"Image": models, code, and papers
Details or Artifacts: A Locally Discriminative Learning Approach to Realistic Image Super-Resolution
Mar 17, 2022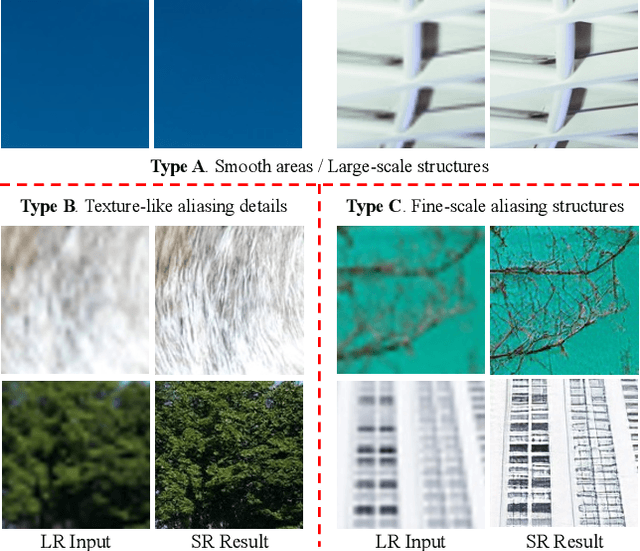
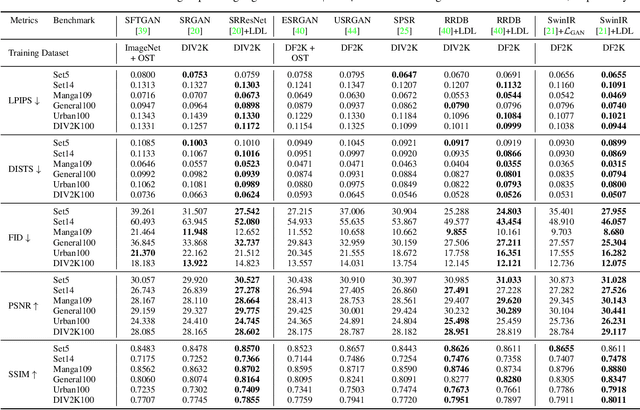
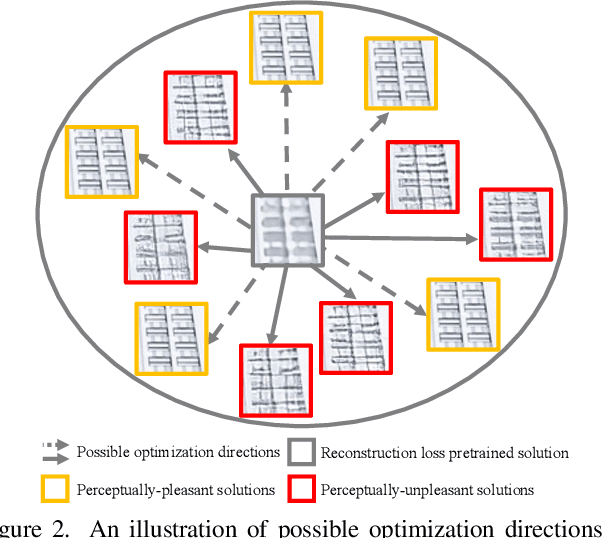
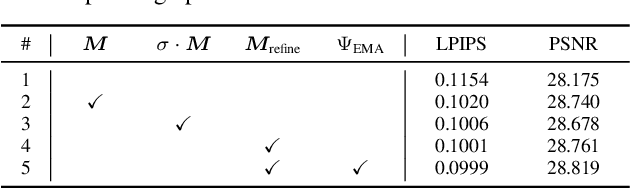
Single image super-resolution (SISR) with generative adversarial networks (GAN) has recently attracted increasing attention due to its potentials to generate rich details. However, the training of GAN is unstable, and it often introduces many perceptually unpleasant artifacts along with the generated details. In this paper, we demonstrate that it is possible to train a GAN-based SISR model which can stably generate perceptually realistic details while inhibiting visual artifacts. Based on the observation that the local statistics (e.g., residual variance) of artifact areas are often different from the areas of perceptually friendly details, we develop a framework to discriminate between GAN-generated artifacts and realistic details, and consequently generate an artifact map to regularize and stabilize the model training process. Our proposed locally discriminative learning (LDL) method is simple yet effective, which can be easily plugged in off-the-shelf SISR methods and boost their performance. Experiments demonstrate that LDL outperforms the state-of-the-art GAN based SISR methods, achieving not only higher reconstruction accuracy but also superior perceptual quality on both synthetic and real-world datasets. Codes and models are available at https://github.com/csjliang/LDL.
Learning Correspondency in Frequency Domain by a Latent-Space Similarity Loss for Multispectral Pansharpening
Jul 18, 2022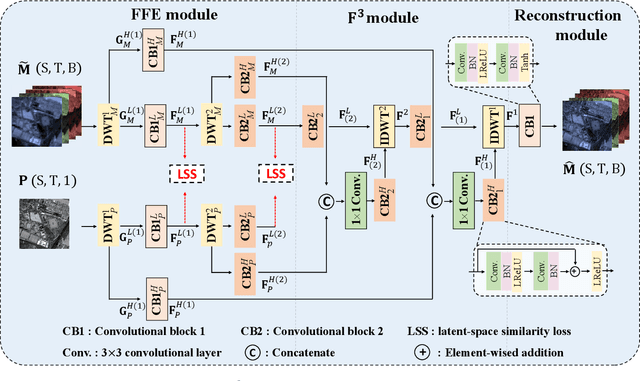
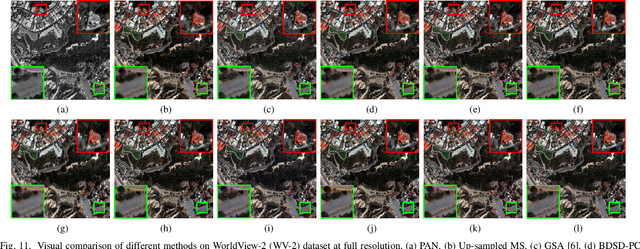


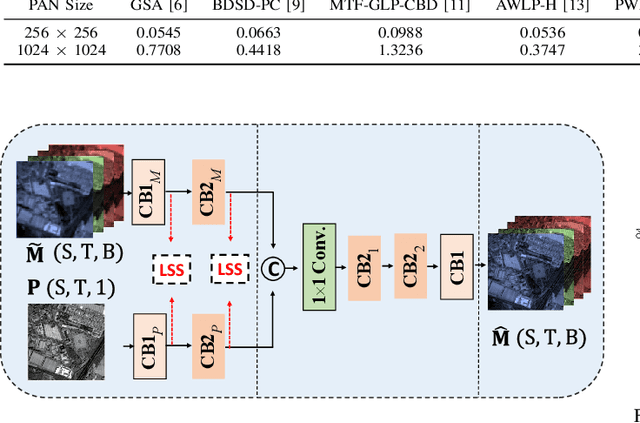
The process of fuse a high spatial resolution (HR) panchromatic (PAN) image and a low spatial resolution (LR) multispectral (MS) image to obtain an HRMS image is known as pansharpening. With the development of convolutional neural networks, the performance of pansharpening methods has been improved, however, the blurry effects and the spectral distortion still exist in their fusion results due to the insufficiency in details learning and the mismatch between the high-frequency (HF) and low-frequency (LF) components. Therefore, the improvements of spatial details at the premise of reducing spectral distortion is still a challenge. In this paper, we propose a frequency-aware network (FAN) together with a novel latent-space similarity loss to address above mentioned problems. FAN is composed of three modules, where the frequency feature extraction module aims to extract features in the frequency domain with the help of discrete wavelet transform (DWT) layers, and the inverse DWT (IDWT) layers are then utilized in the frequency feature fusion module to reconstruct the features. Finally, the fusion results are obtained through the reconstruction module. In order to learn the correspondency, we also propose a latent-space similarity loss to constrain the LF features derived from PAN and MS branches, so that HF features of PAN can reasonably be used to supplement that of MS. Experimental results on three datasets at both reduced- and full-resolution demonstrate the superiority of the proposed method compared with several state-of-the-art pansharpening models, especially for the fusion at full resolution.
Label Propagation for 3D Carotid Vessel Wall Segmentation and Atherosclerosis Diagnosis
Aug 29, 2022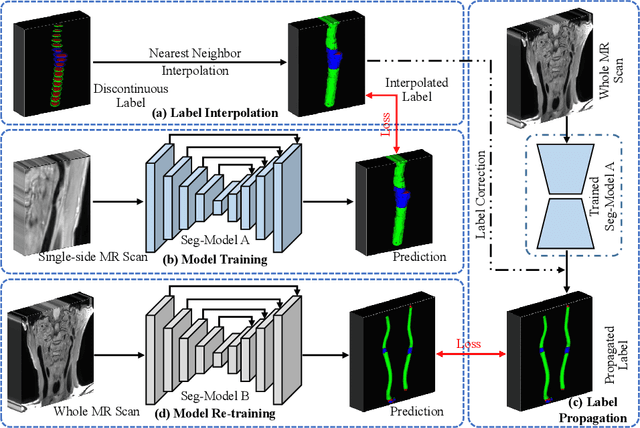
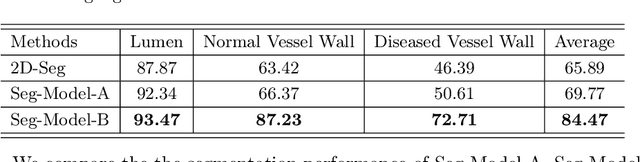
Carotid vessel wall segmentation is a crucial yet challenging task in the computer-aided diagnosis of atherosclerosis. Although numerous deep learning models have achieved remarkable success in many medical image segmentation tasks, accurate segmentation of carotid vessel wall on magnetic resonance (MR) images remains challenging, due to limited annotations and heterogeneous arteries. In this paper, we propose a semi-supervised label propagation framework to segment lumen, normal vessel walls, and atherosclerotic vessel wall on 3D MR images. By interpolating the provided annotations, we get 3D continuous labels for training 3D segmentation model. With the trained model, we generate pseudo labels for unlabeled slices to incorporate them for model training. Then we use the whole MR scans and the propagated labels to re-train the segmentation model and improve its robustness. We evaluated the label propagation framework on the CarOtid vessel wall SegMentation and atherosclerOsis diagnosiS (COSMOS) Challenge dataset and achieved a QuanM score of 83.41\% on the testing dataset, which got the 1-st place on the online evaluation leaderboard. The results demonstrate the effectiveness of the proposed framework.
A Multi-attribute Controllable Generative Model for Histopathology Image Synthesis
Nov 10, 2021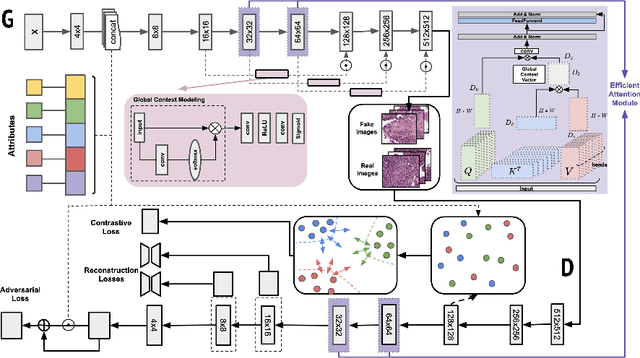
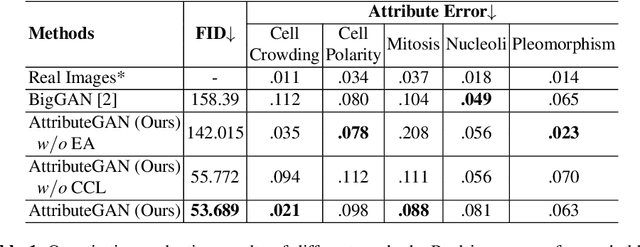
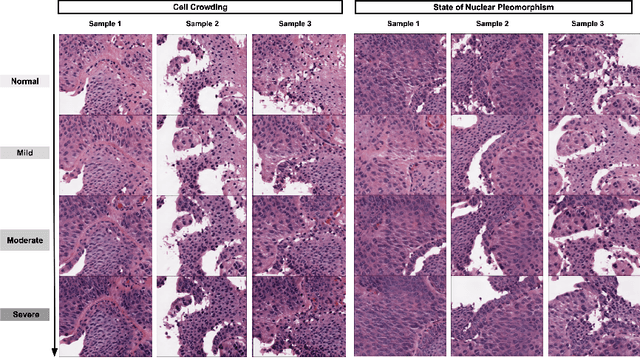
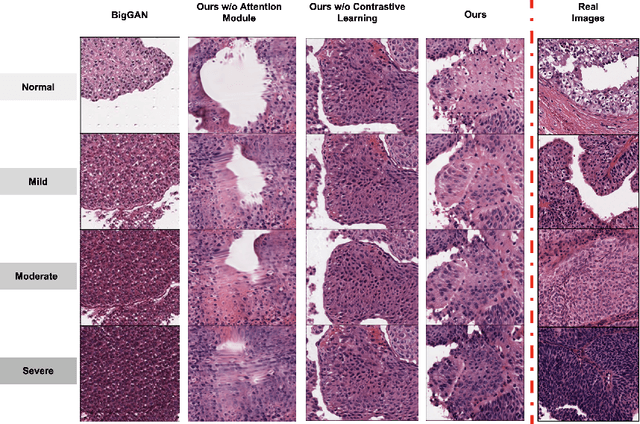
Generative models have been applied in the medical imaging domain for various image recognition and synthesis tasks. However, a more controllable and interpretable image synthesis model is still lacking yet necessary for important applications such as assisting in medical training. In this work, we leverage the efficient self-attention and contrastive learning modules and build upon state-of-the-art generative adversarial networks (GANs) to achieve an attribute-aware image synthesis model, termed AttributeGAN, which can generate high-quality histopathology images based on multi-attribute inputs. In comparison to existing single-attribute conditional generative models, our proposed model better reflects input attributes and enables smoother interpolation among attribute values. We conduct experiments on a histopathology dataset containing stained H&E images of urothelial carcinoma and demonstrate the effectiveness of our proposed model via comprehensive quantitative and qualitative comparisons with state-of-the-art models as well as different variants of our model. Code is available at https://github.com/karenyyy/MICCAI2021AttributeGAN.
Boosting 3D Object Detection by Simulating Multimodality on Point Clouds
Jun 30, 2022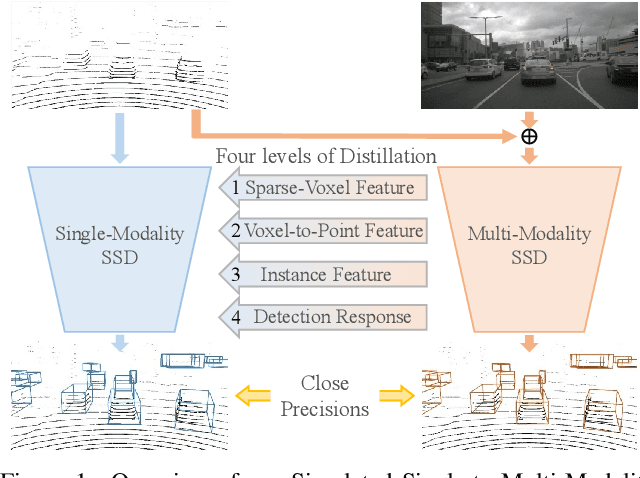

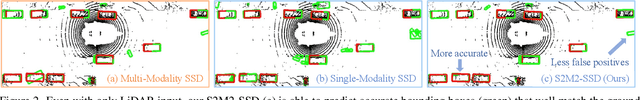
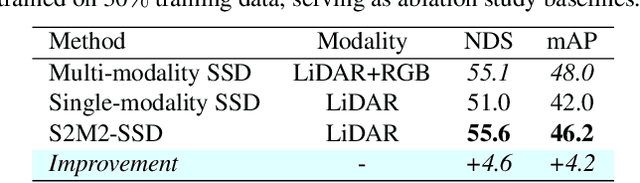
This paper presents a new approach to boost a single-modality (LiDAR) 3D object detector by teaching it to simulate features and responses that follow a multi-modality (LiDAR-image) detector. The approach needs LiDAR-image data only when training the single-modality detector, and once well-trained, it only needs LiDAR data at inference. We design a novel framework to realize the approach: response distillation to focus on the crucial response samples and avoid the background samples; sparse-voxel distillation to learn voxel semantics and relations from the estimated crucial voxels; a fine-grained voxel-to-point distillation to better attend to features of small and distant objects; and instance distillation to further enhance the deep-feature consistency. Experimental results on the nuScenes dataset show that our approach outperforms all SOTA LiDAR-only 3D detectors and even surpasses the baseline LiDAR-image detector on the key NDS metric, filling 72% mAP gap between the single- and multi-modality detectors.
Deep Neural Patchworks: Coping with Large Segmentation Tasks
Jun 07, 2022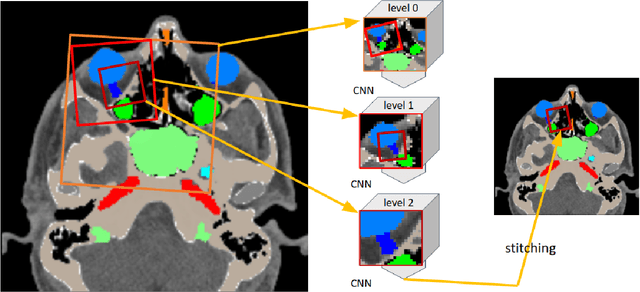
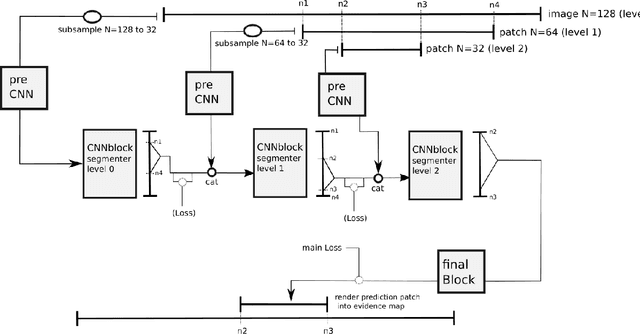
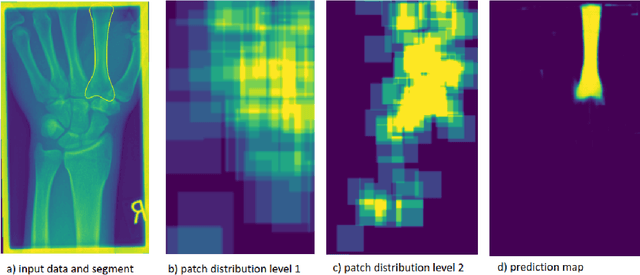
Convolutional neural networks are the way to solve arbitrary image segmentation tasks. However, when images are large, memory demands often exceed the available resources, in particular on a common GPU. Especially in biomedical imaging, where 3D images are common, the problems are apparent. A typical approach to solve this limitation is to break the task into smaller subtasks by dividing images into smaller image patches. Another approach, if applicable, is to look at the 2D image sections separately, and to solve the problem in 2D. Often, the loss of global context makes such approaches less effective; important global information might not be present in the current image patch, or the selected 2D image section. Here, we propose Deep Neural Patchworks (DNP), a segmentation framework that is based on hierarchical and nested stacking of patch-based networks that solves the dilemma between global context and memory limitations.
Generalization in Neural Networks: A Broad Survey
Sep 04, 2022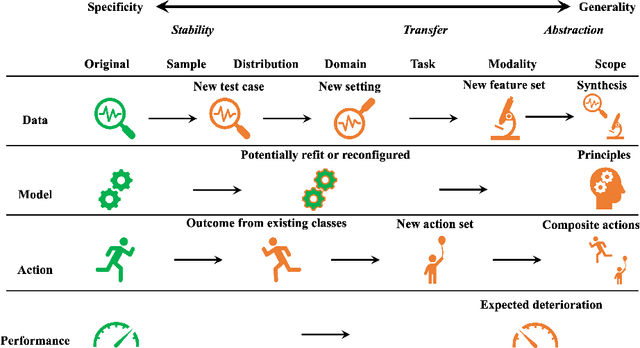
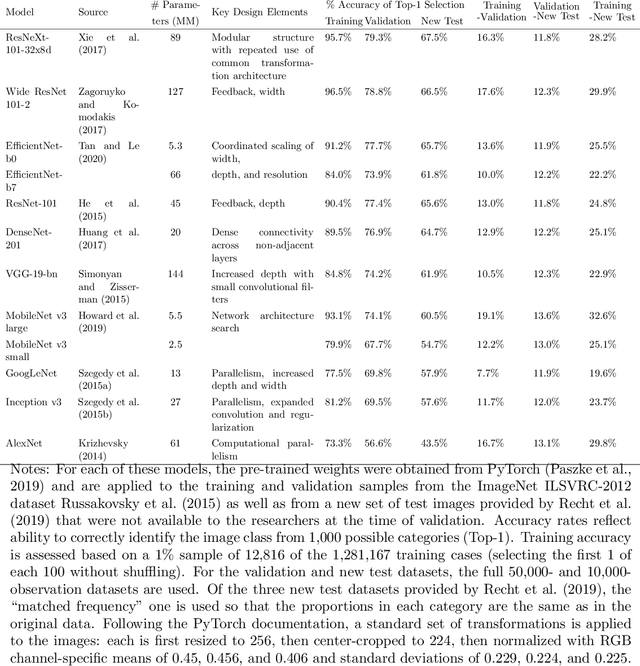
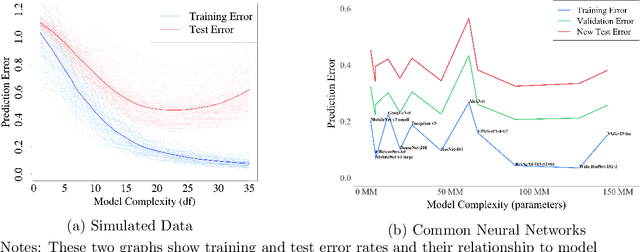
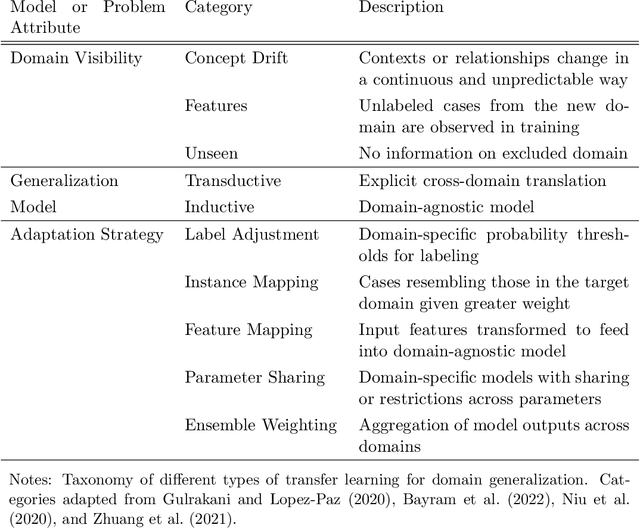
This paper reviews concepts, modeling approaches, and recent findings along a spectrum of different levels of abstraction of neural network models including generalization across (1) Samples, (2) Distributions, (3) Domains, (4) Tasks, (5) Modalities, and (6) Scopes. Results on (1) sample generalization show that, in the case of ImageNet, nearly all the recent improvements reduced training error while overfitting stayed flat; with nearly all the training error eliminated, future progress will require a focus on reducing overfitting. Perspectives from statistics highlight how (2) distribution generalization can be viewed alternately as a change in sample weights or a change in the input-output relationship. Transfer learning approaches to (3) domain generalization are summarized, as are recent advances and the wealth of domain adaptation benchmark datasets available. Recent breakthroughs surveyed in (4) task generalization include few-shot meta-learning approaches and the BERT NLP engine, and recent (5) modality generalization studies are discussed that integrate image and text data and that apply a biologically-inspired network across olfactory, visual, and auditory modalities. Recent (6) scope generalization results are reviewed that embed knowledge graphs into deep NLP approaches. Additionally, concepts from neuroscience are discussed on the modular architecture of brains and the steps by which dopamine-driven conditioning leads to abstract thinking.
SeasoNet: A Seasonal Scene Classification, segmentation and Retrieval dataset for satellite Imagery over Germany
Jul 19, 2022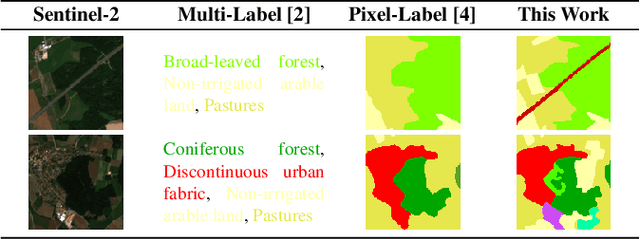
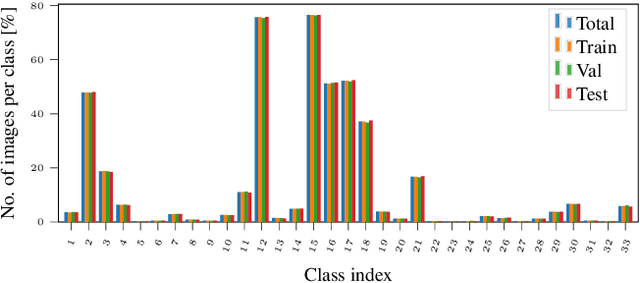
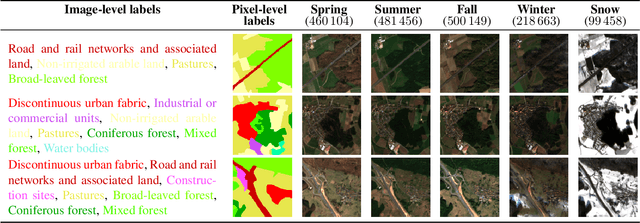
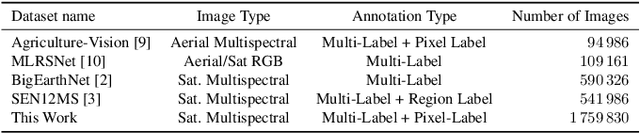
This work presents SeasoNet, a new large-scale multi-label land cover and land use scene understanding dataset. It includes $1\,759\,830$ images from Sentinel-2 tiles, with 12 spectral bands and patch sizes of up to $ 120 \ \mathrm{px} \times 120 \ \mathrm{px}$. Each image is annotated with large scale pixel level labels from the German land cover model LBM-DE2018 with land cover classes based on the CORINE Land Cover database (CLC) 2018 and a five times smaller minimum mapping unit (MMU) than the original CLC maps. We provide pixel synchronous examples from all four seasons, plus an additional snowy set. These properties make SeasoNet the currently most versatile and biggest remote sensing scene understanding dataset with possible applications ranging from scene classification over land cover mapping to content-based cross season image retrieval and self-supervised feature learning. We provide baseline results by evaluating state-of-the-art deep networks on the new dataset in scene classification and semantic segmentation scenarios.
Novel Deep Learning Approach to Derive Cytokeratin Expression and Epithelium Segmentation from DAPI
Aug 16, 2022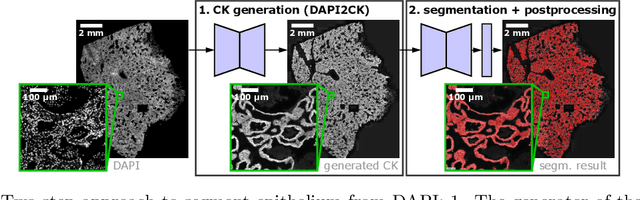
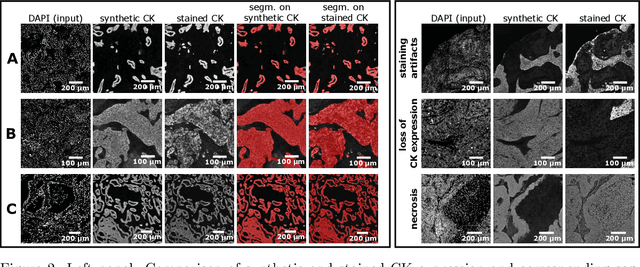

Generative Adversarial Networks (GANs) are state of the art for image synthesis. Here, we present dapi2ck, a novel GAN-based approach to synthesize cytokeratin (CK) staining from immunofluorescent (IF) DAPI staining of nuclei in non-small cell lung cancer (NSCLC) images. We use the synthetic CK to segment epithelial regions, which, compared to expert annotations, yield equally good results as segmentation on stained CK. Considering the limited number of markers in a multiplexed IF (mIF) panel, our approach allows to replace CK by another marker addressing the complexity of the tumor micro-environment (TME) to facilitate patient selection for immunotherapies. In contrast to stained CK, dapi2ck does not suffer from issues like unspecific CK staining or loss of tumoral CK expression.
Transformer-Unet: Raw Image Processing with Unet
Sep 17, 2021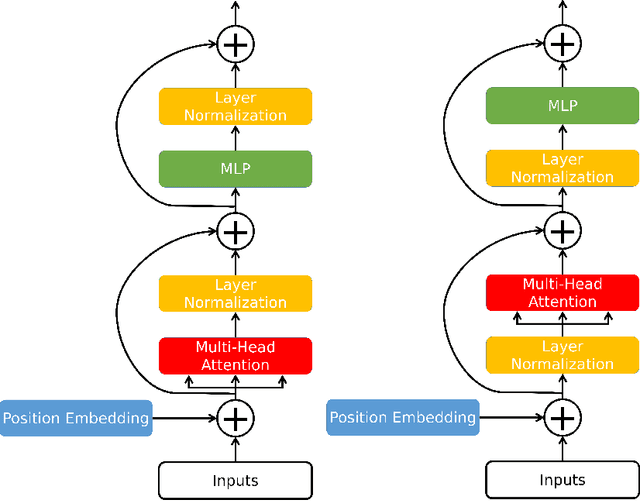
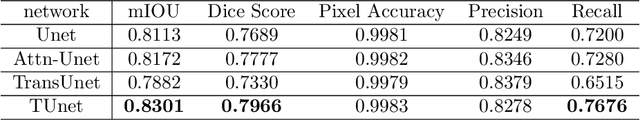
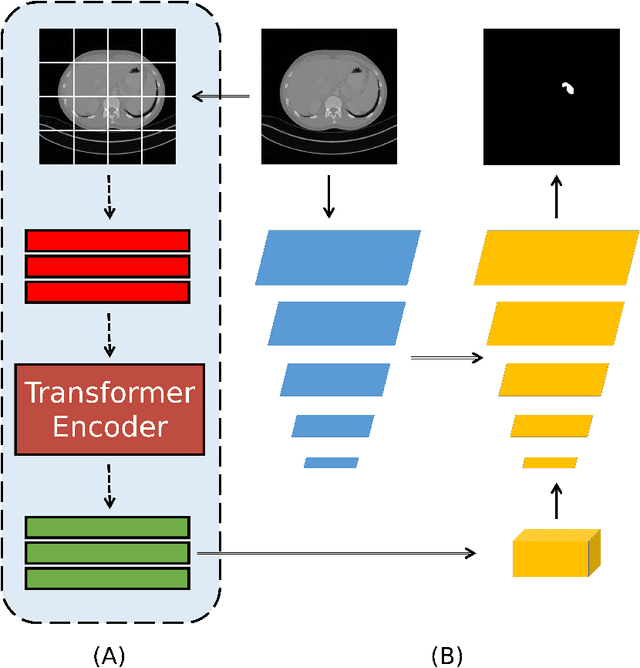
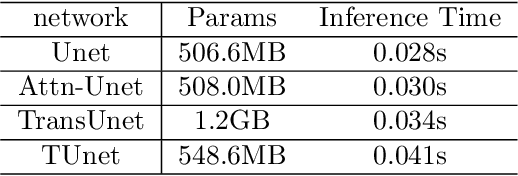
Medical image segmentation have drawn massive attention as it is important in biomedical image analysis. Good segmentation results can assist doctors with their judgement and further improve patients' experience. Among many available pipelines in medical image analysis, Unet is one of the most popular neural networks as it keeps raw features by adding concatenation between encoder and decoder, which makes it still widely used in industrial field. In the mean time, as a popular model which dominates natural language process tasks, transformer is now introduced to computer vision tasks and have seen promising results in object detection, image classification and semantic segmentation tasks. Therefore, the combination of transformer and Unet is supposed to be more efficient than both methods working individually. In this article, we propose Transformer-Unet by adding transformer modules in raw images instead of feature maps in Unet and test our network in CT82 datasets for Pancreas segmentation accordingly. We form an end-to-end network and gain segmentation results better than many previous Unet based algorithms in our experiment. We demonstrate our network and show our experimental results in this paper accordingly.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge