"Image": models, code, and papers
VoLux-GAN: A Generative Model for 3D Face Synthesis with HDRI Relighting
Jan 13, 2022
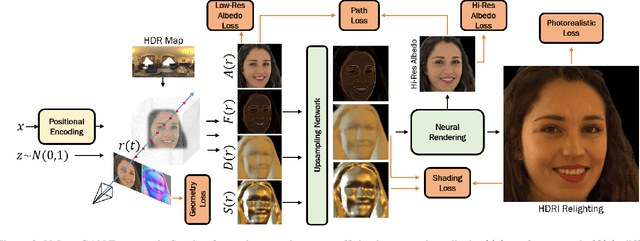
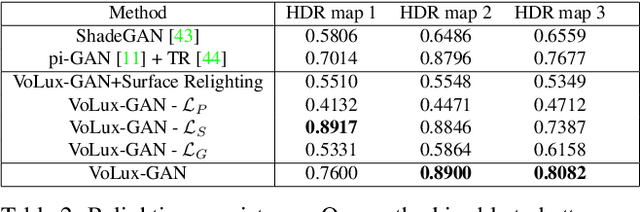

We propose VoLux-GAN, a generative framework to synthesize 3D-aware faces with convincing relighting. Our main contribution is a volumetric HDRI relighting method that can efficiently accumulate albedo, diffuse and specular lighting contributions along each 3D ray for any desired HDR environmental map. Additionally, we show the importance of supervising the image decomposition process using multiple discriminators. In particular, we propose a data augmentation technique that leverages recent advances in single image portrait relighting to enforce consistent geometry, albedo, diffuse and specular components. Multiple experiments and comparisons with other generative frameworks show how our model is a step forward towards photorealistic relightable 3D generative models.
Pneumonia Detection in Chest X-Rays using Neural Networks
Apr 07, 2022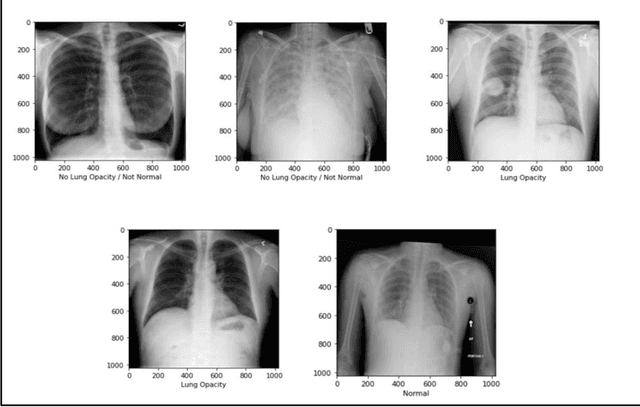
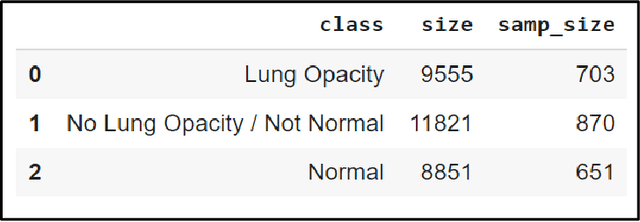
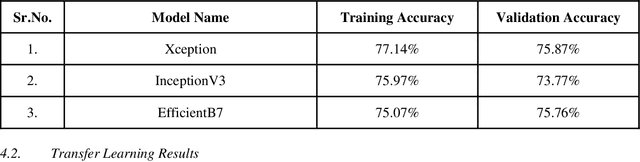
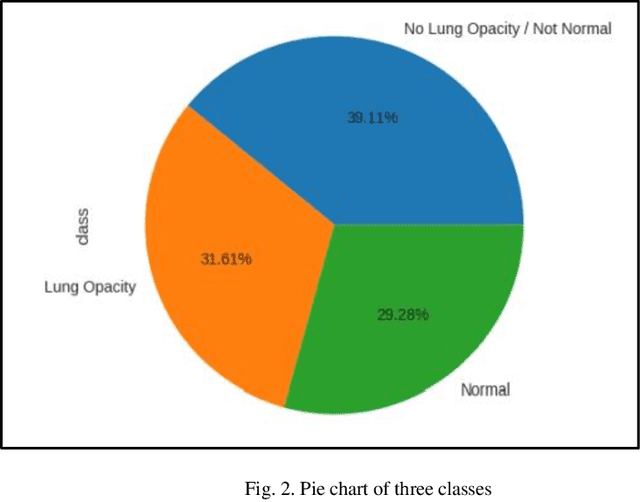

With the advancement in AI, deep learning techniques are widely used to design robust classification models in several areas such as medical diagnosis tasks in which it achieves good performance. In this paper, we have proposed the CNN model (Convolutional Neural Network) for the classification of Chest X-ray images for Radiological Society of North America Pneumonia (RSNA) datasets. The study also tries to achieve the same RSNA benchmark results using the limited computational resources by trying out various approaches to the methodologies that have been implemented in recent years. The proposed method is based on a non-complex CNN and the use of transfer learning algorithms like Xception, InceptionV3/V4, EfficientNetB7. Along with this, the study also tries to achieve the same RSNA benchmark results using the limited computational resources by trying out various approaches to the methodologies that have been implemented in recent years. The RSNA benchmark MAP score is 0.25, but using the Mask RCNN model on a stratified sample of 3017 along with image augmentation gave a MAP score of 0.15. Meanwhile, the YoloV3 without any hyperparameter tuning gave the MAP score of 0.32 but still, the loss keeps decreasing. Running the model for a greater number of iterations can give better results.
I-Nema: A Biological Image Dataset for Nematode Recognition
Mar 15, 2021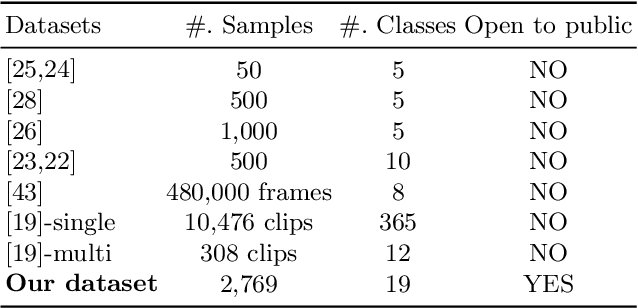
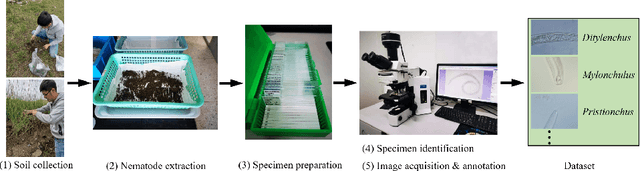
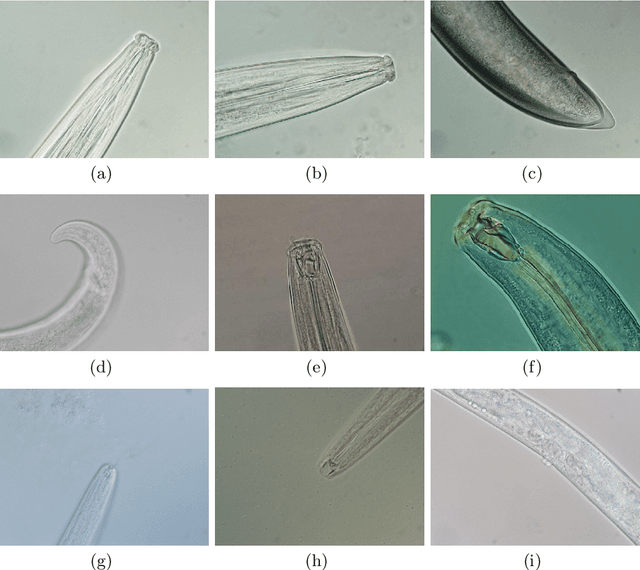
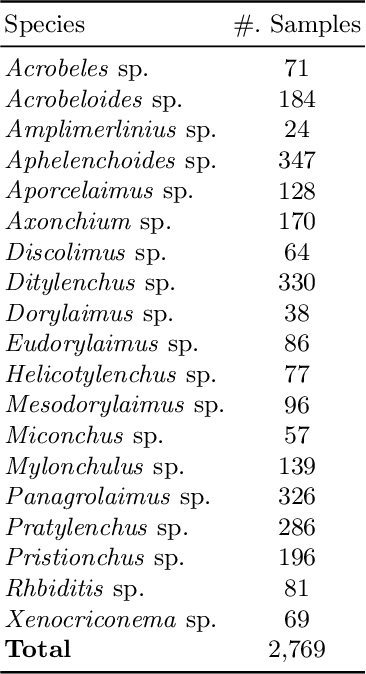
Nematode worms are one of most abundant metazoan groups on the earth, occupying diverse ecological niches. Accurate recognition or identification of nematodes are of great importance for pest control, soil ecology, bio-geography, habitat conservation and against climate changes. Computer vision and image processing have witnessed a few successes in species recognition of nematodes; however, it is still in great demand. In this paper, we identify two main bottlenecks: (1) the lack of a publicly available imaging dataset for diverse species of nematodes (especially the species only found in natural environment) which requires considerable human resources in field work and experts in taxonomy, and (2) the lack of a standard benchmark of state-of-the-art deep learning techniques on this dataset which demands the discipline background in computer science. With these in mind, we propose an image dataset consisting of diverse nematodes (both laboratory cultured and naturally isolated), which, to our knowledge, is the first time in the community. We further set up a species recognition benchmark by employing state-of-the-art deep learning networks on this dataset. We discuss the experimental results, compare the recognition accuracy of different networks, and show the challenges of our dataset. We make our dataset publicly available at: https://github.com/xuequanlu/I-Nema
Category Guided Attention Network for Brain Tumor Segmentation in MRI
Mar 29, 2022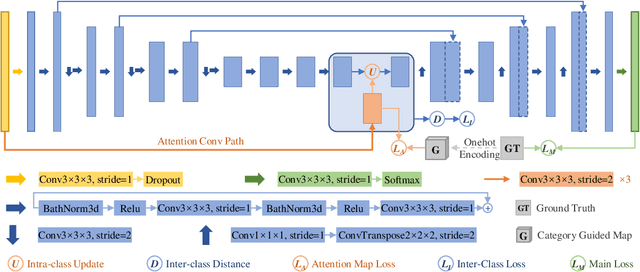
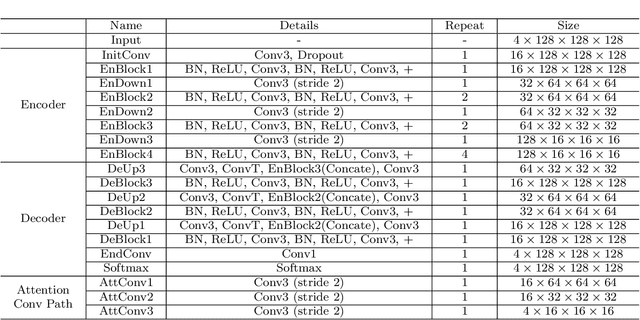
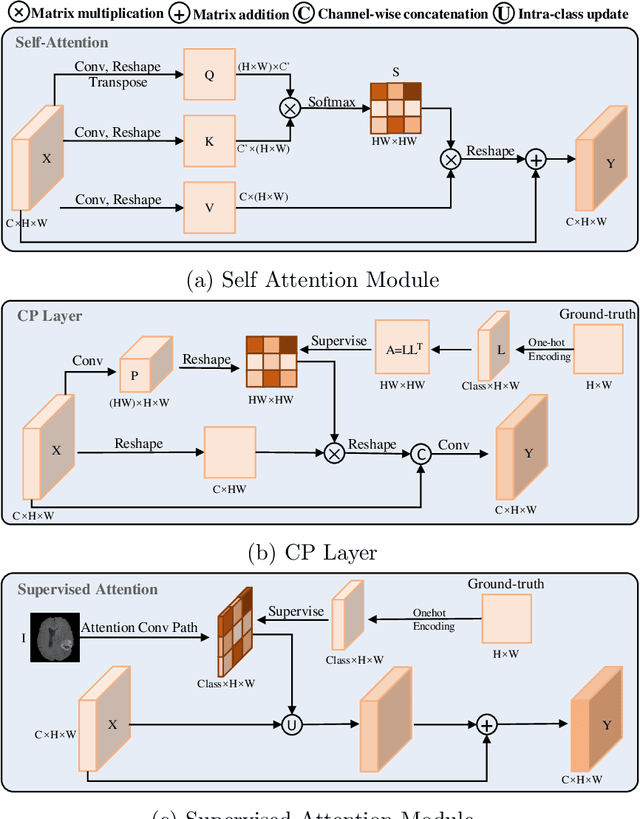
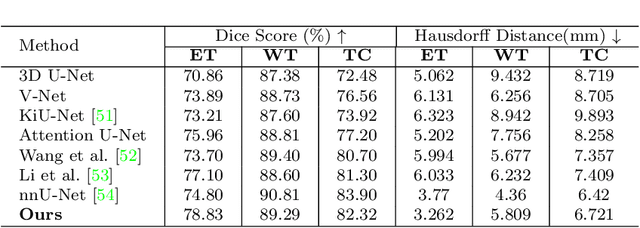
Objective: Magnetic resonance imaging (MRI) has been widely used for the analysis and diagnosis of brain diseases. Accurate and automatic brain tumor segmentation is of paramount importance for radiation treatment. However, low tissue contrast in tumor regions makes it a challenging task.Approach: We propose a novel segmentation network named Category Guided Attention U-Net (CGA U-Net). In this model, we design a Supervised Attention Module (SAM) based on the attention mechanism, which can capture more accurate and stable long-range dependency in feature maps without introducing much computational cost. Moreover, we propose an intra-class update approach to reconstruct feature maps by aggregating pixels of the same category. Main results: Experimental results on the BraTS 2019 datasets show that the proposed method outperformers the state-of-the-art algorithms in both segmentation performance and computational complexity. Significance: The CGA U-Net can effectively capture the global semantic information in the MRI image by using the SAM module, while significantly reducing the computational cost. Code is available at https://github.com/delugewalker/CGA-U-Net.
Super-Efficient Super Resolution for Fast Adversarial Defense at the Edge
Dec 29, 2021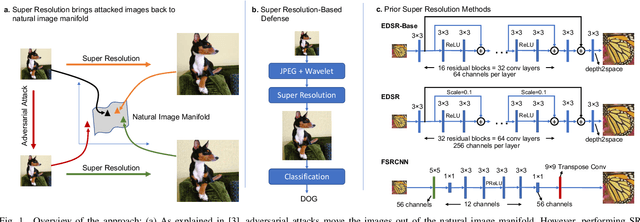
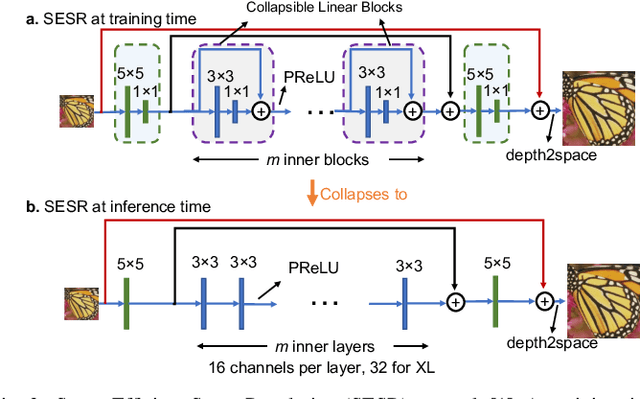
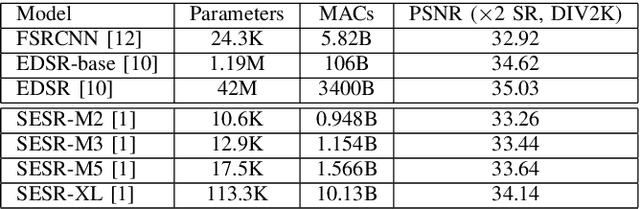
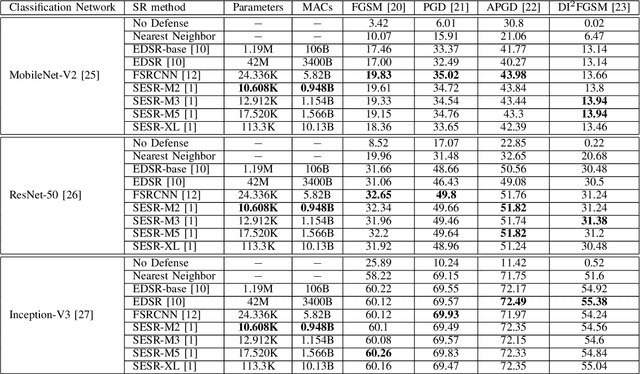
Autonomous systems are highly vulnerable to a variety of adversarial attacks on Deep Neural Networks (DNNs). Training-free model-agnostic defenses have recently gained popularity due to their speed, ease of deployment, and ability to work across many DNNs. To this end, a new technique has emerged for mitigating attacks on image classification DNNs, namely, preprocessing adversarial images using super resolution -- upscaling low-quality inputs into high-resolution images. This defense requires running both image classifiers and super resolution models on constrained autonomous systems. However, super resolution incurs a heavy computational cost. Therefore, in this paper, we investigate the following question: Does the robustness of image classifiers suffer if we use tiny super resolution models? To answer this, we first review a recent work called Super-Efficient Super Resolution (SESR) that achieves similar or better image quality than prior art while requiring 2x to 330x fewer Multiply-Accumulate (MAC) operations. We demonstrate that despite being orders of magnitude smaller than existing models, SESR achieves the same level of robustness as significantly larger networks. Finally, we estimate end-to-end performance of super resolution-based defenses on a commercial Arm Ethos-U55 micro-NPU. Our findings show that SESR achieves nearly 3x higher FPS than a baseline while achieving similar robustness.
Multiclass classification using quantum convolutional neural networks with hybrid quantum-classical learning
Mar 29, 2022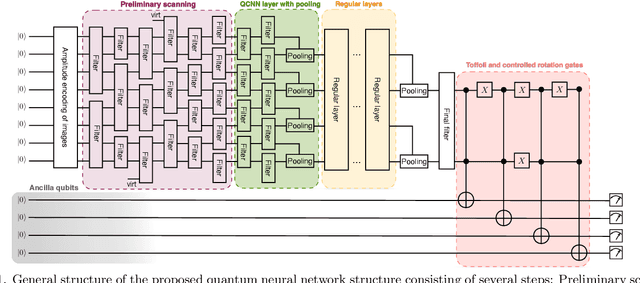
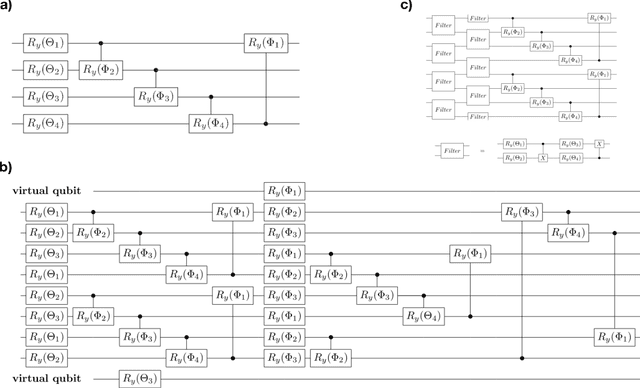
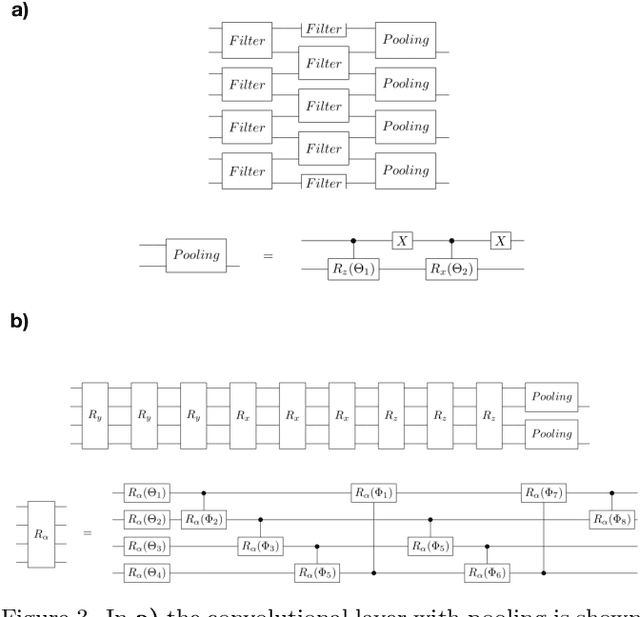
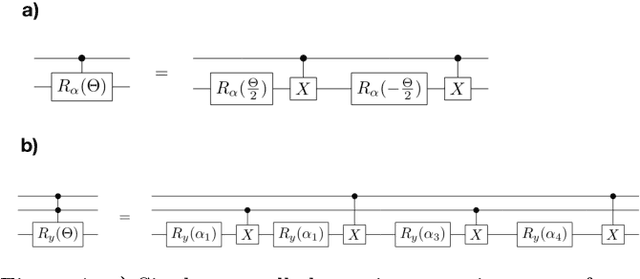
Multiclass classification is of great interest for various machine learning applications, for example, it is a common task in computer vision, where one needs to categorize an image into three or more classes. Here we propose a quantum machine learning approach based on quantum convolutional neural networks for solving this problem. The corresponding learning procedure is implemented via TensorFlowQuantum as a hybrid quantum-classical (variational) model, where quantum output results are fed to softmax cost function with subsequent minimization of it via optimization of parameters of quantum circuit. Our conceptional improvements include a new model for quantum perceptron and optimized structure of the quantum circuit. We use the proposed approach to demonstrate the 4-class classification for the case of the MNIST dataset using eight qubits for data encoding and four acnilla qubits. Our results demonstrate comparable accuracy of our solution with classical convolutional neural networks with comparable numbers of trainable parameters. We expect that our finding provide a new step towards the use of quantum machine learning for solving practically relevant problems in the NISQ era and beyond.
Conditional Image Generation with Score-Based Diffusion Models
Nov 26, 2021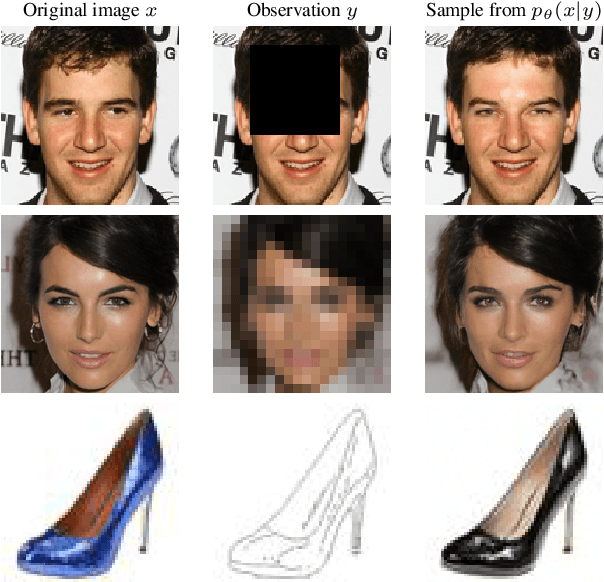
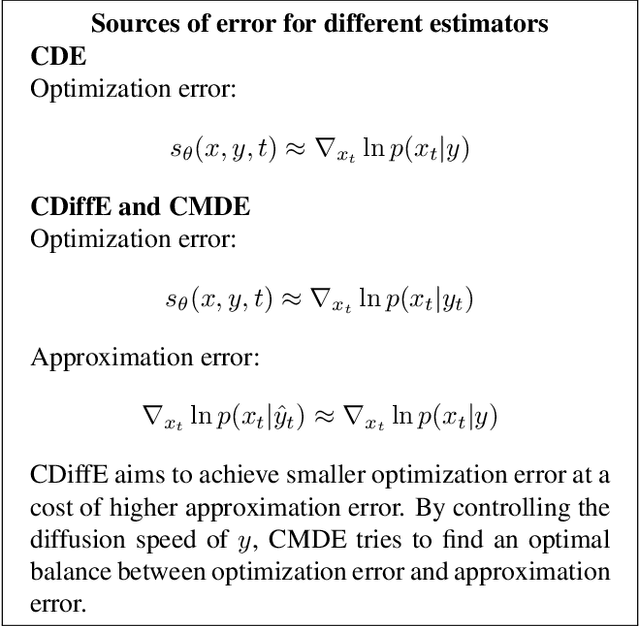
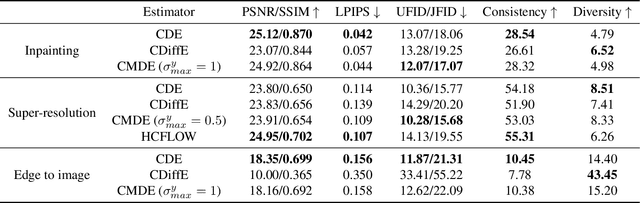

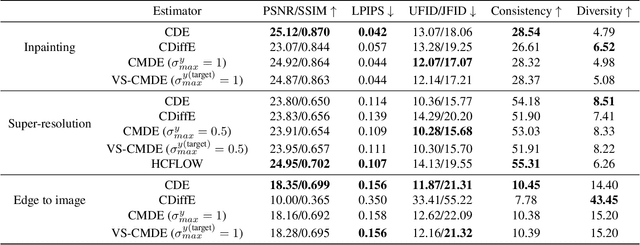
Score-based diffusion models have emerged as one of the most promising frameworks for deep generative modelling. In this work we conduct a systematic comparison and theoretical analysis of different approaches to learning conditional probability distributions with score-based diffusion models. In particular, we prove results which provide a theoretical justification for one of the most successful estimators of the conditional score. Moreover, we introduce a multi-speed diffusion framework, which leads to a new estimator for the conditional score, performing on par with previous state-of-the-art approaches. Our theoretical and experimental findings are accompanied by an open source library MSDiff which allows for application and further research of multi-speed diffusion models.
Quality-Aware Memory Network for Interactive Volumetric Image Segmentation
Jun 20, 2021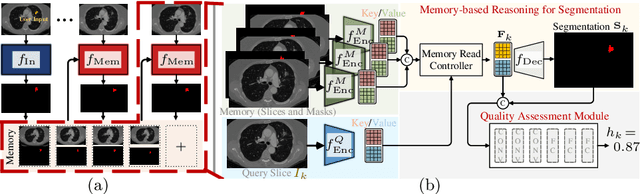
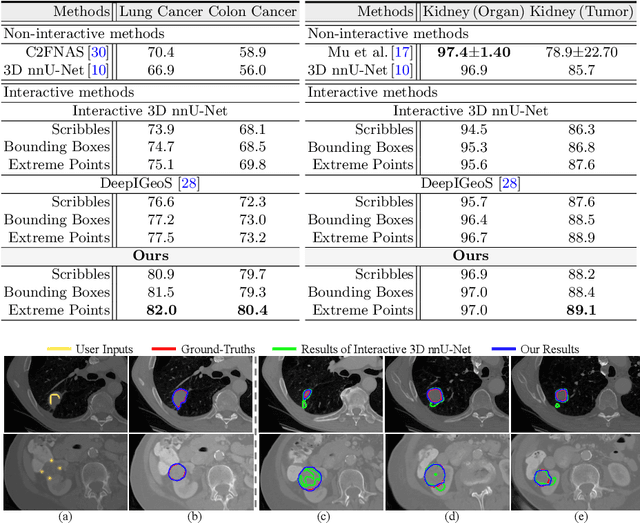
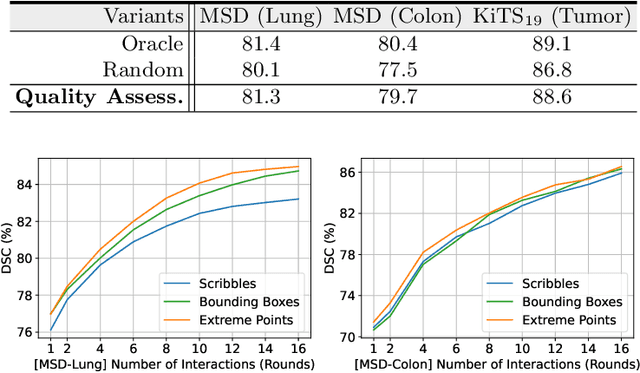
Despite recent progress of automatic medical image segmentation techniques, fully automatic results usually fail to meet the clinical use and typically require further refinement. In this work, we propose a quality-aware memory network for interactive segmentation of 3D medical images. Provided by user guidance on an arbitrary slice, an interaction network is firstly employed to obtain an initial 2D segmentation. The quality-aware memory network subsequently propagates the initial segmentation estimation bidirectionally over the entire volume. Subsequent refinement based on additional user guidance on other slices can be incorporated in the same manner. To further facilitate interactive segmentation, a quality assessment module is introduced to suggest the next slice to segment based on the current segmentation quality of each slice. The proposed network has two appealing characteristics: 1) The memory-augmented network offers the ability to quickly encode past segmentation information, which will be retrieved for the segmentation of other slices; 2) The quality assessment module enables the model to directly estimate the qualities of segmentation predictions, which allows an active learning paradigm where users preferentially label the lowest-quality slice for multi-round refinement. The proposed network leads to a robust interactive segmentation engine, which can generalize well to various types of user annotations (e.g., scribbles, boxes). Experimental results on various medical datasets demonstrate the superiority of our approach in comparison with existing techniques.
A study on the effects of compression on hyperspectral image classification
Apr 01, 2021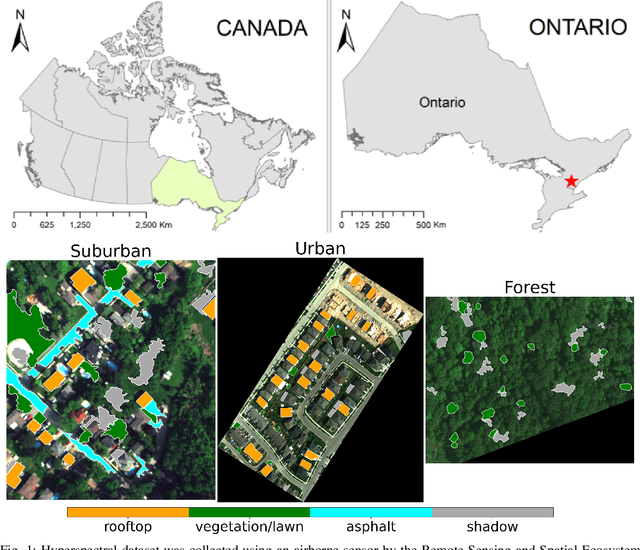
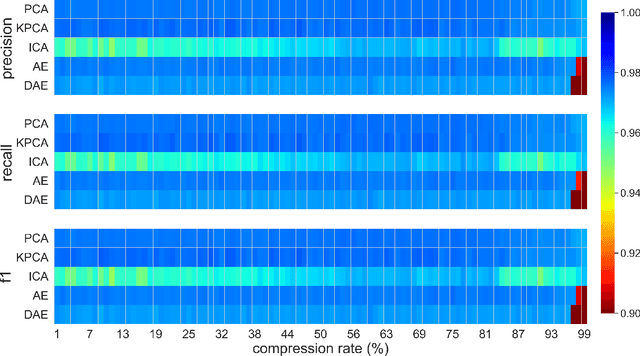
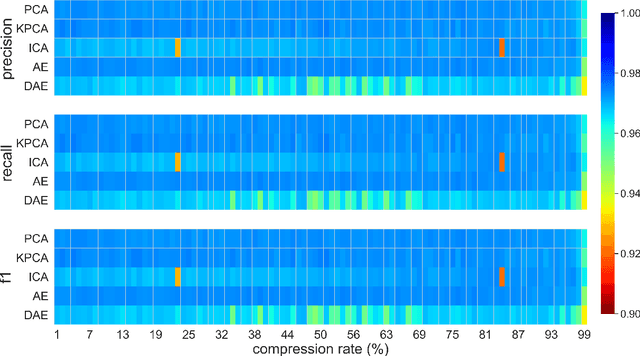
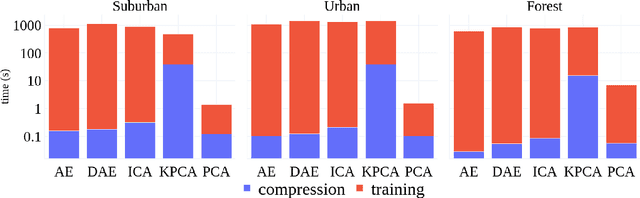
This paper presents a systematic study the effects of compression on hyperspectral pixel classification task. We use five dimensionality reduction methods -- PCA, KPCA, ICA, AE, and DAE -- to compress 301-dimensional hyperspectral pixels. Compressed pixels are subsequently used to perform pixel-based classifications. Pixel classification accuracies together with compression method, compression rates, and reconstruction errors provide a new lens to study the suitability of a compression method for the task of pixel-based classification. We use three high-resolution hyperspectral image datasets, representing three common landscape units (i.e. urban, transitional suburban, and forests) collected by the Remote Sensing and Spatial Ecosystem Modeling laboratory of the University of Toronto. We found that PCA, KPCA, and ICA post greater signal reconstruction capability; however, when compression rate is more than 90\% those methods showed lower classification scores. AE and DAE methods post better classification accuracy at 95\% compression rate, however decreasing again at 97\%, suggesting a sweet-spot at the 95\% mark. Our results demonstrate that the choice of a compression method with the compression rate are important considerations when designing a hyperspectral image classification pipeline.
Sensor fusion in ptychography
Mar 18, 2022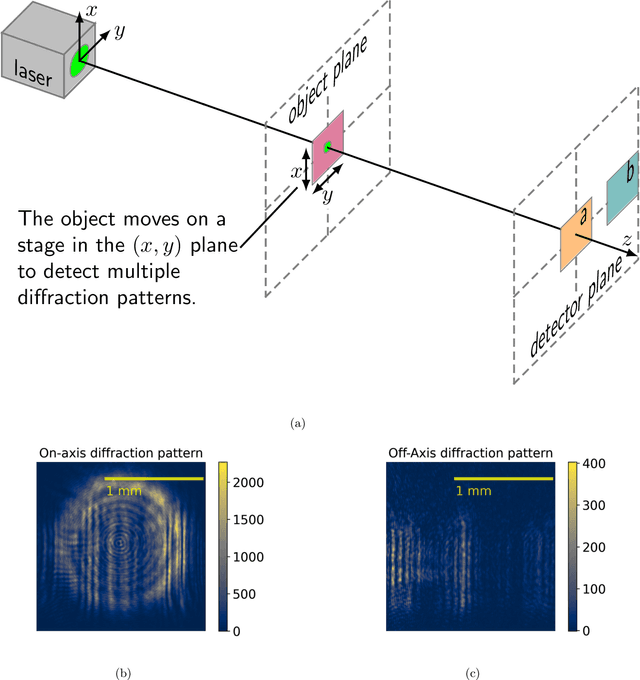
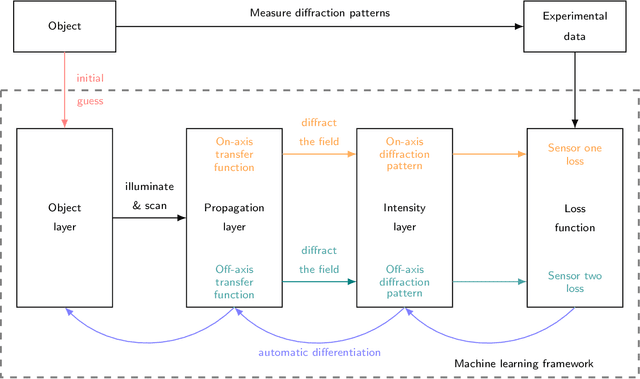
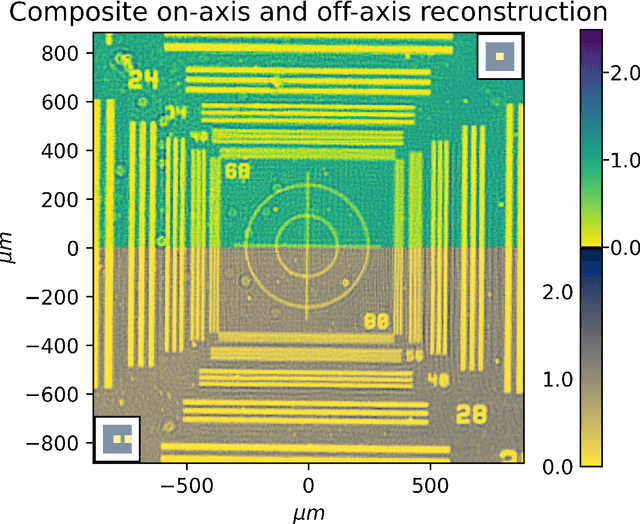
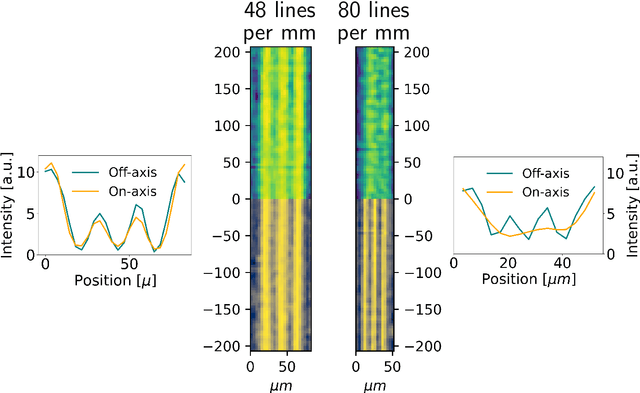
Ptychography is a lensless, computational imaging method that utilises diffraction patterns to determine the amplitude and phase of an object. In transmission ptychography, the diffraction patterns are recorded by a detector positioned along the optical axis downstream of the object. The light scattered at the highest diffraction angle carries information about the finest structures of the object. We present a setup to simultaneously capture a signal near the optical axis and a signal scattered at high diffraction angles. Moreover, we present an algorithm based on a shifted angular spectrum method and automatic differentiation that utilises this recorded signal. By jointly reconstructing the object from the resulting low and high diffraction angle images, the resolution of the reconstructed image is improved remarkably. The effective numerical aperture of the compound sensor is determined by the maximum diffraction angle captured by the off axis sensor.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge