"Image": models, code, and papers
Towards Real-World Video Deblurring by Exploring Blur Formation Process
Aug 28, 2022

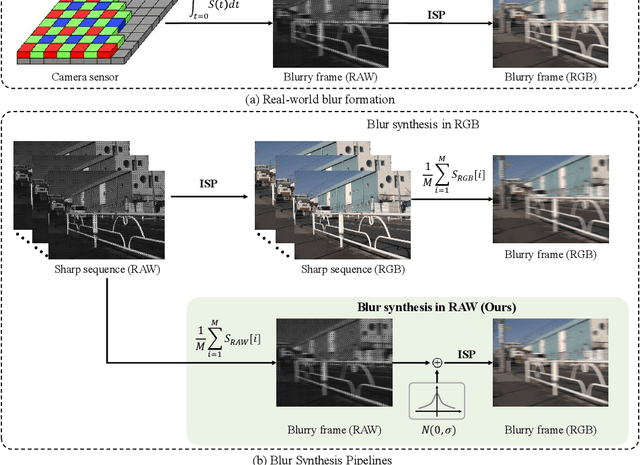
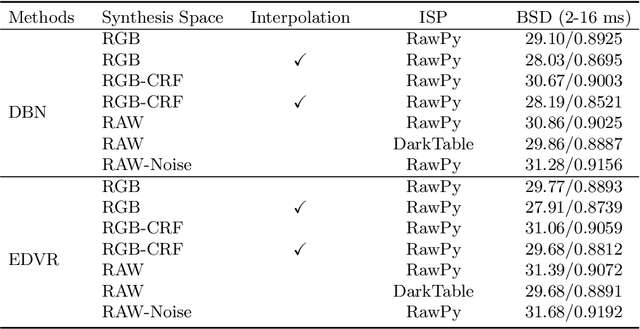
This paper aims at exploring how to synthesize close-to-real blurs that existing video deblurring models trained on them can generalize well to real-world blurry videos. In recent years, deep learning-based approaches have achieved promising success on video deblurring task. However, the models trained on existing synthetic datasets still suffer from generalization problems over real-world blurry scenarios with undesired artifacts. The factors accounting for the failure remain unknown. Therefore, we revisit the classical blur synthesis pipeline and figure out the possible reasons, including shooting parameters, blur formation space, and image signal processor~(ISP). To analyze the effects of these potential factors, we first collect an ultra-high frame-rate (940 FPS) RAW video dataset as the data basis to synthesize various kinds of blurs. Then we propose a novel realistic blur synthesis pipeline termed as RAW-Blur by leveraging blur formation cues. Through numerous experiments, we demonstrate that synthesizing blurs in the RAW space and adopting the same ISP as the real-world testing data can effectively eliminate the negative effects of synthetic data. Furthermore, the shooting parameters of the synthesized blurry video, e.g., exposure time and frame-rate play significant roles in improving the performance of deblurring models. Impressively, the models trained on the blurry data synthesized by the proposed RAW-Blur pipeline can obtain more than 5dB PSNR gain against those trained on the existing synthetic blur datasets. We believe the novel realistic synthesis pipeline and the corresponding RAW video dataset can help the community to easily construct customized blur datasets to improve real-world video deblurring performance largely, instead of laboriously collecting real data pairs.
Texture Generation Using Graph Generative Adversarial Network And Differentiable Rendering
Jun 17, 2022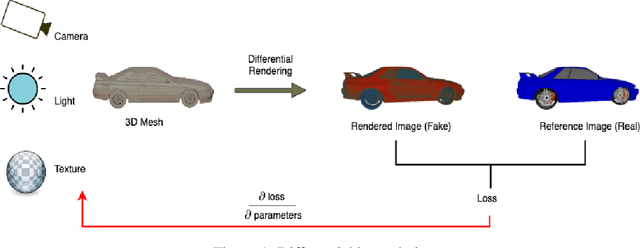
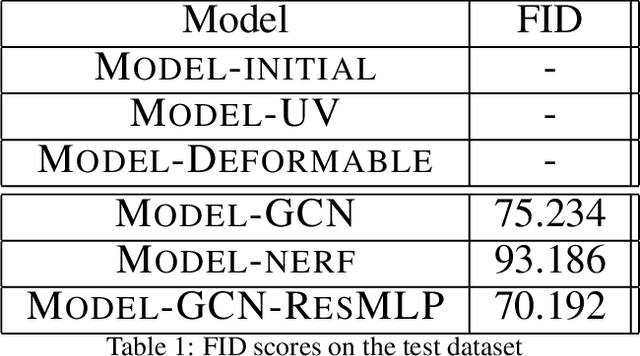
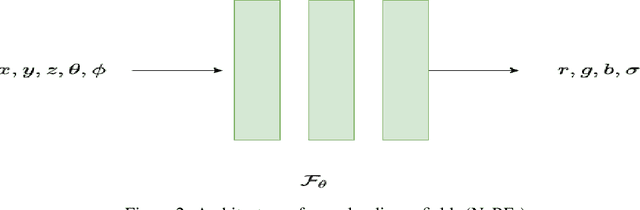
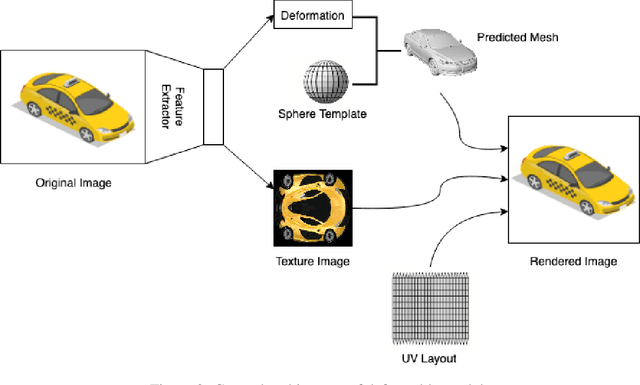
Novel texture synthesis for existing 3D mesh models is an important step towards photo realistic asset generation for existing simulators. But existing methods inherently work in the 2D image space which is the projection of the 3D space from a given camera perspective. These methods take camera angle, 3D model information, lighting information and generate photorealistic 2D image. To generate a photorealistic image from another perspective or lighting, we need to make a computationally expensive forward pass each time we change the parameters. Also, it is hard to generate such images for a simulator that can satisfy the temporal constraints the sequences of images should be similar but only need to change the viewpoint of lighting as desired. The solution can not be directly integrated with existing tools like Blender and Unreal Engine. Manual solution is expensive and time consuming. We thus present a new system called a graph generative adversarial network (GGAN) that can generate textures which can be directly integrated into a given 3D mesh models with tools like Blender and Unreal Engine and can be simulated from any perspective and lighting condition easily.
Few-shot image segmentation for cross-institution male pelvic organs using registration-assisted prototypical learning
Jan 17, 2022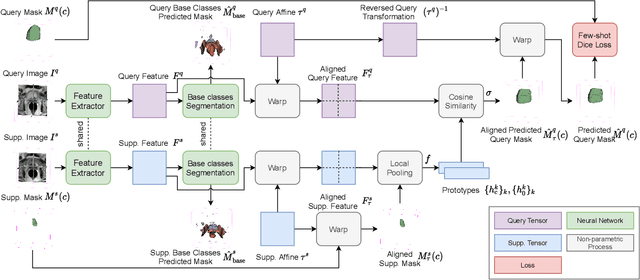
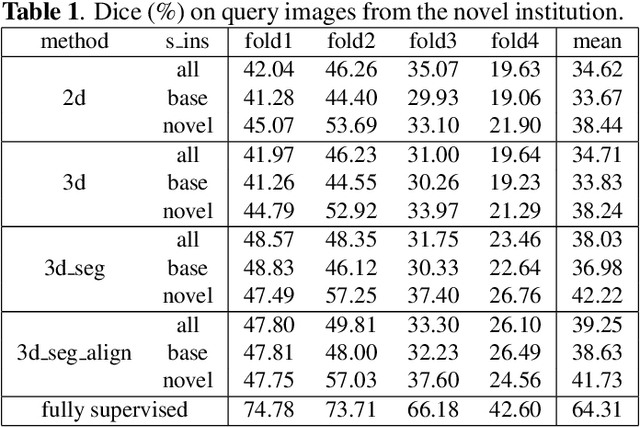
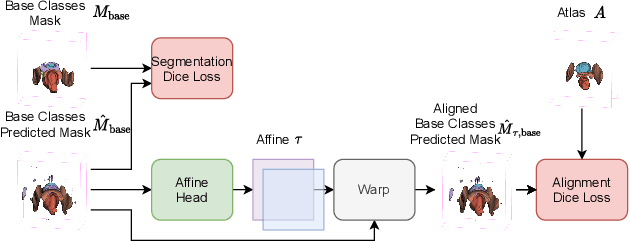
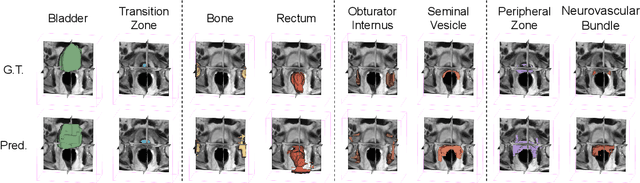
The ability to adapt medical image segmentation networks for a novel class such as an unseen anatomical or pathological structure, when only a few labelled examples of this class are available from local healthcare providers, is sought-after. This potentially addresses two widely recognised limitations in deploying modern deep learning models to clinical practice, expertise-and-labour-intensive labelling and cross-institution generalisation. This work presents the first 3D few-shot interclass segmentation network for medical images, using a labelled multi-institution dataset from prostate cancer patients with eight regions of interest. We propose an image alignment module registering the predicted segmentation of both query and support data, in a standard prototypical learning algorithm, to a reference atlas space. The built-in registration mechanism can effectively utilise the prior knowledge of consistent anatomy between subjects, regardless whether they are from the same institution or not. Experimental results demonstrated that the proposed registration-assisted prototypical learning significantly improved segmentation accuracy (p-values<0.01) on query data from a holdout institution, with varying availability of support data from multiple institutions. We also report the additional benefits of the proposed 3D networks with 75% fewer parameters and an arguably simpler implementation, compared with existing 2D few-shot approaches that segment 2D slices of volumetric medical images.
Sparse Nonnegative Tucker Decomposition and Completion under Noisy Observations
Aug 17, 2022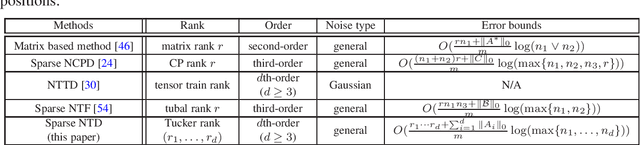
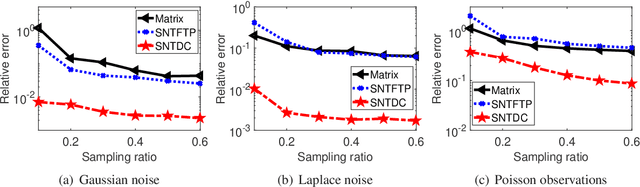
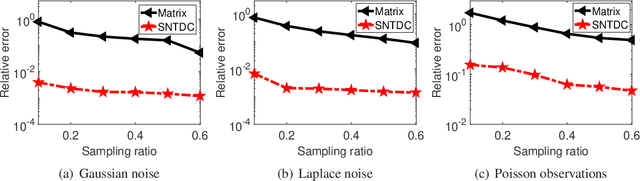
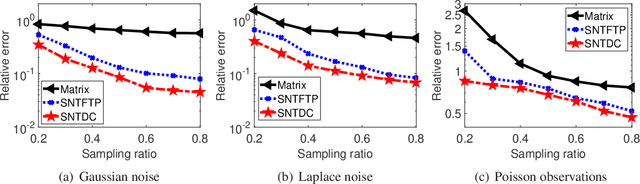
Tensor decomposition is a powerful tool for extracting physically meaningful latent factors from multi-dimensional nonnegative data, and has been an increasing interest in a variety of fields such as image processing, machine learning, and computer vision. In this paper, we propose a sparse nonnegative Tucker decomposition and completion method for the recovery of underlying nonnegative data under noisy observations. Here the underlying nonnegative data tensor is decomposed into a core tensor and several factor matrices with all entries being nonnegative and the factor matrices being sparse. The loss function is derived by the maximum likelihood estimation of the noisy observations, and the $\ell_0$ norm is employed to enhance the sparsity of the factor matrices. We establish the error bound of the estimator of the proposed model under generic noise scenarios, which is then specified to the observations with additive Gaussian noise, additive Laplace noise, and Poisson observations, respectively. Our theoretical results are better than those by existing tensor-based or matrix-based methods. Moreover, the minimax lower bounds are shown to be matched with the derived upper bounds up to logarithmic factors. Numerical examples on both synthetic and real-world data sets demonstrate the superiority of the proposed method for nonnegative tensor data completion.
Leveraging Unsupervised Image Registration for Discovery of Landmark Shape Descriptor
Nov 13, 2021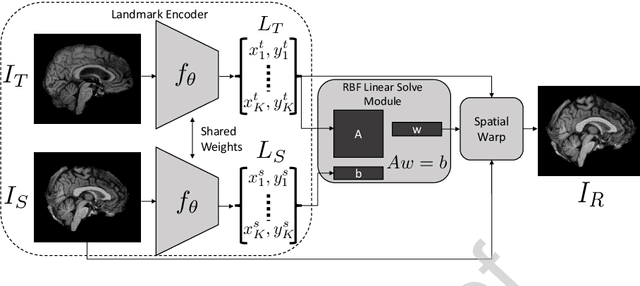
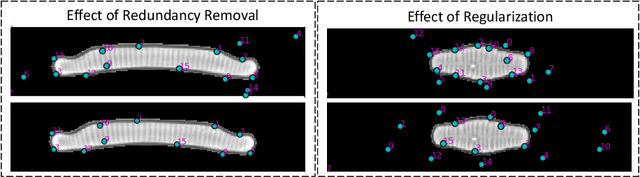
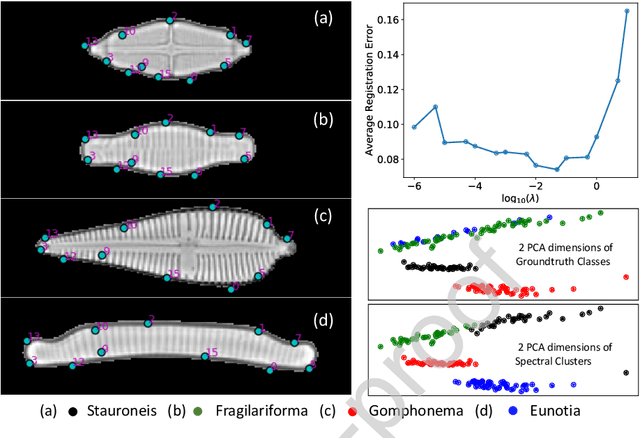
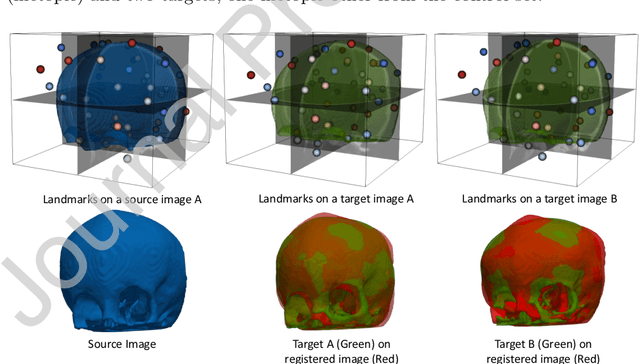
In current biological and medical research, statistical shape modeling (SSM) provides an essential framework for the characterization of anatomy/morphology. Such analysis is often driven by the identification of a relatively small number of geometrically consistent features found across the samples of a population. These features can subsequently provide information about the population shape variation. Dense correspondence models can provide ease of computation and yield an interpretable low-dimensional shape descriptor when followed by dimensionality reduction. However, automatic methods for obtaining such correspondences usually require image segmentation followed by significant preprocessing, which is taxing in terms of both computation as well as human resources. In many cases, the segmentation and subsequent processing require manual guidance and anatomy specific domain expertise. This paper proposes a self-supervised deep learning approach for discovering landmarks from images that can directly be used as a shape descriptor for subsequent analysis. We use landmark-driven image registration as the primary task to force the neural network to discover landmarks that register the images well. We also propose a regularization term that allows for robust optimization of the neural network and ensures that the landmarks uniformly span the image domain. The proposed method circumvents segmentation and preprocessing and directly produces a usable shape descriptor using just 2D or 3D images. In addition, we also propose two variants on the training loss function that allows for prior shape information to be integrated into the model. We apply this framework on several 2D and 3D datasets to obtain their shape descriptors, and analyze their utility for various applications.
End-to-End Supermask Pruning: Learning to Prune Image Captioning Models
Oct 07, 2021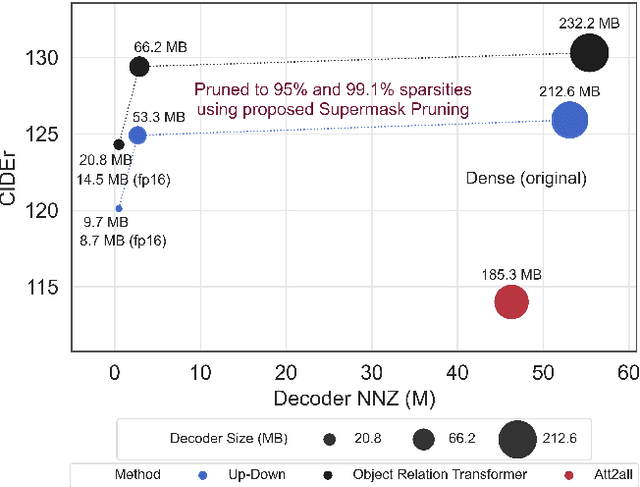
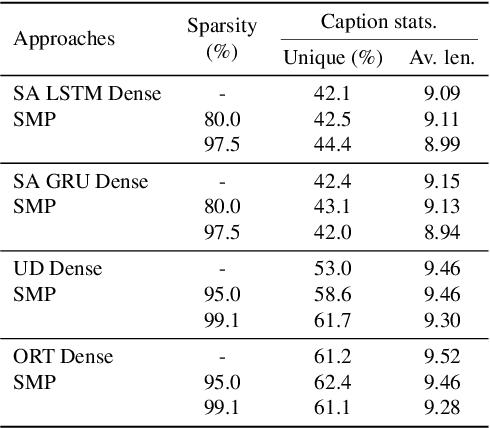
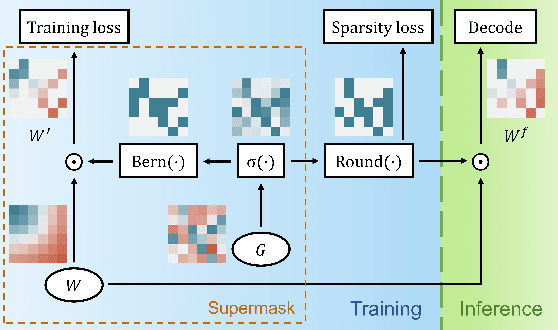
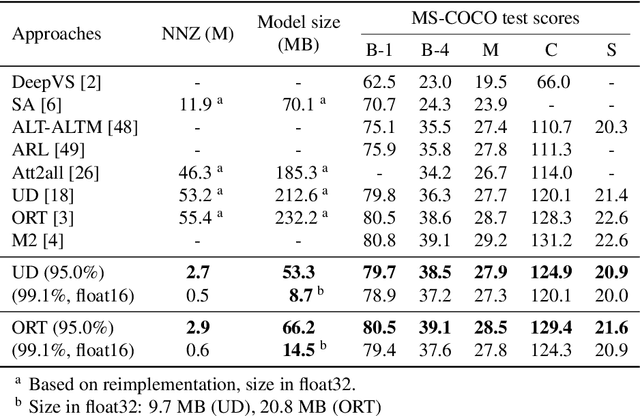
With the advancement of deep models, research work on image captioning has led to a remarkable gain in raw performance over the last decade, along with increasing model complexity and computational cost. However, surprisingly works on compression of deep networks for image captioning task has received little to no attention. For the first time in image captioning research, we provide an extensive comparison of various unstructured weight pruning methods on three different popular image captioning architectures, namely Soft-Attention, Up-Down and Object Relation Transformer. Following this, we propose a novel end-to-end weight pruning method that performs gradual sparsification based on weight sensitivity to the training loss. The pruning schemes are then extended with encoder pruning, where we show that conducting both decoder pruning and training simultaneously prior to the encoder pruning provides good overall performance. Empirically, we show that an 80% to 95% sparse network (up to 75% reduction in model size) can either match or outperform its dense counterpart. The code and pre-trained models for Up-Down and Object Relation Transformer that are capable of achieving CIDEr scores >120 on the MS-COCO dataset but with only 8.7 MB and 14.5 MB in model size (size reduction of 96% and 94% respectively against dense versions) are publicly available at https://github.com/jiahuei/sparse-image-captioning.
BIGRoC: Boosting Image Generation via a Robust Classifier
Aug 08, 2021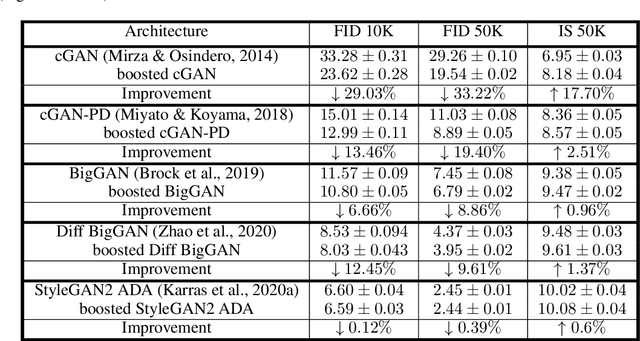
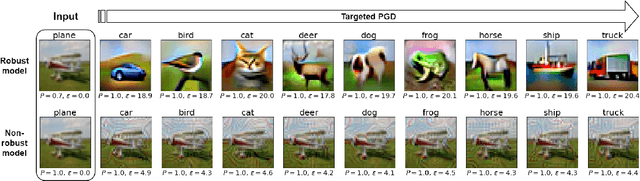
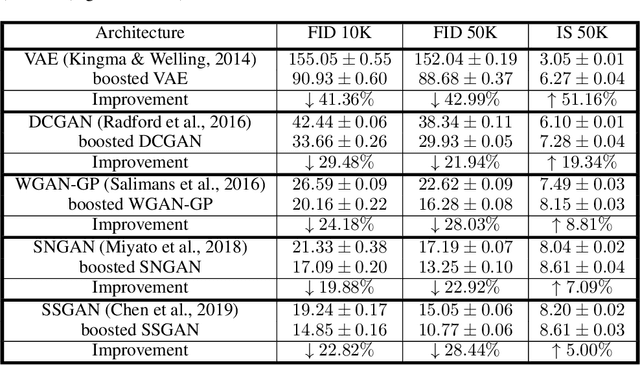
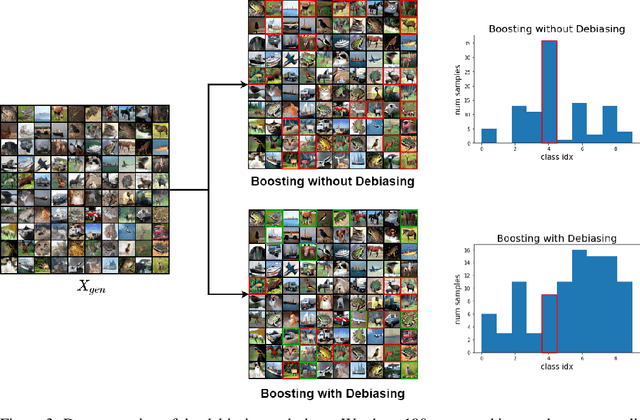
The interest of the machine learning community in image synthesis has grown significantly in recent years, with the introduction of a wide range of deep generative models and means for training them. Such machines' ultimate goal is to match the distributions of the given training images and the synthesized ones. In this work, we propose a general model-agnostic technique for improving the image quality and the distribution fidelity of generated images, obtained by any generative model. Our method, termed BIGRoC (boosting image generation via a robust classifier), is based on a post-processing procedure via the guidance of a given robust classifier and without a need for additional training of the generative model. Given a synthesized image, we propose to update it through projected gradient steps over the robust classifier, in an attempt to refine its recognition. We demonstrate this post-processing algorithm on various image synthesis methods and show a significant improvement of the generated images, both quantitatively and qualitatively.
MLIM: Vision-and-Language Model Pre-training with Masked Language and Image Modeling
Sep 24, 2021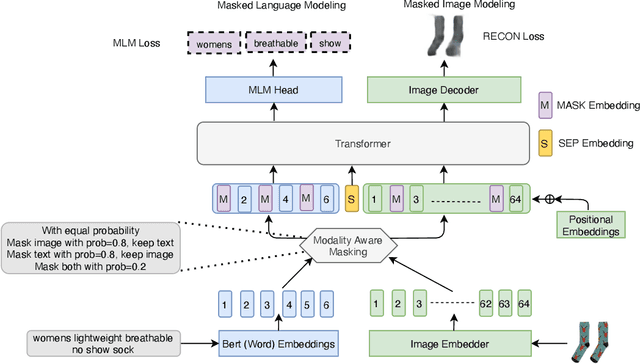
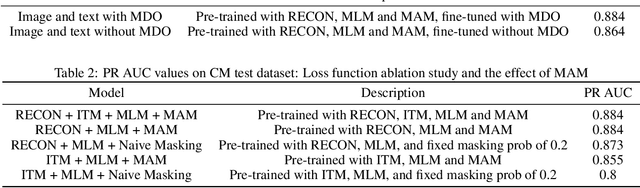

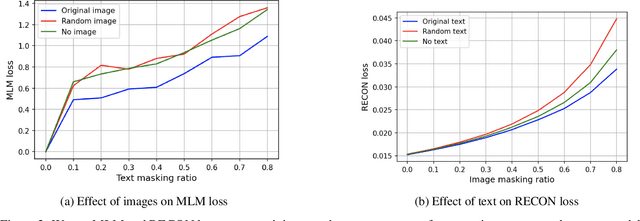
Vision-and-Language Pre-training (VLP) improves model performance for downstream tasks that require image and text inputs. Current VLP approaches differ on (i) model architecture (especially image embedders), (ii) loss functions, and (iii) masking policies. Image embedders are either deep models like ResNet or linear projections that directly feed image-pixels into the transformer. Typically, in addition to the Masked Language Modeling (MLM) loss, alignment-based objectives are used for cross-modality interaction, and RoI feature regression and classification tasks for Masked Image-Region Modeling (MIRM). Both alignment and MIRM objectives mostly do not have ground truth. Alignment-based objectives require pairings of image and text and heuristic objective functions. MIRM relies on object detectors. Masking policies either do not take advantage of multi-modality or are strictly coupled with alignments generated by other models. In this paper, we present Masked Language and Image Modeling (MLIM) for VLP. MLIM uses two loss functions: Masked Language Modeling (MLM) loss and image reconstruction (RECON) loss. We propose Modality Aware Masking (MAM) to boost cross-modality interaction and take advantage of MLM and RECON losses that separately capture text and image reconstruction quality. Using MLM + RECON tasks coupled with MAM, we present a simplified VLP methodology and show that it has better downstream task performance on a proprietary e-commerce multi-modal dataset.
Structure Regularized Attentive Network for Automatic Femoral Head Necrosis Diagnosis and Localization
Aug 23, 2022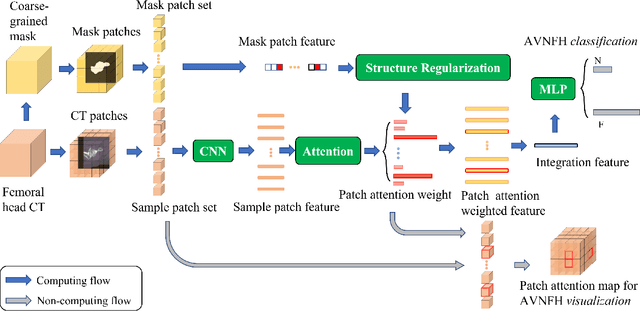
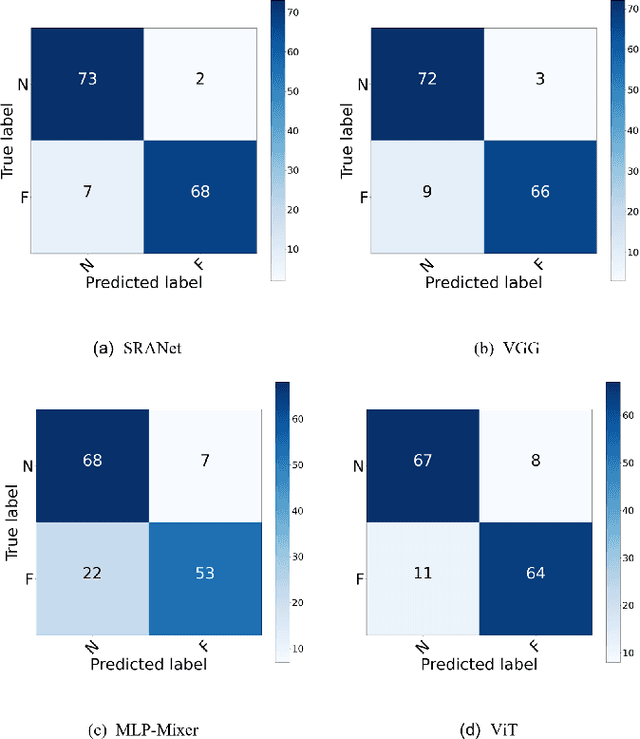
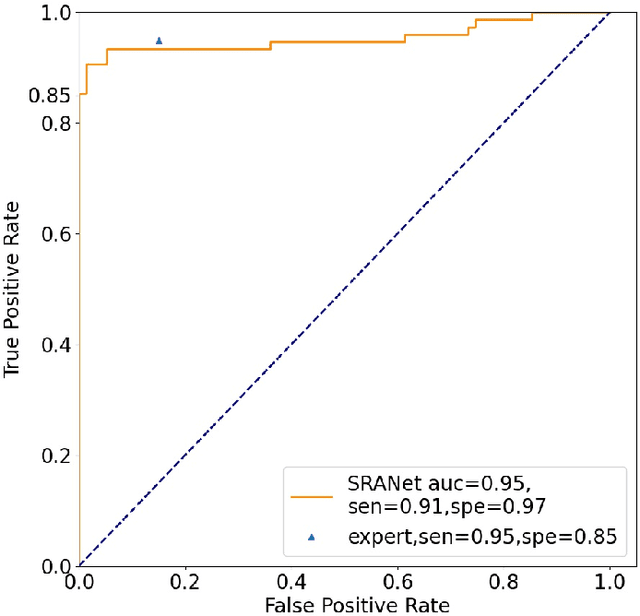
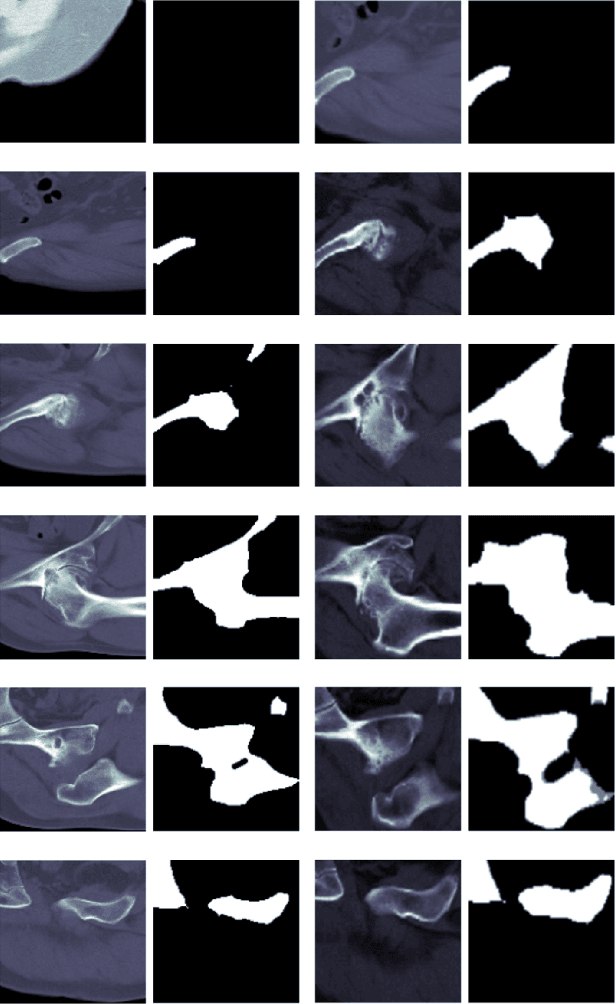
In recent years, several works have adopted the convolutional neural network (CNN) to diagnose the avascular necrosis of the femoral head (AVNFH) based on X-ray images or magnetic resonance imaging (MRI). However, due to the tissue overlap, X-ray images are difficult to provide fine-grained features for early diagnosis. MRI, on the other hand, has a long imaging time, is more expensive, making it impractical in mass screening. Computed tomography (CT) shows layer-wise tissues, is faster to image, and is less costly than MRI. However, to our knowledge, there is no work on CT-based automated diagnosis of AVNFH. In this work, we collected and labeled a large-scale dataset for AVNFH ranking. In addition, existing end-to-end CNNs only yields the classification result and are difficult to provide more information for doctors in diagnosis. To address this issue, we propose the structure regularized attentive network (SRANet), which is able to highlight the necrotic regions during classification based on patch attention. SRANet extracts features in chunks of images, obtains weight via the attention mechanism to aggregate the features, and constrains them by a structural regularizer with prior knowledge to improve the generalization. SRANet was evaluated on our AVNFH-CT dataset. Experimental results show that SRANet is superior to CNNs for AVNFH classification, moreover, it can localize lesions and provide more information to assist doctors in diagnosis. Our codes are made public at https://github.com/tomas-lilingfeng/SRANet.
Weighted Concordance Index Loss-based Multimodal Survival Modeling for Radiation Encephalopathy Assessment in Nasopharyngeal Carcinoma Radiotherapy
Jun 23, 2022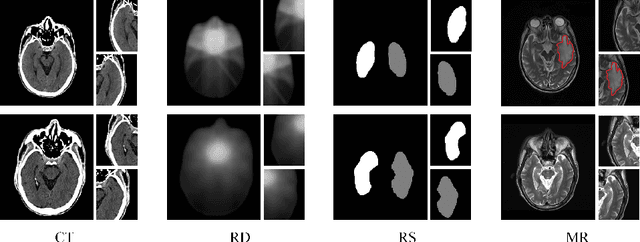
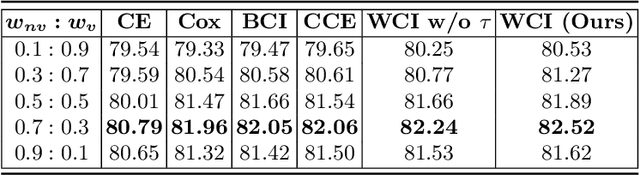
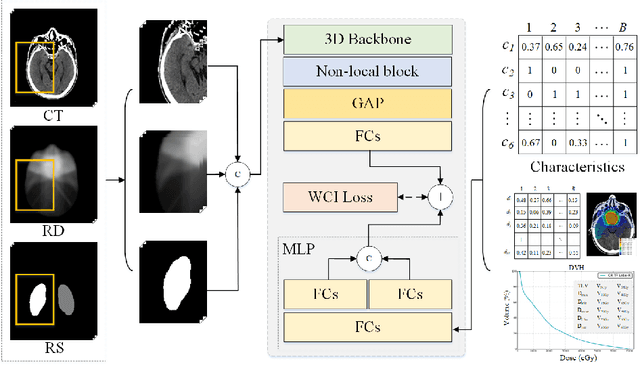
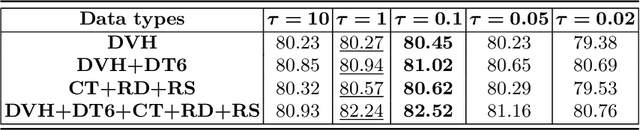
Radiation encephalopathy (REP) is the most common complication for nasopharyngeal carcinoma (NPC) radiotherapy. It is highly desirable to assist clinicians in optimizing the NPC radiotherapy regimen to reduce radiotherapy-induced temporal lobe injury (RTLI) according to the probability of REP onset. To the best of our knowledge, it is the first exploration of predicting radiotherapy-induced REP by jointly exploiting image and non-image data in NPC radiotherapy regimen. We cast REP prediction as a survival analysis task and evaluate the predictive accuracy in terms of the concordance index (CI). We design a deep multimodal survival network (MSN) with two feature extractors to learn discriminative features from multimodal data. One feature extractor imposes feature selection on non-image data, and the other learns visual features from images. Because the priorly balanced CI (BCI) loss function directly maximizing the CI is sensitive to uneven sampling per batch. Hence, we propose a novel weighted CI (WCI) loss function to leverage all REP samples effectively by assigning their different weights with a dual average operation. We further introduce a temperature hyper-parameter for our WCI to sharpen the risk difference of sample pairs to help model convergence. We extensively evaluate our WCI on a private dataset to demonstrate its favourability against its counterparts. The experimental results also show multimodal data of NPC radiotherapy can bring more gains for REP risk prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge