"Image": models, code, and papers
An Effective Approach for Multi-label Classification with Missing Labels
Oct 24, 2022
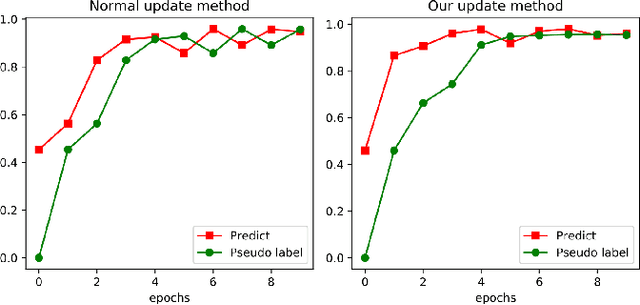
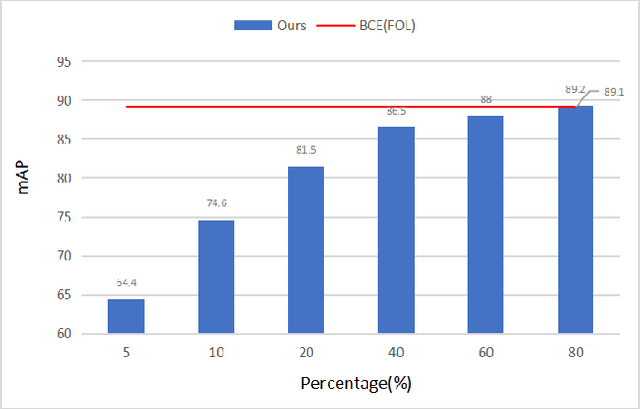
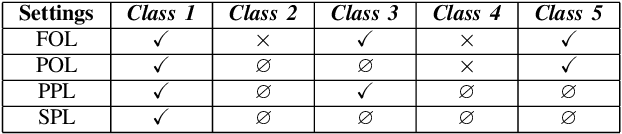
Compared with multi-class classification, multi-label classification that contains more than one class is more suitable in real life scenarios. Obtaining fully labeled high-quality datasets for multi-label classification problems, however, is extremely expensive, and sometimes even infeasible, with respect to annotation efforts, especially when the label spaces are too large. This motivates the research on partial-label classification, where only a limited number of labels are annotated and the others are missing. To address this problem, we first propose a pseudo-label based approach to reduce the cost of annotation without bringing additional complexity to the existing classification networks. Then we quantitatively study the impact of missing labels on the performance of classifier. Furthermore, by designing a novel loss function, we are able to relax the requirement that each instance must contain at least one positive label, which is commonly used in most existing approaches. Through comprehensive experiments on three large-scale multi-label image datasets, i.e. MS-COCO, NUS-WIDE, and Pascal VOC12, we show that our method can handle the imbalance between positive labels and negative labels, while still outperforming existing missing-label learning approaches in most cases, and in some cases even approaches with fully labeled datasets.
LidarAugment: Searching for Scalable 3D LiDAR Data Augmentations
Oct 24, 2022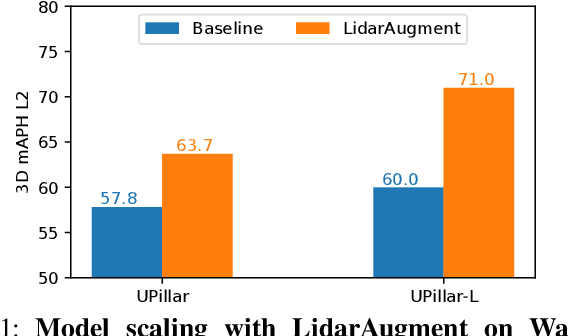

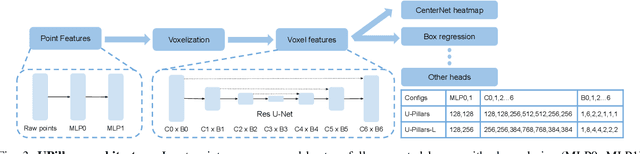

Data augmentations are important in training high-performance 3D object detectors for point clouds. Despite recent efforts on designing new data augmentations, perhaps surprisingly, most state-of-the-art 3D detectors only use a few simple data augmentations. In particular, different from 2D image data augmentations, 3D data augmentations need to account for different representations of input data and require being customized for different models, which introduces significant overhead. In this paper, we resort to a search-based approach, and propose LidarAugment, a practical and effective data augmentation strategy for 3D object detection. Unlike previous approaches where all augmentation policies are tuned in an exponentially large search space, we propose to factorize and align the search space of each data augmentation, which cuts down the 20+ hyperparameters to 2, and significantly reduces the search complexity. We show LidarAugment can be customized for different model architectures with different input representations by a simple 2D grid search, and consistently improve both convolution-based UPillars/StarNet/RSN and transformer-based SWFormer. Furthermore, LidarAugment mitigates overfitting and allows us to scale up 3D detectors to much larger capacity. In particular, by combining with latest 3D detectors, our LidarAugment achieves a new state-of-the-art 74.8 mAPH L2 on Waymo Open Dataset.
SFS-A68: a dataset for the segmentation of space functions in apartment buildings
Sep 13, 2022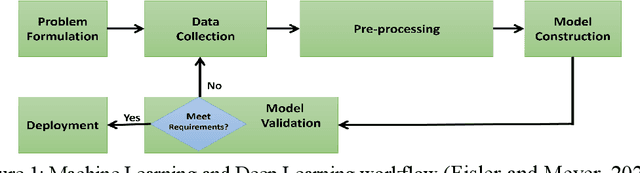
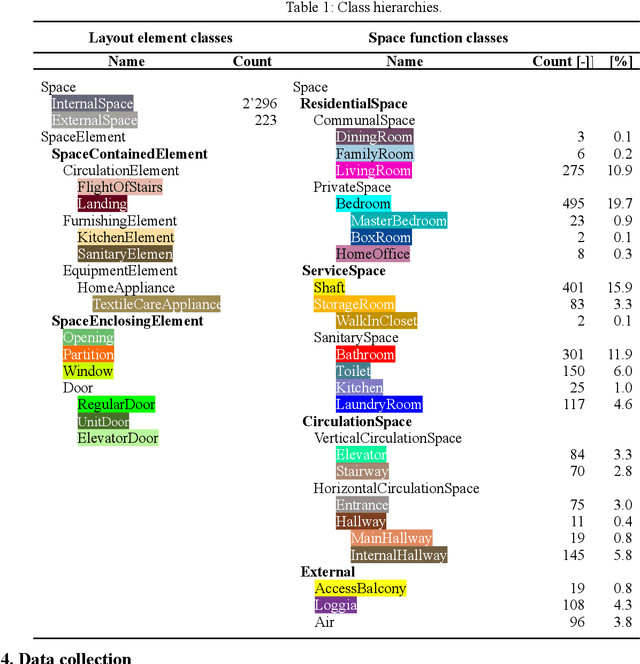
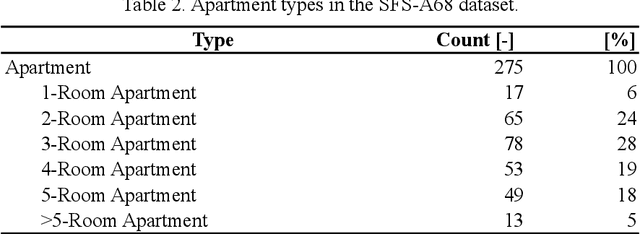

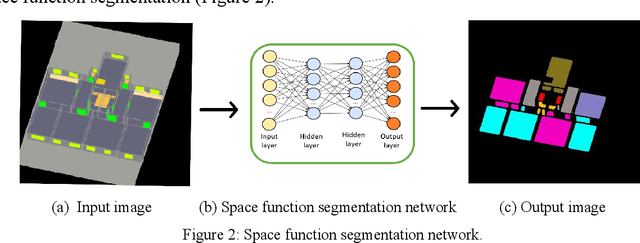

Analyzing building models for usable area, building safety, or energy analysis requires function classification data of spaces and related objects. Automated space function classification is desirable to reduce input model preparation effort and errors. Existing space function classifiers use space feature vectors or space connectivity graphs as input. The application of deep learning (DL) image segmentation methods to space function classification has not been studied. As an initial step towards addressing this gap, we present a dataset, SFS-A68, that consists of input and ground truth images generated from 68 digital 3D models of space layouts of apartment buildings. The dataset is suitable for developing DL models for space function segmentation. We use the dataset to train and evaluate an experimental space function segmentation network based on transfer learning and training from scratch. Test results confirm the applicability of DL image segmentation for space function classification. The code and the dataset of the experiments are publicly available online (https://github.com/A2Amir/SFS-A68).
* Published in proceedings of the 29th International Workshop on Intelligent Computing in Engineering, EG-ICE 2022, Aarhus, Denmark. https://doi.org/10.7146/aul.455.c222
A Deep Neural Network for Multiclass Bridge Element Parsing in Inspection Image Analysis
Sep 05, 2022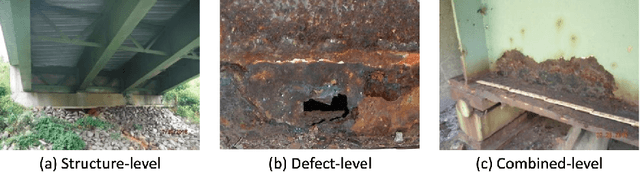
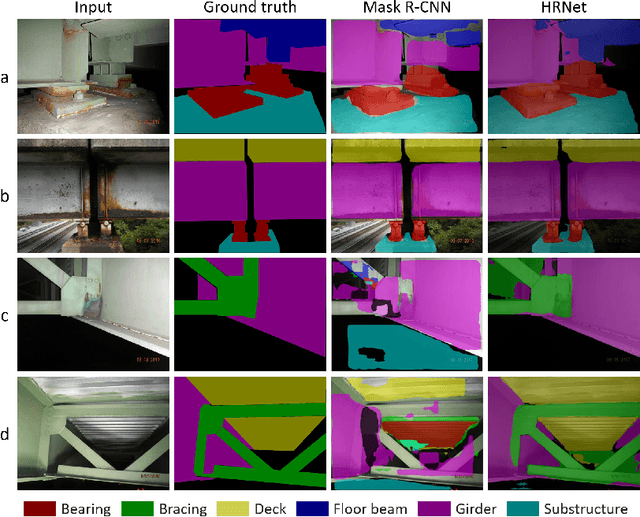
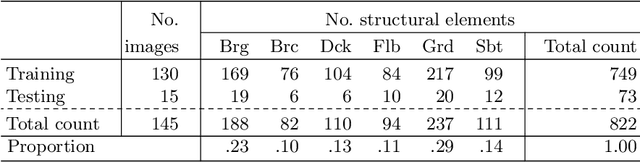
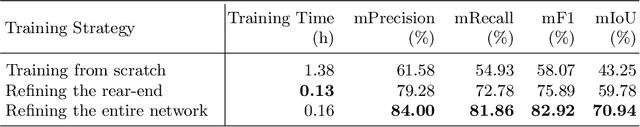

Aerial robots such as drones have been leveraged to perform bridge inspections. Inspection images with both recognizable structural elements and apparent surface defects can be collected by onboard cameras to provide valuable information for the condition assessment. This article aims to determine a suitable deep neural network (DNN) for parsing multiclass bridge elements in inspection images. An extensive set of quantitative evaluations along with qualitative examples show that High-Resolution Net (HRNet) possesses the desired ability. With data augmentation and a training sample of 130 images, a pre-trained HRNet is efficiently transferred to the task of structural element parsing and has achieved a 92.67% mean F1-score and 86.33% mean IoU.
Imitrob: Imitation Learning Dataset for Training and Evaluating 6D Object Pose Estimators
Sep 19, 2022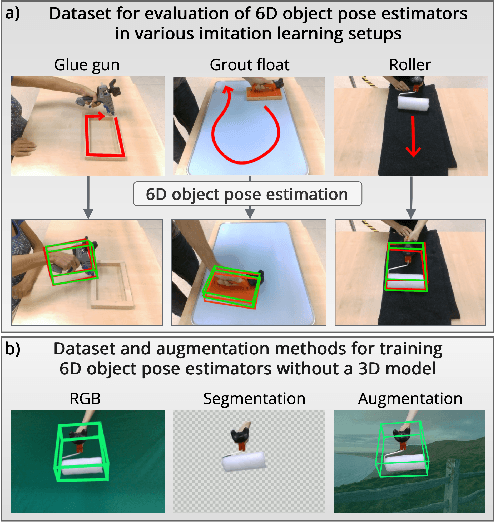
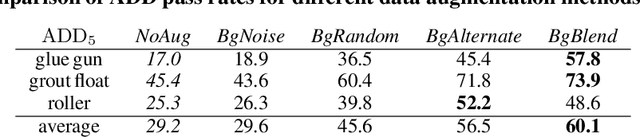
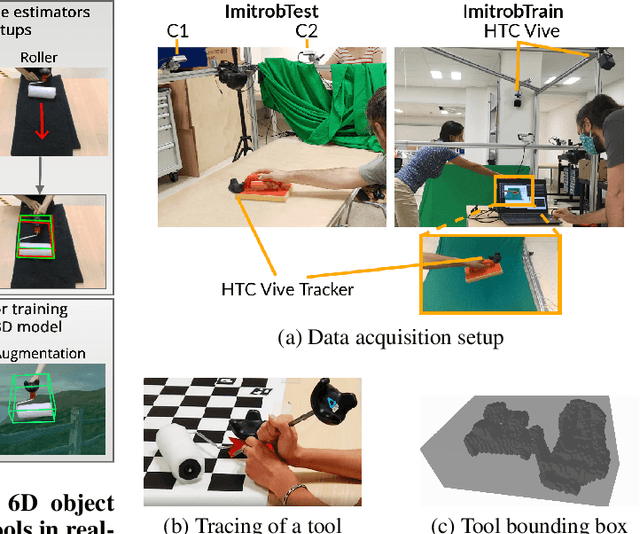


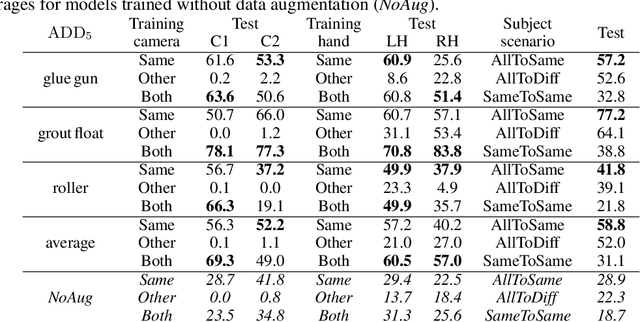
This paper introduces a dataset for training and evaluating methods for 6D pose estimation of hand-held tools in task demonstrations captured by a standard RGB camera. Despite the significant progress of 6D pose estimation methods, their performance is usually limited for heavily occluded objects, which is a common case in imitation learning where the object is typically partially occluded by the manipulating hand. Currently, there is a lack of datasets that would enable the development of robust 6D pose estimation methods for these conditions. To overcome this problem, we collect a new dataset (Imitrob) aimed at 6D pose estimation in imitation learning and other applications where a human holds a tool and performs a task. The dataset contains image sequences of three different tools and six manipulation tasks with two camera viewpoints, four human subjects, and left/right hand. Each image is accompanied by an accurate ground truth measurement of the 6D object pose, obtained by the HTC Vive motion tracking device. The use of the dataset is demonstrated by training and evaluating a recent 6D object pose estimation method (DOPE) in various setups. The dataset and code are publicly available at http://imitrob.ciirc.cvut.cz/imitrobdataset.php.
Improving anatomical plausibility in medical image segmentation via hybrid graph neural networks: applications to chest x-ray analysis
Apr 01, 2022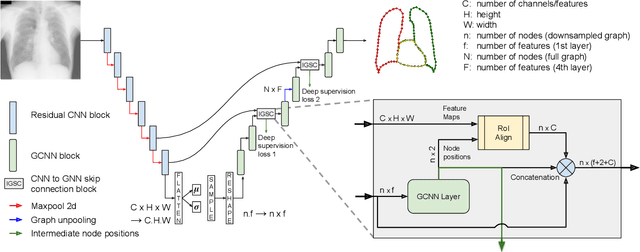
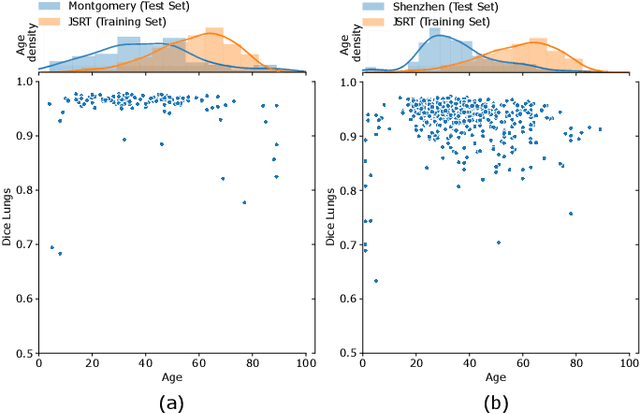
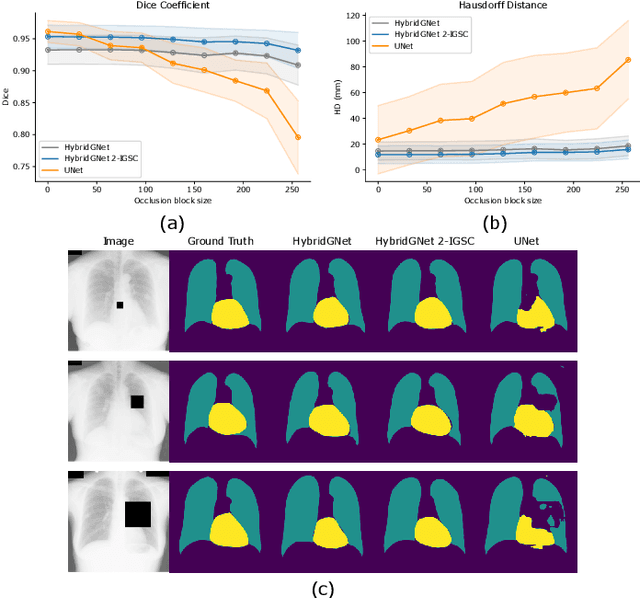
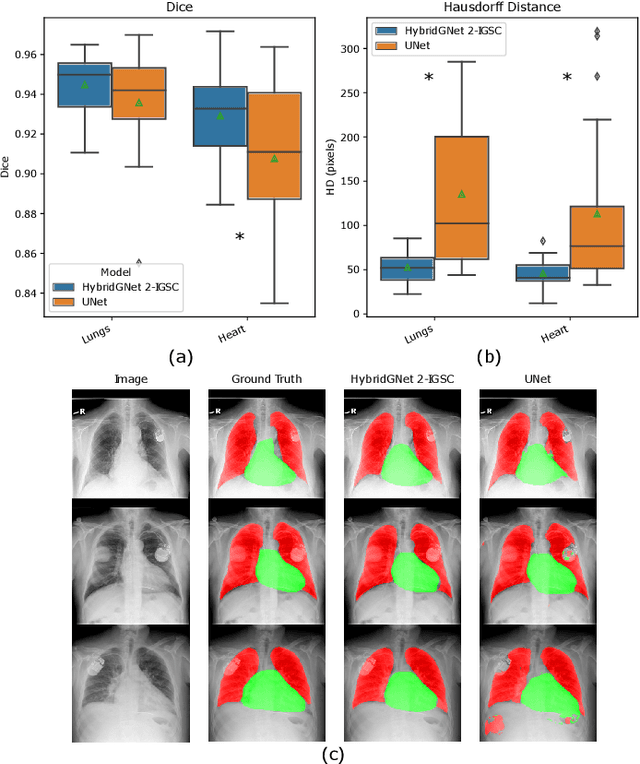
Anatomical segmentation is a fundamental task in medical image computing, generally tackled with fully convolutional neural networks which produce dense segmentation masks. These models are often trained with loss functions such as cross-entropy or Dice, which assume pixels to be independent of each other, thus ignoring topological errors and anatomical inconsistencies. We address this limitation by moving from pixel-level to graph representations, which allow to naturally incorporate anatomical constraints by construction. To this end, we introduce HybridGNet, an encoder-decoder neural architecture that leverages standard convolutions for image feature encoding and graph convolutional neural networks (GCNNs) to decode plausible representations of anatomical structures. We also propose a novel image-to-graph skip connection layer which allows localized features to flow from standard convolutional blocks to GCNN blocks, and show that it improves segmentation accuracy. The proposed architecture is extensively evaluated in a variety of domain shift and image occlusion scenarios, and audited considering different types of demographic domain shift. Our comprehensive experimental setup compares HybridGNet with other landmark and pixel-based models for anatomical segmentation in chest x-ray images, and shows that it produces anatomically plausible results in challenging scenarios where other models tend to fail.
ISA-Net: Improved spatial attention network for PET-CT tumor segmentation
Nov 04, 2022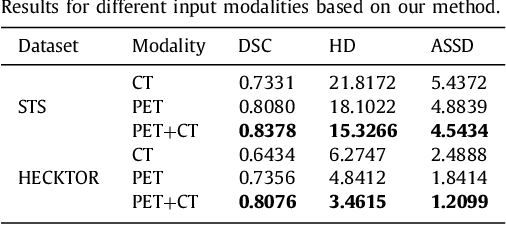
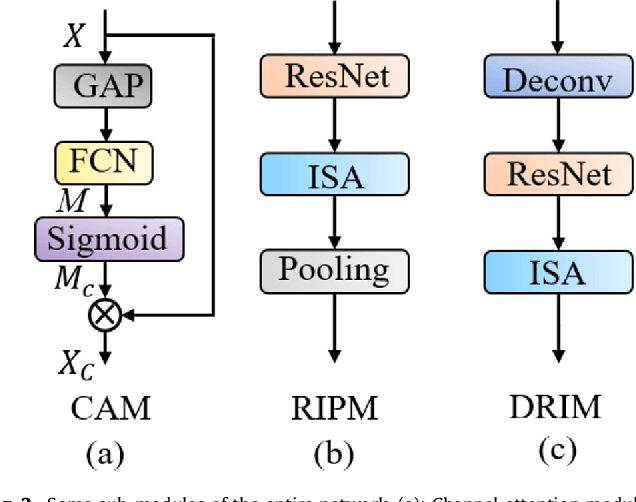
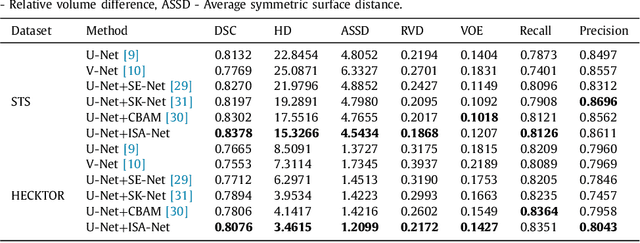
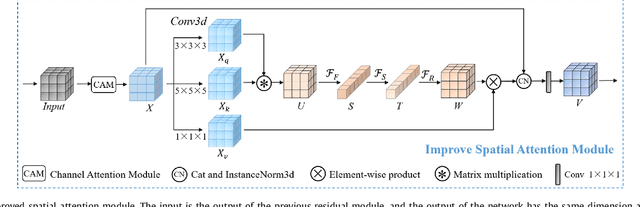
Achieving accurate and automated tumor segmentation plays an important role in both clinical practice and radiomics research. Segmentation in medicine is now often performed manually by experts, which is a laborious, expensive and error-prone task. Manual annotation relies heavily on the experience and knowledge of these experts. In addition, there is much intra- and interobserver variation. Therefore, it is of great significance to develop a method that can automatically segment tumor target regions. In this paper, we propose a deep learning segmentation method based on multimodal positron emission tomography-computed tomography (PET-CT), which combines the high sensitivity of PET and the precise anatomical information of CT. We design an improved spatial attention network(ISA-Net) to increase the accuracy of PET or CT in detecting tumors, which uses multi-scale convolution operation to extract feature information and can highlight the tumor region location information and suppress the non-tumor region location information. In addition, our network uses dual-channel inputs in the coding stage and fuses them in the decoding stage, which can take advantage of the differences and complementarities between PET and CT. We validated the proposed ISA-Net method on two clinical datasets, a soft tissue sarcoma(STS) and a head and neck tumor(HECKTOR) dataset, and compared with other attention methods for tumor segmentation. The DSC score of 0.8378 on STS dataset and 0.8076 on HECKTOR dataset show that ISA-Net method achieves better segmentation performance and has better generalization. Conclusions: The method proposed in this paper is based on multi-modal medical image tumor segmentation, which can effectively utilize the difference and complementarity of different modes. The method can also be applied to other multi-modal data or single-modal data by proper adjustment.
Data Models for Dataset Drift Controls in Machine Learning With Images
Nov 04, 2022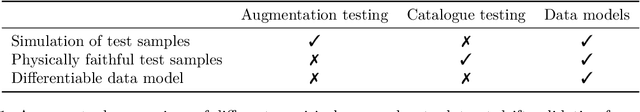


Camera images are ubiquitous in machine learning research. They also play a central role in the delivery of important services spanning medicine and environmental surveying. However, the application of machine learning models in these domains has been limited because of robustness concerns. A primary failure mode are performance drops due to differences between the training and deployment data. While there are methods to prospectively validate the robustness of machine learning models to such dataset drifts, existing approaches do not account for explicit models of the primary object of interest: the data. This makes it difficult to create physically faithful drift test cases or to provide specifications of data models that should be avoided when deploying a machine learning model. In this study, we demonstrate how these shortcomings can be overcome by pairing machine learning robustness validation with physical optics. We examine the role raw sensor data and differentiable data models can play in controlling performance risks related to image dataset drift. The findings are distilled into three applications. First, drift synthesis enables the controlled generation of physically faithful drift test cases. The experiments presented here show that the average decrease in model performance is ten to four times less severe than under post-hoc augmentation testing. Second, the gradient connection between task and data models allows for drift forensics that can be used to specify performance-sensitive data models which should be avoided during deployment of a machine learning model. Third, drift adjustment opens up the possibility for processing adjustments in the face of drift. This can lead to speed up and stabilization of classifier training at a margin of up to 20% in validation accuracy. A guide to access the open code and datasets is available at https://github.com/aiaudit-org/raw2logit.
Deep Learning Neural Network for Lung Cancer Classification: Enhanced Optimization Function
Aug 05, 2022Background and Purpose: Convolutional neural network is widely used for image recognition in the medical area at nowadays. However, overall accuracy in predicting lung tumor is low and the processing time is high as the error occurred while reconstructing the CT image. The aim of this work is to increase the overall prediction accuracy along with reducing processing time by using multispace image in pooling layer of convolution neural network. Methodology: The proposed method has the autoencoder system to improve the overall accuracy, and to predict lung cancer by using multispace image in pooling layer of convolution neural network and Adam Algorithm for optimization. First, the CT images were pre-processed by feeding image to the convolution filter and down sampled by using max pooling. Then, features are extracted using the autoencoder model based on convolutional neural network and multispace image reconstruction technique is used to reduce error while reconstructing the image which then results improved accuracy to predict lung nodule. Finally, the reconstructed images are taken as input for SoftMax classifier to classify the CT images. Results: The state-of-art and proposed solutions were processed in Python Tensor Flow and It provides significant increase in accuracy in classification of lung cancer to 99.5 from 98.9 and decrease in processing time from 10 frames/second to 12 seconds/second. Conclusion: The proposed solution provides high classification accuracy along with less processing time compared to the state of art. For future research, large dataset can be implemented, and low pixel image can be processed to evaluate the classification
* 22pages
Caption supervision enables robust learners
Oct 13, 2022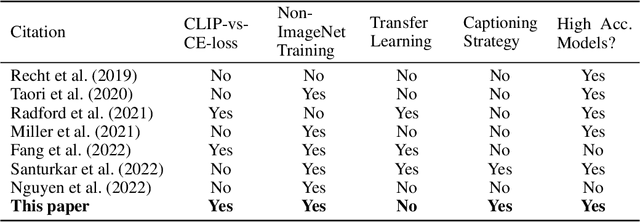
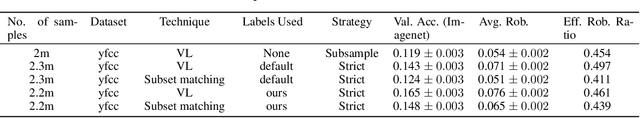
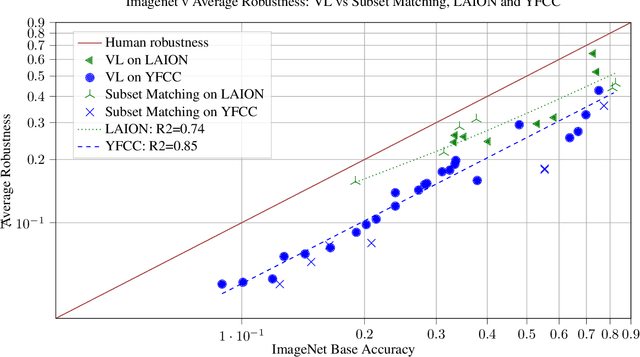


Vision language models like CLIP are robust to natural distribution shifts, in part because CLIP learns on unstructured data using a technique called caption supervision; the model inteprets image-linked texts as ground-truth labels. In a carefully controlled comparison study, we show that CNNs trained on a standard cross-entropy loss can also benefit from caption supervision, in some cases even more than VL models, on the same data. To facilitate future experiments with high-accuracy caption-supervised models, we introduce CaptionNet (https://github.com/penfever/CaptionNet/), which includes a class-balanced, fully supervised dataset with over 50,000 new human-labeled ImageNet-compliant samples which includes web-scraped captions. In a series of experiments on CaptionNet, we show how the choice of loss function, data filtration and supervision strategy enable robust computer vision. We also provide the codebase necessary to reproduce our experiments at https://github.com/penfever/vlhub/
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge