"Image": models, code, and papers
Entropy Regularized Iterative Weighted Shrinkage-Thresholding Algorithm (ERIWSTA): An Application to CT Image Restoration
Dec 22, 2021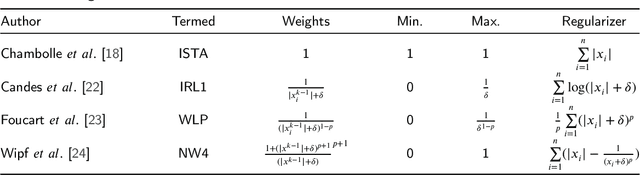
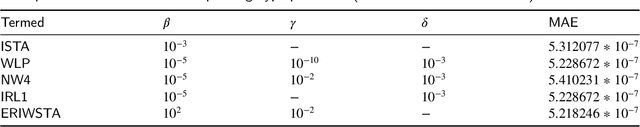
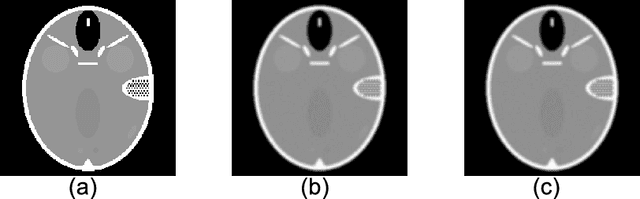
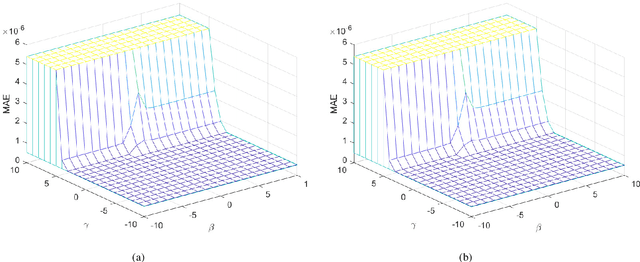

The iterative weighted shrinkage-thresholding algorithm (IWSTA) has shown superiority to the classic unweighted iterative shrinkage-thresholding algorithm (ISTA) for solving linear inverse problems, which address the attributes differently. This paper proposes a new entropy regularized IWSTA (ERIWSTA) that adds an entropy regularizer to the cost function to measure the uncertainty of the weights to stimulate attributes to participate in problem solving. Then, the weights are solved with a Lagrange multiplier method to obtain a simple iterative update. The weights can be explained as the probability of the contribution of an attribute to the problem solution. Experimental results on CT image restoration show that the proposed method has better performance in terms of convergence speed and restoration accuracy than the existing methods.
Development of Automatic Endotracheal Tube and Carina Detection on Portable Supine Chest Radiographs using Artificial Intelligence
Jun 07, 2022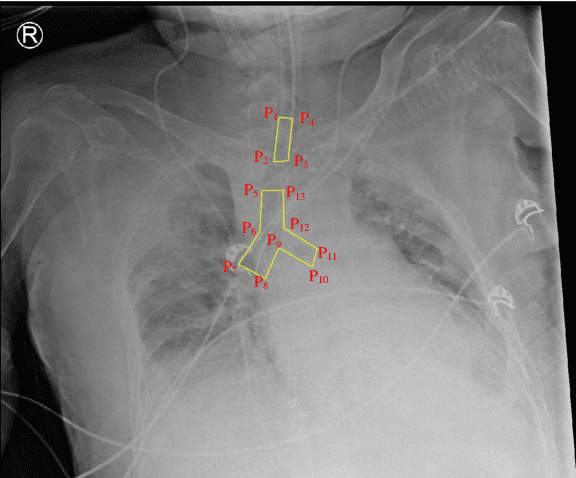
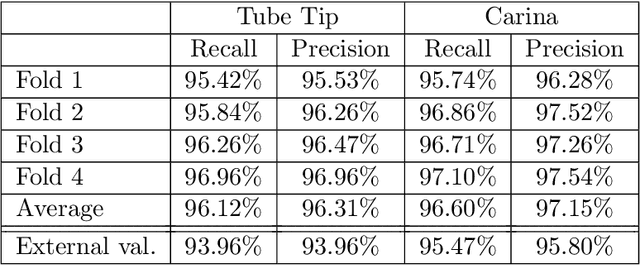
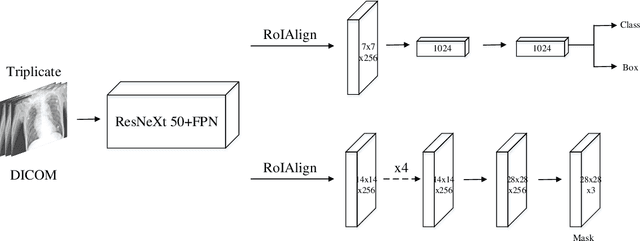
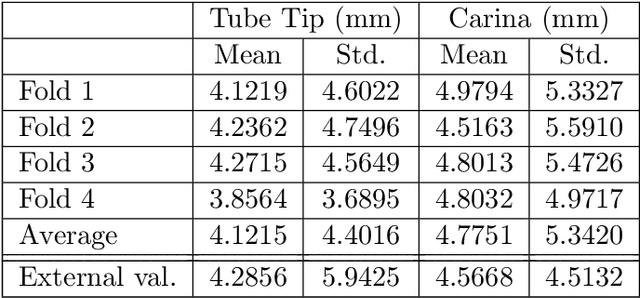
The image quality of portable supine chest radiographs is inherently poor due to low contrast and high noise. The endotracheal intubation detection requires the locations of the endotracheal tube (ETT) tip and carina. The goal is to find the distance between the ETT tip and the carina in chest radiography. To overcome such a problem, we propose a feature extraction method with Mask R-CNN. The Mask R-CNN predicts a tube and a tracheal bifurcation in an image. Then, the feature extraction method is used to find the feature point of the ETT tip and that of the carina. Therefore, the ETT-carina distance can be obtained. In our experiments, our results can exceed 96\% in terms of recall and precision. Moreover, the object error is less than $4.7751\pm 5.3420$ mm, and the ETT-carina distance errors are less than $5.5432\pm 6.3100$ mm. The external validation shows that the proposed method is a high-robustness system. According to the Pearson correlation coefficient, we have a strong correlation between the board-certified intensivists and our result in terms of ETT-carina distance.
3D Clothed Human Reconstruction in the Wild
Jul 20, 2022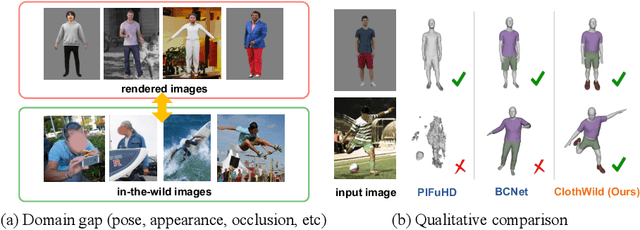
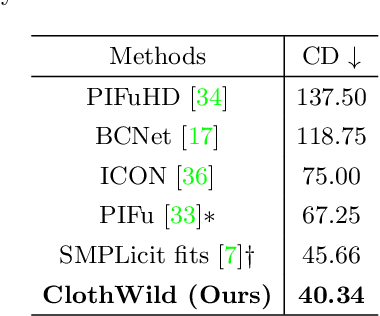
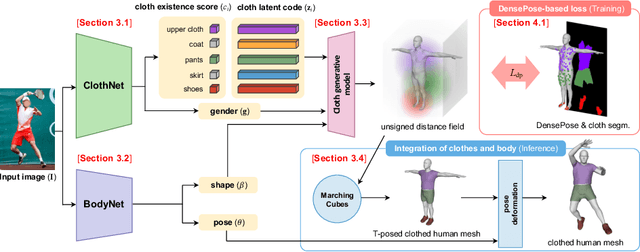
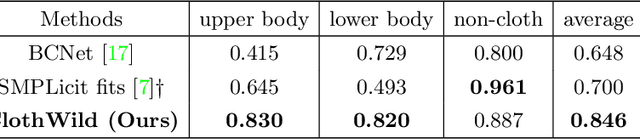
Although much progress has been made in 3D clothed human reconstruction, most of the existing methods fail to produce robust results from in-the-wild images, which contain diverse human poses and appearances. This is mainly due to the large domain gap between training datasets and in-the-wild datasets. The training datasets are usually synthetic ones, which contain rendered images from GT 3D scans. However, such datasets contain simple human poses and less natural image appearances compared to those of real in-the-wild datasets, which makes generalization of it to in-the-wild images extremely challenging. To resolve this issue, in this work, we propose ClothWild, a 3D clothed human reconstruction framework that firstly addresses the robustness on in-thewild images. First, for the robustness to the domain gap, we propose a weakly supervised pipeline that is trainable with 2D supervision targets of in-the-wild datasets. Second, we design a DensePose-based loss function to reduce ambiguities of the weak supervision. Extensive empirical tests on several public in-the-wild datasets demonstrate that our proposed ClothWild produces much more accurate and robust results than the state-of-the-art methods. The codes are available in here: https://github.com/hygenie1228/ClothWild_RELEASE.
3D-Aware Video Generation
Jun 29, 2022
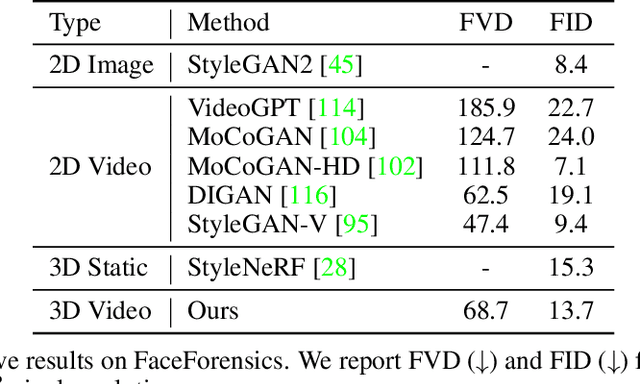
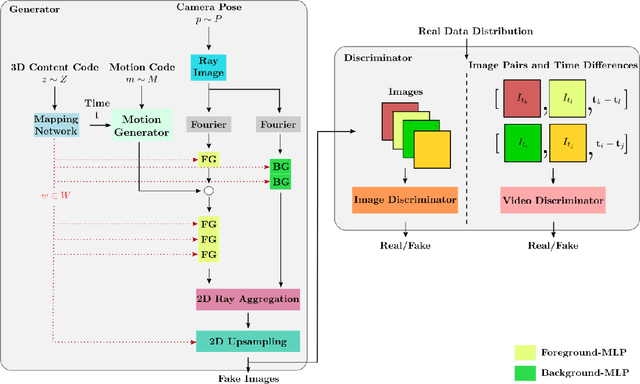

Generative models have emerged as an essential building block for many image synthesis and editing tasks. Recent advances in this field have also enabled high-quality 3D or video content to be generated that exhibits either multi-view or temporal consistency. With our work, we explore 4D generative adversarial networks (GANs) that learn unconditional generation of 3D-aware videos. By combining neural implicit representations with time-aware discriminator, we develop a GAN framework that synthesizes 3D video supervised only with monocular videos. We show that our method learns a rich embedding of decomposable 3D structures and motions that enables new visual effects of spatio-temporal renderings while producing imagery with quality comparable to that of existing 3D or video GANs.
A Spatial Guided Self-supervised Clustering Network for Medical Image Segmentation
Jul 11, 2021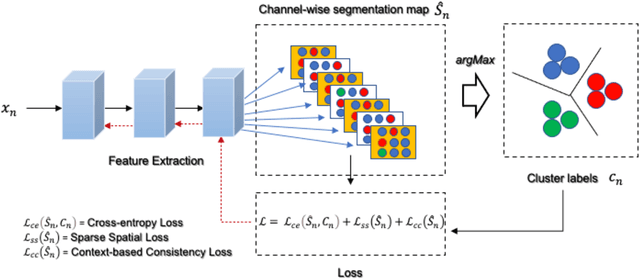
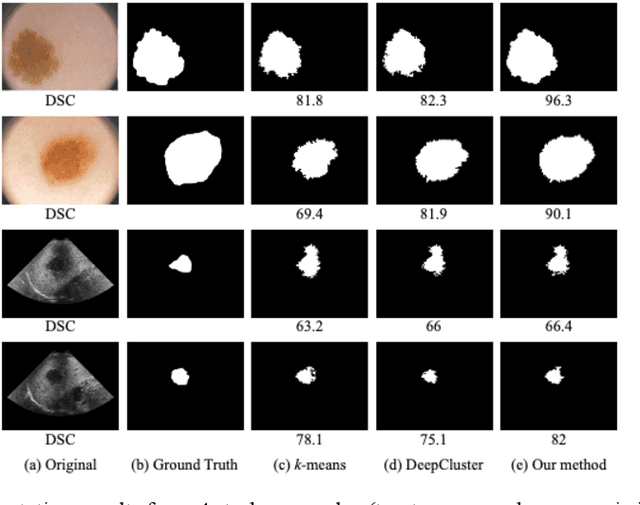
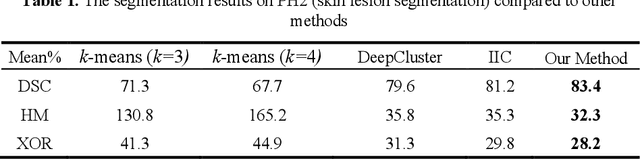
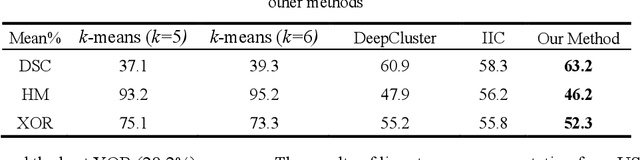

The segmentation of medical images is a fundamental step in automated clinical decision support systems. Existing medical image segmentation methods based on supervised deep learning, however, remain problematic because of their reliance on large amounts of labelled training data. Although medical imaging data repositories continue to expand, there has not been a commensurate increase in the amount of annotated data. Hence, we propose a new spatial guided self-supervised clustering network (SGSCN) for medical image segmentation, where we introduce multiple loss functions designed to aid in grouping image pixels that are spatially connected and have similar feature representations. It iteratively learns feature representations and clustering assignment of each pixel in an end-to-end fashion from a single image. We also propose a context-based consistency loss that better delineates the shape and boundaries of image regions. It enforces all the pixels belonging to a cluster to be spatially close to the cluster centre. We evaluated our method on 2 public medical image datasets and compared it to existing conventional and self-supervised clustering methods. Experimental results show that our method was most accurate for medical image segmentation.
Image Based Reconstruction of Liquids from 2D Surface Detections
Nov 22, 2021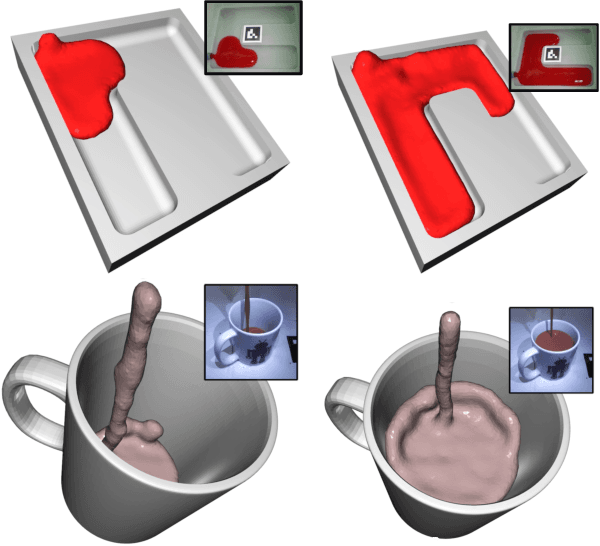
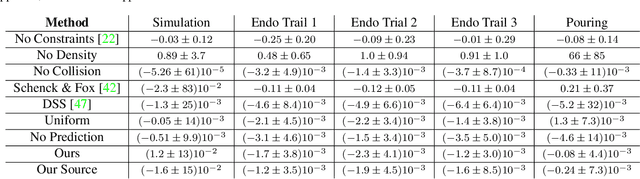
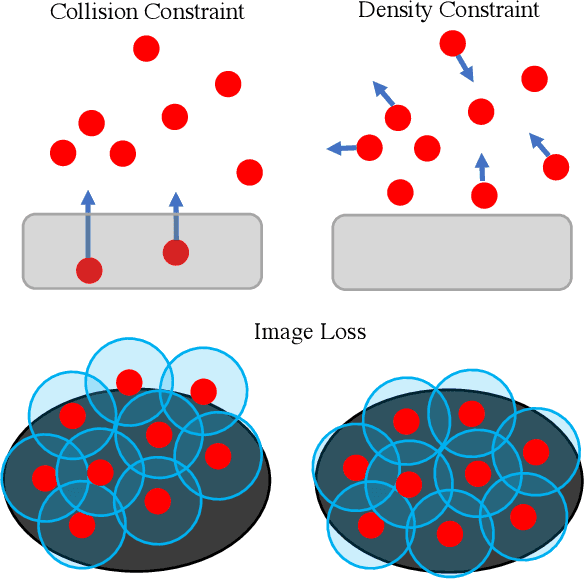
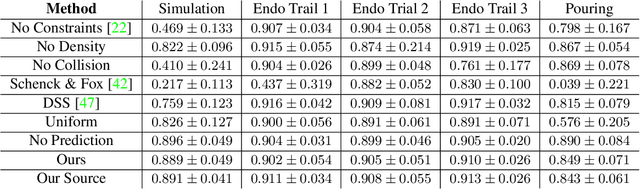
In this work, we present a solution to the challenging problem of reconstructing liquids from image data. The challenges in reconstructing liquids, which is not faced in previous reconstruction works on rigid and deforming surfaces, lies in the inability to use depth sensing and color features due the variable index of refraction, opacity, and environmental reflections. Therefore, we limit ourselves to only surface detections (i.e. binary mask) of liquids as observations and do not assume any prior knowledge on the liquids properties. A novel optimization problem is posed which reconstructs the liquid as particles by minimizing the error between a rendered surface from the particles and the surface detections while satisfying liquid constraints. Our solvers to this optimization problem are presented and no training data is required to apply them. We also propose a dynamic prediction to seed the reconstruction optimization from the previous time-step. We test our proposed methods in simulation and on two new liquid datasets which we open source so the broader research community can continue developing in this under explored area.
A Comparative Study of Confidence Calibration in Deep Learning: From Computer Vision to Medical Imaging
Jun 17, 2022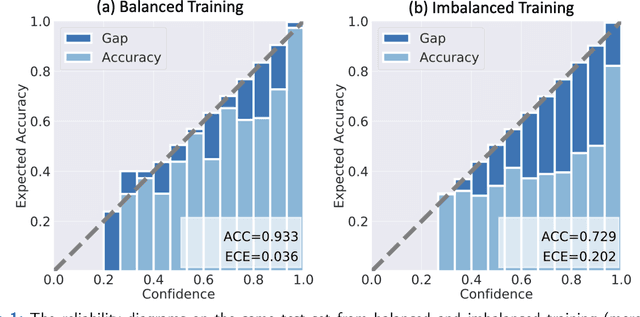
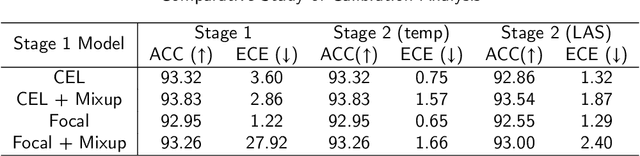
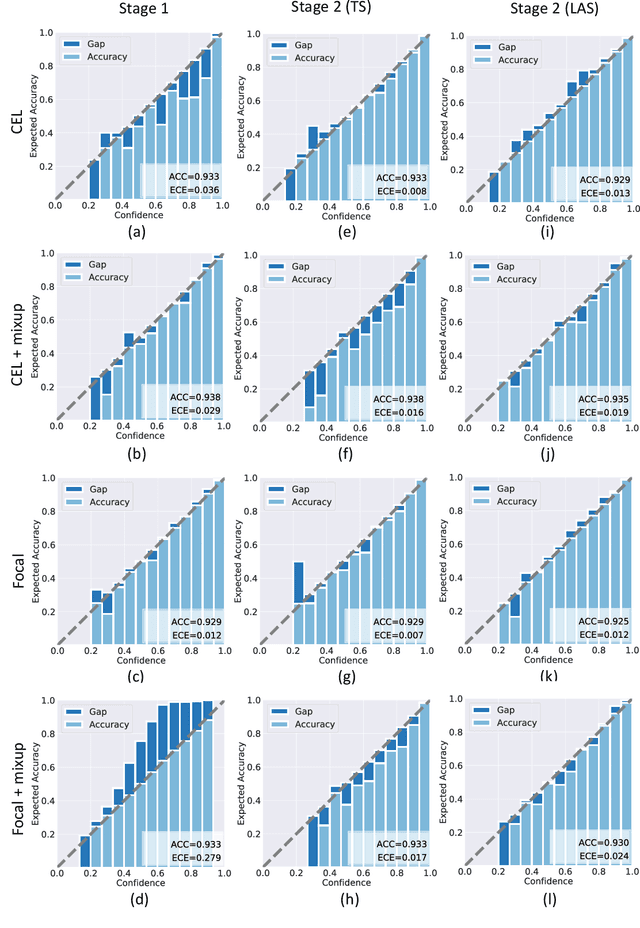
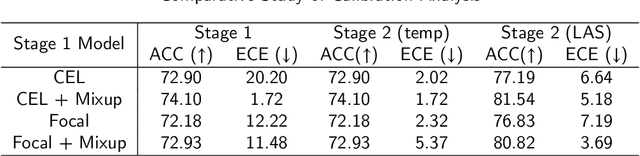
Although deep learning prediction models have been successful in the discrimination of different classes, they can often suffer from poor calibration across challenging domains including healthcare. Moreover, the long-tail distribution poses great challenges in deep learning classification problems including clinical disease prediction. There are approaches proposed recently to calibrate deep prediction in computer vision, but there are no studies found to demonstrate how the representative models work in different challenging contexts. In this paper, we bridge the confidence calibration from computer vision to medical imaging with a comparative study of four high-impact calibration models. Our studies are conducted in different contexts (natural image classification and lung cancer risk estimation) including in balanced vs. imbalanced training sets and in computer vision vs. medical imaging. Our results support key findings: (1) We achieve new conclusions which are not studied under different learning contexts, e.g., combining two calibration models that both mitigate the overconfident prediction can lead to under-confident prediction, and simpler calibration models from the computer vision domain tend to be more generalizable to medical imaging. (2) We highlight the gap between general computer vision tasks and medical imaging prediction, e.g., calibration methods ideal for general computer vision tasks may in fact damage the calibration of medical imaging prediction. (3) We also reinforce previous conclusions in natural image classification settings. We believe that this study has merits to guide readers to choose calibration models and understand gaps between general computer vision and medical imaging domains.
Lossless Image Compression Using a Multi-Scale Progressive Statistical Model
Aug 24, 2021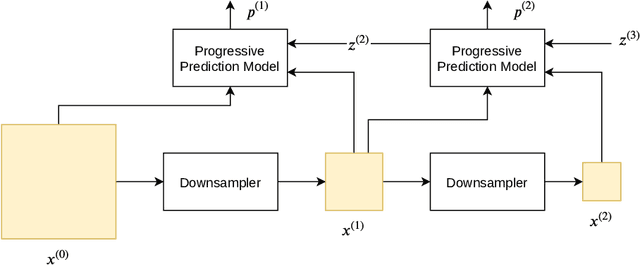

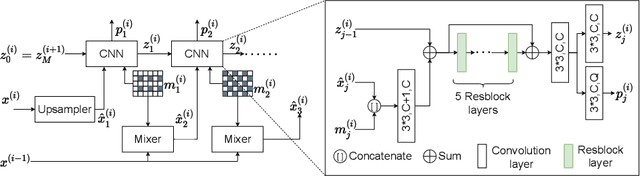
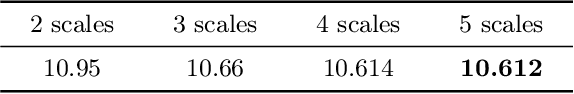
Lossless image compression is an important technique for image storage and transmission when information loss is not allowed. With the fast development of deep learning techniques, deep neural networks have been used in this field to achieve a higher compression rate. Methods based on pixel-wise autoregressive statistical models have shown good performance. However, the sequential processing way prevents these methods to be used in practice. Recently, multi-scale autoregressive models have been proposed to address this limitation. Multi-scale approaches can use parallel computing systems efficiently and build practical systems. Nevertheless, these approaches sacrifice compression performance in exchange for speed. In this paper, we propose a multi-scale progressive statistical model that takes advantage of the pixel-wise approach and the multi-scale approach. We developed a flexible mechanism where the processing order of the pixels can be adjusted easily. Our proposed method outperforms the state-of-the-art lossless image compression methods on two large benchmark datasets by a significant margin without degrading the inference speed dramatically.
Uncertainty-based Cross-Modal Retrieval with Probabilistic Representations
Apr 20, 2022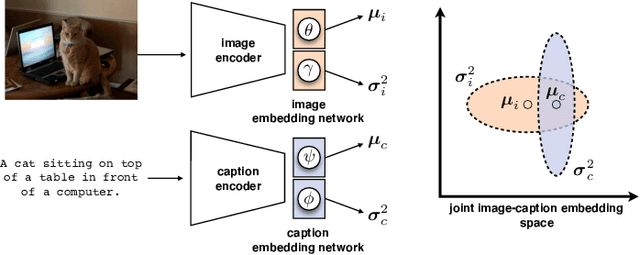
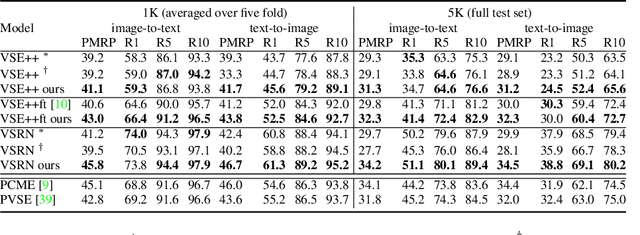
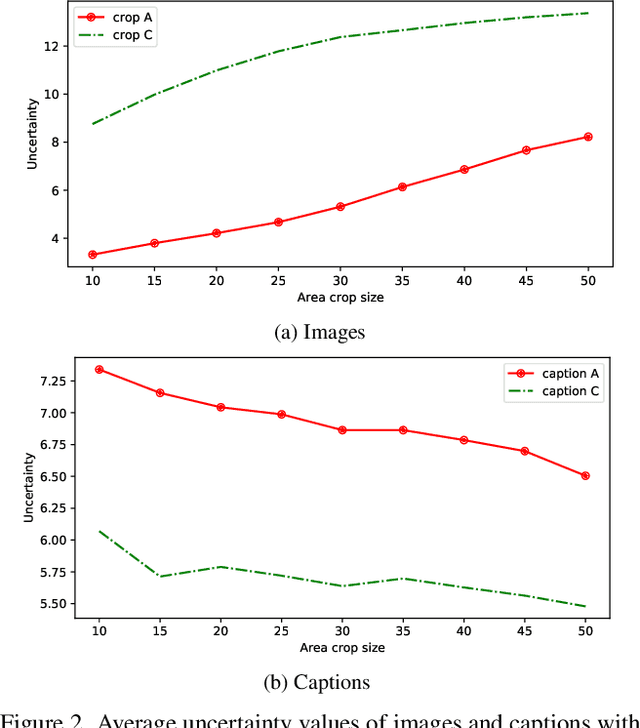
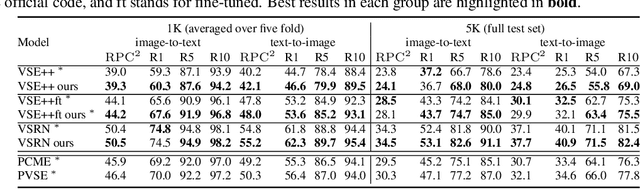
Probabilistic embeddings have proven useful for capturing polysemous word meanings, as well as ambiguity in image matching. In this paper, we study the advantages of probabilistic embeddings in a cross-modal setting (i.e., text and images), and propose a simple approach that replaces the standard vector point embeddings in extant image-text matching models with probabilistic distributions that are parametrically learned. Our guiding hypothesis is that the uncertainty encoded in the probabilistic embeddings captures the cross-modal ambiguity in the input instances, and that it is through capturing this uncertainty that the probabilistic models can perform better at downstream tasks, such as image-to-text or text-to-image retrieval. Through extensive experiments on standard and new benchmarks, we show a consistent advantage for probabilistic representations in cross-modal retrieval, and validate the ability of our embeddings to capture uncertainty.
Visual Enhanced 3D Point Cloud Reconstruction from A Single Image
Aug 17, 2021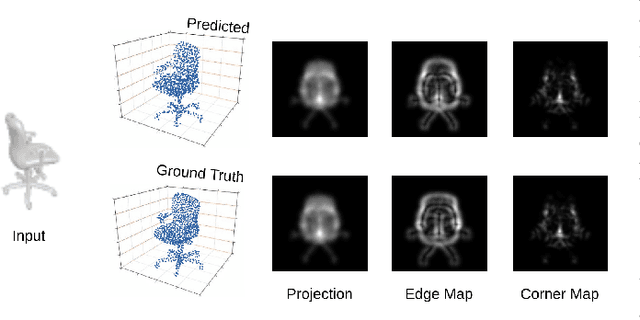
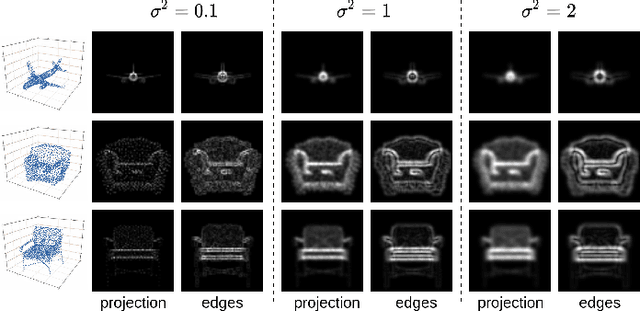
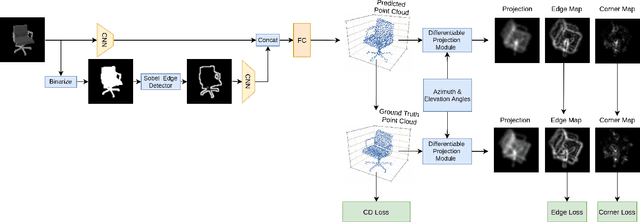
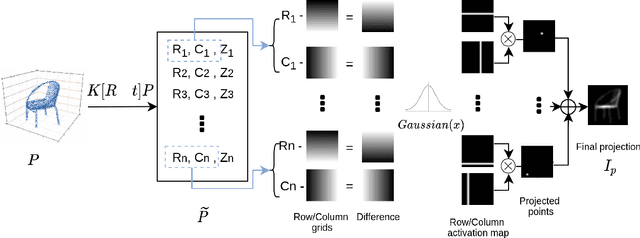

Solving the challenging problem of 3D object reconstruction from a single image appropriately gives existing technologies the ability to perform with a single monocular camera rather than requiring depth sensors. In recent years, thanks to the development of deep learning, 3D reconstruction of a single image has demonstrated impressive progress. Existing researches use Chamfer distance as a loss function to guide the training of the neural network. However, the Chamfer loss will give equal weights to all points inside the 3D point clouds. It tends to sacrifice fine-grained and thin structures to avoid incurring a high loss, which will lead to visually unsatisfactory results. This paper proposes a framework that can recover a detailed three-dimensional point cloud from a single image by focusing more on boundaries (edge and corner points). Experimental results demonstrate that the proposed method outperforms existing techniques significantly, both qualitatively and quantitatively, and has fewer training parameters.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge