"Image": models, code, and papers
VinDr-PCXR: An open, large-scale chest radiograph dataset for interpretation of common thoracic diseases in children
Mar 20, 2022
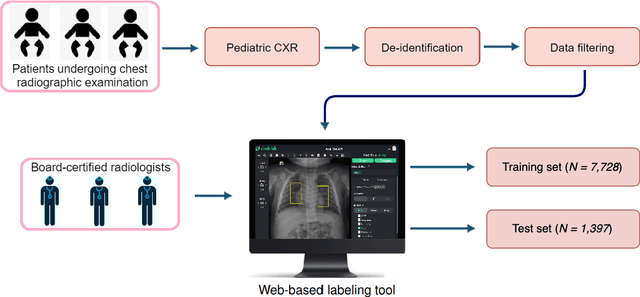
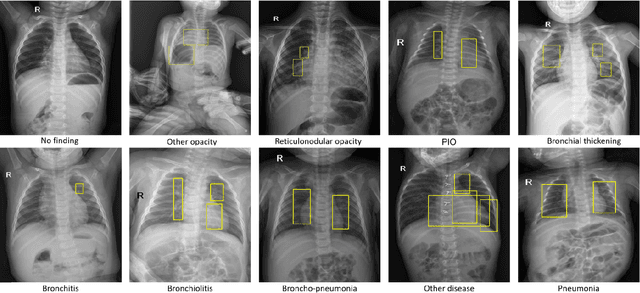
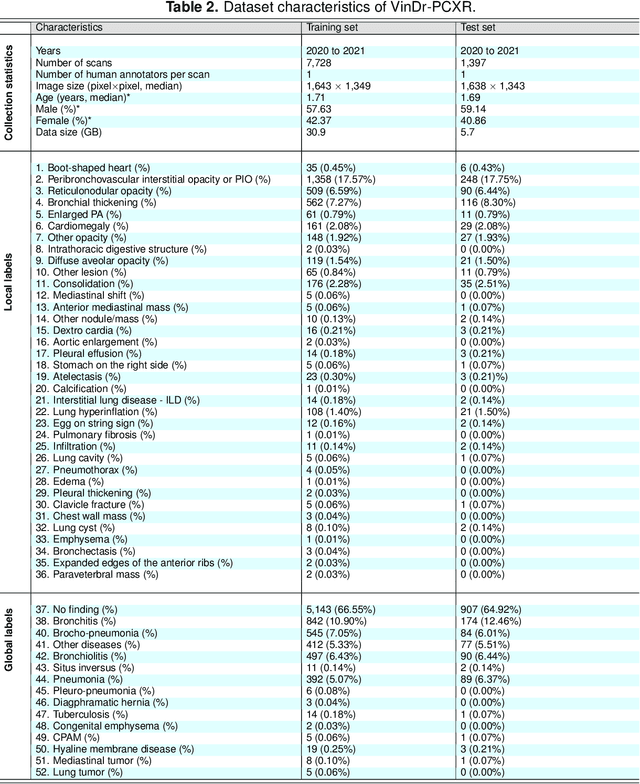
Computer-aided diagnosis systems in adult chest radiography (CXR) have recently achieved great success thanks to the availability of large-scale, annotated datasets and the advent of high-performance supervised learning algorithms. However, the development of diagnostic models for detecting and diagnosing pediatric diseases in CXR scans is undertaken due to the lack of high-quality physician-annotated datasets. To overcome this challenge, we introduce and release VinDr-PCXR, a new pediatric CXR dataset of 9,125 studies retrospectively collected from a major pediatric hospital in Vietnam between 2020 and 2021. Each scan was manually annotated by a pediatric radiologist who has more than ten years of experience. The dataset was labeled for the presence of 36 critical findings and 15 diseases. In particular, each abnormal finding was identified via a rectangle bounding box on the image. To the best of our knowledge, this is the first and largest pediatric CXR dataset containing lesion-level annotations and image-level labels for the detection of multiple findings and diseases. For algorithm development, the dataset was divided into a training set of 7,728 and a test set of 1,397. To encourage new advances in pediatric CXR interpretation using data-driven approaches, we provide a detailed description of the VinDr-PCXR data sample and make the dataset publicly available on https://physionet.org/.
Fast and Automatic Object Registration for Human-Robot Collaboration in Industrial Manufacturing
Apr 01, 2022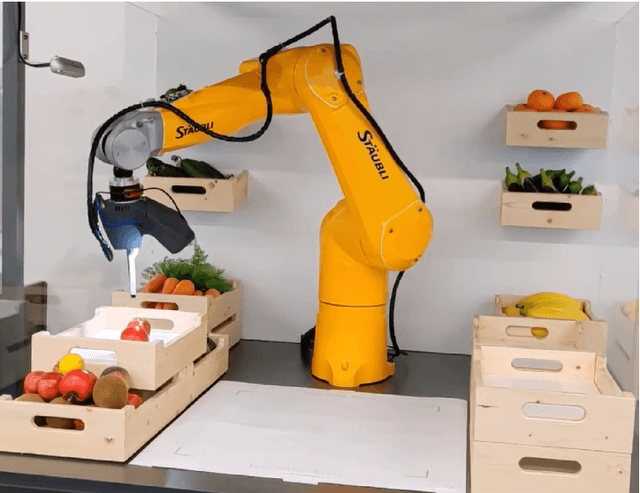
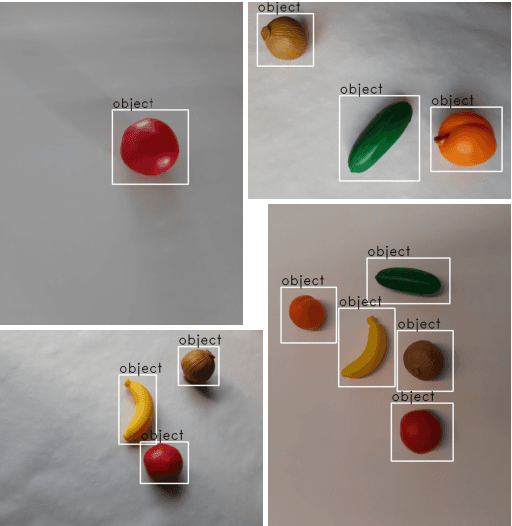
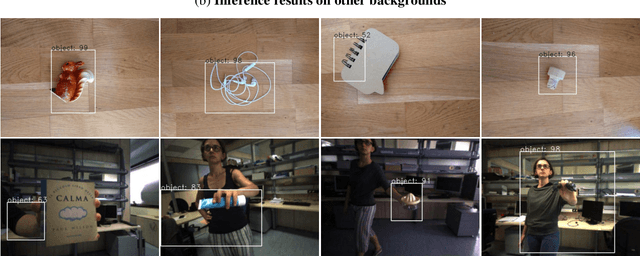
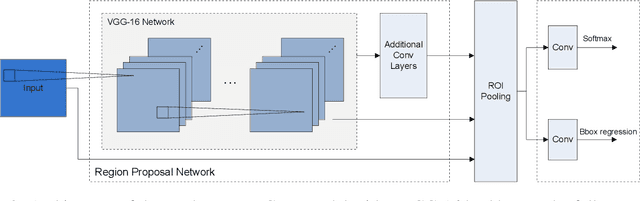


We present an end-to-end framework for fast retraining of object detection models in human-robot-collaboration. Our Faster R-CNN based setup covers the whole workflow of automatic image generation and labeling, model retraining on-site as well as inference on a FPGA edge device. The intervention of a human operator reduces to providing the new object together with its label and starting the training process. Moreover, we present a new loss, the intraspread-objectosphere loss, to tackle the problem of open world recognition. Though it fails to completely solve the problem, it significantly reduces the number of false positive detections of unknown objects.
Generative Convolution Layer for Image Generation
Nov 30, 2021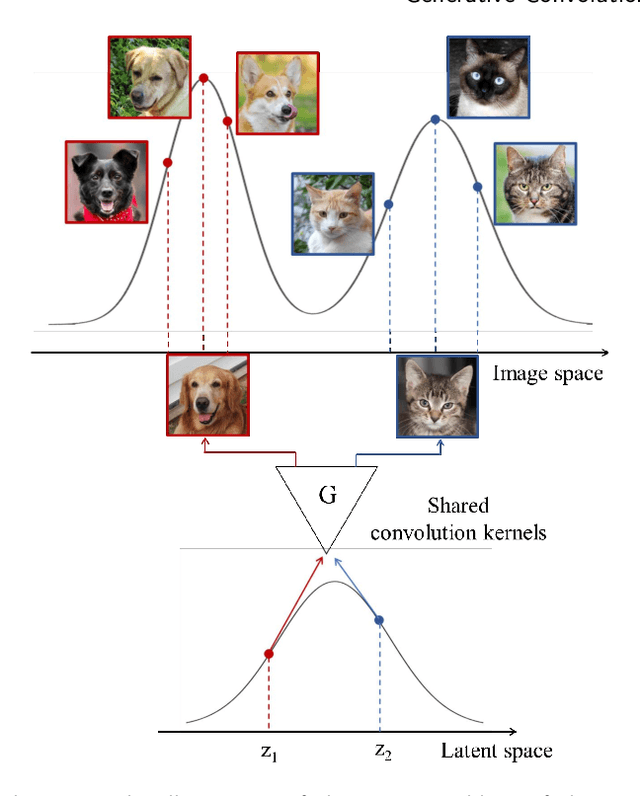
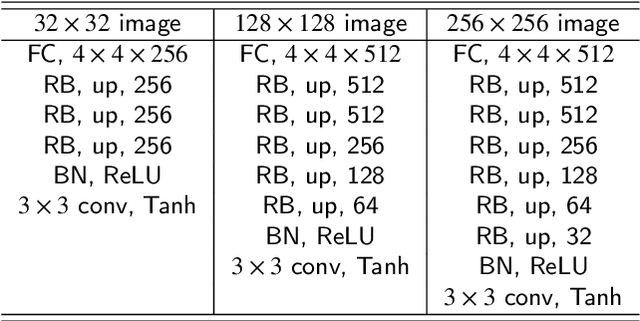
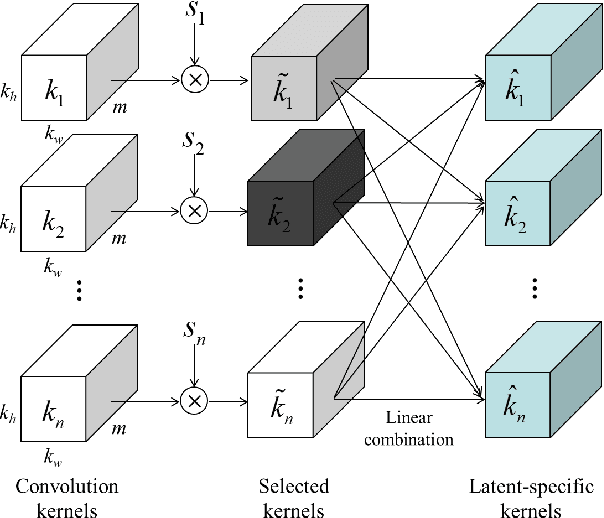
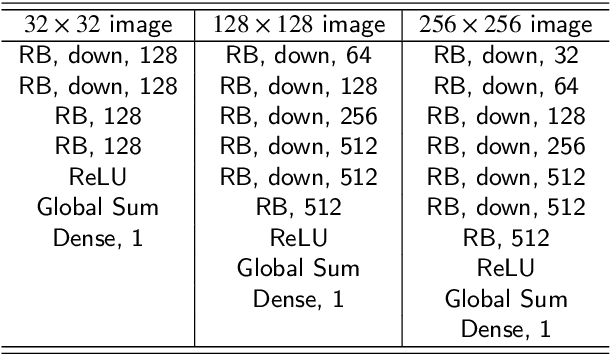
This paper introduces a novel convolution method, called generative convolution (GConv), which is simple yet effective for improving the generative adversarial network (GAN) performance. Unlike the standard convolution, GConv first selects useful kernels compatible with the given latent vector, and then linearly combines the selected kernels to make latent-specific kernels. Using the latent-specific kernels, the proposed method produces the latent-specific features which encourage the generator to produce high-quality images. This approach is simple but surprisingly effective. First, the GAN performance is significantly improved with a little additional hardware cost. Second, GConv can be employed to the existing state-of-the-art generators without modifying the network architecture. To reveal the superiority of GConv, this paper provides extensive experiments using various standard datasets including CIFAR-10, CIFAR-100, LSUN-Church, CelebA, and tiny-ImageNet. Quantitative evaluations prove that GConv significantly boosts the performances of the unconditional and conditional GANs in terms of Inception score (IS) and Frechet inception distance (FID). For example, the proposed method improves both FID and IS scores on the tiny-ImageNet dataset from 35.13 to 29.76 and 20.23 to 22.64, respectively.
Assessing hierarchies by their consistent segmentations
Apr 11, 2022
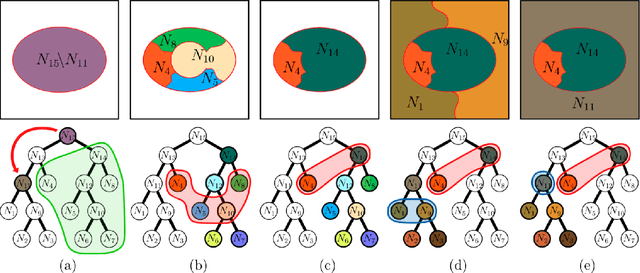
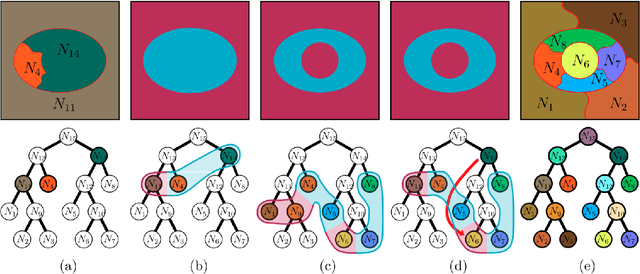
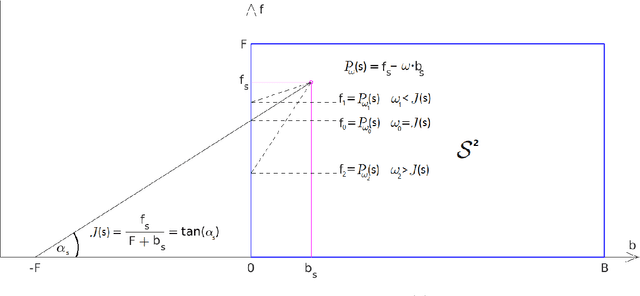
Recent segmentation approaches start by creating a hierarchy of nested image partitions, and then specify a segmentation from it, usually, by choosing one horizontal cut. Our first contribution is to describe several different ways, some of them new, for specifying segmentations using the hierarchy regions. Then we consider the best hierarchy-induced segmentation, in which the segments are specified by a limited number, k, of hierarchy nodes/regions. The number of hierarchy-induced segmentations grows exponentially with the hierarchy size, implying that exhaustive search is unfeasible. We focus on a common quality measure, the Jaccard index (known also as IoU). Optimizing the Jaccard index is highly nontrivial. Yet, we propose an efficient optimization * This work was done when the first author was with the Math dept. Technion, Israel.
Metal Artifact Reduction with Intra-Oral Scan Data for 3D Low Dose Maxillofacial CBCT Modeling
Feb 08, 2022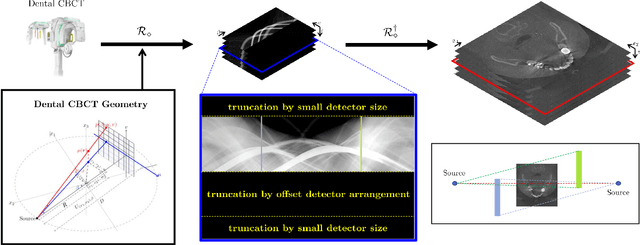
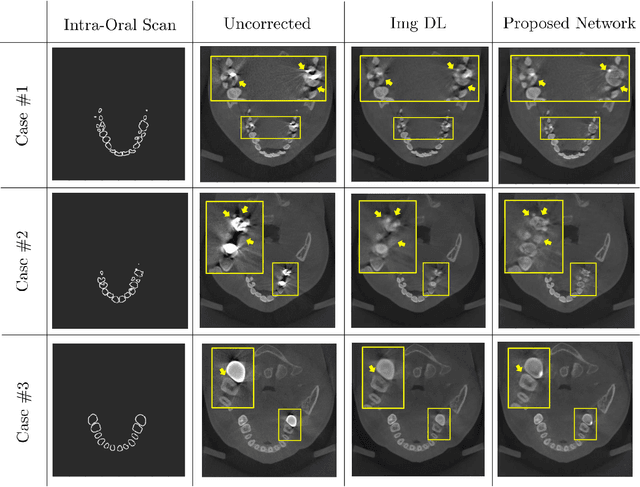
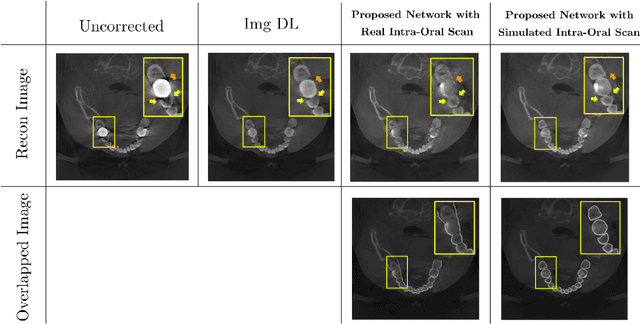
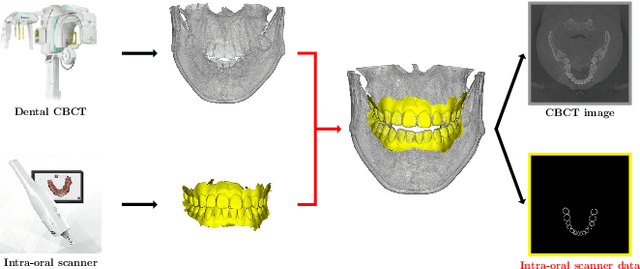
Low-dose dental cone beam computed tomography (CBCT) has been increasingly used for maxillofacial modeling. However, the presence of metallic inserts, such as implants, crowns, and dental filling, causes severe streaking and shading artifacts in a CBCT image and loss of the morphological structures of the teeth, which consequently prevents accurate segmentation of bones. A two-stage metal artifact reduction method is proposed for accurate 3D low-dose maxillofacial CBCT modeling, where a key idea is to utilize explicit tooth shape prior information from intra-oral scan data whose acquisition does not require any extra radiation exposure. In the first stage, an image-to-image deep learning network is employed to mitigate metal-related artifacts. To improve the learning ability, the proposed network is designed to take advantage of the intra-oral scan data as side-inputs and perform multi-task learning of auxiliary tooth segmentation. In the second stage, a 3D maxillofacial model is constructed by segmenting the bones from the dental CBCT image corrected in the first stage. For accurate bone segmentation, weighted thresholding is applied, wherein the weighting region is determined depending on the geometry of the intra-oral scan data. Because acquiring a paired training dataset of metal-artifact-free and metal artifact-affected dental CBCT images is challenging in clinical practice, an automatic method of generating a realistic dataset according to the CBCT physics model is introduced. Numerical simulations and clinical experiments show the feasibility of the proposed method, which takes advantage of tooth surface information from intra-oral scan data in 3D low dose maxillofacial CBCT modeling.
CryoAI: Amortized Inference of Poses for Ab Initio Reconstruction of 3D Molecular Volumes from Real Cryo-EM Images
Mar 16, 2022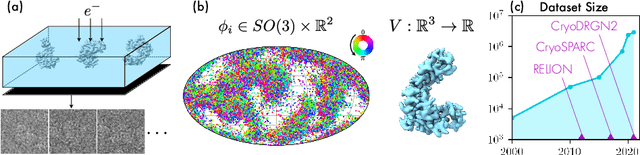
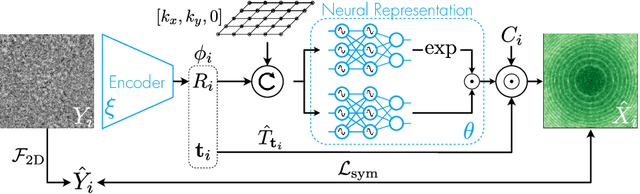
Cryo-electron microscopy (cryo-EM) has become a tool of fundamental importance in structural biology, helping us understand the basic building blocks of life. The algorithmic challenge of cryo-EM is to jointly estimate the unknown 3D poses and the 3D electron scattering potential of a biomolecule from millions of extremely noisy 2D images. Existing reconstruction algorithms, however, cannot easily keep pace with the rapidly growing size of cryo-EM datasets due to their high computational and memory cost. We introduce cryoAI, an ab initio reconstruction algorithm for homogeneous conformations that uses direct gradient-based optimization of particle poses and the electron scattering potential from single-particle cryo-EM data. CryoAI combines a learned encoder that predicts the poses of each particle image with a physics-based decoder to aggregate each particle image into an implicit representation of the scattering potential volume. This volume is stored in the Fourier domain for computational efficiency and leverages a modern coordinate network architecture for memory efficiency. Combined with a symmetrized loss function, this framework achieves results of a quality on par with state-of-the-art cryo-EM solvers for both simulated and experimental data, one order of magnitude faster for large datasets and with significantly lower memory requirements than existing methods.
CBANet: Towards Complexity and Bitrate Adaptive Deep Image Compression using a Single Network
May 26, 2021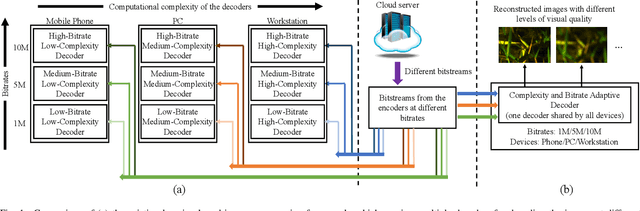
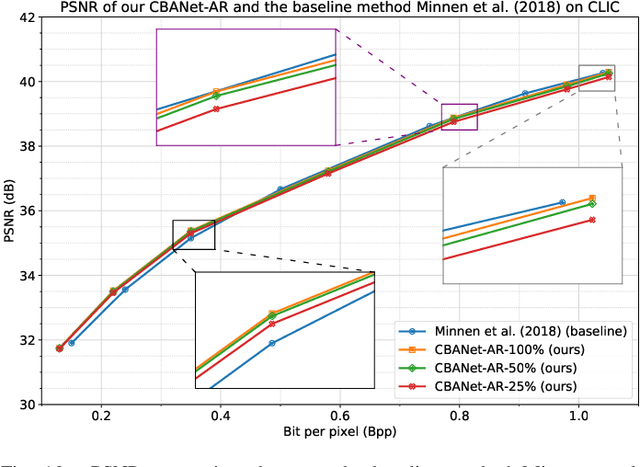
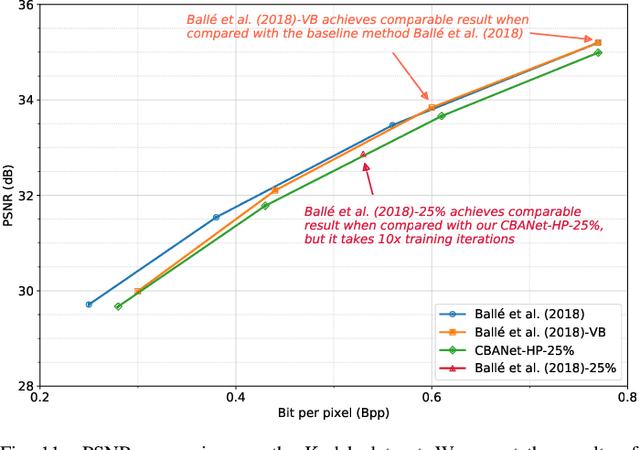
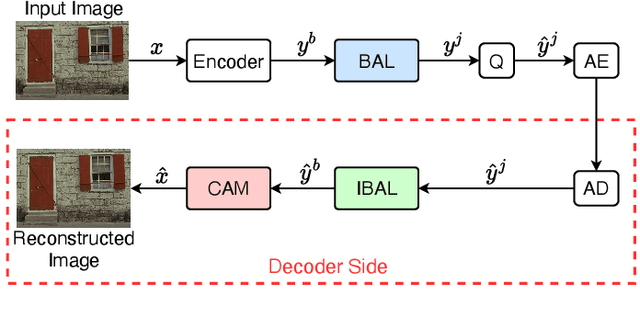
In this paper, we propose a new deep image compression framework called Complexity and Bitrate Adaptive Network (CBANet), which aims to learn one single network to support variable bitrate coding under different computational complexity constraints. In contrast to the existing state-of-the-art learning based image compression frameworks that only consider the rate-distortion trade-off without introducing any constraint related to the computational complexity, our CBANet considers the trade-off between the rate and distortion under dynamic computational complexity constraints. Specifically, to decode the images with one single decoder under various computational complexity constraints, we propose a new multi-branch complexity adaptive module, in which each branch only takes a small portion of the computational budget of the decoder. The reconstructed images with different visual qualities can be readily generated by using different numbers of branches. Furthermore, to achieve variable bitrate decoding with one single decoder, we propose a bitrate adaptive module to project the representation from a base bitrate to the expected representation at a target bitrate for transmission. Then it will project the transmitted representation at the target bitrate back to that at the base bitrate for the decoding process. The proposed bit adaptive module can significantly reduce the storage requirement for deployment platforms. As a result, our CBANet enables one single codec to support multiple bitrate decoding under various computational complexity constraints. Comprehensive experiments on two benchmark datasets demonstrate the effectiveness of our CBANet for deep image compression.
Cross-Domain Object Detection with Mean-Teacher Transformer
May 03, 2022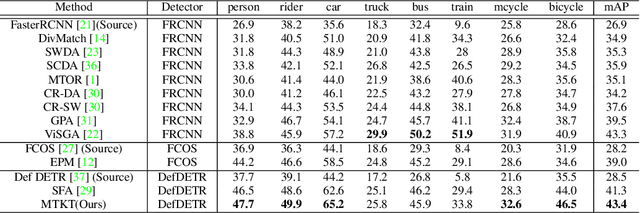
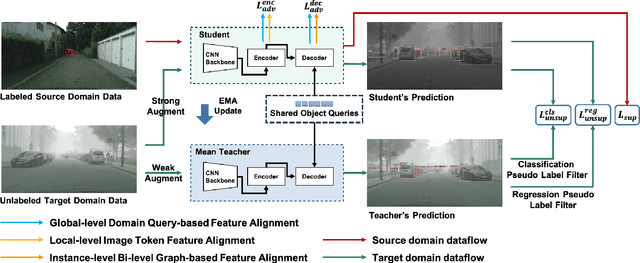
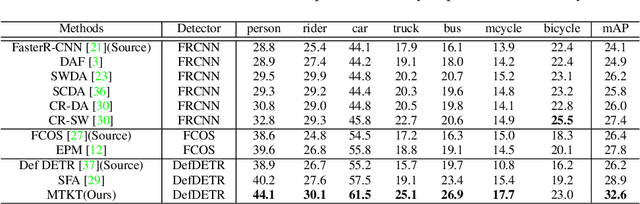
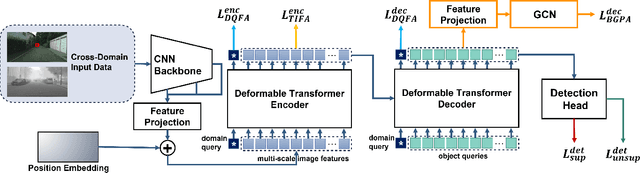
Recently, DEtection TRansformer (DETR), an end-to-end object detection pipeline, has achieved promising performance. However, it requires large-scale labeled data and suffers from domain shift, especially when no labeled data is available in the target domain. To solve this problem, we propose an end-to-end cross-domain detection transformer based on the mean teacher knowledge transfer (MTKT), which transfers knowledge between domains via pseudo labels. To improve the quality of pseudo labels in the target domain, which is a crucial factor for better domain adaptation, we design three levels of source-target feature alignment strategies based on the architecture of the Transformer, including domain query-based feature alignment (DQFA), bi-level-graph-based prototype alignment (BGPA), and token-wise image feature alignment (TIFA). These three levels of feature alignment match the global, local, and instance features between source and target, respectively. With these strategies, more accurate pseudo labels can be obtained, and knowledge can be better transferred from source to target, thus improving the cross-domain capability of the detection transformer. Extensive experiments demonstrate that our proposed method achieves state-of-the-art performance on three domain adaptation scenarios, especially the result of Sim10k to Cityscapes scenario is remarkably improved from 52.6 mAP to 57.9 mAP. Code will be released.
Few Shot Generative Model Adaption via Relaxed Spatial Structural Alignment
Mar 06, 2022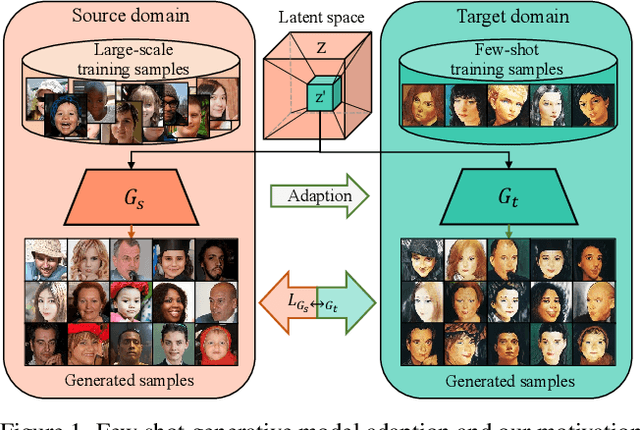
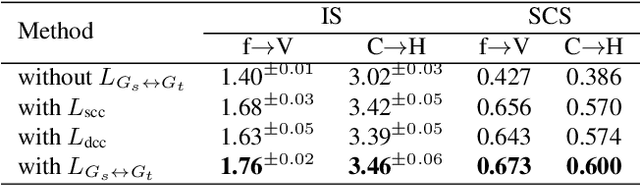
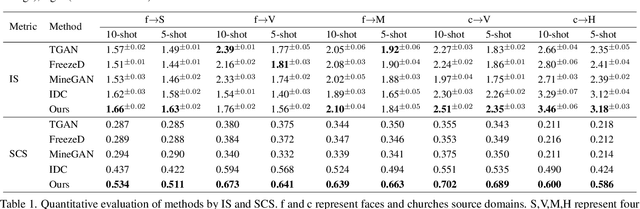
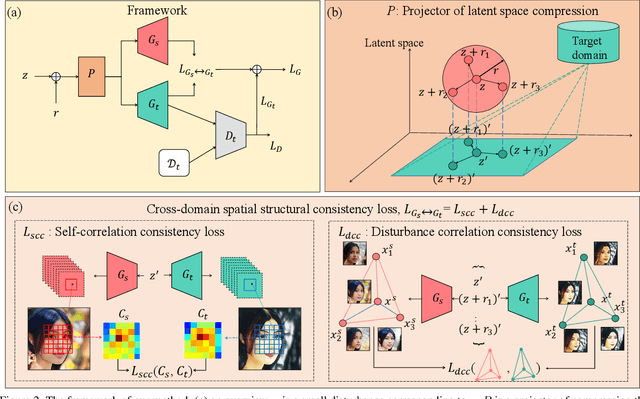

Training a generative adversarial network (GAN) with limited data has been a challenging task. A feasible solution is to start with a GAN well-trained on a large scale source domain and adapt it to the target domain with a few samples, termed as few shot generative model adaption. However, existing methods are prone to model overfitting and collapse in extremely few shot setting (less than 10). To solve this problem, we propose a relaxed spatial structural alignment method to calibrate the target generative models during the adaption. We design a cross-domain spatial structural consistency loss comprising the self-correlation and disturbance correlation consistency loss. It helps align the spatial structural information between the synthesis image pairs of the source and target domains. To relax the cross-domain alignment, we compress the original latent space of generative models to a subspace. Image pairs generated from the subspace are pulled closer. Qualitative and quantitative experiments show that our method consistently surpasses the state-of-the-art methods in few shot setting.
MisMatch: Learning to Change Predictive Confidences with Attention for Consistency-Based, Semi-Supervised Medical Image Segmentation
Oct 23, 2021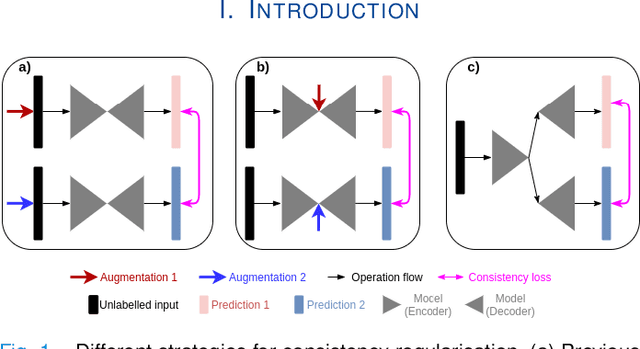
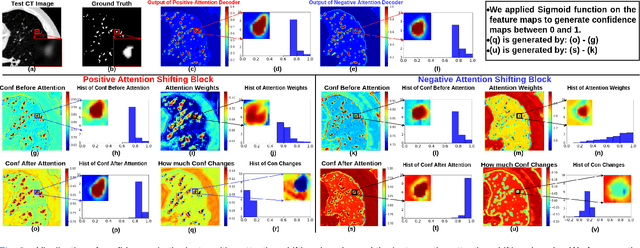

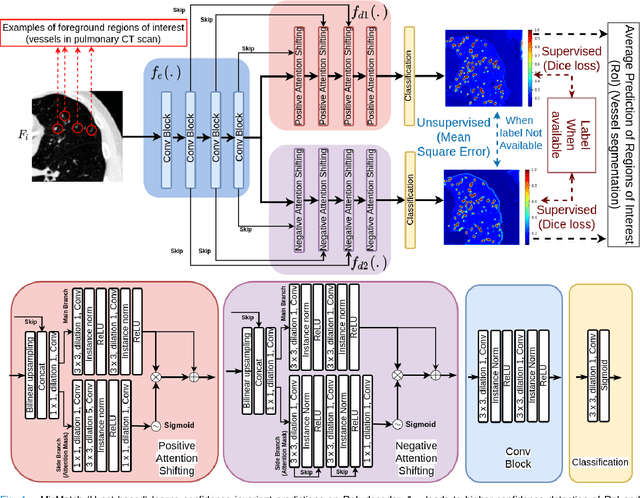
The lack of labels is one of the fundamental constraints in deep learning based methods for image classification and segmentation, especially in applications such as medical imaging. Semi-supervised learning (SSL) is a promising method to address the challenge of labels carcity. The state-of-the-art SSL methods utilise consistency regularisation to learn unlabelled predictions which are invariant to perturbations on the prediction confidence. However, such SSL approaches rely on hand-crafted augmentation techniques which could be sub-optimal. In this paper, we propose MisMatch, a novel consistency based semi-supervised segmentation method. MisMatch automatically learns to produce paired predictions with increasedand decreased confidences. MisMatch consists of an encoder and two decoders. One decoder learns positive attention for regions of interest (RoI) on unlabelled data thereby generating higher confidence predictions of RoI. The other decoder learns negative attention for RoI on the same unlabelled data thereby generating lower confidence predictions. We then apply a consistency regularisation between the paired predictions of the decoders. For evaluation, we first perform extensive cross-validation on a CT-based pulmonary vessel segmentation task and show that MisMatch statistically outperforms state-of-the-art semi-supervised methods when only 6.25% of the total labels are used. Furthermore MisMatch performance using 6.25% ofthe total labels is comparable to state-of-the-art methodsthat utilise all available labels. In a second experiment, MisMatch outperforms state-of-the-art methods on an MRI-based brain tumour segmentation task.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge