"Image": models, code, and papers
FedMed-ATL: Misaligned Unpaired Brain Image Synthesis via Affine Transform Loss
Jan 29, 2022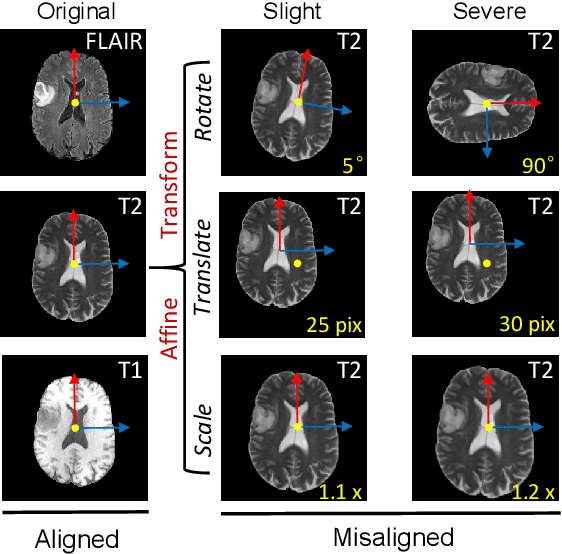
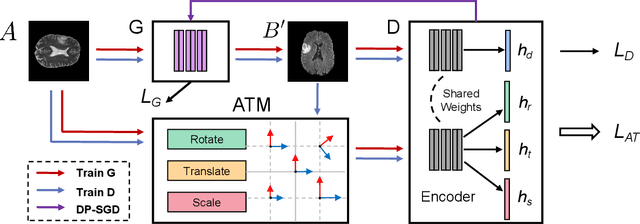
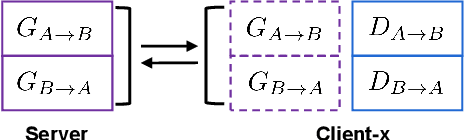
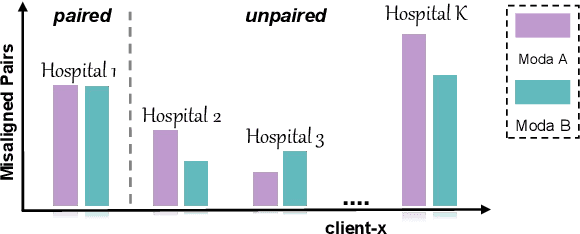
The existence of completely aligned and paired multi-modal neuroimaging data has proved its effectiveness in the diagnosis of brain diseases. However, collecting the full set of well-aligned and paired data is impractical or even luxurious, since the practical difficulties may include high cost, long time acquisition, image corruption, and privacy issues. Previously, the misaligned unpaired neuroimaging data (termed as MUD) are generally treated as noisy label. However, such a noisy label-based method could not work very well when misaligned data occurs distortions severely, for example, different angles of rotation. In this paper, we propose a novel federated self-supervised learning (FedMed) for brain image synthesis. An affine transform loss (ATL) was formulated to make use of severely distorted images without violating privacy legislation for the hospital. We then introduce a new data augmentation procedure for self-supervised training and fed it into three auxiliary heads, namely auxiliary rotation, auxiliary translation, and auxiliary scaling heads. The proposed method demonstrates advanced performance in both the quality of synthesized results under a severely misaligned and unpaired data setting, and better stability than other GAN-based algorithms. The proposed method also reduces the demand for deformable registration while encouraging to realize the usage of those misaligned and unpaired data. Experimental results verify the outstanding ability of our learning paradigm compared to other state-of-the-art approaches. Our code is available on the website: https://github.com/FedMed-Meta/FedMed-ATL
Image quality assessment by overlapping task-specific and task-agnostic measures: application to prostate multiparametric MR images for cancer segmentation
Feb 20, 2022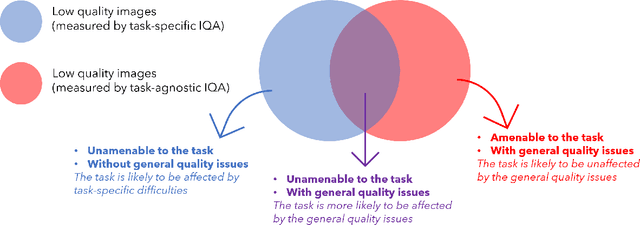
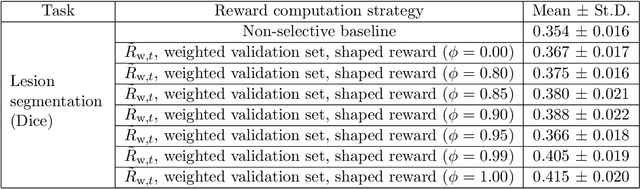

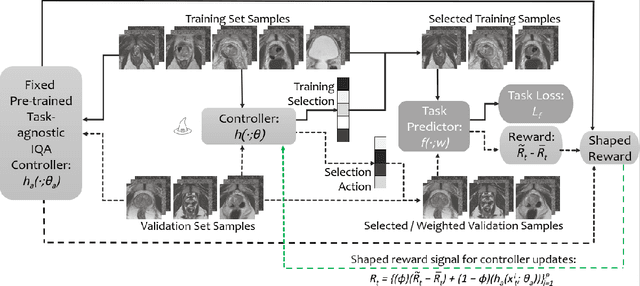
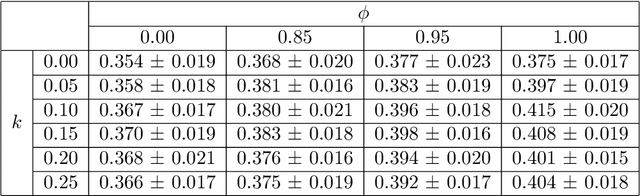
Image quality assessment (IQA) in medical imaging can be used to ensure that downstream clinical tasks can be reliably performed. Quantifying the impact of an image on the specific target tasks, also named as task amenability, is needed. A task-specific IQA has recently been proposed to learn an image-amenability-predicting controller simultaneously with a target task predictor. This allows for the trained IQA controller to measure the impact an image has on the target task performance, when this task is performed using the predictor, e.g. segmentation and classification neural networks in modern clinical applications. In this work, we propose an extension to this task-specific IQA approach, by adding a task-agnostic IQA based on auto-encoding as the target task. Analysing the intersection between low-quality images, deemed by both the task-specific and task-agnostic IQA, may help to differentiate the underpinning factors that caused the poor target task performance. For example, common imaging artefacts may not adversely affect the target task, which would lead to a low task-agnostic quality and a high task-specific quality, whilst individual cases considered clinically challenging, which can not be improved by better imaging equipment or protocols, is likely to result in a high task-agnostic quality but a low task-specific quality. We first describe a flexible reward shaping strategy which allows for the adjustment of weighting between task-agnostic and task-specific quality scoring. Furthermore, we evaluate the proposed algorithm using a clinically challenging target task of prostate tumour segmentation on multiparametric magnetic resonance (mpMR) images, from 850 patients. The proposed reward shaping strategy, with appropriately weighted task-specific and task-agnostic qualities, successfully identified samples that need re-acquisition due to defected imaging process.
Single Image Dehazing with An Independent Detail-Recovery Network
Sep 22, 2021
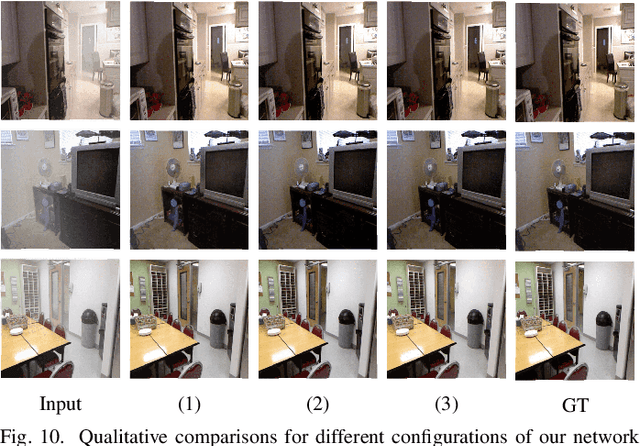

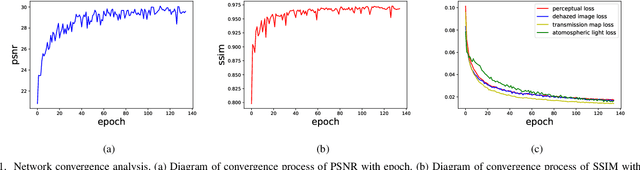
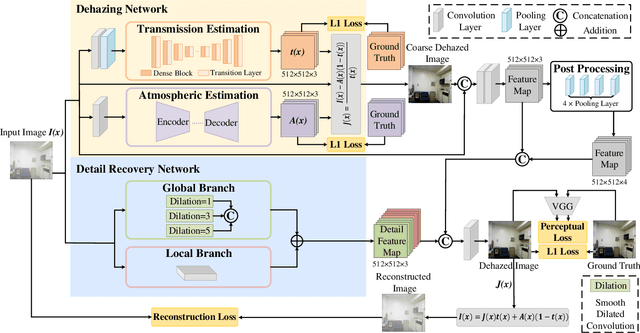
Single image dehazing is a prerequisite which affects the performance of many computer vision tasks and has attracted increasing attention in recent years. However, most existing dehazing methods emphasize more on haze removal but less on the detail recovery of the dehazed images. In this paper, we propose a single image dehazing method with an independent Detail Recovery Network (DRN), which considers capturing the details from the input image over a separate network and then integrates them into a coarse dehazed image. The overall network consists of two independent networks, named DRN and the dehazing network respectively. Specifically, the DRN aims to recover the dehazed image details through local and global branches respectively. The local branch can obtain local detail information through the convolution layer and the global branch can capture more global information by the Smooth Dilated Convolution (SDC). The detail feature map is fused into the coarse dehazed image to obtain the dehazed image with rich image details. Besides, we integrate the DRN, the physical-model-based dehazing network and the reconstruction loss into an end-to-end joint learning framework. Extensive experiments on the public image dehazing datasets (RESIDE-Indoor, RESIDE-Outdoor and the TrainA-TestA) illustrate the effectiveness of the modules in the proposed method and show that our method outperforms the state-of-the-art dehazing methods both quantitatively and qualitatively. The code is released in https://github.com/YanLi-LY/Dehazing-DRN.
DocEnTr: An End-to-End Document Image Enhancement Transformer
Jan 25, 2022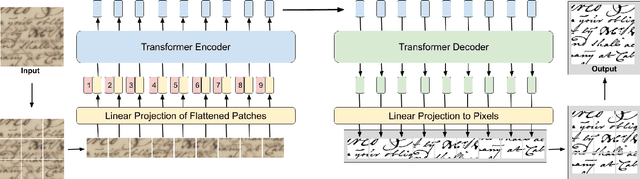
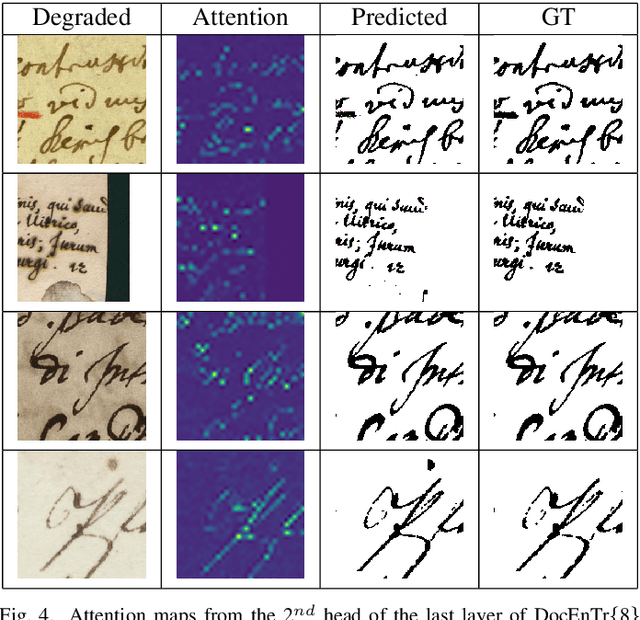



Document images can be affected by many degradation scenarios, which cause recognition and processing difficulties. In this age of digitization, it is important to denoise them for proper usage. To address this challenge, we present a new encoder-decoder architecture based on vision transformers to enhance both machine-printed and handwritten document images, in an end-to-end fashion. The encoder operates directly on the pixel patches with their positional information without the use of any convolutional layers, while the decoder reconstructs a clean image from the encoded patches. Conducted experiments show a superiority of the proposed model compared to the state-of the-art methods on several DIBCO benchmarks. Code and models will be publicly available at: \url{https://github.com/dali92002/DocEnTR}.
Towards Sparsification of Graph Neural Networks
Sep 11, 2022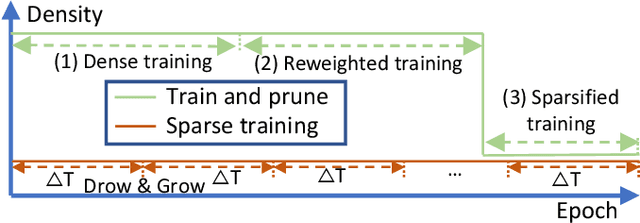
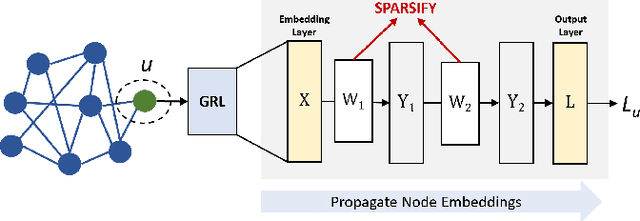
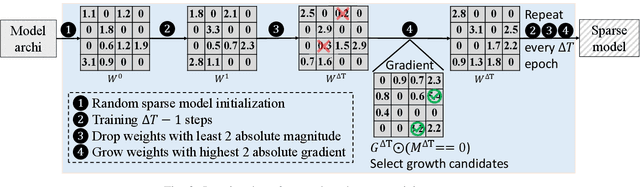
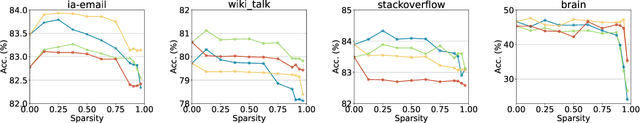
As real-world graphs expand in size, larger GNN models with billions of parameters are deployed. High parameter count in such models makes training and inference on graphs expensive and challenging. To reduce the computational and memory costs of GNNs, optimization methods such as pruning the redundant nodes and edges in input graphs have been commonly adopted. However, model compression, which directly targets the sparsification of model layers, has been mostly limited to traditional Deep Neural Networks (DNNs) used for tasks such as image classification and object detection. In this paper, we utilize two state-of-the-art model compression methods (1) train and prune and (2) sparse training for the sparsification of weight layers in GNNs. We evaluate and compare the efficiency of both methods in terms of accuracy, training sparsity, and training FLOPs on real-world graphs. Our experimental results show that on the ia-email, wiki-talk, and stackoverflow datasets for link prediction, sparse training with much lower training FLOPs achieves a comparable accuracy with the train and prune method. On the brain dataset for node classification, sparse training uses a lower number FLOPs (less than 1/7 FLOPs of train and prune method) and preserves a much better accuracy performance under extreme model sparsity.
Revisiting RCAN: Improved Training for Image Super-Resolution
Jan 27, 2022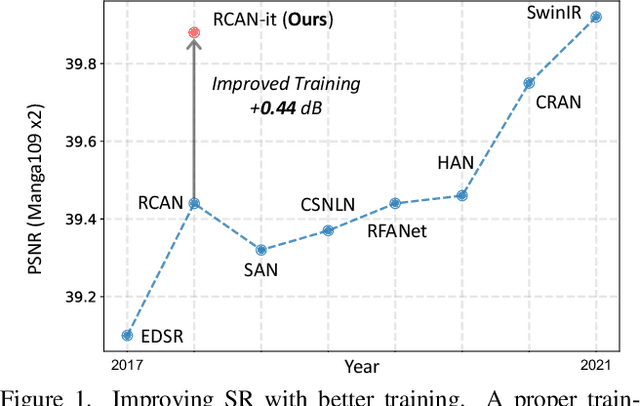
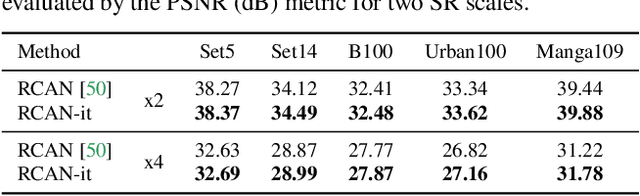
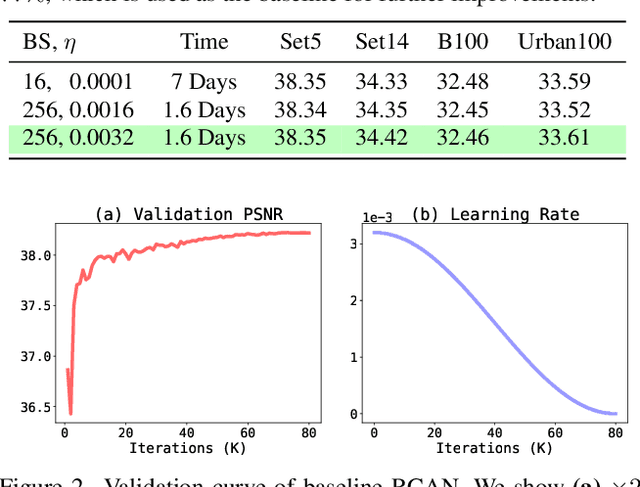
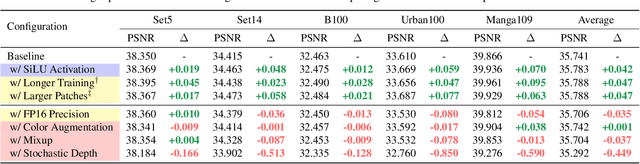
Image super-resolution (SR) is a fast-moving field with novel architectures attracting the spotlight. However, most SR models were optimized with dated training strategies. In this work, we revisit the popular RCAN model and examine the effect of different training options in SR. Surprisingly (or perhaps as expected), we show that RCAN can outperform or match nearly all the CNN-based SR architectures published after RCAN on standard benchmarks with a proper training strategy and minimal architecture change. Besides, although RCAN is a very large SR architecture with more than four hundred convolutional layers, we draw a notable conclusion that underfitting is still the main problem restricting the model capability instead of overfitting. We observe supportive evidence that increasing training iterations clearly improves the model performance while applying regularization techniques generally degrades the predictions. We denote our simply revised RCAN as RCAN-it and recommend practitioners to use it as baselines for future research. Code is publicly available at https://github.com/zudi-lin/rcan-it.
Instance-Aware Observer Network for Out-of-Distribution Object Segmentation
Jul 18, 2022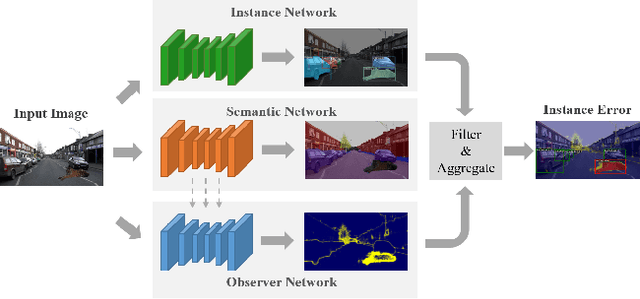
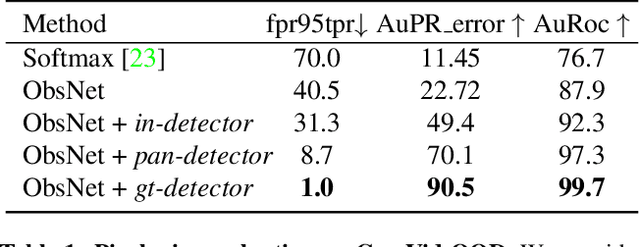

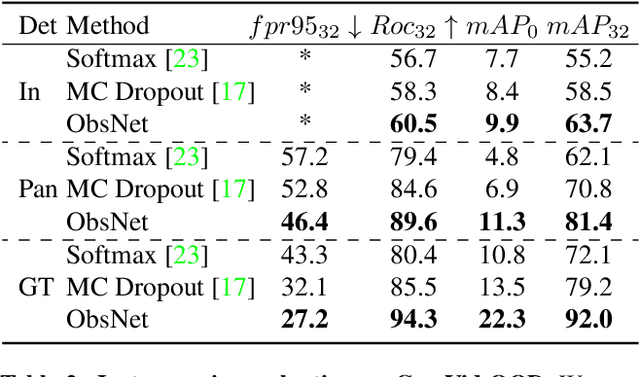
Recent work on Observer Network has shown promising results on Out-Of-Distribution (OOD) detection for semantic segmentation. These methods have difficulty in precisely locating the point of interest in the image, i.e, the anomaly. This limitation is due to the difficulty of fine-grained prediction at the pixel level. To address this issue, we provide instance knowledge to the observer. We extend the approach of ObsNet by harnessing an instance-wise mask prediction. We use an additional, class agnostic, object detector to filter and aggregate observer predictions. Finally, we predict an unique anomaly score for each instance in the image. We show that our proposed method accurately disentangle in-distribution objects from Out-Of-Distribution objects on three datasets.
Interactive Medical Image Segmentation with Self-Adaptive Confidence Calibration
Nov 15, 2021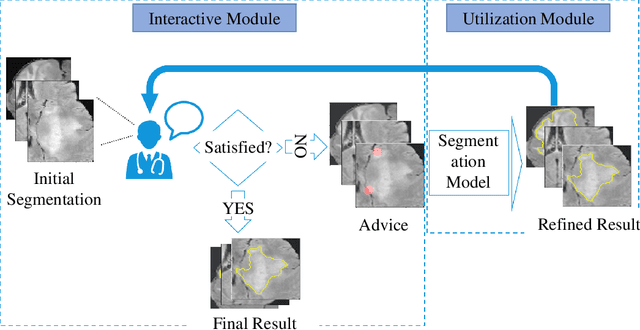
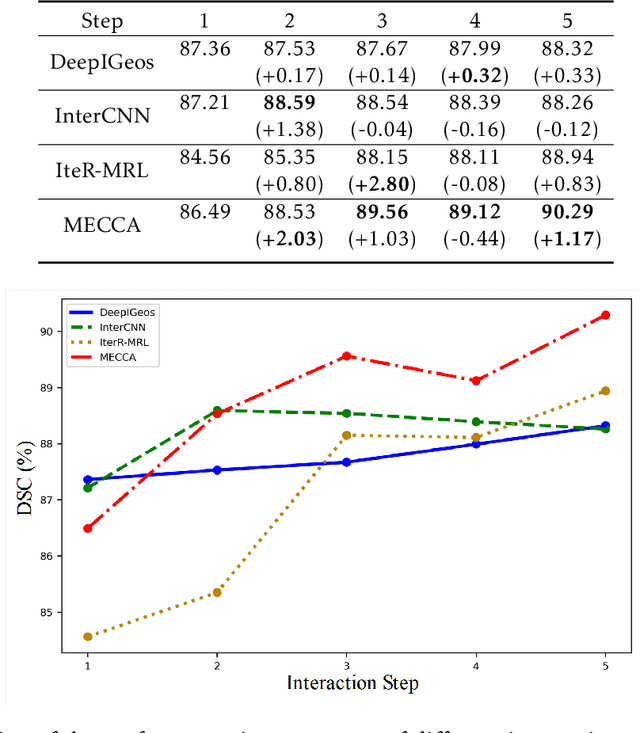
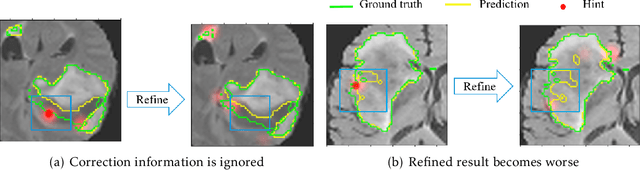
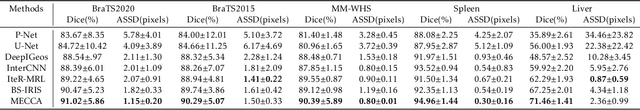
Medical image segmentation is one of the fundamental problems for artificial intelligence-based clinical decision systems. Current automatic medical image segmentation methods are often failed to meet clinical requirements. As such, a series of interactive segmentation algorithms are proposed to utilize expert correction information. However, existing methods suffer from some segmentation refining failure problems after long-term interactions and some cost problems from expert annotation, which hinder clinical applications. This paper proposes an interactive segmentation framework, called interactive MEdical segmentation with self-adaptive Confidence CAlibration (MECCA), by introducing the corrective action evaluation, which combines the action-based confidence learning and multi-agent reinforcement learning (MARL). The evaluation is established through a novel action-based confidence network, and the corrective actions are obtained from MARL. Based on the confidential information, a self-adaptive reward function is designed to provide more detailed feedback, and a simulated label generation mechanism is proposed on unsupervised data to reduce over-reliance on labeled data. Experimental results on various medical image datasets have shown the significant performance of the proposed algorithm.
Semi-Automatic Labeling and Semantic Segmentation of Gram-Stained Microscopic Images from DIBaS Dataset
Aug 23, 2022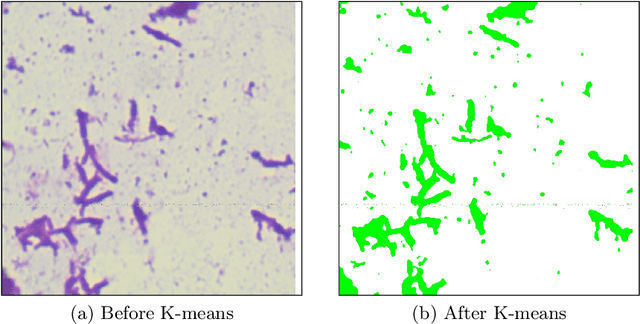
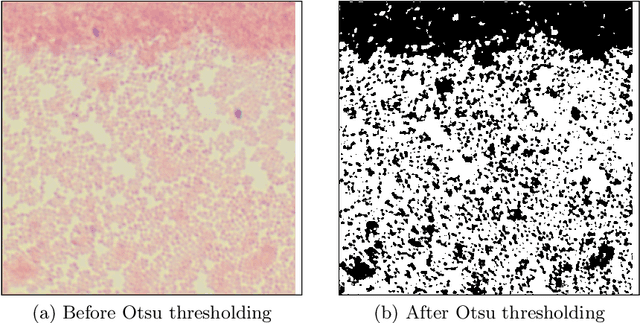
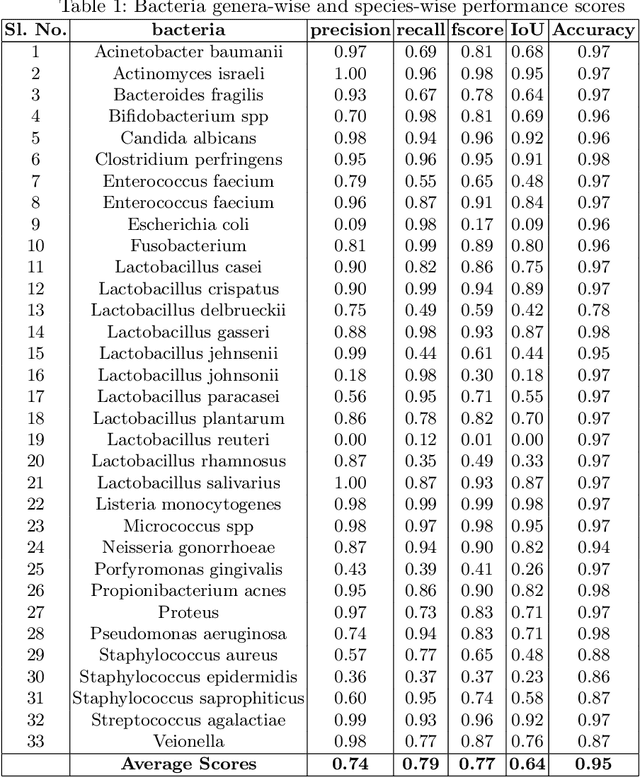

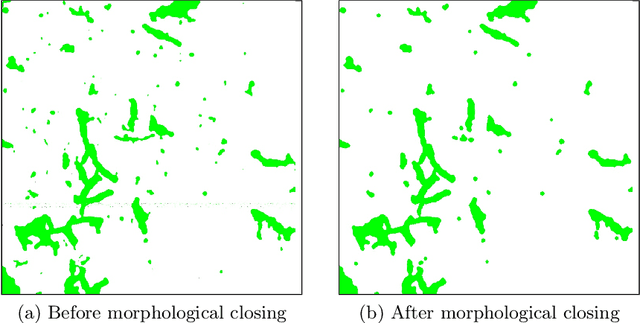
In this paper, a semi-automatic annotation of bacteria genera and species from DIBaS dataset is implemented using clustering and thresholding algorithms. A Deep learning model is trained to achieve the semantic segmentation and classification of the bacteria species. Classification accuracy of 95% is achieved. Deep learning models find tremendous applications in biomedical image processing. Automatic segmentation of bacteria from gram-stained microscopic images is essential to diagnose respiratory and urinary tract infections, detect cancers, etc. Deep learning will aid the biologists to get reliable results in less time. Additionally, a lot of human intervention can be reduced. This work can be helpful to detect bacteria from urinary smear images, sputum smear images, etc to diagnose urinary tract infections, tuberculosis, pneumonia, etc.
A constrained Shannon-Fano entropy coder for image storage in synthetic DNA
Mar 18, 2022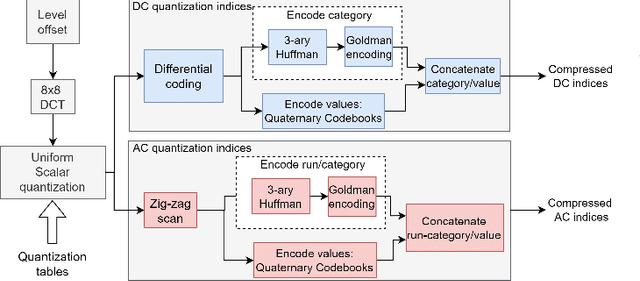
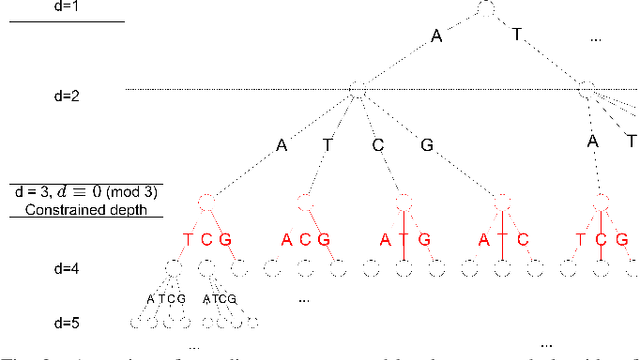
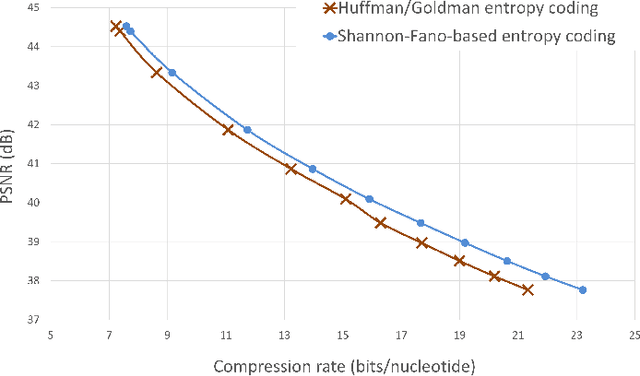
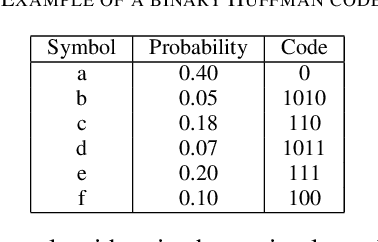
The exponentially increasing demand for data storage has been facing more and more challenges during the past years. The energy costs that it represents are also increasing, and the availability of the storage hardware is not able to follow the storage demand's trend. The short lifespan of conventional storage media -- 10 to 20 years - forces the duplication of the hardware and worsens the situation. The majority of this storage demand concerns "cold" data, data very rarely accessed but that has to be kept for long periods of time. The coding abilities of synthetic DNA, and its long durability (several hundred years), make it a serious candidate as an alternative storage media for "cold" data. In this paper, we propose a variable-length coding algorithm adapted to DNA data storage with improved performance. The proposed algorithm is based on a modified Shannon-Fano code that respects some biochemichal constraints imposed by the synthesis chemistry. We have inserted this code in a JPEG compression algorithm adapted to DNA image storage and we highlighted an improvement of the compression ratio ranging from 0.5 up to 2 bits per nucleotide compared to the state-of-the-art solution, without affecting the reconstruction quality.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge