"Image": models, code, and papers
A Novel End-To-End Network for Reconstruction of Non-Regularly Sampled Image Data Using Locally Fully Connected Layers
Mar 17, 2022
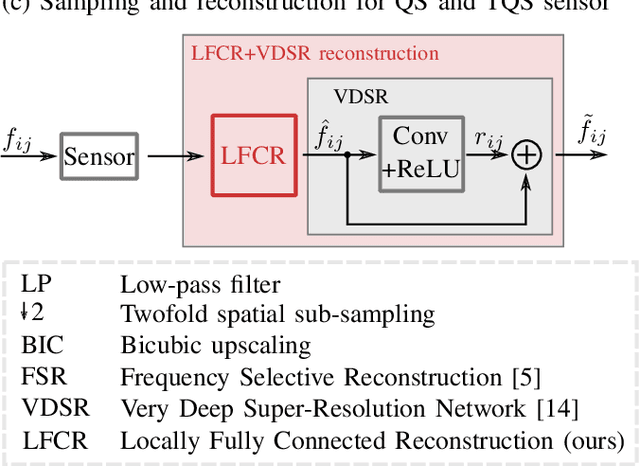
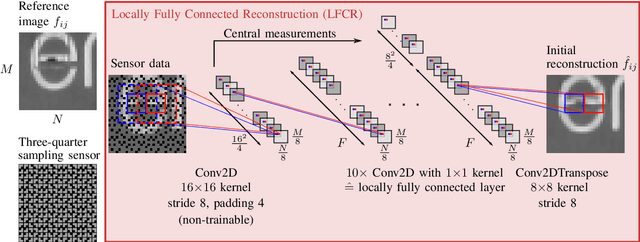
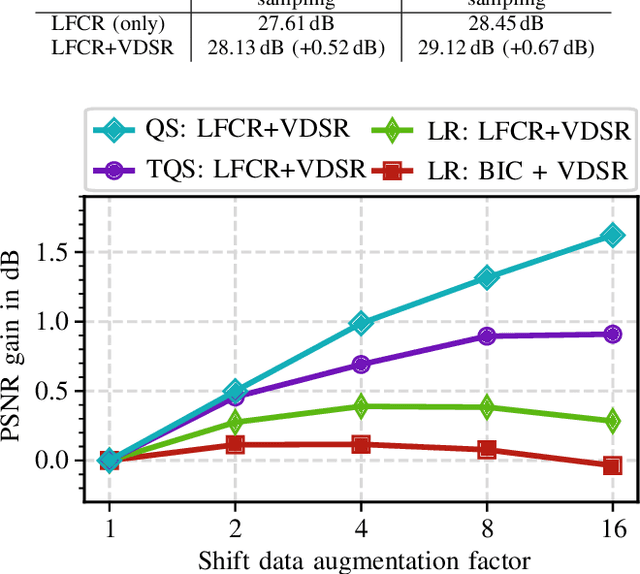
Quarter sampling and three-quarter sampling are novel sensor concepts that enable the acquisition of higher resolution images without increasing the number of pixels. This is achieved by non-regularly covering parts of each pixel of a low-resolution sensor such that only one quadrant or three quadrants of the sensor area of each pixel is sensitive to light. Combining a properly designed mask and a high-quality reconstruction algorithm, a higher image quality can be achieved than using a low-resolution sensor and subsequent upsampling. For the latter case, the image quality can be further enhanced using super resolution algorithms such as the very deep super resolution network (VDSR). In this paper, we propose a novel end-to-end neural network to reconstruct high resolution images from non-regularly sampled sensor data. The network is a concatenation of a locally fully connected reconstruction network (LFCR) and a standard VDSR network. Altogether, using a three-quarter sampling sensor with our novel neural network layout, the image quality in terms of PSNR for the Urban100 dataset can be increased by 2.96 dB compared to the state-of-the-art approach. Compared to a low-resolution sensor with VDSR, a gain of 1.11 dB is achieved.
Jigsaw-ViT: Learning Jigsaw Puzzles in Vision Transformer
Jul 25, 2022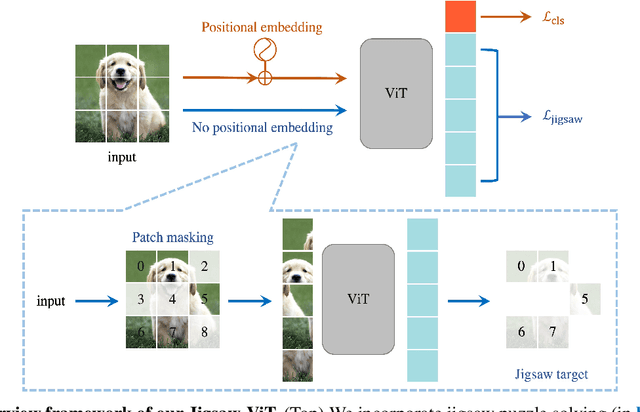
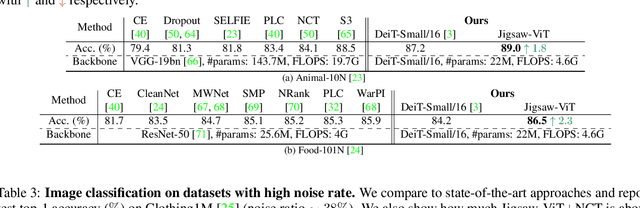
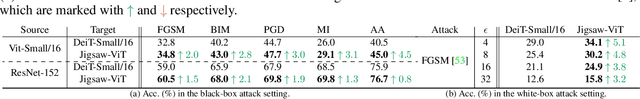


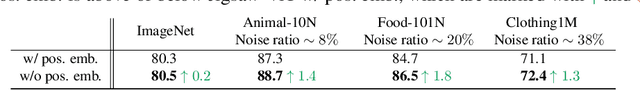
The success of Vision Transformer (ViT) in various computer vision tasks has promoted the ever-increasing prevalence of this convolution-free network. The fact that ViT works on image patches makes it potentially relevant to the problem of jigsaw puzzle solving, which is a classical self-supervised task aiming at reordering shuffled sequential image patches back to their natural form. Despite its simplicity, solving jigsaw puzzle has been demonstrated to be helpful for diverse tasks using Convolutional Neural Networks (CNNs), such as self-supervised feature representation learning, domain generalization, and fine-grained classification. In this paper, we explore solving jigsaw puzzle as a self-supervised auxiliary loss in ViT for image classification, named Jigsaw-ViT. We show two modifications that can make Jigsaw-ViT superior to standard ViT: discarding positional embeddings and masking patches randomly. Yet simple, we find that Jigsaw-ViT is able to improve both in generalization and robustness over the standard ViT, which is usually rather a trade-off. Experimentally, we show that adding the jigsaw puzzle branch provides better generalization than ViT on large-scale image classification on ImageNet. Moreover, the auxiliary task also improves robustness to noisy labels on Animal-10N, Food-101N, and Clothing1M as well as adversarial examples. Our implementation is available at https://yingyichen-cyy.github.io/Jigsaw-ViT/.
Improved Abdominal Multi-Organ Segmentation via 3D Boundary-Constrained Deep Neural Networks
Oct 09, 2022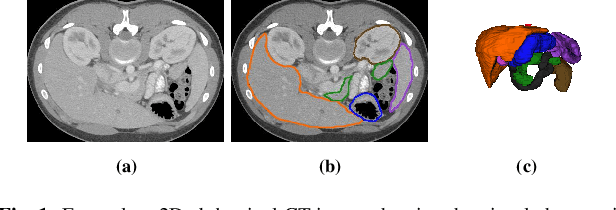
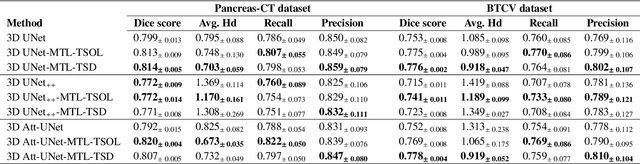
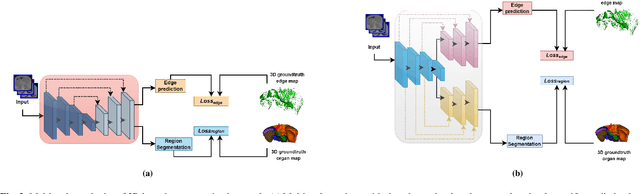
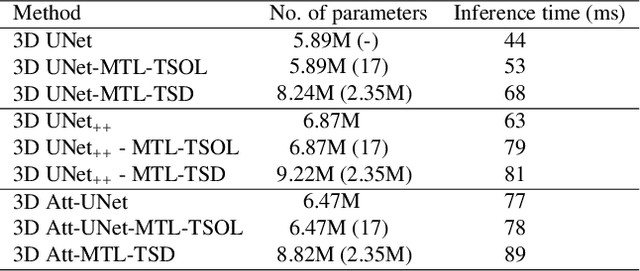
Quantitative assessment of the abdominal region from clinically acquired CT scans requires the simultaneous segmentation of abdominal organs. Thanks to the availability of high-performance computational resources, deep learning-based methods have resulted in state-of-the-art performance for the segmentation of 3D abdominal CT scans. However, the complex characterization of organs with fuzzy boundaries prevents the deep learning methods from accurately segmenting these anatomical organs. Specifically, the voxels on the boundary of organs are more vulnerable to misprediction due to the highly-varying intensity of inter-organ boundaries. This paper investigates the possibility of improving the abdominal image segmentation performance of the existing 3D encoder-decoder networks by leveraging organ-boundary prediction as a complementary task. To address the problem of abdominal multi-organ segmentation, we train the 3D encoder-decoder network to simultaneously segment the abdominal organs and their corresponding boundaries in CT scans via multi-task learning. The network is trained end-to-end using a loss function that combines two task-specific losses, i.e., complete organ segmentation loss and boundary prediction loss. We explore two different network topologies based on the extent of weights shared between the two tasks within a unified multi-task framework. To evaluate the utilization of complementary boundary prediction task in improving the abdominal multi-organ segmentation, we use three state-of-the-art encoder-decoder networks: 3D UNet, 3D UNet++, and 3D Attention-UNet. The effectiveness of utilizing the organs' boundary information for abdominal multi-organ segmentation is evaluated on two publically available abdominal CT datasets. A maximum relative improvement of 3.5% and 3.6% is observed in Mean Dice Score for Pancreas-CT and BTCV datasets, respectively.
Initialization and Alignment for Adversarial Texture Optimization
Jul 28, 2022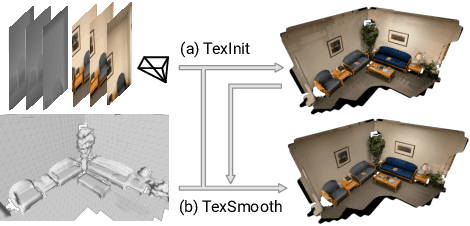
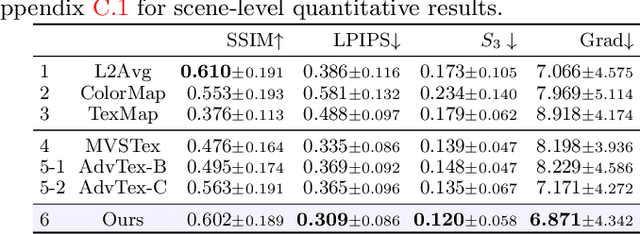

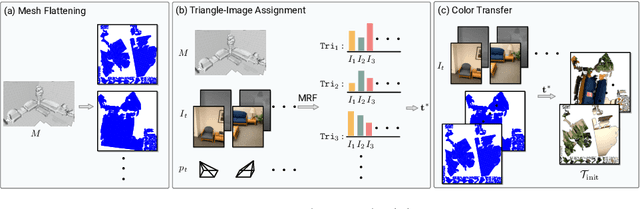
While recovery of geometry from image and video data has received a lot of attention in computer vision, methods to capture the texture for a given geometry are less mature. Specifically, classical methods for texture generation often assume clean geometry and reasonably well-aligned image data. While very recent methods, e.g., adversarial texture optimization, better handle lower-quality data obtained from hand-held devices, we find them to still struggle frequently. To improve robustness, particularly of recent adversarial texture optimization, we develop an explicit initialization and an alignment procedure. It deals with complex geometry due to a robust mapping of the geometry to the texture map and a hard-assignment-based initialization. It deals with misalignment of geometry and images by integrating fast image-alignment into the texture refinement optimization. We demonstrate efficacy of our texture generation on a dataset of 11 scenes with a total of 2807 frames, observing 7.8% and 11.1% relative improvements regarding perceptual and sharpness measurements.
Class-Level Logit Perturbation
Sep 26, 2022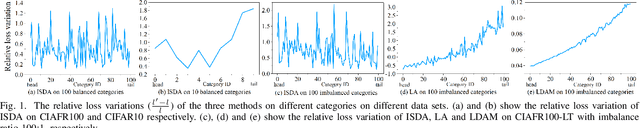
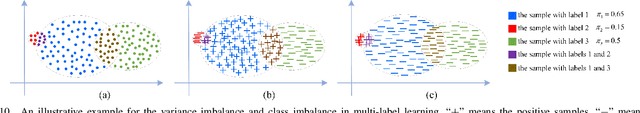
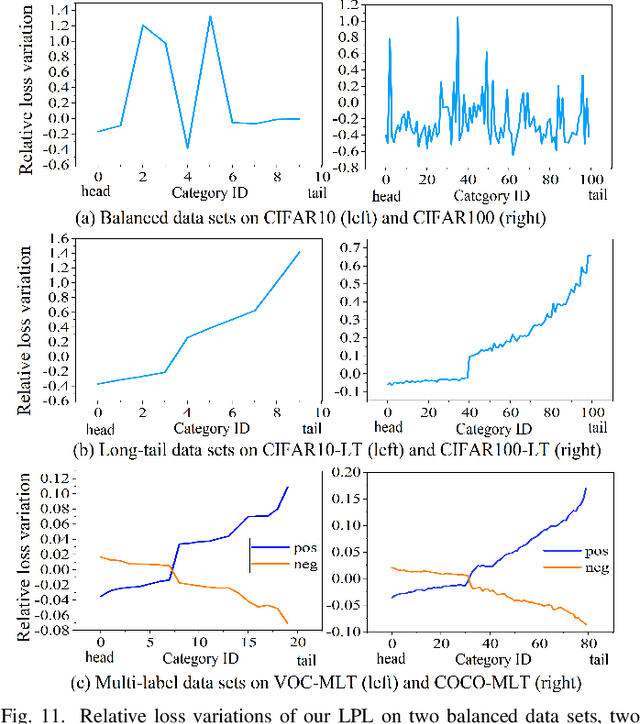
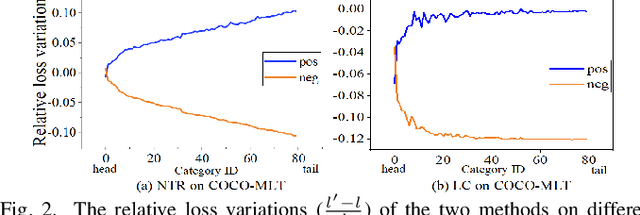
Features, logits, and labels are the three primary data when a sample passes through a deep neural network. Feature perturbation and label perturbation receive increasing attention in recent years. They have been proven to be useful in various deep learning approaches. For example, (adversarial) feature perturbation can improve the robustness or even generalization capability of learned models. However, limited studies have explicitly explored for the perturbation of logit vectors. This work discusses several existing methods related to class-level logit perturbation. A unified viewpoint between positive/negative data augmentation and loss variations incurred by logit perturbation is established. A theoretical analysis is provided to illuminate why class-level logit perturbation is useful. Accordingly, new methodologies are proposed to explicitly learn to perturb logits for both single-label and multi-label classification tasks. Extensive experiments on benchmark image classification data sets and their long-tail versions indicated the competitive performance of our learning method. As it only perturbs on logit, it can be used as a plug-in to fuse with any existing classification algorithms. All the codes are available at https://github.com/limengyang1992/lpl.
BLUE at Memotion 2.0 2022: You have my Image, my Text and my Transformer
Feb 28, 2022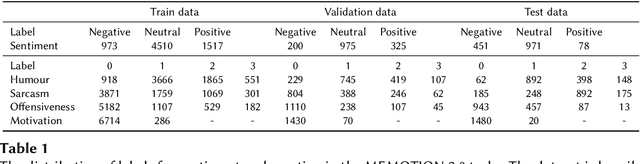

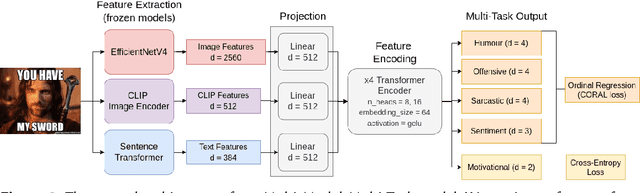
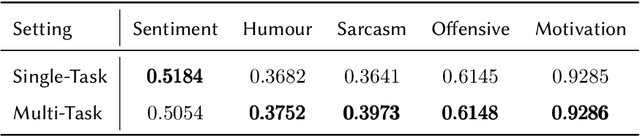
Memes are prevalent on the internet and continue to grow and evolve alongside our culture. An automatic understanding of memes propagating on the internet can shed light on the general sentiment and cultural attitudes of people. In this work, we present team BLUE's solution for the second edition of the MEMOTION competition. We showcase two approaches for meme classification (i.e. sentiment, humour, offensive, sarcasm and motivation levels) using a text-only method using BERT, and a Multi-Modal-Multi-Task transformer network that operates on both the meme image and its caption to output the final scores. In both approaches, we leverage state-of-the-art pretrained models for text (BERT, Sentence Transformer) and image processing (EfficientNetV4, CLIP). Through our efforts, we obtain first place in task A, second place in task B and third place in task C. In addition, our team obtained the highest average score for all three tasks.
Negative Evidence Matters in Interpretable Histology Image Classification
Jan 07, 2022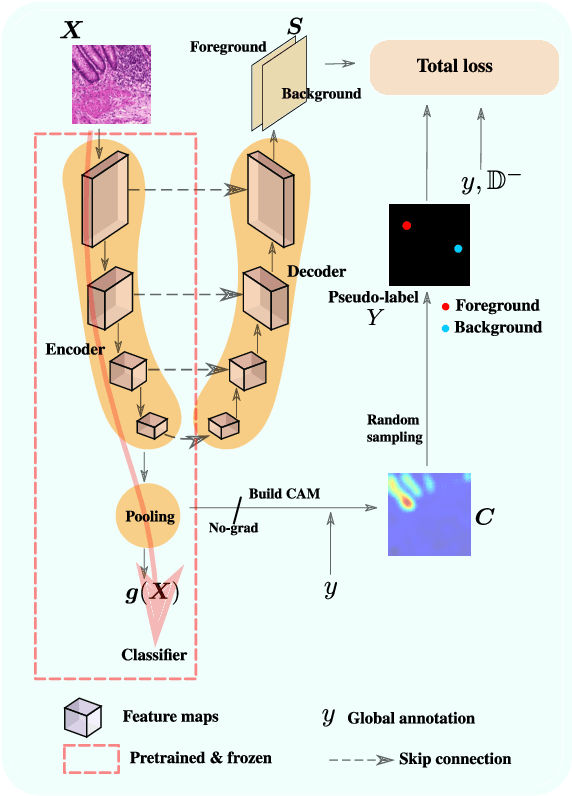
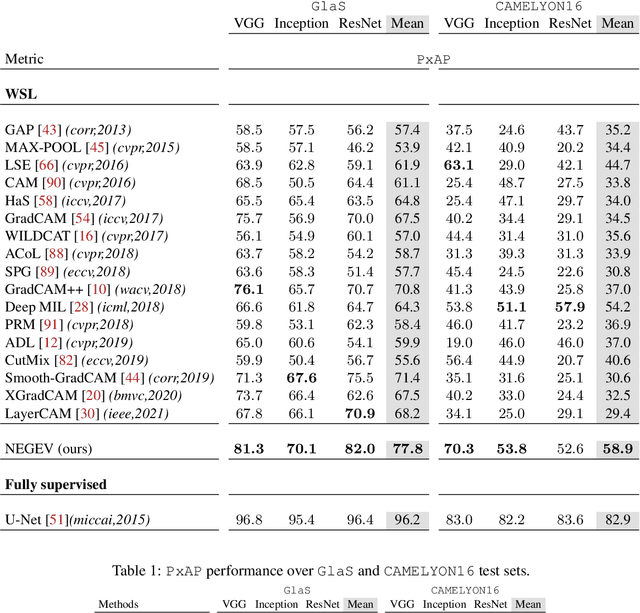
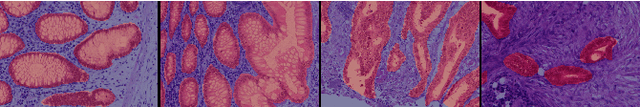
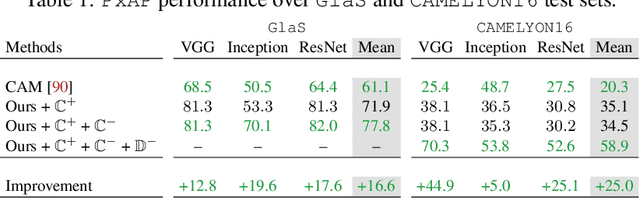
Using only global annotations such as the image class labels, weakly-supervised learning methods allow CNN classifiers to jointly classify an image, and yield the regions of interest associated with the predicted class. However, without any guidance at the pixel level, such methods may yield inaccurate regions. This problem is known to be more challenging with histology images than with natural ones, since objects are less salient, structures have more variations, and foreground and background regions have stronger similarities. Therefore, methods in computer vision literature for visual interpretation of CNNs may not directly apply. In this work, we propose a simple yet efficient method based on a composite loss function that leverages information from the fully negative samples. Our new loss function contains two complementary terms: the first exploits positive evidence collected from the CNN classifier, while the second leverages the fully negative samples from the training dataset. In particular, we equip a pre-trained classifier with a decoder that allows refining the regions of interest. The same classifier is exploited to collect both the positive and negative evidence at the pixel level to train the decoder. This enables to take advantages of the fully negative samples that occurs naturally in the data, without any additional supervision signals and using only the image class as supervision. Compared to several recent related methods, over the public benchmark GlaS for colon cancer and a Camelyon16 patch-based benchmark for breast cancer using three different backbones, we show the substantial improvements introduced by our method. Our results shows the benefits of using both negative and positive evidence, ie, the one obtained from a classifier and the one naturally available in datasets. We provide an ablation study of both terms. Our code is publicly available.
L^3U-net: Low-Latency Lightweight U-net Based Image Segmentation Model for Parallel CNN Processors
Mar 30, 2022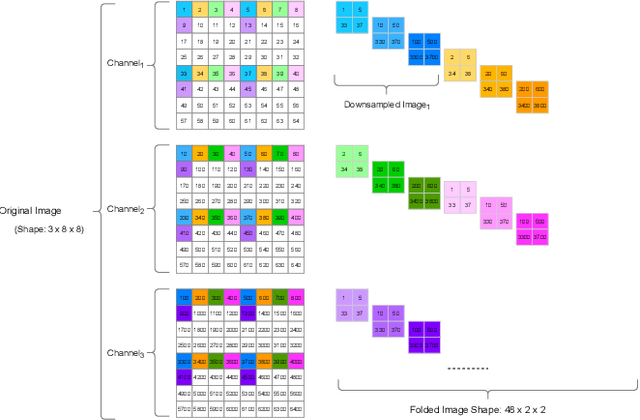
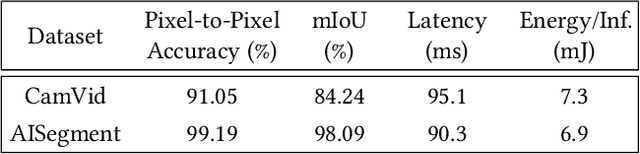
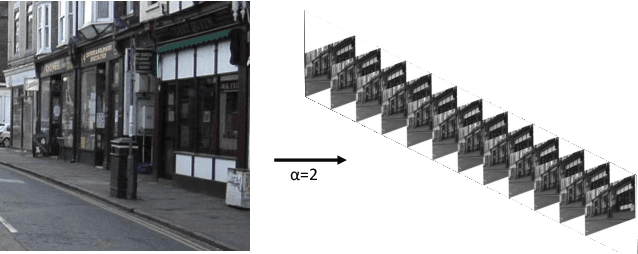
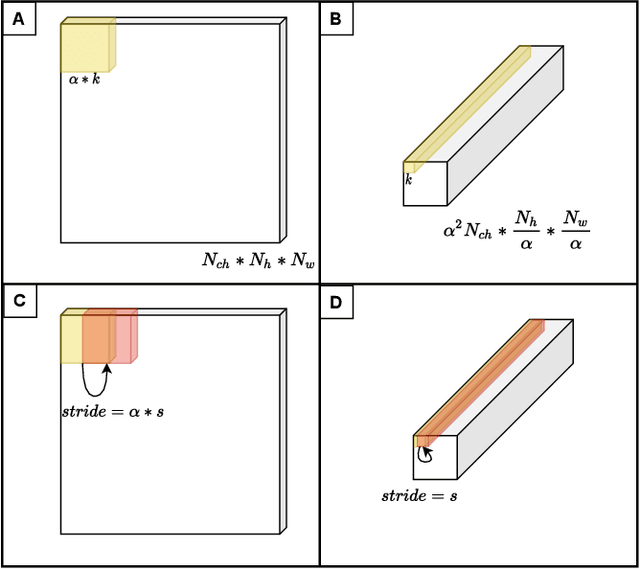
In this research, we propose a tiny image segmentation model, L^3U-net, that works on low-resource edge devices in real-time. We introduce a data folding technique that reduces inference latency by leveraging the parallel convolutional layer processing capability of the CNN accelerators. We also deploy the proposed model to such a device, MAX78000, and the results show that L^3U-net achieves more than 90% accuracy over two different segmentation datasets with 10 fps.
CIRCA: comprehensible online system in support of chest X-rays-based COVID-19 diagnosis
Oct 11, 2022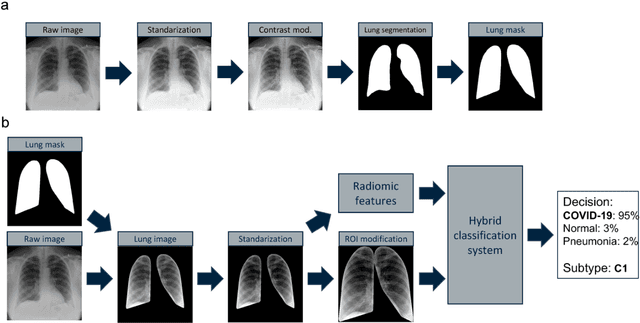
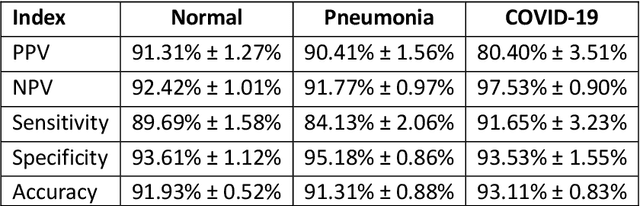
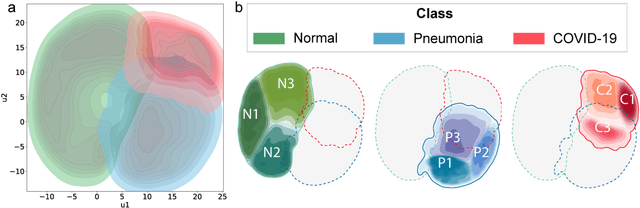
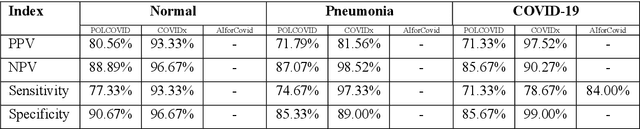
Due to the large accumulation of patients requiring hospitalization, the COVID-19 pandemic disease caused a high overload of health systems, even in developed countries. Deep learning techniques based on medical imaging data can help in the faster detection of COVID-19 cases and monitoring of disease progression. Regardless of the numerous proposed solutions for lung X-rays, none of them is a product that can be used in the clinic. Five different datasets (POLCOVID, AIforCOVID, COVIDx, NIH, and artificially generated data) were used to construct a representative dataset of 23 799 CXRs for model training; 1 050 images were used as a hold-out test set, and 44 247 as independent test set (BIMCV database). A U-Net-based model was developed to identify a clinically relevant region of the CXR. Each image class (normal, pneumonia, and COVID-19) was divided into 3 subtypes using a 2D Gaussian mixture model. A decision tree was used to aggregate predictions from the InceptionV3 network based on processed CXRs and a dense neural network on radiomic features. The lung segmentation model gave the Sorensen-Dice coefficient of 94.86% in the validation dataset, and 93.36% in the testing dataset. In 5-fold cross-validation, the accuracy for all classes ranged from 91% to 93%, keeping slightly higher specificity than sensitivity and NPV than PPV. In the hold-out test set, the balanced accuracy ranged between 68% and 100%. The highest performance was obtained for the subtypes N1, P1, and C1. A similar performance was obtained on the independent dataset for normal and COVID-19 class subtypes. Seventy-six percent of COVID-19 patients wrongly classified as normal cases were annotated by radiologists as with no signs of disease. Finally, we developed the online service (https://circa.aei.polsl.pl) to provide access to fast diagnosis support tools.
Data-free Dense Depth Distillation
Aug 26, 2022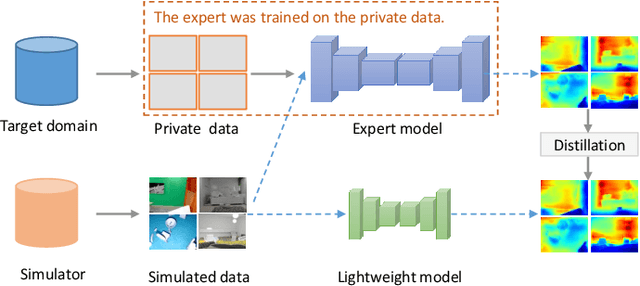
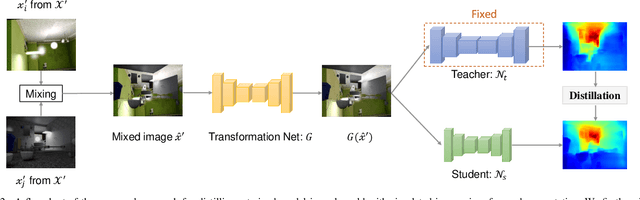
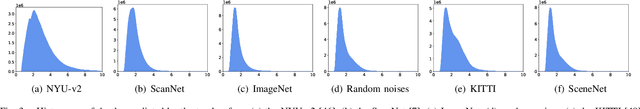
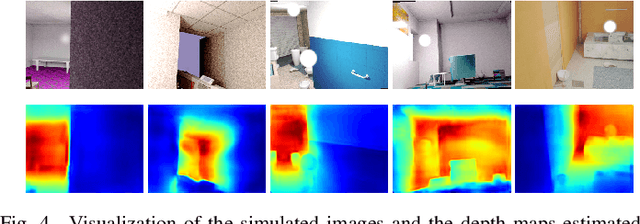
We study data-free knowledge distillation (KD) for monocular depth estimation (MDE), which learns a lightweight network for real-world depth perception by compressing from a trained expert model under the teacher-student framework while lacking training data in the target domain. Owing to the essential difference between dense regression and image recognition, previous methods of data-free KD are not applicable to MDE. To strengthen the applicability in the real world, in this paper, we seek to apply KD with out-of-distribution simulated images. The major challenges are i) lacking prior information about object distribution of the original training data; ii) the domain shift between the real world and the simulation. To cope with the first difficulty, we apply object-wise image mixing to generate new training samples for maximally covering distributed patterns of objects in the target domain. To tackle the second difficulty, we propose to utilize a transformation network that efficiently learns to fit the simulated data to the feature distribution of the teacher model. We evaluate the proposed approach for various depth estimation models and two different datasets. As a result, our method outperforms the baseline KD by a good margin and even achieves slightly better performance with as few as $1/6$ images, demonstrating a clear superiority.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge