"Image": models, code, and papers
Exploring Resolution and Degradation Clues as Self-supervised Signal for Low Quality Object Detection
Aug 05, 2022
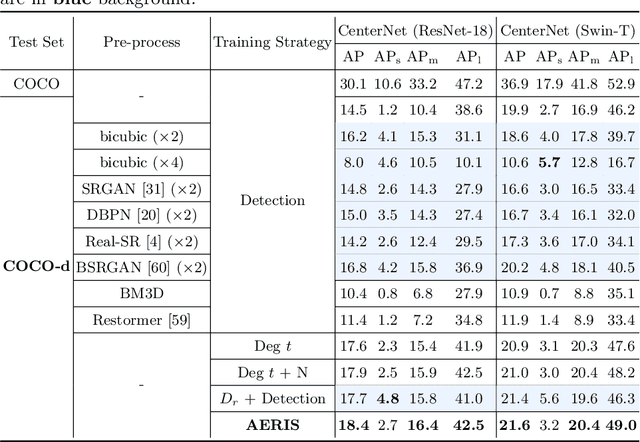
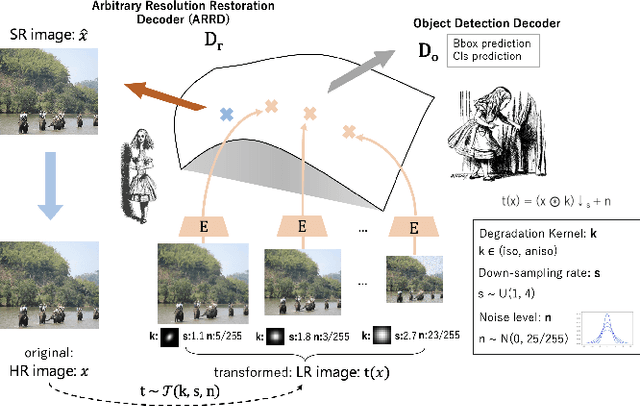
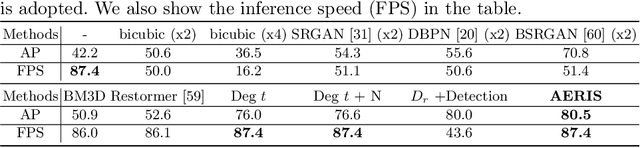
Image restoration algorithms such as super resolution (SR) are indispensable pre-processing modules for object detection in low quality images. Most of these algorithms assume the degradation is fixed and known a priori. However, in practical, either the real degradation or optimal up-sampling ratio rate is unknown or differs from assumption, leading to a deteriorating performance for both the pre-processing module and the consequent high-level task such as object detection. Here, we propose a novel self-supervised framework to detect objects in degraded low resolution images. We utilizes the downsampling degradation as a kind of transformation for self-supervised signals to explore the equivariant representation against various resolutions and other degradation conditions. The Auto Encoding Resolution in Self-supervision (AERIS) framework could further take the advantage of advanced SR architectures with an arbitrary resolution restoring decoder to reconstruct the original correspondence from the degraded input image. Both the representation learning and object detection are optimized jointly in an end-to-end training fashion. The generic AERIS framework could be implemented on various mainstream object detection architectures with different backbones. The extensive experiments show that our methods has achieved superior performance compared with existing methods when facing variant degradation situations. Code would be released at https://github.com/cuiziteng/ECCV_AERIS.
GAF-NAU: Gramian Angular Field encoded Neighborhood Attention U-Net for Pixel-Wise Hyperspectral Image Classification
Apr 21, 2022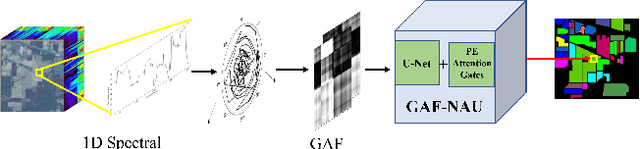
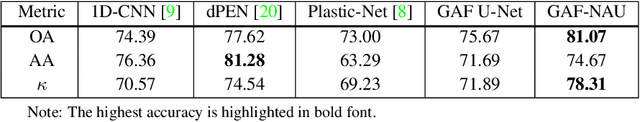
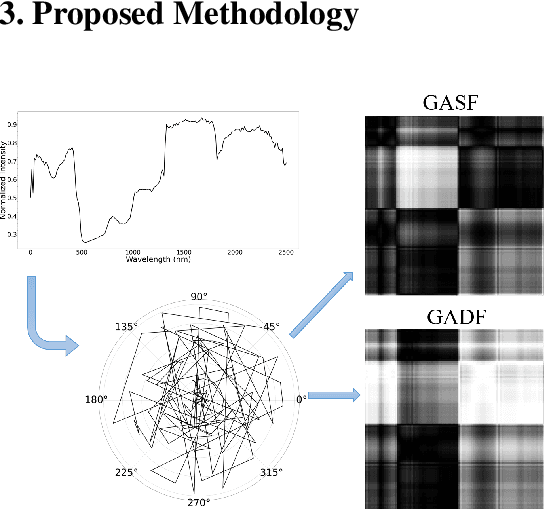
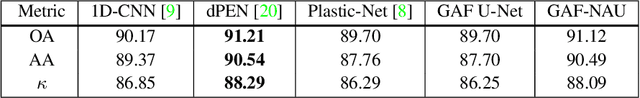
Hyperspectral image (HSI) classification is the most vibrant area of research in the hyperspectral community due to the rich spectral information contained in HSI can greatly aid in identifying objects of interest. However, inherent non-linearity between materials and the corresponding spectral profiles brings two major challenges in HSI classification: interclass similarity and intraclass variability. Many advanced deep learning methods have attempted to address these issues from the perspective of a region/patch-based approach, instead of a pixel-based alternate. However, the patch-based approaches hypothesize that neighborhood pixels of a target pixel in a fixed spatial window belong to the same class. And this assumption is not always true. To address this problem, we herein propose a new deep learning architecture, namely Gramian Angular Field encoded Neighborhood Attention U-Net (GAF-NAU), for pixel-based HSI classification. The proposed method does not require regions or patches centered around a raw target pixel to perform 2D-CNN based classification, instead, our approach transforms 1D pixel vector in HSI into 2D angular feature space using Gramian Angular Field (GAF) and then embed it to a new neighborhood attention network to suppress irrelevant angular feature while emphasizing on pertinent features useful for HSI classification task. Evaluation results on three publicly available HSI datasets demonstrate the superior performance of the proposed model.
A hierarchical semantic segmentation framework for computer vision-based bridge damage detection
Jul 18, 2022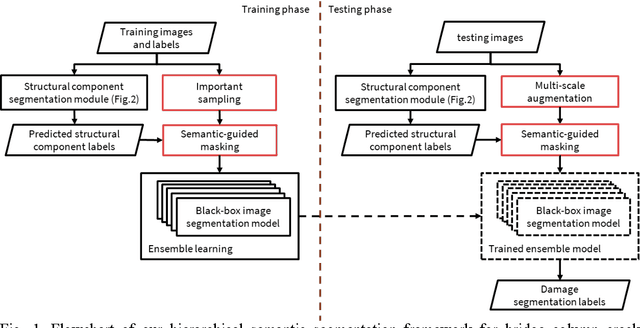
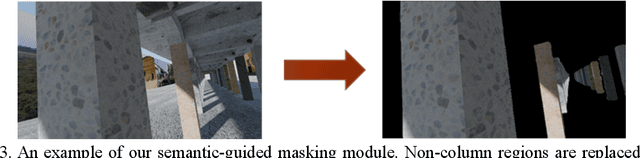
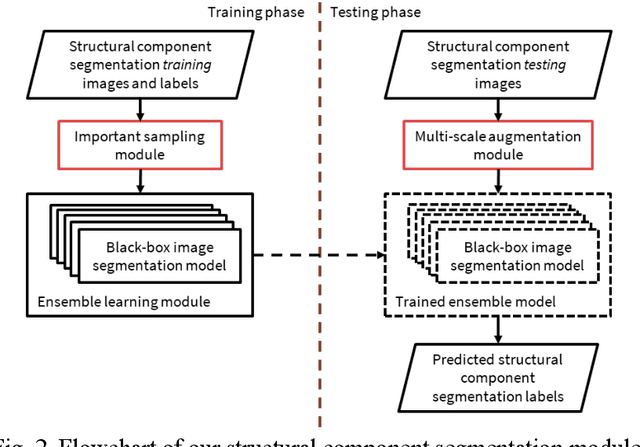

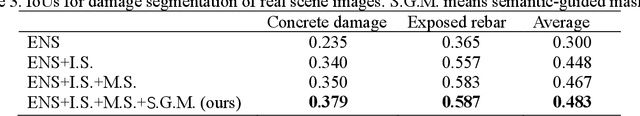
Computer vision-based damage detection using remote cameras and unmanned aerial vehicles (UAVs) enables efficient and low-cost bridge health monitoring that reduces labor costs and the needs for sensor installation and maintenance. By leveraging recent semantic image segmentation approaches, we are able to find regions of critical structural components and recognize damage at the pixel level using images as the only input. However, existing methods perform poorly when detecting small damages (e.g., cracks and exposed rebars) and thin objects with limited image samples, especially when the components of interest are highly imbalanced. To this end, this paper introduces a semantic segmentation framework that imposes the hierarchical semantic relationship between component category and damage types. For example, certain concrete cracks only present on bridge columns and therefore the non-column region will be masked out when detecting such damages. In this way, the damage detection model could focus on learning features from possible damaged regions only and avoid the effects of other irrelevant regions. We also utilize multi-scale augmentation that provides views with different scales that preserves contextual information of each image without losing the ability of handling small and thin objects. Furthermore, the proposed framework employs important sampling that repeatedly samples images containing rare components (e.g., railway sleeper and exposed rebars) to provide more data samples, which addresses the imbalanced data challenge.
Causality-inspired Single-source Domain Generalization for Medical Image Segmentation
Nov 26, 2021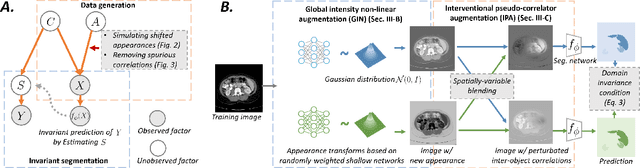
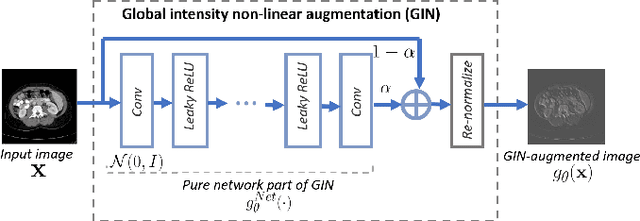
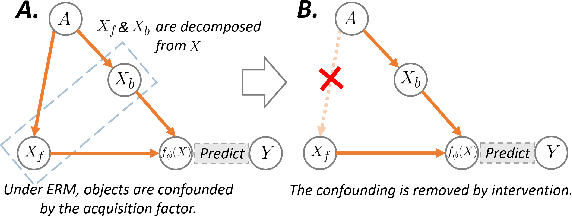
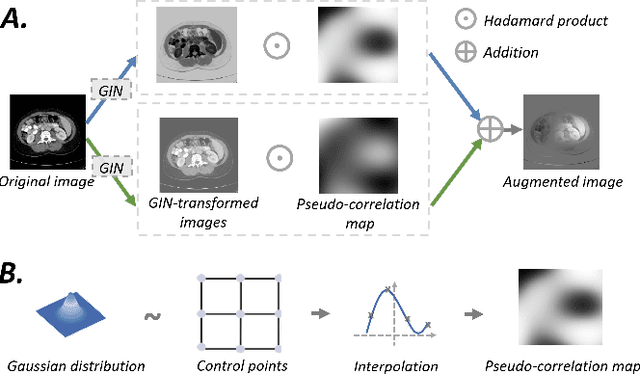
Deep learning models usually suffer from domain shift issues, where models trained on one source domain do not generalize well to other unseen domains. In this work, we investigate the single-source domain generalization problem: training a deep network that is robust to unseen domains, under the condition that training data is only available from one source domain, which is common in medical imaging applications. We tackle this problem in the context of cross-domain medical image segmentation. Under this scenario, domain shifts are mainly caused by different acquisition processes. We propose a simple causality-inspired data augmentation approach to expose a segmentation model to synthesized domain-shifted training examples. Specifically, 1) to make the deep model robust to discrepancies in image intensities and textures, we employ a family of randomly-weighted shallow networks. They augment training images using diverse appearance transformations. 2) Further we show that spurious correlations among objects in an image are detrimental to domain robustness. These correlations might be taken by the network as domain-specific clues for making predictions, and they may break on unseen domains. We remove these spurious correlations via causal intervention. This is achieved by resampling the appearances of potentially correlated objects independently. The proposed approach is validated on three cross-domain segmentation tasks: cross-modality (CT-MRI) abdominal image segmentation, cross-sequence (bSSFP-LGE) cardiac MRI segmentation, and cross-center prostate MRI segmentation. The proposed approach yields consistent performance gains compared with competitive methods when tested on unseen domains.
Learning Incrementally to Segment Multiple Organs in a CT Image
Mar 04, 2022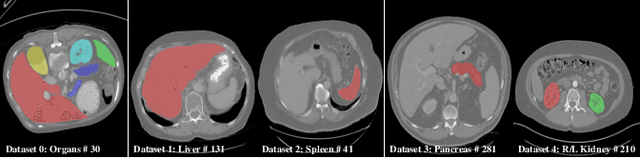
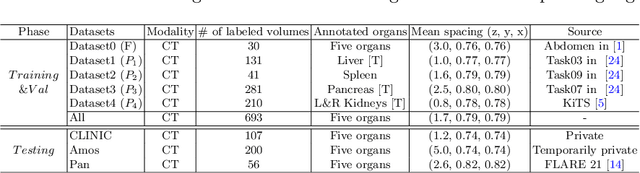
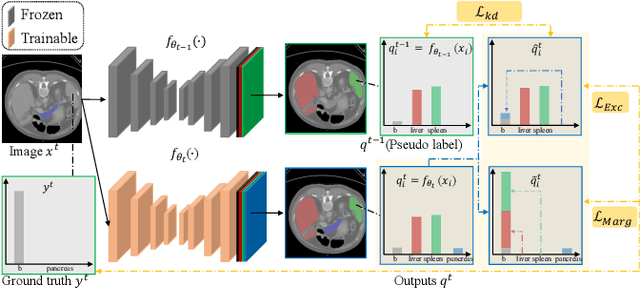
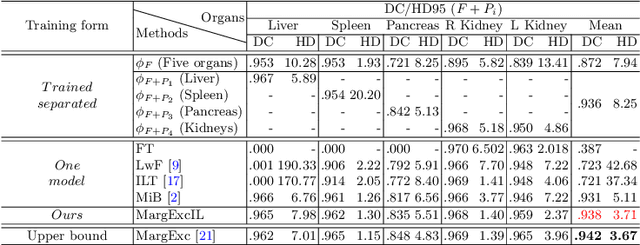
There exists a large number of datasets for organ segmentation, which are partially annotated and sequentially constructed. A typical dataset is constructed at a certain time by curating medical images and annotating the organs of interest. In other words, new datasets with annotations of new organ categories are built over time. To unleash the potential behind these partially labeled, sequentially-constructed datasets, we propose to incrementally learn a multi-organ segmentation model. In each incremental learning (IL) stage, we lose the access to previous data and annotations, whose knowledge is assumingly captured by the current model, and gain the access to a new dataset with annotations of new organ categories, from which we learn to update the organ segmentation model to include the new organs. While IL is notorious for its `catastrophic forgetting' weakness in the context of natural image analysis, we experimentally discover that such a weakness mostly disappears for CT multi-organ segmentation. To further stabilize the model performance across the IL stages, we introduce a light memory module and some loss functions to restrain the representation of different categories in feature space, aggregating feature representation of the same class and separating feature representation of different classes. Extensive experiments on five open-sourced datasets are conducted to illustrate the effectiveness of our method.
Spectral image clustering on dual-energy CT scans using functional regression mixtures
Jan 31, 2022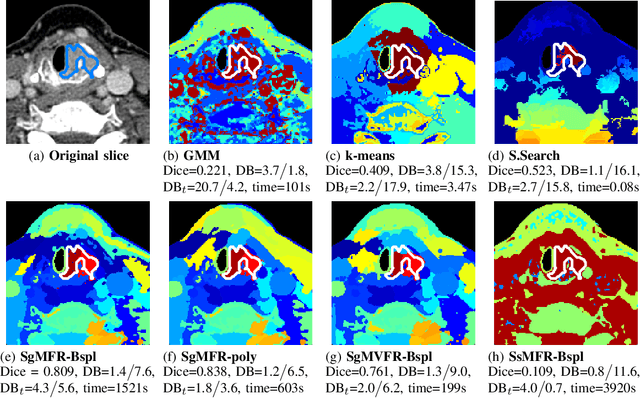
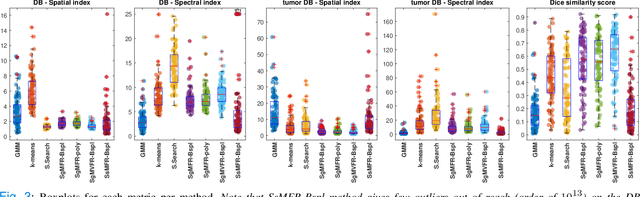
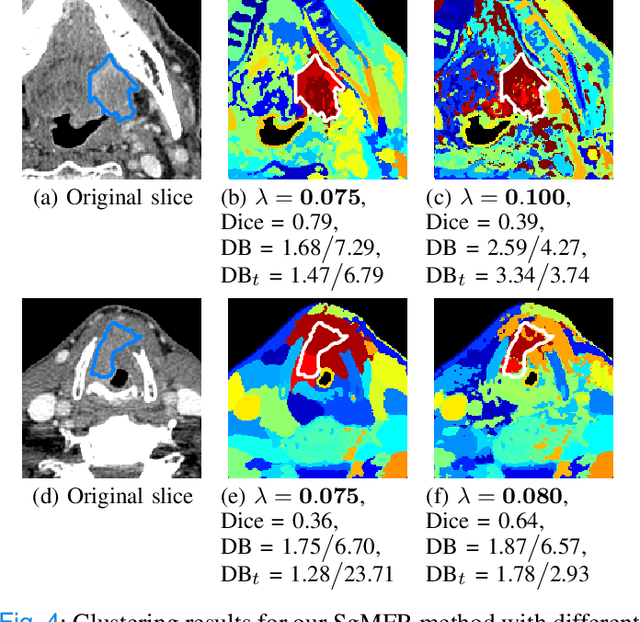
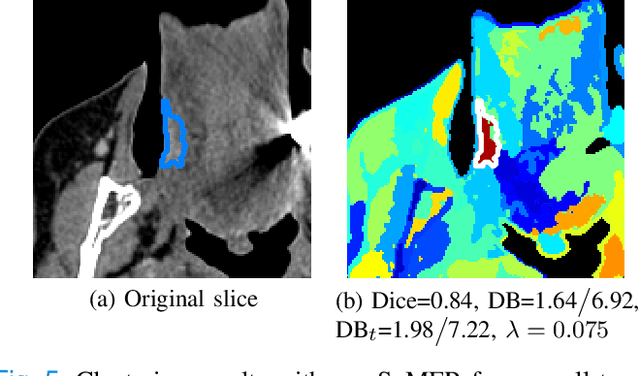
Dual-energy computed tomography (DECT) is an advanced CT scanning technique enabling material characterization not possible with conventional CT scans. It allows the reconstruction of energy decay curves at each 3D image voxel, representing varying image attenuation at different effective scanning energy levels. In this paper, we develop novel functional data analysis (FDA) techniques and adapt them to the analysis of DECT decay curves. More specifically, we construct functional mixture models that integrate spatial context in mixture weights, with mixture component densities being constructed upon the energy decay curves as functional observations. We design unsupervised clustering algorithms by developing dedicated expectation maximization (EM) algorithms for the maximum likelihood estimation of the model parameters. To our knowledge, this is the first article to adapt statistical FDA tools and model-based clustering to take advantage of the full spectral information provided by DECT. We evaluate our methods on 91 head and neck cancer DECT scans. We compare our unsupervised clustering results to tumor contours traced manually by radiologists, as well as to several baseline algorithms. Given the inter-rater variability even among experts at delineating head and neck tumors, and given the potential importance of tissue reactions surrounding the tumor itself, our proposed methodology has the potential to add value in downstream machine learning applications for clinical outcome prediction based on DECT data in head and neck cancer.
Masked Sinogram Model with Transformer for ill-Posed Computed Tomography Reconstruction: a Preliminary Study
Sep 03, 2022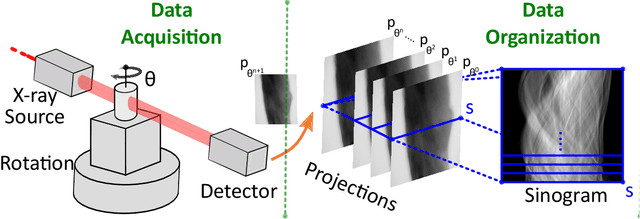
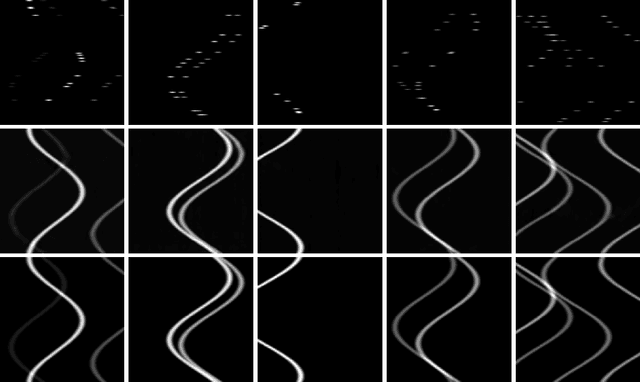
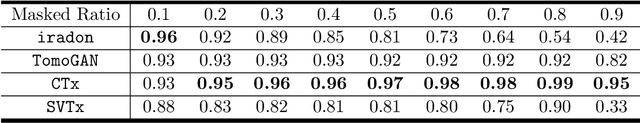
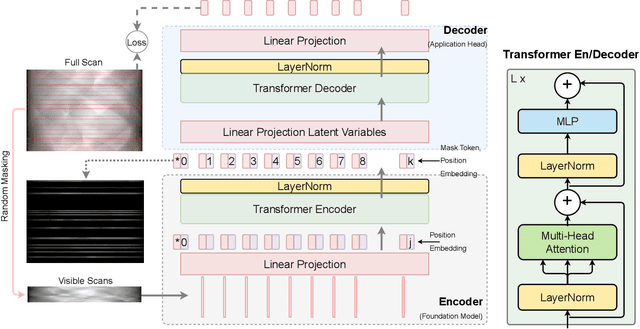

Computed Tomography (CT) is an imaging technique where information about an object are collected at different angles (called projections or scans). Then the cross-sectional image showing the internal structure of the slice is produced by solving an inverse problem. Limited by certain factors such as radiation dosage, projection angles, the produced images can be noisy or contain artifacts. Inspired by the success of transformer for natural language processing, the core idea of this preliminary study is to consider a projection of tomography as a word token, and the whole scan of the cross-section (A.K.A. sinogram) as a sentence in the context of natural language processing. Then we explore the idea of foundation model by training a masked sinogram model (MSM) and fine-tune MSM for various downstream applications including CT reconstruction under data collections restriction (e.g., photon-budget) and a data-driven solution to approximate solutions of the inverse problem for CT reconstruction. Models and data used in this study are available at https://github.com/lzhengchun/TomoTx.
Learning 6D Pose Estimation from Synthetic RGBD Images for Robotic Applications
Aug 30, 2022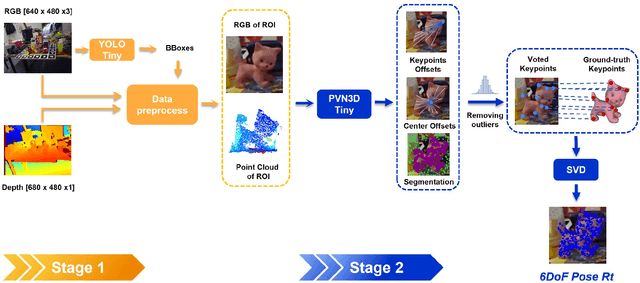
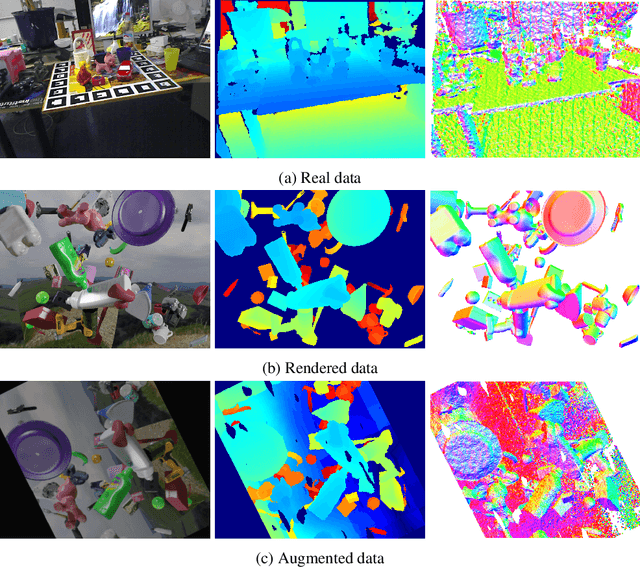
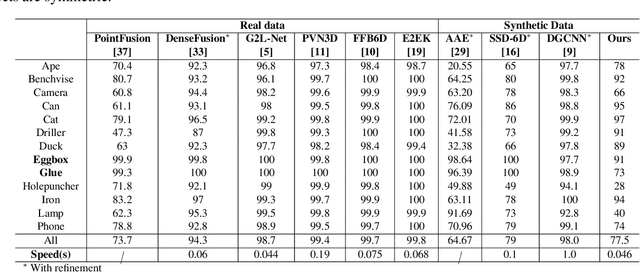

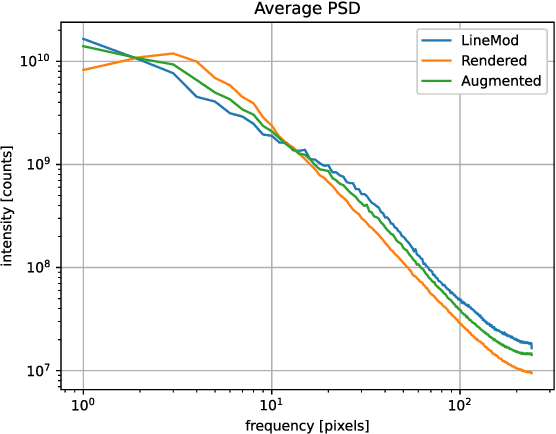
In this work, we propose a data generation pipeline by leveraging the 3D suite Blender to produce synthetic RGBD image datasets with 6D poses for robotic picking. The proposed pipeline can efficiently generate large amounts of photo-realistic RGBD images for the object of interest. In addition, a collection of domain randomization techniques is introduced to bridge the gap between real and synthetic data. Furthermore, we develop a real-time two-stage 6D pose estimation approach by integrating the object detector YOLO-V4-tiny and the 6D pose estimation algorithm PVN3D for time sensitive robotics applications. With the proposed data generation pipeline, our pose estimation approach can be trained from scratch using only synthetic data without any pre-trained models. The resulting network shows competitive performance compared to state-of-the-art methods when evaluated on LineMod dataset. We also demonstrate the proposed approach in a robotic experiment, grasping a household object from cluttered background under different lighting conditions.
ContraReg: Contrastive Learning of Multi-modality Unsupervised Deformable Image Registration
Jun 27, 2022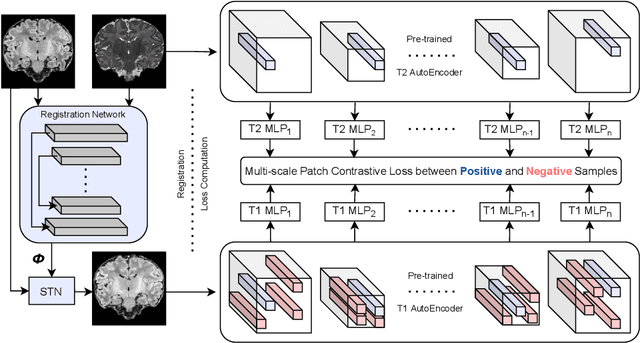
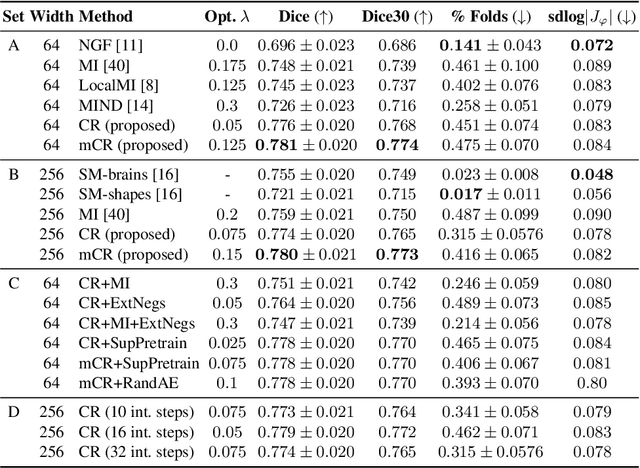
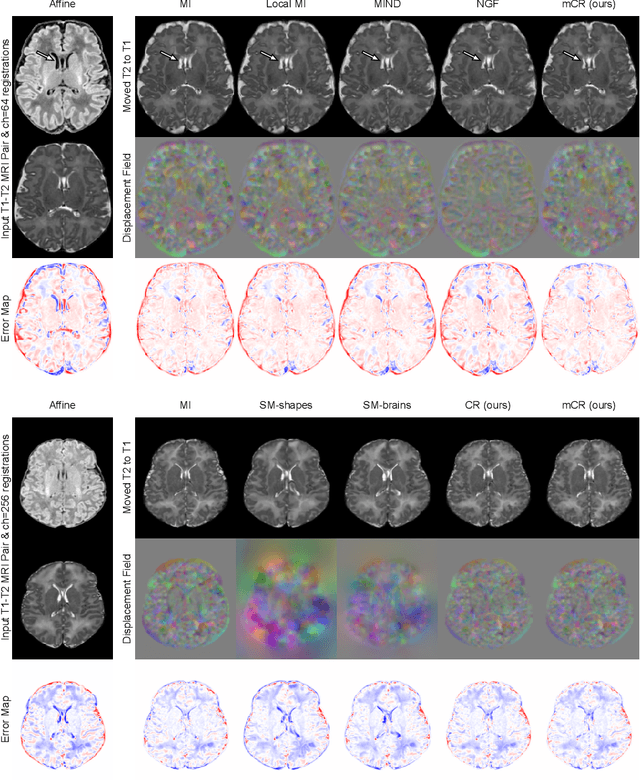
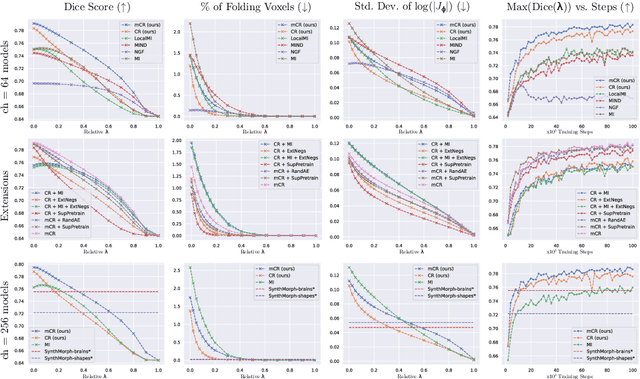
Establishing voxelwise semantic correspondence across distinct imaging modalities is a foundational yet formidable computer vision task. Current multi-modality registration techniques maximize hand-crafted inter-domain similarity functions, are limited in modeling nonlinear intensity-relationships and deformations, and may require significant re-engineering or underperform on new tasks, datasets, and domain pairs. This work presents ContraReg, an unsupervised contrastive representation learning approach to multi-modality deformable registration. By projecting learned multi-scale local patch features onto a jointly learned inter-domain embedding space, ContraReg obtains representations useful for non-rigid multi-modality alignment. Experimentally, ContraReg achieves accurate and robust results with smooth and invertible deformations across a series of baselines and ablations on a neonatal T1-T2 brain MRI registration task with all methods validated over a wide range of deformation regularization strengths.
Rarity Score : A New Metric to Evaluate the Uncommonness of Synthesized Images
Jun 26, 2022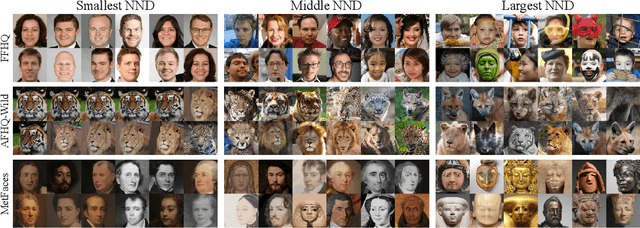
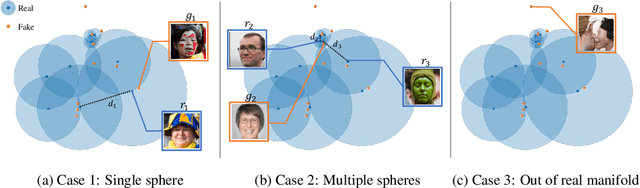
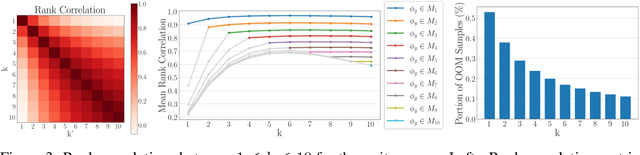

Evaluation metrics in image synthesis play a key role to measure performances of generative models. However, most metrics mainly focus on image fidelity. Existing diversity metrics are derived by comparing distributions, and thus they cannot quantify the diversity or rarity degree of each generated image. In this work, we propose a new evaluation metric, called `rarity score', to measure the individual rarity of each image synthesized by generative models. We first show empirical observation that common samples are close to each other and rare samples are far from each other in nearest-neighbor distances of feature space. We then use our metric to demonstrate that the extent to which different generative models produce rare images can be effectively compared. We also propose a method to compare rarities between datasets that share the same concept such as CelebA-HQ and FFHQ. Finally, we analyze the use of metrics in different designs of feature spaces to better understand the relationship between feature spaces and resulting sparse images. Code will be publicly available online for the research community.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge