"Image": models, code, and papers
Transfer Learning from Synthetic In-vitro Soybean Pods Dataset for In-situ Segmentation of On-branch Soybean Pod
Apr 22, 2022
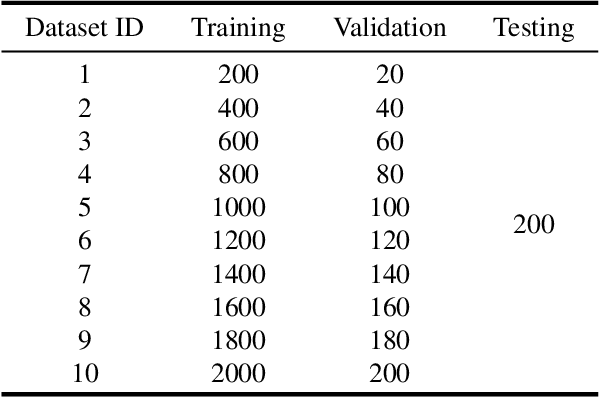
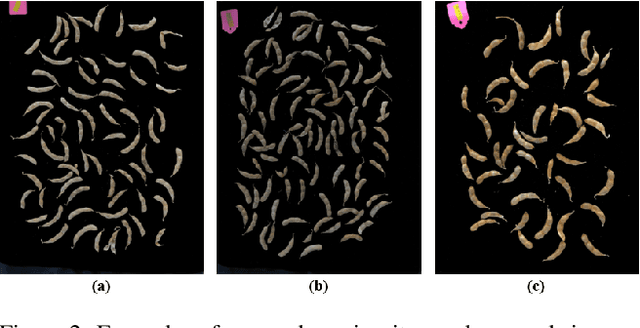
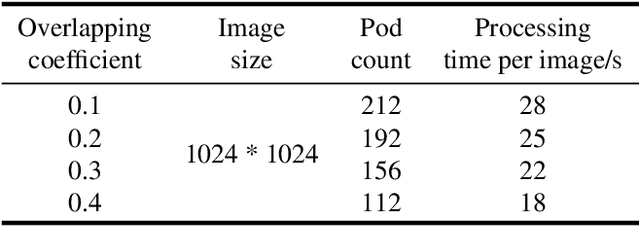
The mature soybean plants are of complex architecture with pods frequently touching each other, posing a challenge for in-situ segmentation of on-branch soybean pods. Deep learning-based methods can achieve accurate training and strong generalization capabilities, but it demands massive labeled data, which is often a limitation, especially for agricultural applications. As lacking the labeled data to train an in-situ segmentation model for on-branch soybean pods, we propose a transfer learning from synthetic in-vitro soybean pods. First, we present a novel automated image generation method to rapidly generate a synthetic in-vitro soybean pods dataset with plenty of annotated samples. The in-vitro soybean pods samples are overlapped to simulate the frequently physically touching of on-branch soybean pods. Then, we design a two-step transfer learning. In the first step, we finetune an instance segmentation network pretrained by a source domain (MS COCO dataset) with a synthetic target domain (in-vitro soybean pods dataset). In the second step, transferring from simulation to reality is performed by finetuning on a few real-world mature soybean plant samples. The experimental results show the effectiveness of the proposed two-step transfer learning method, such that AP$_{50}$ was 0.80 for the real-world mature soybean plant test dataset, which is higher than that of direct adaptation and its AP$_{50}$ was 0.77. Furthermore, the visualizations of in-situ segmentation results of on-branch soybean pods show that our method performs better than other methods, especially when soybean pods overlap densely.
Sampling Bias Correction for Supervised Machine Learning: A Bayesian Inference Approach with Practical Applications
Mar 11, 2022Given a supervised machine learning problem where the training set has been subject to a known sampling bias, how can a model be trained to fit the original dataset? We achieve this through the Bayesian inference framework by altering the posterior distribution to account for the sampling function. We then apply this solution to binary logistic regression, and discuss scenarios where a dataset might be subject to intentional sample bias such as label imbalance. This technique is widely applicable for statistical inference on big data, from the medical sciences to image recognition to marketing. Familiarity with it will give the practitioner tools to improve their inference pipeline from data collection to model selection.
A study on the effects of compression on hyperspectral image classification
Apr 01, 2021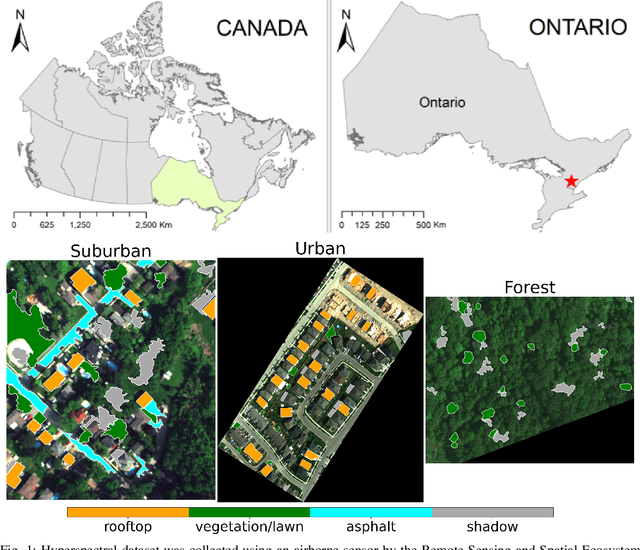
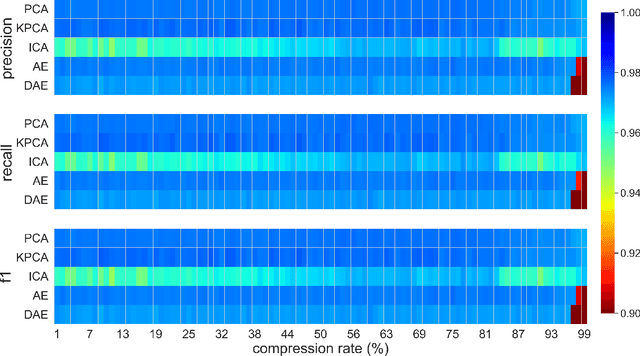
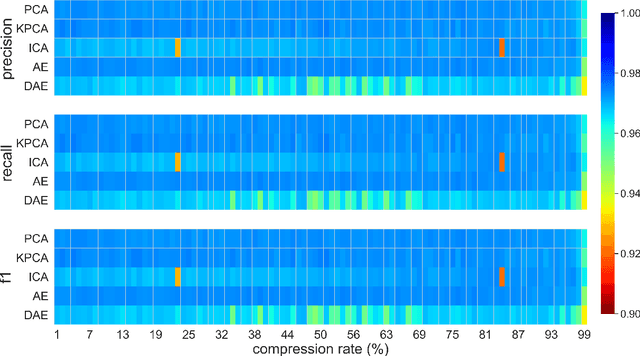
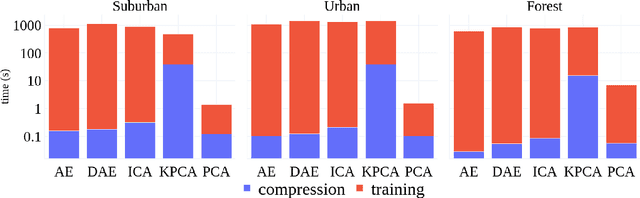
This paper presents a systematic study the effects of compression on hyperspectral pixel classification task. We use five dimensionality reduction methods -- PCA, KPCA, ICA, AE, and DAE -- to compress 301-dimensional hyperspectral pixels. Compressed pixels are subsequently used to perform pixel-based classifications. Pixel classification accuracies together with compression method, compression rates, and reconstruction errors provide a new lens to study the suitability of a compression method for the task of pixel-based classification. We use three high-resolution hyperspectral image datasets, representing three common landscape units (i.e. urban, transitional suburban, and forests) collected by the Remote Sensing and Spatial Ecosystem Modeling laboratory of the University of Toronto. We found that PCA, KPCA, and ICA post greater signal reconstruction capability; however, when compression rate is more than 90\% those methods showed lower classification scores. AE and DAE methods post better classification accuracy at 95\% compression rate, however decreasing again at 97\%, suggesting a sweet-spot at the 95\% mark. Our results demonstrate that the choice of a compression method with the compression rate are important considerations when designing a hyperspectral image classification pipeline.
Light-weight Document Image Cleanup using Perceptual Loss
May 19, 2021
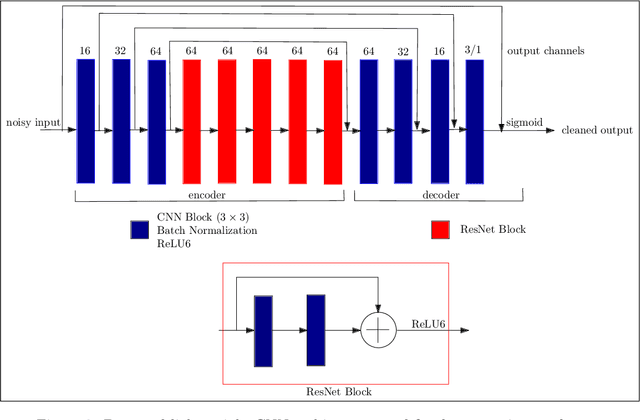
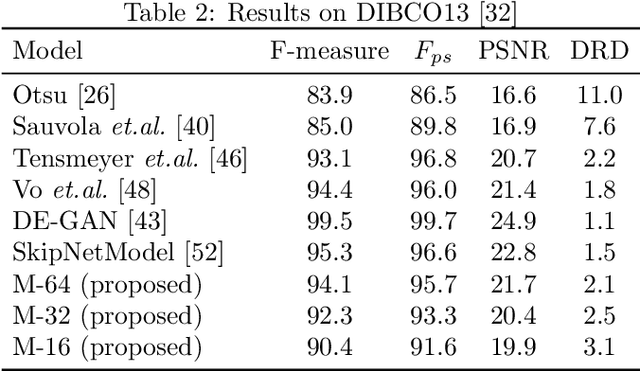

Smartphones have enabled effortless capturing and sharing of documents in digital form. The documents, however, often undergo various types of degradation due to aging, stains, or shortcoming of capturing environment such as shadow, non-uniform lighting, etc., which reduces the comprehensibility of the document images. In this work, we consider the problem of document image cleanup on embedded applications such as smartphone apps, which usually have memory, energy, and latency limitations due to the device and/or for best human user experience. We propose a light-weight encoder decoder based convolutional neural network architecture for removing the noisy elements from document images. To compensate for generalization performance with a low network capacity, we incorporate the perceptual loss for knowledge transfer from pre-trained deep CNN network in our loss function. In terms of the number of parameters and product-sum operations, our models are 65-1030 and 3-27 times, respectively, smaller than existing state-of-the-art document enhancement models. Overall, the proposed models offer a favorable resource versus accuracy trade-off and we empirically illustrate the efficacy of our approach on several real-world benchmark datasets.
CAiD: Context-Aware Instance Discrimination for Self-supervised Learning in Medical Imaging
Apr 15, 2022
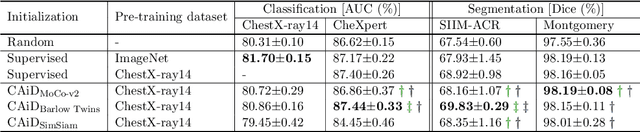
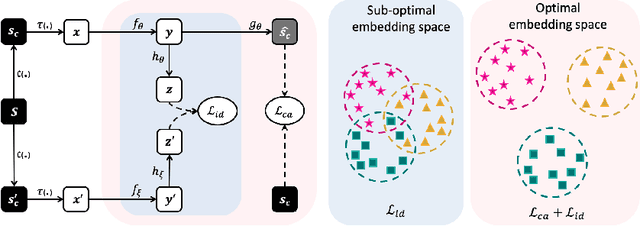
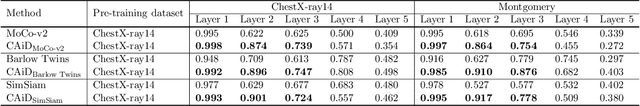
Recently, self-supervised instance discrimination methods have achieved significant success in learning visual representations from unlabeled photographic images. However, given the marked differences between photographic and medical images, the efficacy of instance-based objectives, focusing on learning the most discriminative global features in the image (i.e., wheels in bicycle), remains unknown in medical imaging. Our preliminary analysis showed that high global similarity of medical images in terms of anatomy hampers instance discrimination methods for capturing a set of distinct features, negatively impacting their performance on medical downstream tasks. To alleviate this limitation, we have developed a simple yet effective self-supervised framework, called Context-Aware instance Discrimination (CAiD). CAiD aims to improve instance discrimination learning by providing finer and more discriminative information encoded from a diverse local context of unlabeled medical images. We conduct a systematic analysis to investigate the utility of the learned features from a three-pronged perspective: (i) generalizability and transferability, (ii) separability in the embedding space, and (iii) reusability. Our extensive experiments demonstrate that CAiD (1) enriches representations learned from existing instance discrimination methods; (2) delivers more discriminative features by adequately capturing finer contextual information from individual medial images; and (3) improves reusability of low/mid-level features compared to standard instance discriminative methods. As open science, all codes and pre-trained models are available on our GitHub page: https://github.com/JLiangLab/CAiD.
Label uncertainty-guided multi-stream model for disease screening
Jan 28, 2022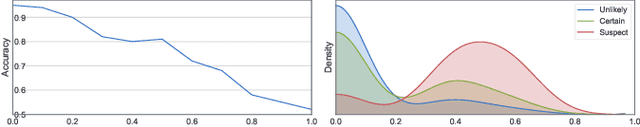
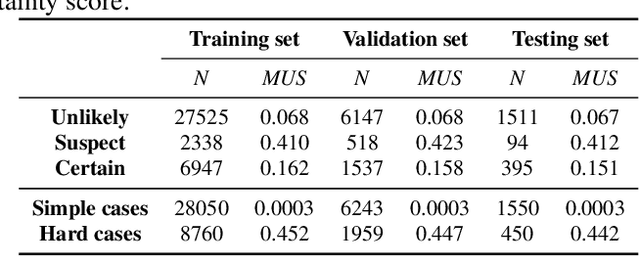
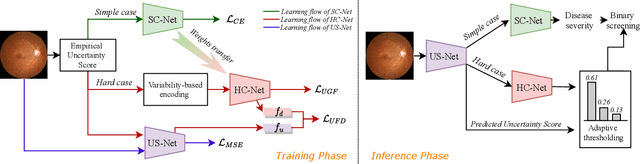
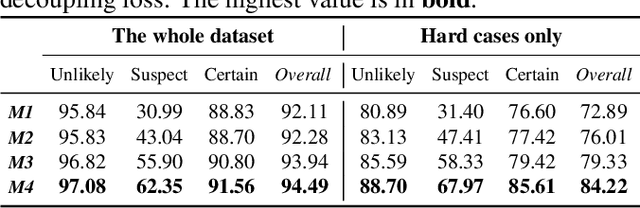
The annotation of disease severity for medical image datasets often relies on collaborative decisions from multiple human graders. The intra-observer variability derived from individual differences always persists in this process, yet the influence is often underestimated. In this paper, we cast the intra-observer variability as an uncertainty problem and incorporate the label uncertainty information as guidance into the disease screening model to improve the final decision. The main idea is dividing the images into simple and hard cases by uncertainty information, and then developing a multi-stream network to deal with different cases separately. Particularly, for hard cases, we strengthen the network's capacity in capturing the correct disease features and resisting the interference of uncertainty. Experiments on a fundus image-based glaucoma screening case study show that the proposed model outperforms several baselines, especially in screening hard cases.
Effective Mutation Rate Adaptation through Group Elite Selection
Apr 11, 2022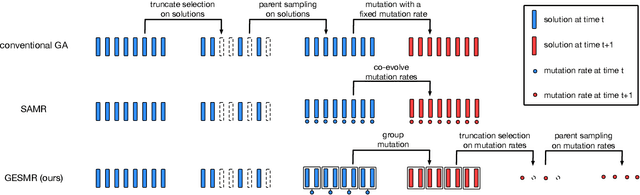
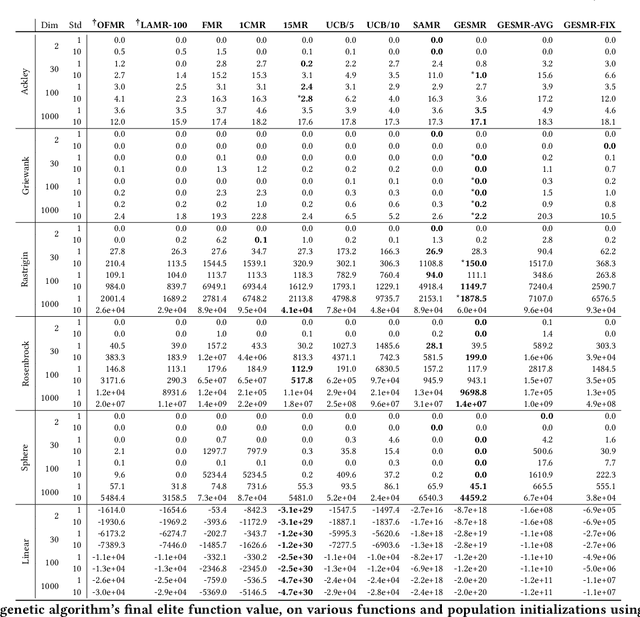
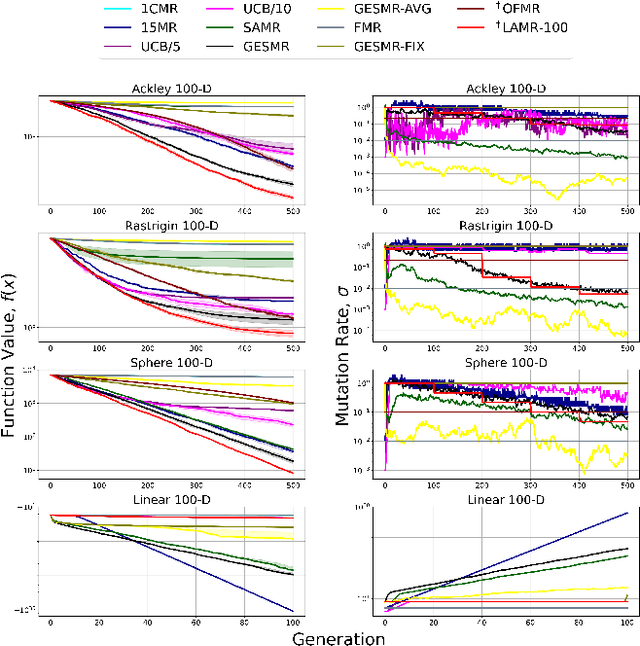
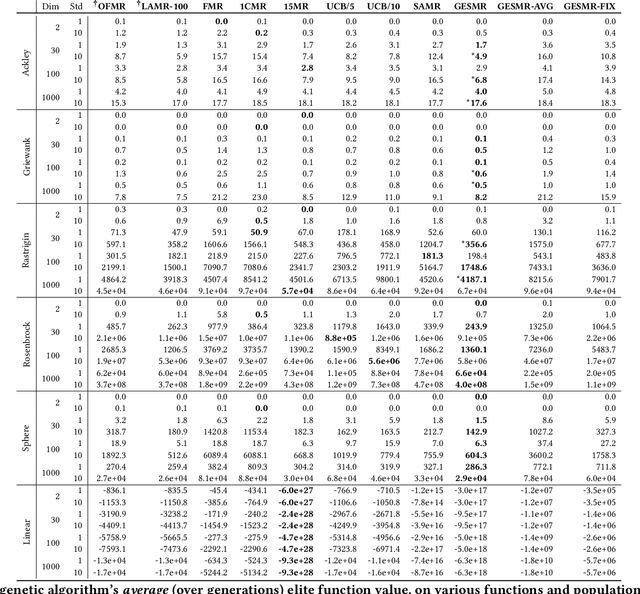
Evolutionary algorithms are sensitive to the mutation rate (MR); no single value of this parameter works well across domains. Self-adaptive MR approaches have been proposed but they tend to be brittle: Sometimes they decay the MR to zero, thus halting evolution. To make self-adaptive MR robust, this paper introduces the Group Elite Selection of Mutation Rates (GESMR) algorithm. GESMR co-evolves a population of solutions and a population of MRs, such that each MR is assigned to a group of solutions. The resulting best mutational change in the group, instead of average mutational change, is used for MR selection during evolution, thus avoiding the vanishing MR problem. With the same number of function evaluations and with almost no overhead, GESMR converges faster and to better solutions than previous approaches on a wide range of continuous test optimization problems. GESMR also scales well to high-dimensional neuroevolution for supervised image-classification tasks and for reinforcement learning control tasks. Remarkably, GESMR produces MRs that are optimal in the long-term, as demonstrated through a comprehensive look-ahead grid search. Thus, GESMR and its theoretical and empirical analysis demonstrate how self-adaptation can be harnessed to improve performance in several applications of evolutionary computation.
On the Neural Tangent Kernel Analysis of Randomly Pruned Wide Neural Networks
Apr 06, 2022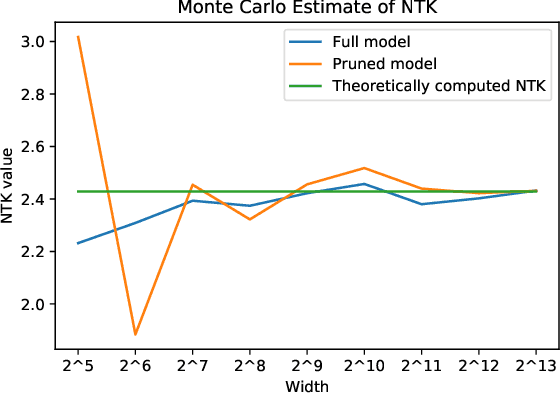

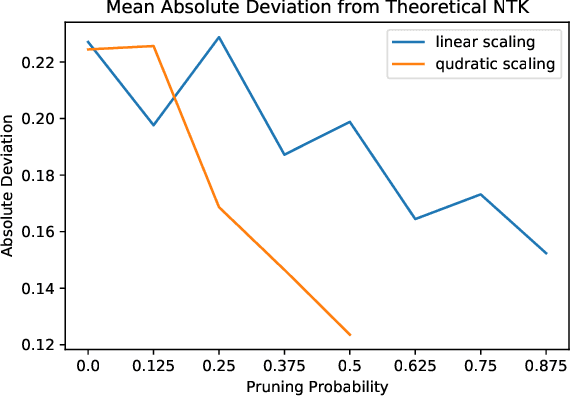
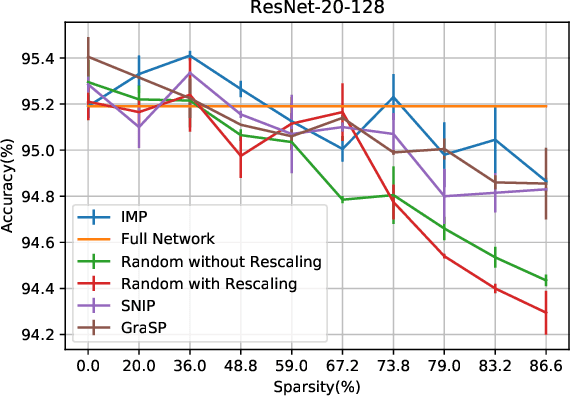
We study the behavior of ultra-wide neural networks when their weights are randomly pruned at the initialization, through the lens of neural tangent kernels (NTKs). We show that for fully-connected neural networks when the network is pruned randomly at the initialization, as the width of each layer grows to infinity, the empirical NTK of the pruned neural network converges to that of the original (unpruned) network with some extra scaling factor. Further, if we apply some appropriate scaling after pruning at the initialization, the empirical NTK of the pruned network converges to the exact NTK of the original network, and we provide a non-asymptotic bound on the approximation error in terms of pruning probability. Moreover, when we apply our result to an unpruned network (i.e., we set the probability of pruning a given weight to be zero), our analysis is optimal up to a logarithmic factor in width compared with the result in \cite{arora2019exact}. We conduct experiments to validate our theoretical results. We further test our theory by evaluating random pruning across different architectures via image classification on MNIST and CIFAR-10 and compare its performance with other pruning strategies.
FrequencyLowCut Pooling -- Plug & Play against Catastrophic Overfitting
Apr 01, 2022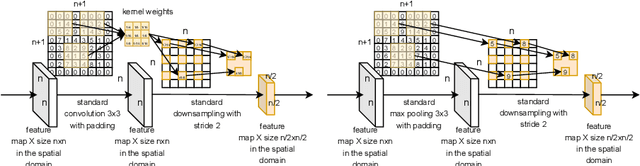
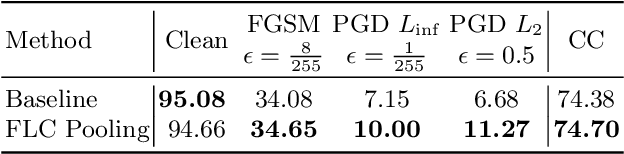

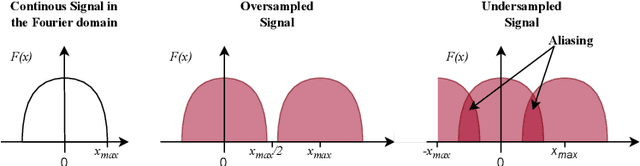
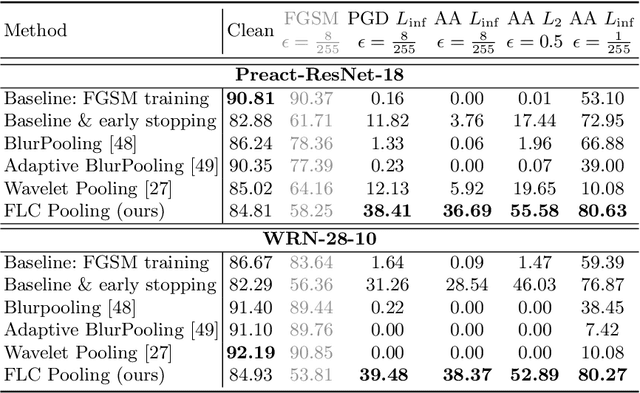
Over the last years, Convolutional Neural Networks (CNNs) have been the dominating neural architecture in a wide range of computer vision tasks. From an image and signal processing point of view, this success might be a bit surprising as the inherent spatial pyramid design of most CNNs is apparently violating basic signal processing laws, i.e. Sampling Theorem in their down-sampling operations. However, since poor sampling appeared not to affect model accuracy, this issue has been broadly neglected until model robustness started to receive more attention. Recent work [17] in the context of adversarial attacks and distribution shifts, showed after all, that there is a strong correlation between the vulnerability of CNNs and aliasing artifacts induced by poor down-sampling operations. This paper builds on these findings and introduces an aliasing free down-sampling operation which can easily be plugged into any CNN architecture: FrequencyLowCut pooling. Our experiments show, that in combination with simple and fast FGSM adversarial training, our hyper-parameter free operator significantly improves model robustness and avoids catastrophic overfitting.
Ensembles of Vision Transformers as a New Paradigm for Automated Classification in Ecology
Mar 03, 2022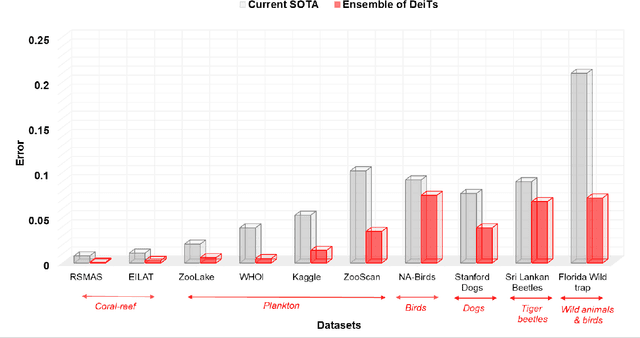
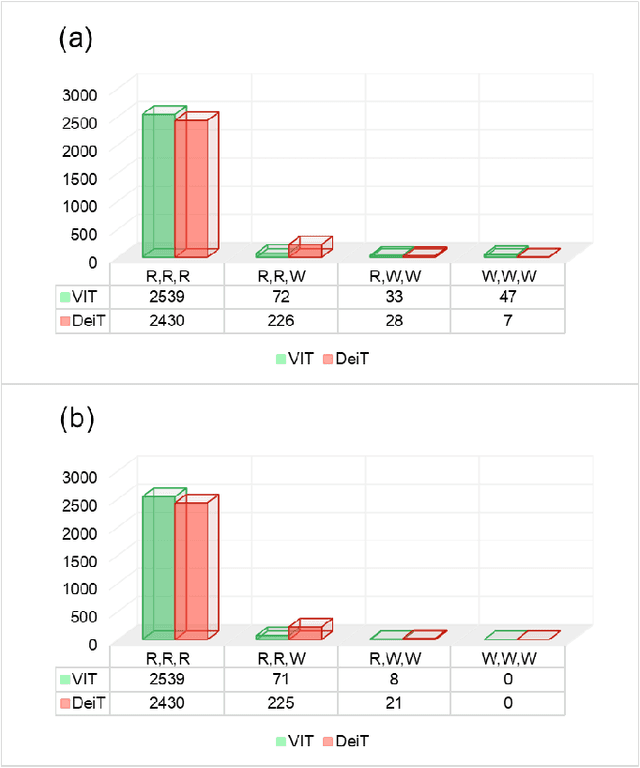
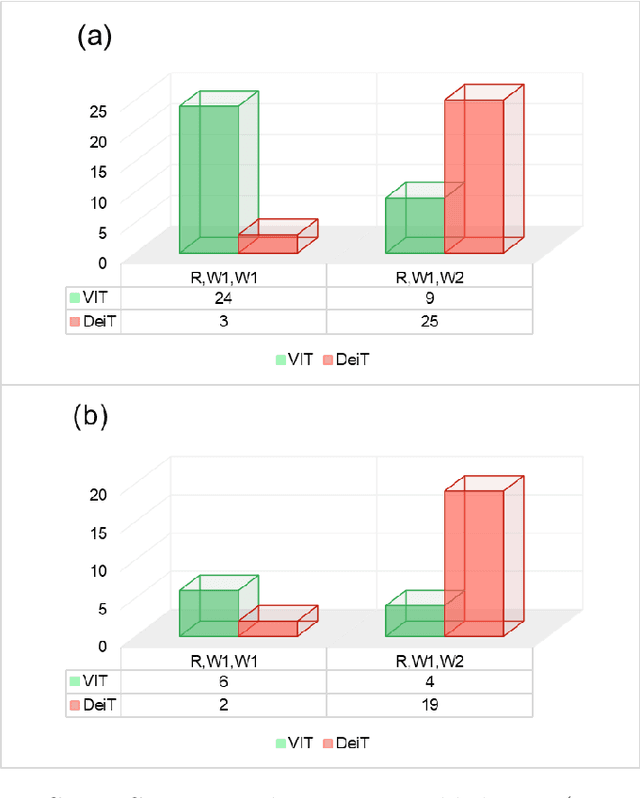
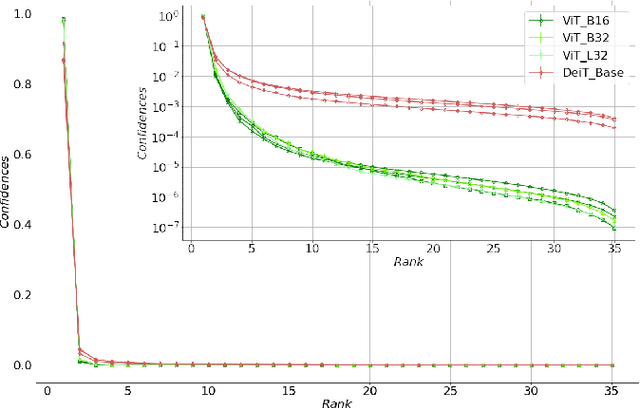
Monitoring biodiversity is paramount to manage and protect natural resources, particularly in times of global change. Collecting images of organisms over large temporal or spatial scales is a promising practice to monitor and study biodiversity change of natural ecosystems, providing large amounts of data with minimal interference with the environment. Deep learning models are currently used to automate classification of organisms into taxonomic units. However, imprecision in these classifiers introduce a measurement noise that is difficult to control and can significantly hinder the analysis and interpretation of data. In our study, we show that this limitation can be overcome by ensembles of Data-efficient image Transformers (DeiTs), which significantly outperform the previous state of the art (SOTA). We validate our results on a large number of ecological imaging datasets of diverse origin, and organisms of study ranging from plankton to insects, birds, dog breeds, animals in the wild, and corals. On all the data sets we test, we achieve a new SOTA, with a reduction of the error with respect to the previous SOTA ranging from 18.48% to 87.50%, depending on the data set, and often achieving performances very close to perfect classification. The main reason why ensembles of DeiTs perform better is not due to the single-model performance of DeiTs, but rather to the fact that predictions by independent models have a smaller overlap, and this maximizes the profit gained by ensembling. This positions DeiT ensembles as the best candidate for image classification in biodiversity monitoring.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge