"Image": models, code, and papers
Improving Semantic Segmentation in Transformers using Hierarchical Inter-Level Attention
Jul 05, 2022
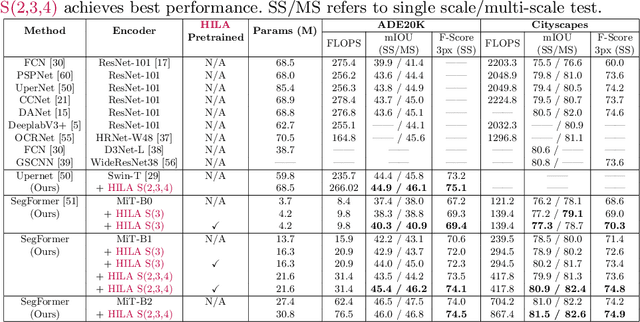
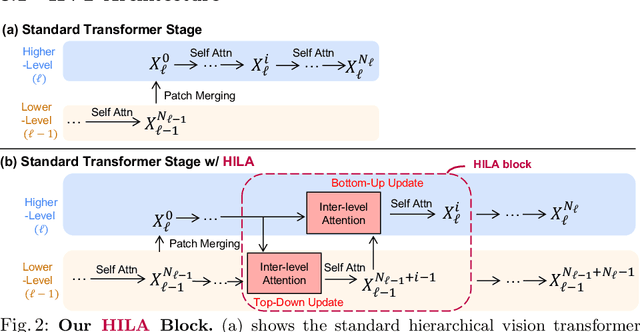
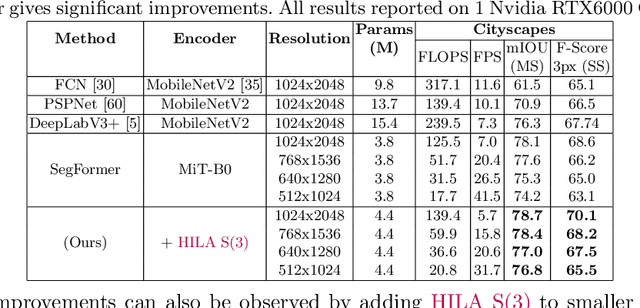
Existing transformer-based image backbones typically propagate feature information in one direction from lower to higher-levels. This may not be ideal since the localization ability to delineate accurate object boundaries, is most prominent in the lower, high-resolution feature maps, while the semantics that can disambiguate image signals belonging to one object vs. another, typically emerges in a higher level of processing. We present Hierarchical Inter-Level Attention (HILA), an attention-based method that captures Bottom-Up and Top-Down Updates between features of different levels. HILA extends hierarchical vision transformer architectures by adding local connections between features of higher and lower levels to the backbone encoder. In each iteration, we construct a hierarchy by having higher-level features compete for assignments to update lower-level features belonging to them, iteratively resolving object-part relationships. These improved lower-level features are then used to re-update the higher-level features. HILA can be integrated into the majority of hierarchical architectures without requiring any changes to the base model. We add HILA into SegFormer and the Swin Transformer and show notable improvements in accuracy in semantic segmentation with fewer parameters and FLOPS. Project website and code: https://www.cs.toronto.edu/~garyleung/hila/
LAConv: Local Adaptive Convolution for Image Fusion
Jul 24, 2021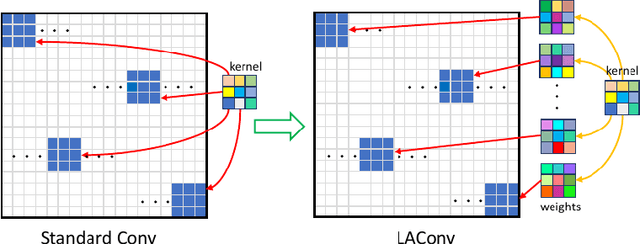
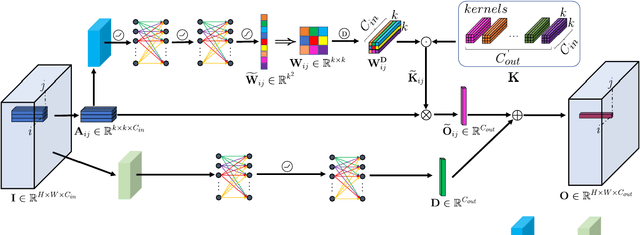
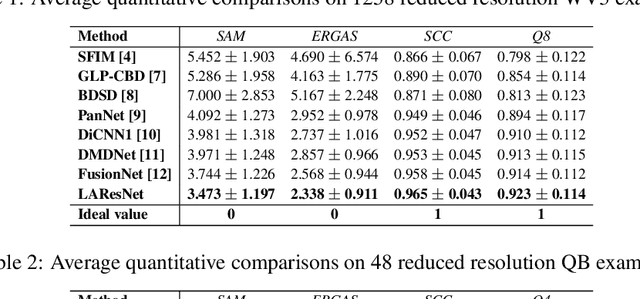
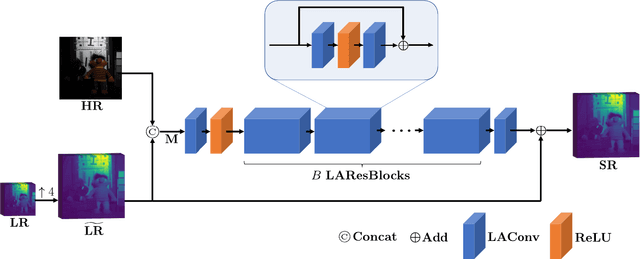
The convolution operation is a powerful tool for feature extraction and plays a prominent role in the field of computer vision. However, when targeting the pixel-wise tasks like image fusion, it would not fully perceive the particularity of each pixel in the image if the uniform convolution kernel is used on different patches. In this paper, we propose a local adaptive convolution (LAConv), which is dynamically adjusted to different spatial locations. LAConv enables the network to pay attention to every specific local area in the learning process. Besides, the dynamic bias (DYB) is introduced to provide more possibilities for the depiction of features and make the network more flexible. We further design a residual structure network equipped with the proposed LAConv and DYB modules, and apply it to two image fusion tasks. Experiments for pansharpening and hyperspectral image super-resolution (HISR) demonstrate the superiority of our method over other state-of-the-art methods. It is worth mentioning that LAConv can also be competent for other super-resolution tasks with less computation effort.
Detecting Pulmonary Embolism from Computed Tomography Using Convolutional Neural Network
Jun 03, 2022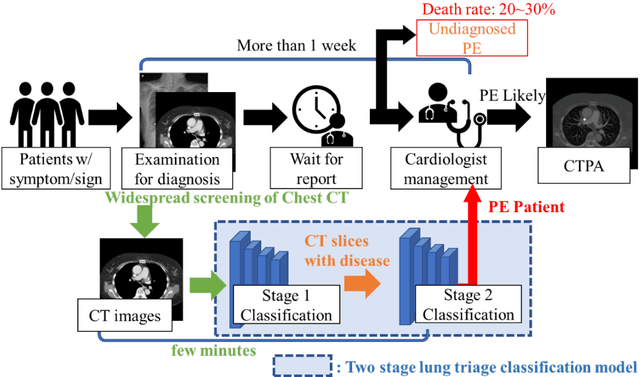
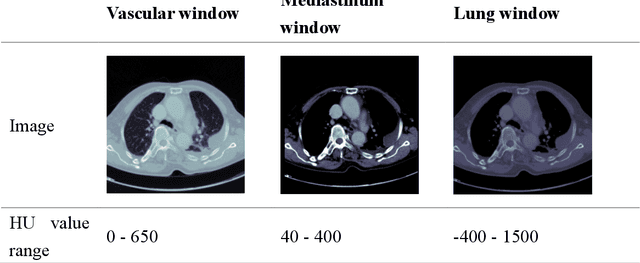
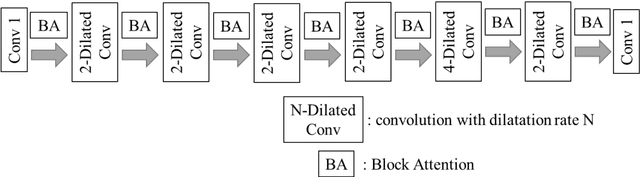
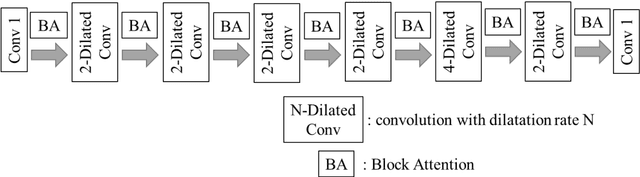
The clinical symptoms of pulmonary embolism (PE) are very diverse and non-specific, which makes it difficult to diagnose. In addition, pulmonary embolism has multiple triggers and is one of the major causes of vascular death. Therefore, if it can be detected and treated quickly, it can significantly reduce the risk of death in hospitalized patients. In the detection process, the cost of computed tomography pulmonary angiography (CTPA) is high, and angiography requires the injection of contrast agents, which increase the risk of damage to the patient. Therefore, this study will use a deep learning approach to detect pulmonary embolism in all patients who take a CT image of the chest using a convolutional neural network. With the proposed pulmonary embolism detection system, we can detect the possibility of pulmonary embolism at the same time as the patient's first CT image, and schedule the CTPA test immediately, saving more than a week of CT image screening time and providing timely diagnosis and treatment to the patient.
Scaling Vision Transformers to Gigapixel Images via Hierarchical Self-Supervised Learning
Jun 06, 2022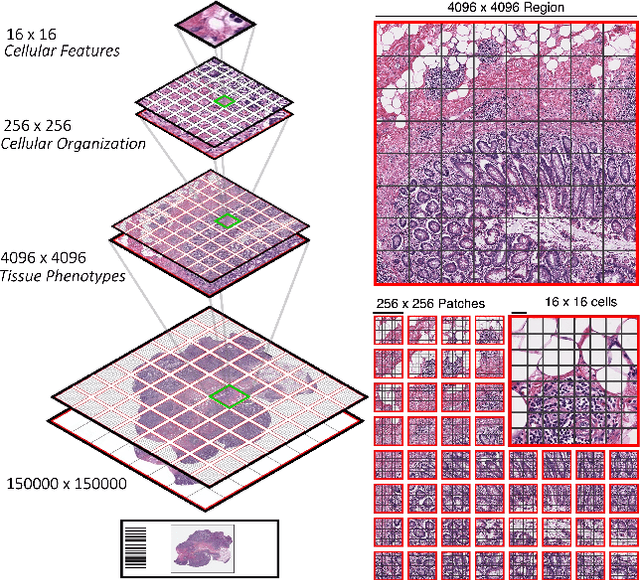
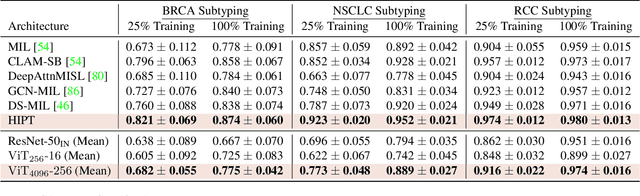
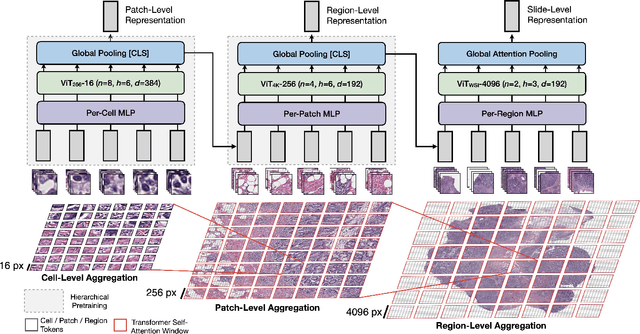
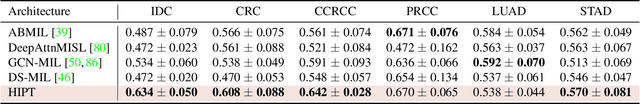
Vision Transformers (ViTs) and their multi-scale and hierarchical variations have been successful at capturing image representations but their use has been generally studied for low-resolution images (e.g. - 256x256, 384384). For gigapixel whole-slide imaging (WSI) in computational pathology, WSIs can be as large as 150000x150000 pixels at 20X magnification and exhibit a hierarchical structure of visual tokens across varying resolutions: from 16x16 images capture spatial patterns among cells, to 4096x4096 images characterizing interactions within the tissue microenvironment. We introduce a new ViT architecture called the Hierarchical Image Pyramid Transformer (HIPT), which leverages the natural hierarchical structure inherent in WSIs using two levels of self-supervised learning to learn high-resolution image representations. HIPT is pretrained across 33 cancer types using 10,678 gigapixel WSIs, 408,218 4096x4096 images, and 104M 256x256 images. We benchmark HIPT representations on 9 slide-level tasks, and demonstrate that: 1) HIPT with hierarchical pretraining outperforms current state-of-the-art methods for cancer subtyping and survival prediction, 2) self-supervised ViTs are able to model important inductive biases about the hierarchical structure of phenotypes in the tumor microenvironment.
AvatarCap: Animatable Avatar Conditioned Monocular Human Volumetric Capture
Jul 05, 2022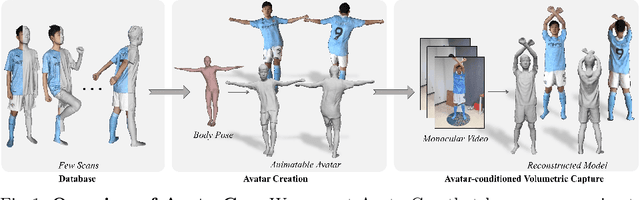

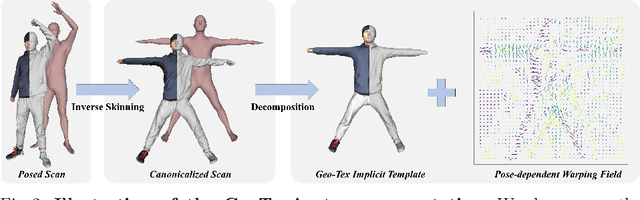
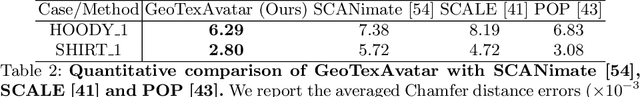
To address the ill-posed problem caused by partial observations in monocular human volumetric capture, we present AvatarCap, a novel framework that introduces animatable avatars into the capture pipeline for high-fidelity reconstruction in both visible and invisible regions. Our method firstly creates an animatable avatar for the subject from a small number (~20) of 3D scans as a prior. Then given a monocular RGB video of this subject, our method integrates information from both the image observation and the avatar prior, and accordingly recon-structs high-fidelity 3D textured models with dynamic details regardless of the visibility. To learn an effective avatar for volumetric capture from only few samples, we propose GeoTexAvatar, which leverages both geometry and texture supervisions to constrain the pose-dependent dynamics in a decomposed implicit manner. An avatar-conditioned volumetric capture method that involves a canonical normal fusion and a reconstruction network is further proposed to integrate both image observations and avatar dynamics for high-fidelity reconstruction in both observed and invisible regions. Overall, our method enables monocular human volumetric capture with detailed and pose-dependent dynamics, and the experiments show that our method outperforms state of the art. Code is available at https://github.com/lizhe00/AvatarCap.
Mutually improved endoscopic image synthesis and landmark detection in unpaired image-to-image translation
Jul 14, 2021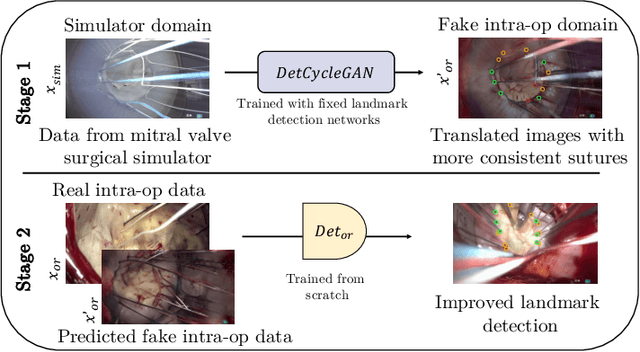
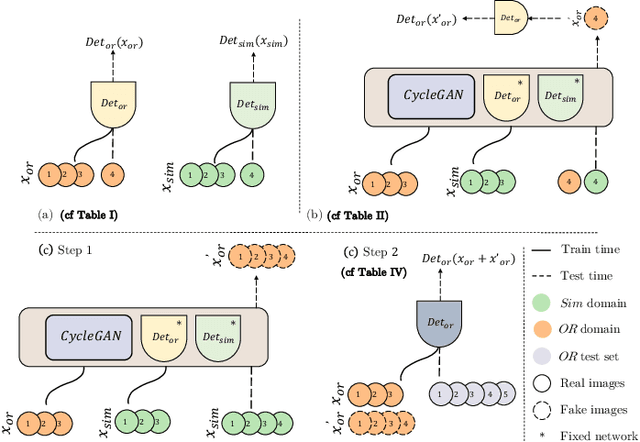
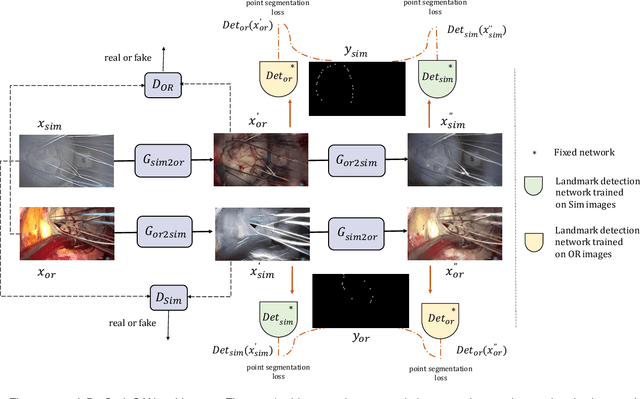
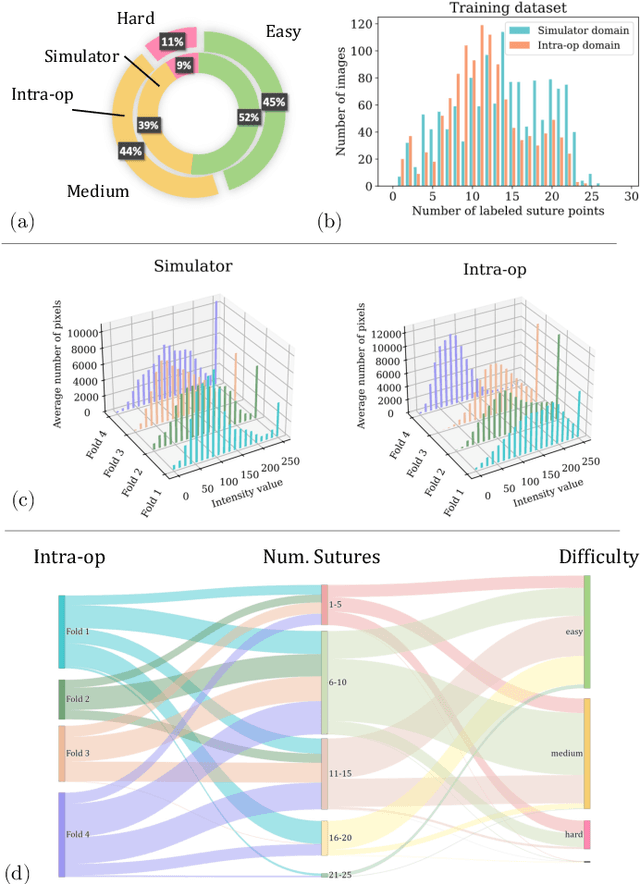
The CycleGAN framework allows for unsupervised image-to-image translation of unpaired data. In a scenario of surgical training on a physical surgical simulator, this method can be used to transform endoscopic images of phantoms into images which more closely resemble the intra-operative appearance of the same surgical target structure. This can be viewed as a novel augmented reality approach, which we coined Hyperrealism in previous work. In this use case, it is of paramount importance to display objects like needles, sutures or instruments consistent in both domains while altering the style to a more tissue-like appearance. Segmentation of these objects would allow for a direct transfer, however, contouring of these, partly tiny and thin foreground objects is cumbersome and perhaps inaccurate. Instead, we propose to use landmark detection on the points when sutures pass into the tissue. This objective is directly incorporated into a CycleGAN framework by treating the performance of pre-trained detector models as an additional optimization goal. We show that a task defined on these sparse landmark labels improves consistency of synthesis by the generator network in both domains. Comparing a baseline CycleGAN architecture to our proposed extension (DetCycleGAN), mean precision (PPV) improved by +61.32, mean sensitivity (TPR) by +37.91, and mean F1 score by +0.4743. Furthermore, it could be shown that by dataset fusion, generated intra-operative images can be leveraged as additional training data for the detection network itself. The data is released within the scope of the AdaptOR MICCAI Challenge 2021 at https://adaptor2021.github.io/, and code at https://github.com/Cardio-AI/detcyclegan_pytorch.
Knowledge Distillation with Representative Teacher Keys Based on Attention Mechanism for Image Classification Model Compression
Jun 26, 2022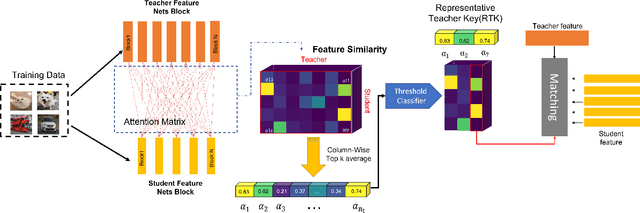
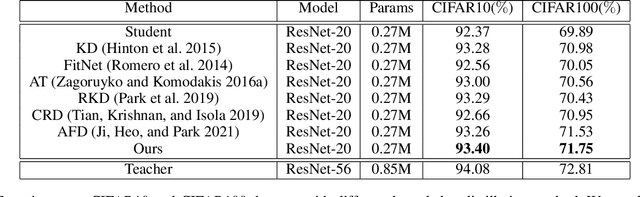
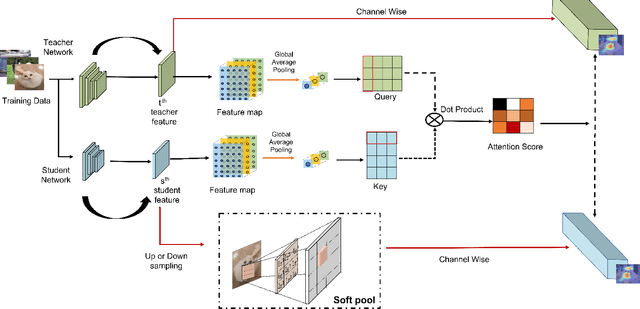
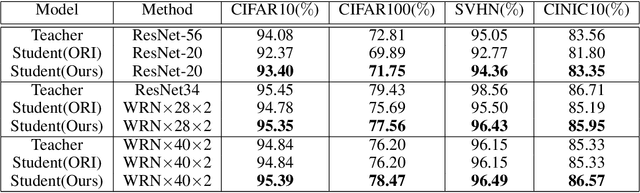
With the improvement of AI chips (e.g., GPU, TPU, and NPU) and the fast development of internet of things (IoTs), some powerful deep neural networks (DNNs) are usually composed of millions or even hundreds of millions of parameters, which may not be suitable to be directly deployed on low computation and low capacity units (e.g., edge devices). Recently, knowledge distillation (KD) has been recognized as one of the effective method of model compression to decrease the model parameters. The main concept of KD is to extract useful information from the feature maps of a large model (i.e., teacher model) as a reference to successfully train a small model (i.e., student model) which model size is much smaller than the teacher one. Although many KD-based methods have been proposed to utilize the information from the feature maps of intermediate layers in teacher model, however, most of them did not consider the similarity of feature maps between teacher model and student model, which may let student model learn useless information. Inspired by attention mechanism, we propose a novel KD method called representative teacher key (RTK) that not only consider the similarity of feature maps but also filter out the useless information to improve the performance of the target student model. In the experiments, we validate our proposed method with several backbone networks (e.g., ResNet and WideResNet) and datasets (e.g., CIFAR10, CIFAR100, SVHN, and CINIC10). The results show that our proposed RTK can effectively improve the classification accuracy of the state-of-the-art attention-based KD method.
An Efficient Multi-Scale Fusion Network for 3D Organ at Risk (OAR) Segmentation
Aug 15, 2022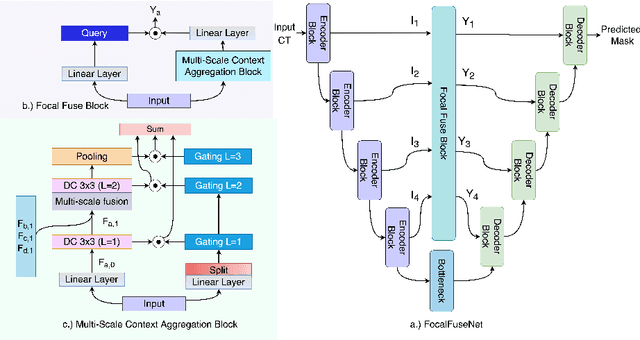

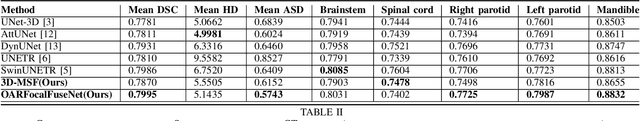
Accurate segmentation of organs-at-risks (OARs) is a precursor for optimizing radiation therapy planning. Existing deep learning-based multi-scale fusion architectures have demonstrated a tremendous capacity for 2D medical image segmentation. The key to their success is aggregating global context and maintaining high resolution representations. However, when translated into 3D segmentation problems, existing multi-scale fusion architectures might underperform due to their heavy computation overhead and substantial data diet. To address this issue, we propose a new OAR segmentation framework, called OARFocalFuseNet, which fuses multi-scale features and employs focal modulation for capturing global-local context across multiple scales. Each resolution stream is enriched with features from different resolution scales, and multi-scale information is aggregated to model diverse contextual ranges. As a result, feature representations are further boosted. The comprehensive comparisons in our experimental setup with OAR segmentation as well as multi-organ segmentation show that our proposed OARFocalFuseNet outperforms the recent state-of-the-art methods on publicly available OpenKBP datasets and Synapse multi-organ segmentation. Both of the proposed methods (3D-MSF and OARFocalFuseNet) showed promising performance in terms of standard evaluation metrics. Our best performing method (OARFocalFuseNet) obtained a dice coefficient of 0.7995 and hausdorff distance of 5.1435 on OpenKBP datasets and dice coefficient of 0.8137 on Synapse multi-organ segmentation dataset.
A differentiable forward model for the concurrent, multi-peak Bragg coherent x-ray diffraction imaging problem
Aug 01, 2022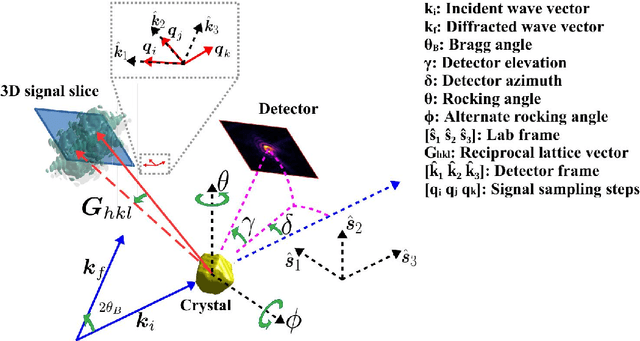
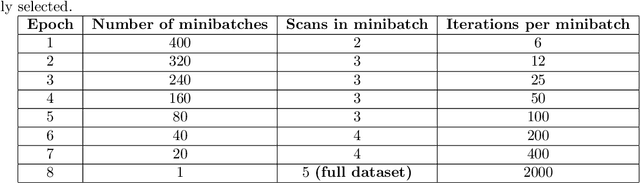
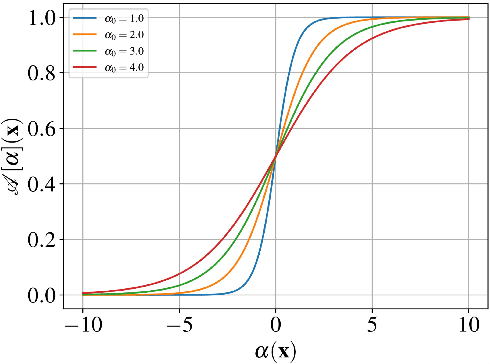
We present a general analytic approach to spatially resolve the nano-scale lattice distortion field of strained and defected compact crystals with Bragg coherent x-ray diffraction imaging (BCDI). Our approach relies on fitting a differentiable forward model simultaneously to multiple BCDI datasets corresponding to independent Bragg reflections from the same single crystal. It is designed to be faithful to heterogeneities that potentially manifest as phase discontinuities in the coherently diffracted wave, such as lattice dislocations in an imperfect crystal. We retain fidelity to such small features in the reconstruction process through a Fourier transform -based resampling algorithm designed to largely avoid the point spread tendencies of commonly employed interpolation methods. The reconstruction model defined in this manner brings BCDI reconstruction into the scope of explicit optimization driven by automatic differentiation. With results from simulations and experimental diffraction data, we demonstrate significant improvement in the final image quality compared to conventional phase retrieval, enabled by explicitly coupling multiple BCDI datasets into the reconstruction loss function.
What do Deep Neural Networks Learn in Medical Images?
Aug 01, 2022
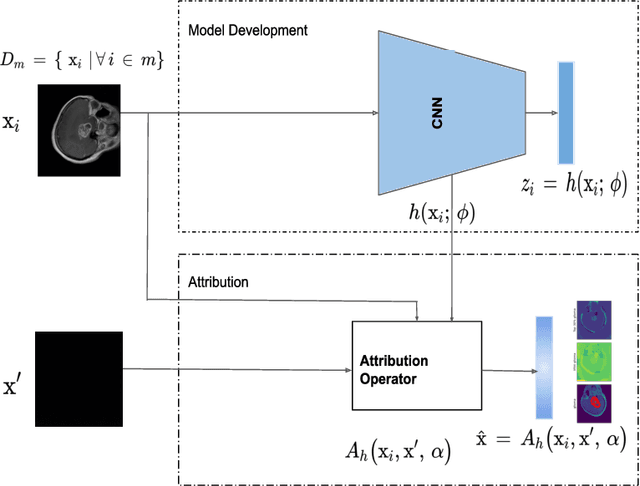
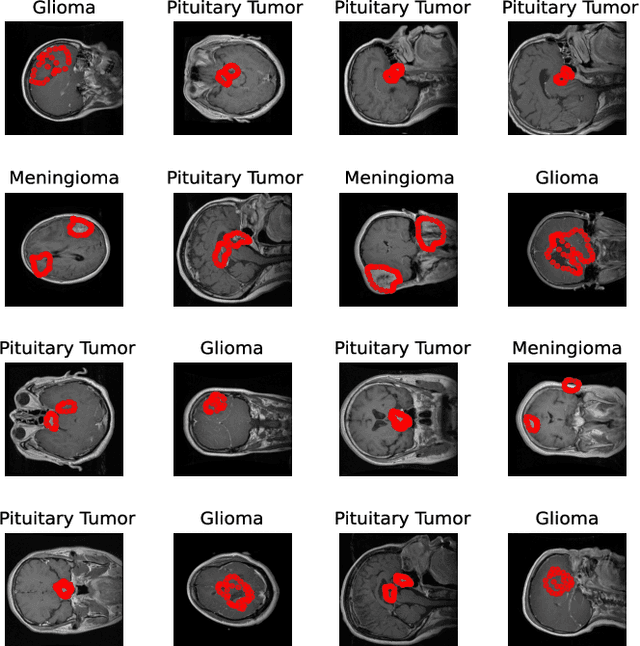
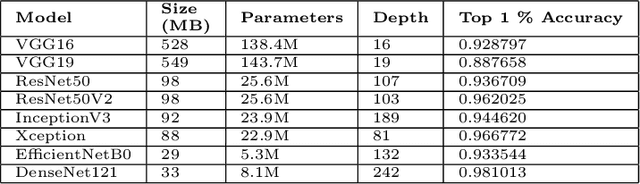
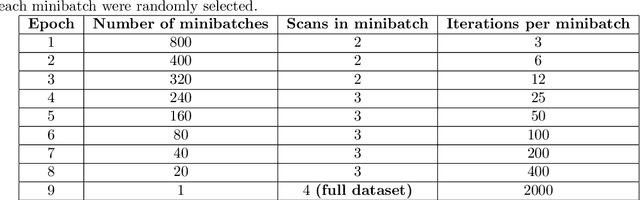
Deep learning is increasingly gaining rapid adoption in healthcare to help improve patient outcomes. This is more so in medical image analysis which requires extensive training to gain the requisite expertise to become a trusted practitioner. However, while deep learning techniques have continued to provide state-of-the-art predictive performance, one of the primary challenges that stands to hinder this progress in healthcare is the opaque nature of the inference mechanism of these models. So, attribution has a vital role in building confidence in stakeholders for the predictions made by deep learning models to inform clinical decisions. This work seeks to answer the question: what do deep neural network models learn in medical images? In that light, we present a novel attribution framework using adaptive path-based gradient integration techniques. Results show a promising direction of building trust in domain experts to improve healthcare outcomes by allowing them to understand the input-prediction correlative structures, discover new bio-markers, and reveal potential model biases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge