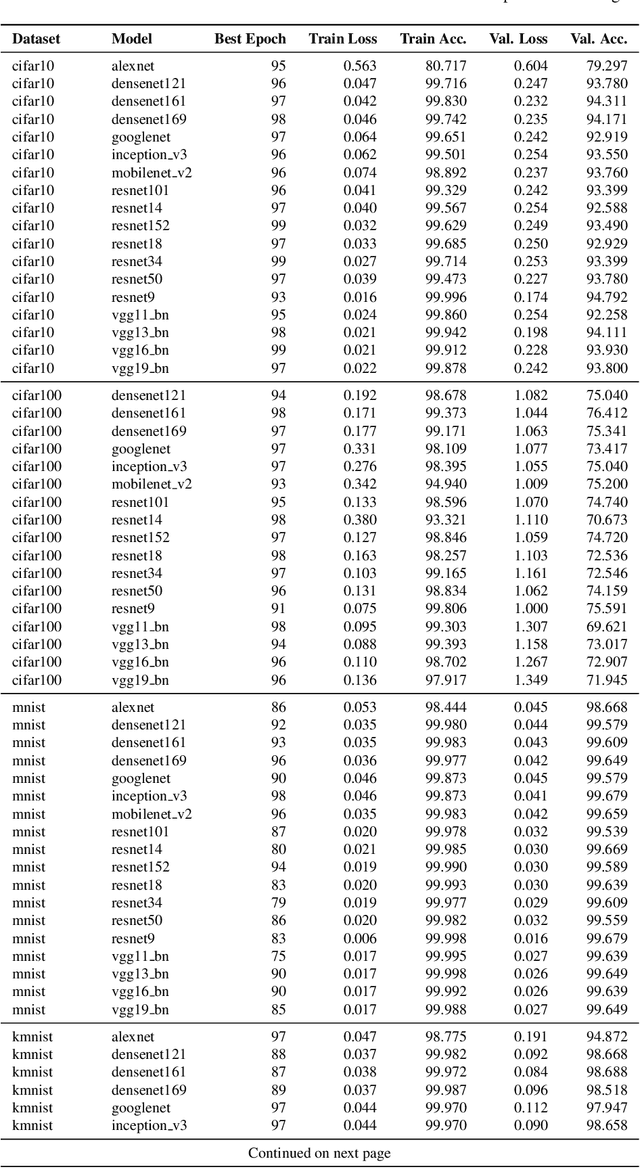
Currently, many theoretical as well as practically relevant questions towards the transferability and robustness of Convolutional Neural Networks (CNNs) remain unsolved. While ongoing research efforts are engaging these problems from various angles, in most computer vision related cases these approaches can be generalized to investigations of the effects of distribution shifts in image data. In this context, we propose to study the shifts in the learned weights of trained CNN models. Here we focus on the properties of the distributions of dominantly used 3x3 convolution filter kernels. We collected and publicly provide a dataset with over 1.4 billion filters from hundreds of trained CNNs, using a wide range of datasets, architectures, and vision tasks. In a first use case of the proposed dataset, we can show highly relevant properties of many publicly available pre-trained models for practical applications: I) We analyze distribution shifts (or the lack thereof) between trained filters along different axes of meta-parameters, like visual category of the dataset, task, architecture, or layer depth. Based on these results, we conclude that model pre-training can succeed on arbitrary datasets if they meet size and variance conditions. II) We show that many pre-trained models contain degenerated filters which make them less robust and less suitable for fine-tuning on target applications. Data & Project website: https://github.com/paulgavrikov/cnn-filter-db
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge