"Image": models, code, and papers
DeepSTI: Towards Tensor Reconstruction using Fewer Orientations in Susceptibility Tensor Imaging
Sep 09, 2022
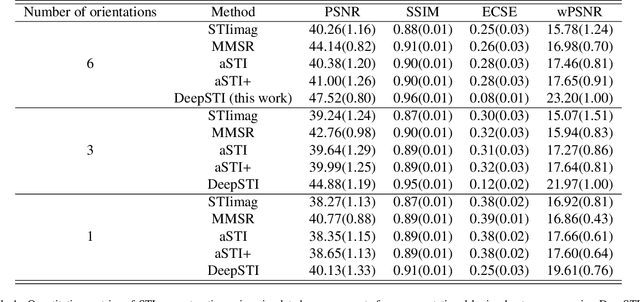
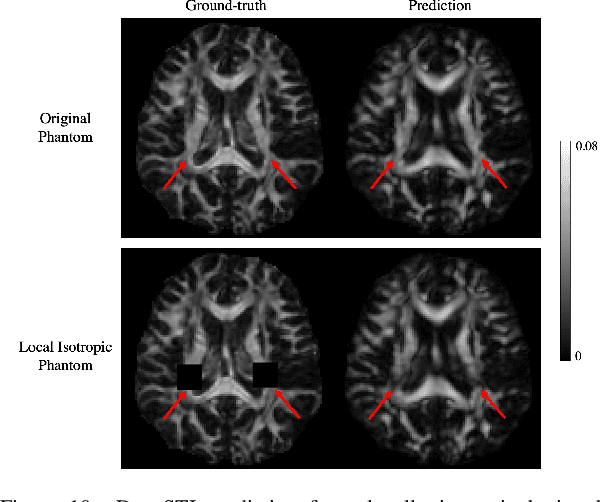
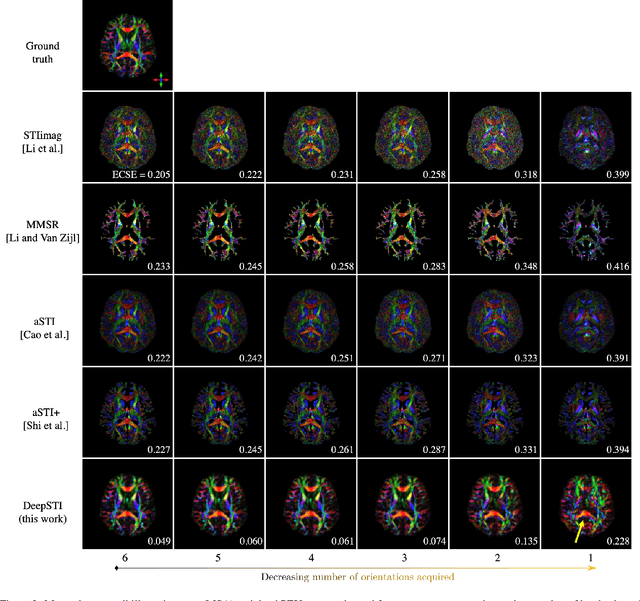
Susceptibility tensor imaging (STI) is an emerging magnetic resonance imaging technique that characterizes the anisotropic tissue magnetic susceptibility with a second-order tensor model. STI has the potential to provide information for both the reconstruction of white matter fiber pathways and detection of myelin changes in the brain at mm resolution or less, which would be of great value for understanding brain structure and function in healthy and diseased brain. However, the application of STI in vivo has been hindered by its cumbersome and time-consuming acquisition requirement of measuring susceptibility induced MR phase changes at multiple (usually more than six) head orientations. This complexity is enhanced by the limitation in head rotation angles due to physical constraints of the head coil. As a result, STI has not yet been widely applied in human studies in vivo. In this work, we tackle these issues by proposing an image reconstruction algorithm for STI that leverages data-driven priors. Our method, called DeepSTI, learns the data prior implicitly via a deep neural network that approximates the proximal operator of a regularizer function for STI. The dipole inversion problem is then solved iteratively using the learned proximal network. Experimental results using both simulation and in vivo human data demonstrate great improvement over state-of-the-art algorithms in terms of the reconstructed tensor image, principal eigenvector maps and tractography results, while allowing for tensor reconstruction with MR phase measured at much less than six different orientations. Notably, promising reconstruction results are achieved by our method from only one orientation in human in vivo, and we demonstrate a potential application of this technique for estimating lesion susceptibility anisotropy in patients with multiple sclerosis.
Improving Computed Tomography (CT) Reconstruction via 3D Shape Induction
Aug 23, 2022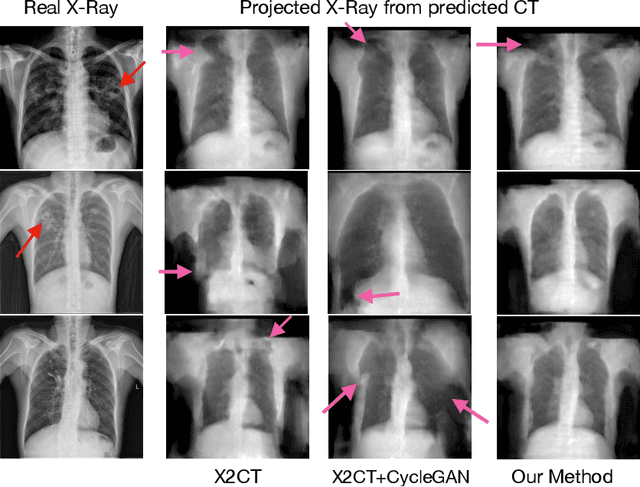

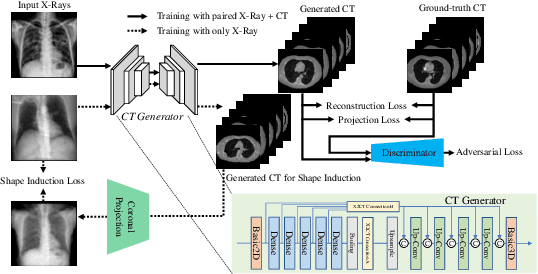

Chest computed tomography (CT) imaging adds valuable insight in the diagnosis and management of pulmonary infectious diseases, like tuberculosis (TB). However, due to the cost and resource limitations, only X-ray images may be available for initial diagnosis or follow up comparison imaging during treatment. Due to their projective nature, X-rays images may be more difficult to interpret by clinicians. The lack of publicly available paired X-ray and CT image datasets makes it challenging to train a 3D reconstruction model. In addition, Chest X-ray radiology may rely on different device modalities with varying image quality and there may be variation in underlying population disease spectrum that creates diversity in inputs. We propose shape induction, that is, learning the shape of 3D CT from X-ray without CT supervision, as a novel technique to incorporate realistic X-ray distributions during training of a reconstruction model. Our experiments demonstrate that this process improves both the perceptual quality of generated CT and the accuracy of down-stream classification of pulmonary infectious diseases.
Multiplex-detection Based Multiple Instance Learning Network for Whole Slide Image Classification
Aug 06, 2022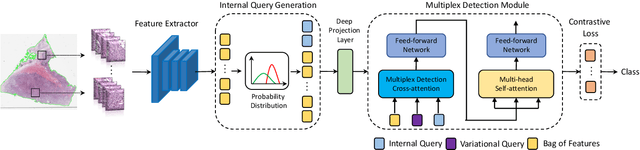
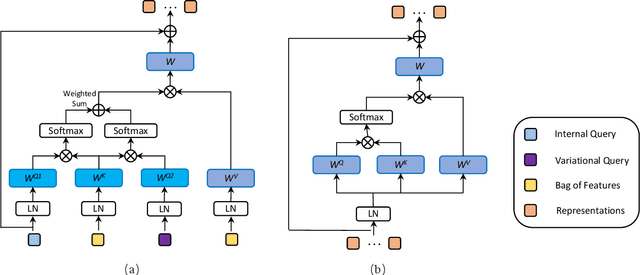
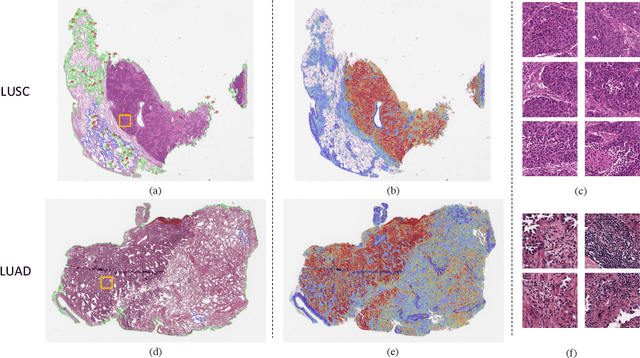
Multiple instance learning (MIL) is a powerful approach to classify whole slide images (WSIs) for diagnostic pathology. A fundamental challenge of MIL on WSI classification is to discover the \textit{critical instances} that trigger the bag label. However, previous methods are primarily designed under the independent and identical distribution hypothesis (\textit{i.i.d}), ignoring either the correlations between instances or heterogeneity of tumours. In this paper, we propose a novel multiplex-detection-based multiple instance learning (MDMIL) to tackle the issues above. Specifically, MDMIL is constructed by the internal query generation module (IQGM) and the multiplex detection module (MDM) and assisted by the memory-based contrastive loss during training. Firstly, IQGM gives the probability of instances and generates the internal query (IQ) for the subsequent MDM by aggregating highly reliable features after the distribution analysis. Secondly, the multiplex-detection cross-attention (MDCA) and multi-head self-attention (MHSA) in MDM cooperate to generate the final representations for the WSI. In this process, the IQ and trainable variational query (VQ) successfully build up the connections between instances and significantly improve the model's robustness toward heterogeneous tumours. At last, to further enforce constraints in the feature space and stabilize the training process, we adopt a memory-based contrastive loss, which is practicable for WSI classification even with a single sample as input in each iteration. We conduct experiments on three computational pathology datasets, e.g., CAMELYON16, TCGA-NSCLC, and TCGA-RCC datasets. The superior accuracy and AUC demonstrate the superiority of our proposed MDMIL over other state-of-the-art methods.
ITA: Image-Text Alignments for Multi-Modal Named Entity Recognition
Dec 13, 2021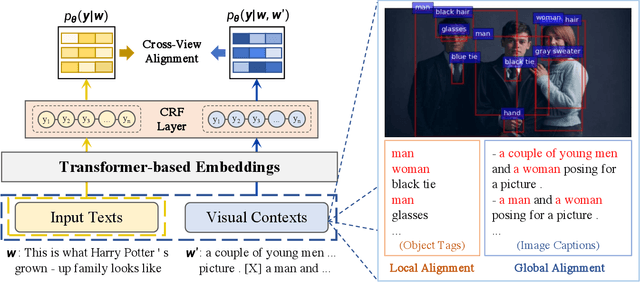
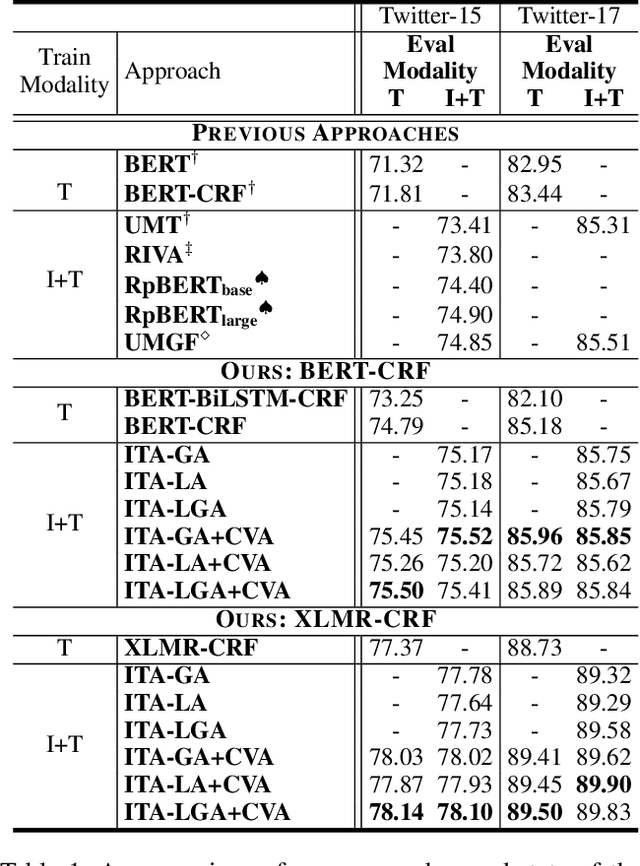
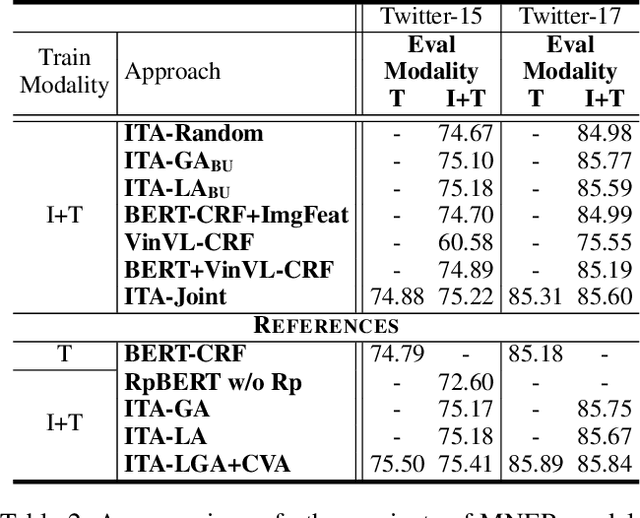
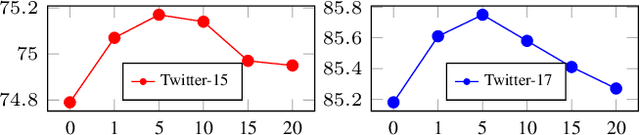
Recently, Multi-modal Named Entity Recognition (MNER) has attracted a lot of attention. Most of the work utilizes image information through region-level visual representations obtained from a pretrained object detector and relies on an attention mechanism to model the interactions between image and text representations. However, it is difficult to model such interactions as image and text representations are trained separately on the data of their respective modality and are not aligned in the same space. As text representations take the most important role in MNER, in this paper, we propose {\bf I}mage-{\bf t}ext {\bf A}lignments (ITA) to align image features into the textual space, so that the attention mechanism in transformer-based pretrained textual embeddings can be better utilized. ITA first locally and globally aligns regional object tags and image-level captions as visual contexts, concatenates them with the input texts as a new cross-modal input, and then feeds it into a pretrained textual embedding model. This makes it easier for the attention module of a pretrained textual embedding model to model the interaction between the two modalities since they are both represented in the textual space. ITA further aligns the output distributions predicted from the cross-modal input and textual input views so that the MNER model can be more practical and robust to noises from images. In our experiments, we show that ITA models can achieve state-of-the-art accuracy on multi-modal Named Entity Recognition datasets, even without image information.
There is a Time and Place for Reasoning Beyond the Image
Mar 01, 2022
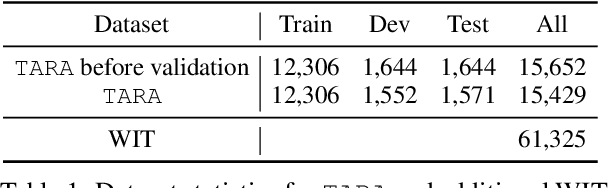

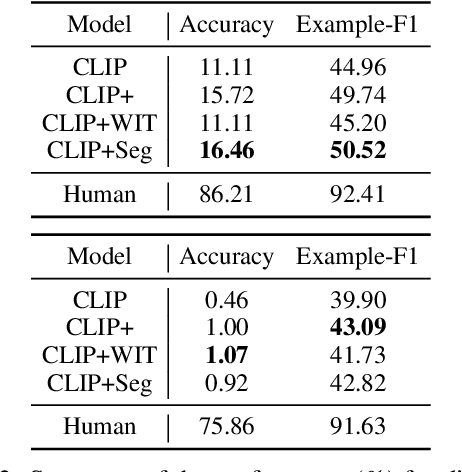
Images are often more significant than only the pixels to human eyes, as we can infer, associate, and reason with contextual information from other sources to establish a more complete picture. For example, in Figure 1, we can find a way to identify the news articles related to the picture through segment-wise understandings on the signs, the buildings, the crowds, and more. This tells us the time when and the location where the image is taken, which will help us in subsequent tasks, such as evidence retrieval for criminal activities, automatic storyline construction, and upper-stream processing such as image clustering. In this work, we formulate this problem and introduce TARA: a dataset with 16k images with their associated news, time and location automatically extracted from New York Times (NYT), and an additional 61k examples as distant supervision from WIT. On top of the extractions, we present a crowdsourced subset in which images are believed to be feasible to find their spatio-temporal information for evaluation purpose. We show that there exists a 70% gap between a state-of-the-art joint model and human performance, which is slightly filled by our proposed model that uses segment-wise reasoning, motivating higher-level vision-language joint models that can conduct open-ended reasoning with world knowledge.
Evons: A Dataset for Fake and Real News Virality Analysis and Prediction
Sep 16, 2022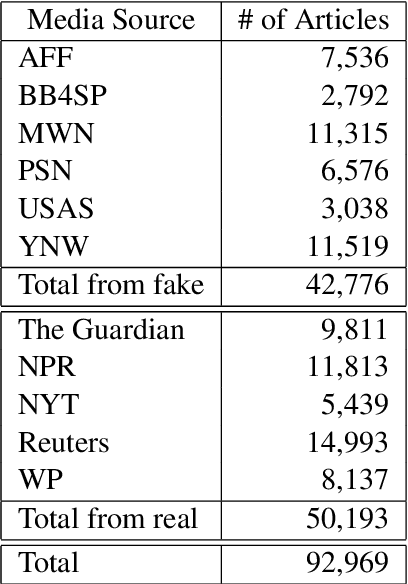
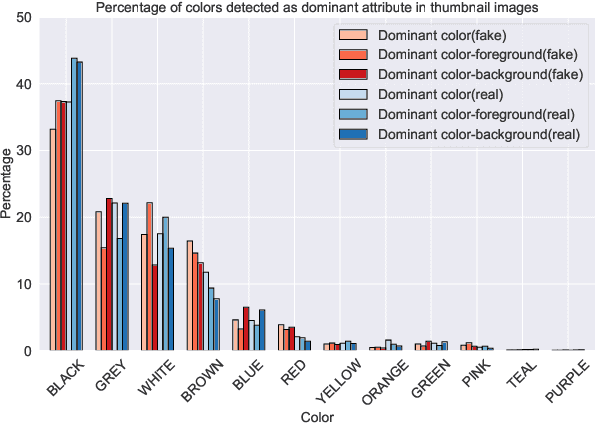

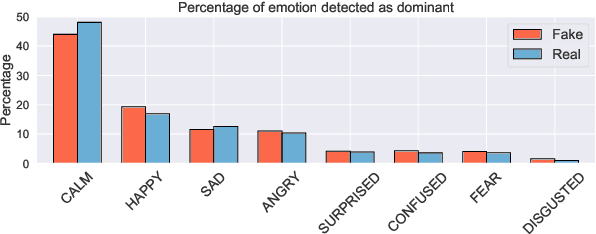
We present a novel collection of news articles originating from fake and real news media sources for the analysis and prediction of news virality. Unlike existing fake news datasets which either contain claims or news article headline and body, in this collection each article is supported with a Facebook engagement count which we consider as an indicator of the article virality. In addition we also provide the article description and thumbnail image with which the article was shared on Facebook. These images were automatically annotated with object tags and color attributes. Using cloud based vision analysis tools, thumbnail images were also analyzed for faces and detected faces were annotated with facial attributes. We empirically investigate the use of this collection on an example task of article virality prediction.
Diffusion Adversarial Representation Learning for Self-supervised Vessel Segmentation
Sep 29, 2022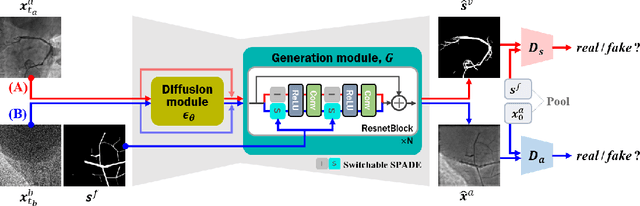
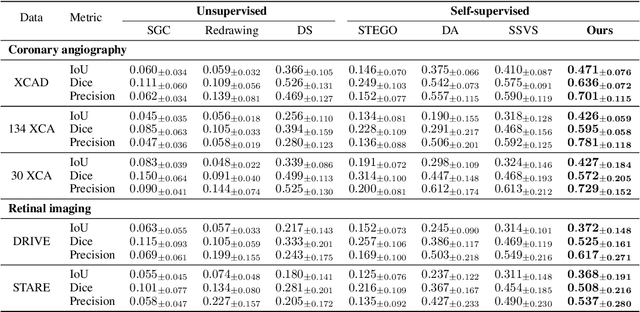
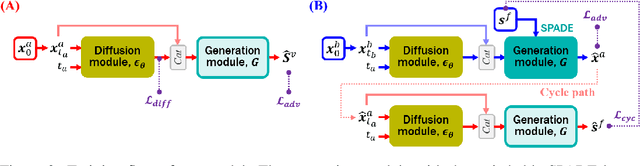
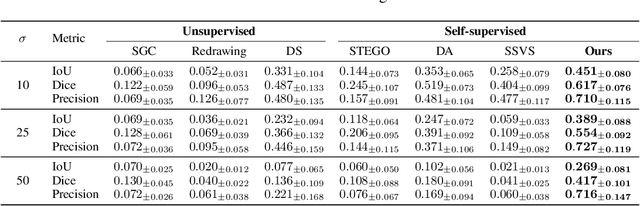
Vessel segmentation in medical images is one of the important tasks in the diagnosis of vascular diseases and therapy planning. Although learning-based segmentation approaches have been extensively studied, a large amount of ground-truth labels are required in supervised methods and confusing background structures make neural networks hard to segment vessels in an unsupervised manner. To address this, here we introduce a novel diffusion adversarial representation learning (DARL) model that leverages a denoising diffusion probabilistic model with adversarial learning, and apply it for vessel segmentation. In particular, for self-supervised vessel segmentation, DARL learns background image distribution using a diffusion module, which lets a generation module effectively provide vessel representations. Also, by adversarial learning based on the proposed switchable spatially-adaptive denormalization, our model estimates synthetic fake vessel images as well as vessel segmentation masks, which further makes the model capture vessel-relevant semantic information. Once the proposed model is trained, the model generates segmentation masks by one step and can be applied to general vascular structure segmentation of coronary angiography and retinal images. Experimental results on various datasets show that our method significantly outperforms existing unsupervised and self-supervised methods in vessel segmentation.
Deep Cervix Model Development from Heterogeneous and Partially Labeled Image Datasets
Jan 18, 2022Cervical cancer is the fourth most common cancer in women worldwide. The availability of a robust automated cervical image classification system can augment the clinical care provider's limitation in traditional visual inspection with acetic acid (VIA). However, there are a wide variety of cervical inspection objectives which impact the labeling criteria for criteria-specific prediction model development. Moreover, due to the lack of confirmatory test results and inter-rater labeling variation, many images are left unlabeled. Motivated by these challenges, we propose a self-supervised learning (SSL) based approach to produce a pre-trained cervix model from unlabeled cervical images. The developed model is further fine-tuned to produce criteria-specific classification models with the available labeled images. We demonstrate the effectiveness of the proposed approach using two cervical image datasets. Both datasets are partially labeled and labeling criteria are different. The experimental results show that the SSL-based initialization improves classification performance (Accuracy: 2.5% min) and the inclusion of images from both datasets during SSL further improves the performance (Accuracy: 1.5% min). Further, considering data-sharing restrictions, we experimented with the effectiveness of Federated SSL and find that it can improve performance over the SSL model developed with just its images. This justifies the importance of SSL-based cervix model development. We believe that the present research shows a novel direction in developing criteria-specific custom deep models for cervical image classification by combining images from different sources unlabeled and/or labeled with varying criteria, and addressing image access restrictions.
Tunable Image Quality Control of 3-D Ultrasound using Switchable CycleGAN
Dec 06, 2021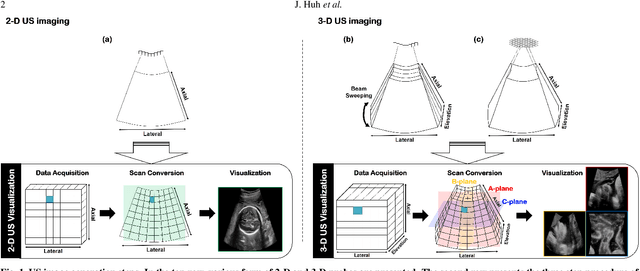
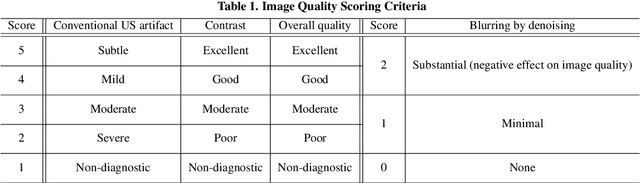
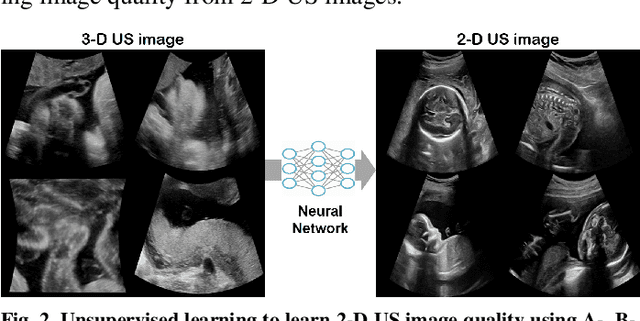
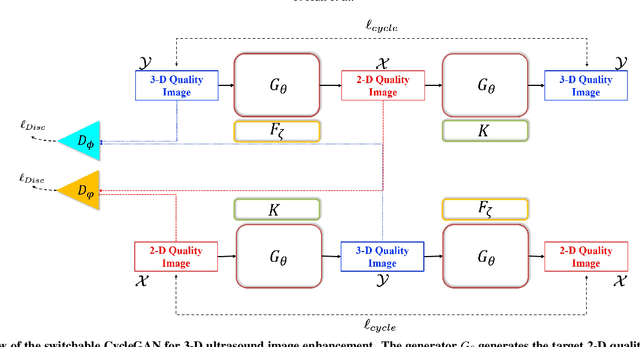
In contrast to 2-D ultrasound (US) for uniaxial plane imaging, a 3-D US imaging system can visualize a volume along three axial planes. This allows for a full view of the anatomy, which is useful for gynecological (GYN) and obstetrical (OB) applications. Unfortunately, the 3-D US has an inherent limitation in resolution compared to the 2-D US. In the case of 3-D US with a 3-D mechanical probe, for example, the image quality is comparable along the beam direction, but significant deterioration in image quality is often observed in the other two axial image planes. To address this, here we propose a novel unsupervised deep learning approach to improve 3-D US image quality. In particular, using {\em unmatched} high-quality 2-D US images as a reference, we trained a recently proposed switchable CycleGAN architecture so that every mapping plane in 3-D US can learn the image quality of 2-D US images. Thanks to the switchable architecture, our network can also provide real-time control of image enhancement level based on user preference, which is ideal for a user-centric scanner setup. Extensive experiments with clinical evaluation confirm that our method offers significantly improved image quality as well user-friendly flexibility.
Semi-supervised Body Parsing and Pose Estimation for Enhancing Infant General Movement Assessment
Oct 14, 2022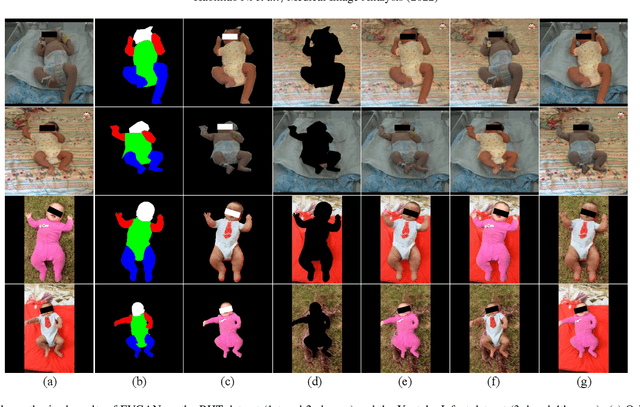
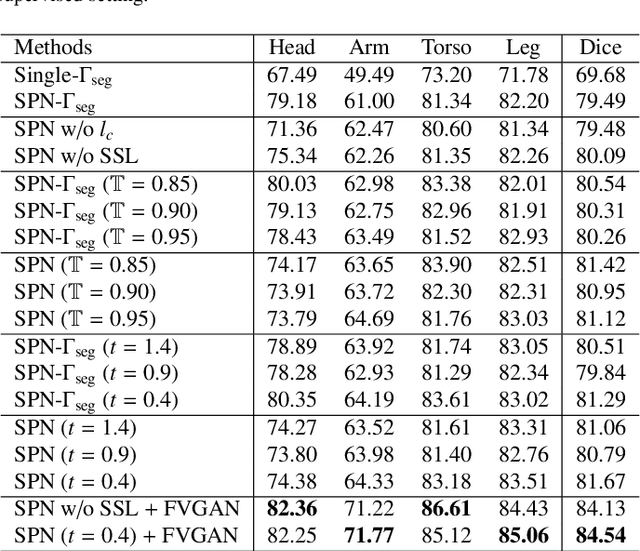
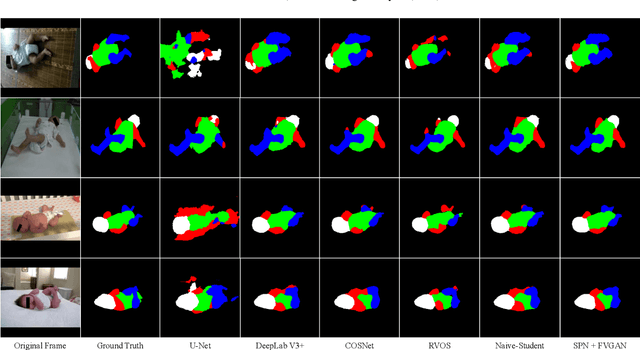
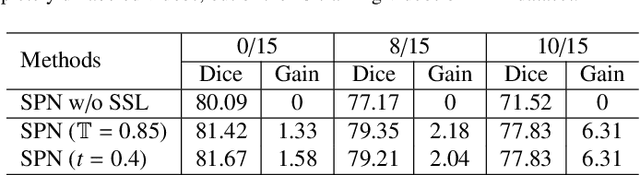
General movement assessment (GMA) of infant movement videos (IMVs) is an effective method for early detection of cerebral palsy (CP) in infants. We demonstrate in this paper that end-to-end trainable neural networks for image sequence recognition can be applied to achieve good results in GMA, and more importantly, augmenting raw video with infant body parsing and pose estimation information can significantly improve performance. To solve the problem of efficiently utilizing partially labeled IMVs for body parsing, we propose a semi-supervised model, termed SiamParseNet (SPN), which consists of two branches, one for intra-frame body parts segmentation and another for inter-frame label propagation. During training, the two branches are jointly trained by alternating between using input pairs of only labeled frames and input of both labeled and unlabeled frames. We also investigate training data augmentation by proposing a factorized video generative adversarial network (FVGAN) to synthesize novel labeled frames for training. When testing, we employ a multi-source inference mechanism, where the final result for a test frame is either obtained via the segmentation branch or via propagation from a nearby key frame. We conduct extensive experiments for body parsing using SPN on two infant movement video datasets, where SPN coupled with FVGAN achieves state-of-the-art performance. We further demonstrate that SPN can be easily adapted to the infant pose estimation task with superior performance. Last but not least, we explore the clinical application of our method for GMA. We collected a new clinical IMV dataset with GMA annotations, and our experiments show that SPN models for body parsing and pose estimation trained on the first two datasets generalize well to the new clinical dataset and their results can significantly boost the CRNN-based GMA prediction performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge