"Image": models, code, and papers
SeaDroneSim: Simulation of Aerial Images for Detection of Objects Above Water
Oct 31, 2022
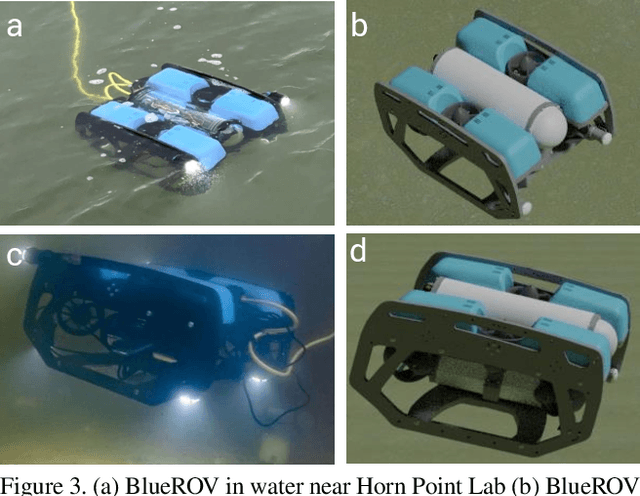
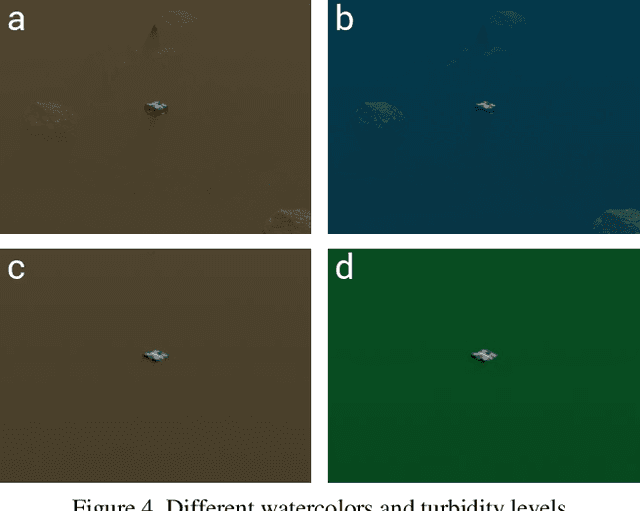



Unmanned Aerial Vehicles (UAVs) are known for their fast and versatile applicability. With UAVs' growth in availability and applications, they are now of vital importance in serving as technological support in search-and-rescue(SAR) operations in marine environments. High-resolution cameras and GPUs can be equipped on the UAVs to provide effective and efficient aid to emergency rescue operations. With modern computer vision algorithms, we can detect objects for aiming such rescue missions. However, these modern computer vision algorithms are dependent on numerous amounts of training data from UAVs, which is time-consuming and labor-intensive for maritime environments. To this end, we present a new benchmark suite, SeaDroneSim, that can be used to create photo-realistic aerial image datasets with the ground truth for segmentation masks of any given object. Utilizing only the synthetic data generated from SeaDroneSim, we obtain 71 mAP on real aerial images for detecting BlueROV as a feasibility study. This result from the new simulation suit also serves as a baseline for the detection of BlueROV.
TW-BAG: Tensor-wise Brain-aware Gate Network for Inpainting Disrupted Diffusion Tensor Imaging
Oct 31, 2022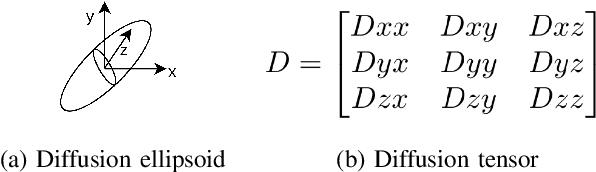
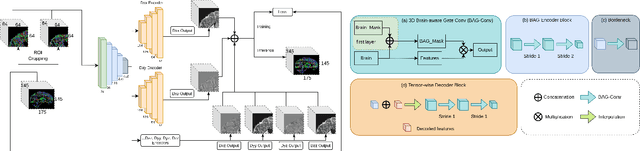
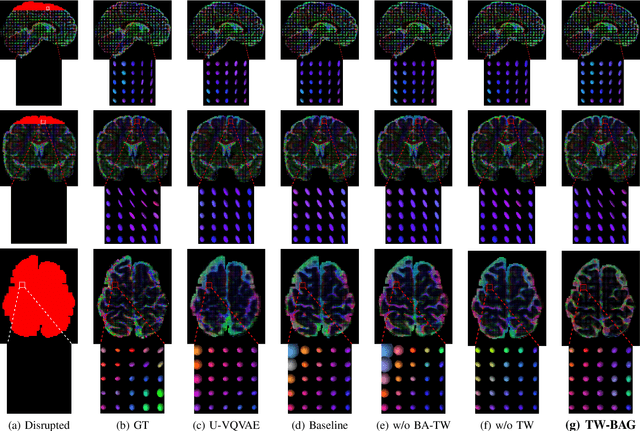
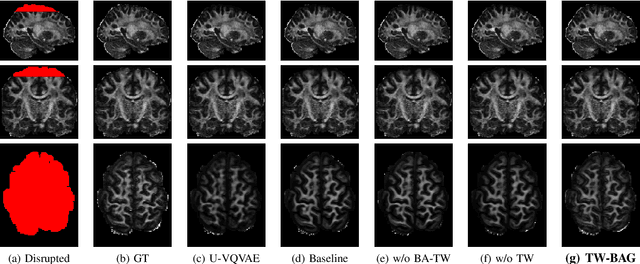
Diffusion Weighted Imaging (DWI) is an advanced imaging technique commonly used in neuroscience and neurological clinical research through a Diffusion Tensor Imaging (DTI) model. Volumetric scalar metrics including fractional anisotropy, mean diffusivity, and axial diffusivity can be derived from the DTI model to summarise water diffusivity and other quantitative microstructural information for clinical studies. However, clinical practice constraints can lead to sub-optimal DWI acquisitions with missing slices (either due to a limited field of view or the acquisition of disrupted slices). To avoid discarding valuable subjects for group-wise studies, we propose a novel 3D Tensor-Wise Brain-Aware Gate network (TW-BAG) for inpainting disrupted DTIs. The proposed method is tailored to the problem with a dynamic gate mechanism and independent tensor-wise decoders. We evaluated the proposed method on the publicly available Human Connectome Project (HCP) dataset using common image similarity metrics derived from the predicted tensors and scalar DTI metrics. Our experimental results show that the proposed approach can reconstruct the original brain DTI volume and recover relevant clinical imaging information.
Is Facial Recognition Biased at Near-Infrared Spectrum As Well?
Oct 31, 2022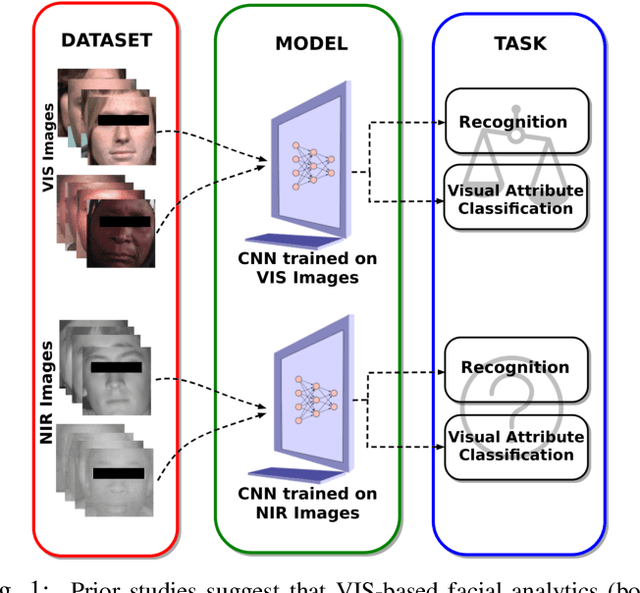
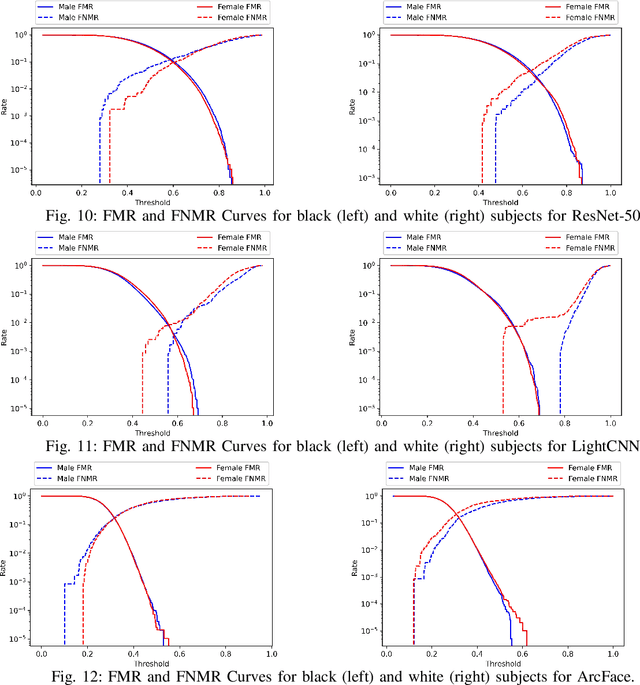
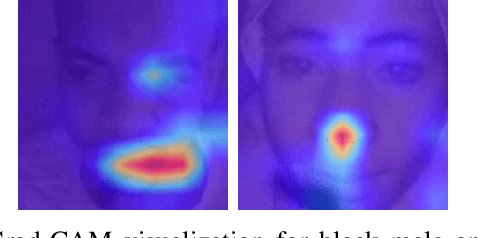
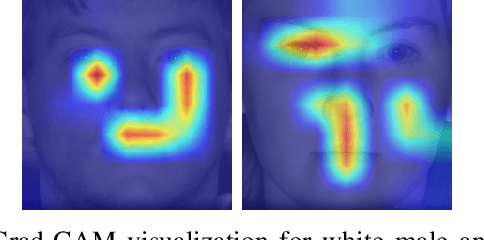
Published academic research and media articles suggest face recognition is biased across demographics. Specifically, unequal performance is obtained for women, dark-skinned people, and older adults. However, these published studies have examined the bias of facial recognition in the visible spectrum (VIS). Factors such as facial makeup, facial hair, skin color, and illumination variation have been attributed to the bias of this technology at the VIS. The near-infrared (NIR) spectrum offers an advantage over the VIS in terms of robustness to factors such as illumination changes, facial makeup, and skin color. Therefore, it is worthwhile to investigate the bias of facial recognition at the near-infrared spectrum (NIR). This first study investigates the bias of the face recognition systems at the NIR spectrum. To this aim, two popular NIR facial image datasets namely, CASIA-Face-Africa and Notre-Dame-NIVL consisting of African and Caucasian subjects, respectively, are used to investigate the bias of facial recognition technology across gender and race. Interestingly, experimental results suggest equitable face recognition performance across gender and race at the NIR spectrum.
Towards a Guideline for Evaluation Metrics in Medical Image Segmentation
Feb 10, 2022
In the last decade, research on artificial intelligence has seen rapid growth with deep learning models, especially in the field of medical image segmentation. Various studies demonstrated that these models have powerful prediction capabilities and achieved similar results as clinicians. However, recent studies revealed that the evaluation in image segmentation studies lacks reliable model performance assessment and showed statistical bias by incorrect metric implementation or usage. Thus, this work provides an overview and interpretation guide on the following metrics for medical image segmentation evaluation in binary as well as multi-class problems: Dice similarity coefficient, Jaccard, Sensitivity, Specificity, Rand index, ROC curves, Cohen's Kappa, and Hausdorff distance. As a summary, we propose a guideline for standardized medical image segmentation evaluation to improve evaluation quality, reproducibility, and comparability in the research field.
Enhancement of Novel View Synthesis Using Omnidirectional Image Completion
Mar 18, 2022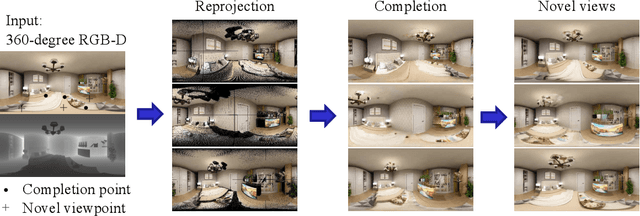
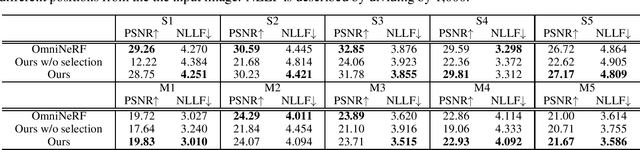
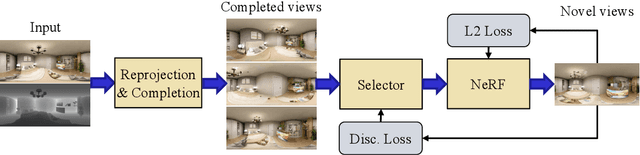
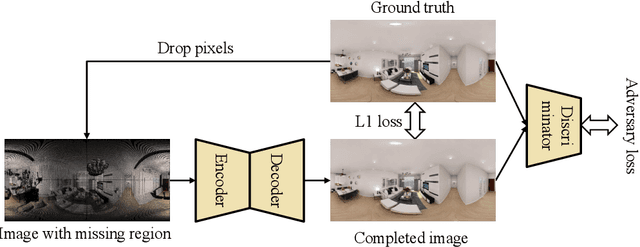
We present a method for synthesizing novel views from a single 360-degree image based on the neural radiance field (NeRF) . Prior studies rely on the neighborhood interpolation capability of multi-layer perceptrons to complete missing regions caused by occlusion and zooming, and this leads to artifacts. In the proposed method, the input image is reprojected to 360-degree images at other camera positions, the missing regions of the reprojected images are completed by a self-supervised trained generative model, and the completed images are utilized to train the NeRF. Because multiple completed images contain inconsistencies in 3D, we introduce a method to train NeRF while dynamically selecting a sparse set of completed images, to reduce the discrimination error of the synthesized views with real images. Experiments indicate that the proposed method can synthesize plausible novel views while preserving the features of the scene for both artificial and real-world data.
HealthyGAN: Learning from Unannotated Medical Images to Detect Anomalies Associated with Human Disease
Sep 05, 2022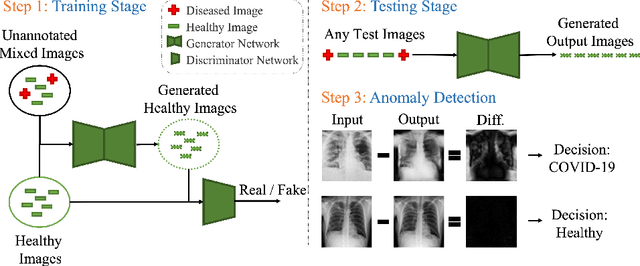
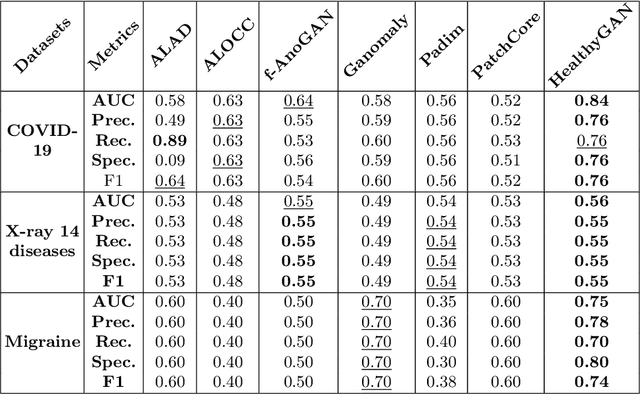
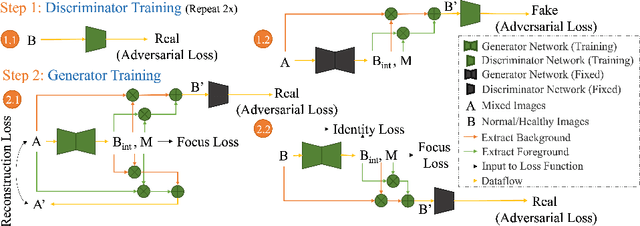
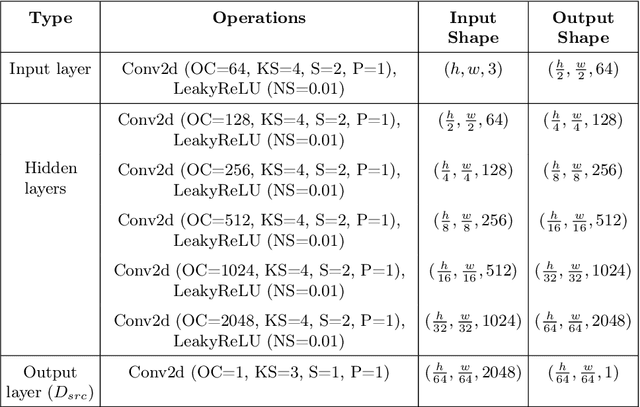
Automated anomaly detection from medical images, such as MRIs and X-rays, can significantly reduce human effort in disease diagnosis. Owing to the complexity of modeling anomalies and the high cost of manual annotation by domain experts (e.g., radiologists), a typical technique in the current medical imaging literature has focused on deriving diagnostic models from healthy subjects only, assuming the model will detect the images from patients as outliers. However, in many real-world scenarios, unannotated datasets with a mix of both healthy and diseased individuals are abundant. Therefore, this paper poses the research question of how to improve unsupervised anomaly detection by utilizing (1) an unannotated set of mixed images, in addition to (2) the set of healthy images as being used in the literature. To answer the question, we propose HealthyGAN, a novel one-directional image-to-image translation method, which learns to translate the images from the mixed dataset to only healthy images. Being one-directional, HealthyGAN relaxes the requirement of cycle consistency of existing unpaired image-to-image translation methods, which is unattainable with mixed unannotated data. Once the translation is learned, we generate a difference map for any given image by subtracting its translated output. Regions of significant responses in the difference map correspond to potential anomalies (if any). Our HealthyGAN outperforms the conventional state-of-the-art methods by significant margins on two publicly available datasets: COVID-19 and NIH ChestX-ray14, and one institutional dataset collected from Mayo Clinic. The implementation is publicly available at https://github.com/mahfuzmohammad/HealthyGAN.
MicroISP: Processing 32MP Photos on Mobile Devices with Deep Learning
Nov 08, 2022
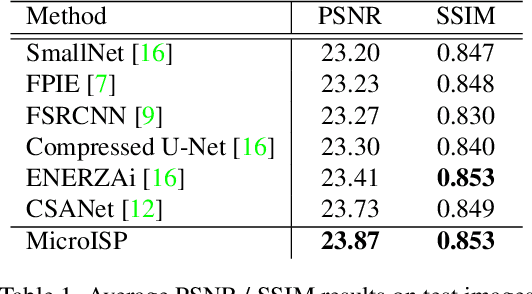
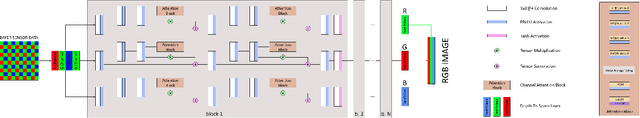
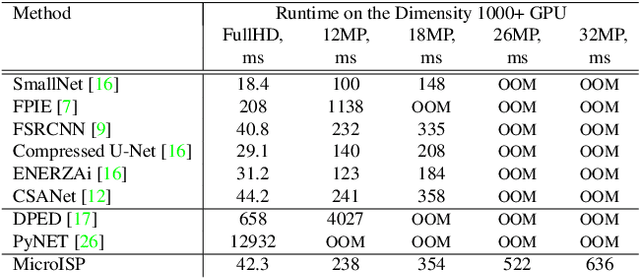
While neural networks-based photo processing solutions can provide a better image quality compared to the traditional ISP systems, their application to mobile devices is still very limited due to their very high computational complexity. In this paper, we present a novel MicroISP model designed specifically for edge devices, taking into account their computational and memory limitations. The proposed solution is capable of processing up to 32MP photos on recent smartphones using the standard mobile ML libraries and requiring less than 1 second to perform the inference, while for FullHD images it achieves real-time performance. The architecture of the model is flexible, allowing to adjust its complexity to devices of different computational power. To evaluate the performance of the model, we collected a novel Fujifilm UltraISP dataset consisting of thousands of paired photos captured with a normal mobile camera sensor and a professional 102MP medium-format FujiFilm GFX100 camera. The experiments demonstrated that, despite its compact size, the MicroISP model is able to provide comparable or better visual results than the traditional mobile ISP systems, while outperforming the previously proposed efficient deep learning based solutions. Finally, this model is also compatible with the latest mobile AI accelerators, achieving good runtime and low power consumption on smartphone NPUs and APUs. The code, dataset and pre-trained models are available on the project website: https://people.ee.ethz.ch/~ihnatova/microisp.html
Rethinking Hierarchies in Pre-trained Plain Vision Transformer
Nov 08, 2022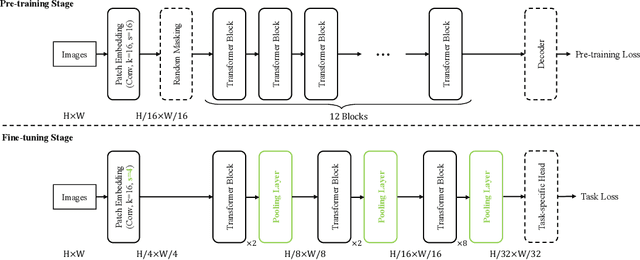

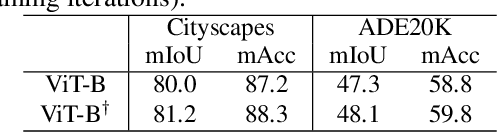
Self-supervised pre-training vision transformer (ViT) via masked image modeling (MIM) has been proven very effective. However, customized algorithms should be carefully designed for the hierarchical ViTs, e.g., GreenMIM, instead of using the vanilla and simple MAE for the plain ViT. More importantly, since these hierarchical ViTs cannot reuse the off-the-shelf pre-trained weights of the plain ViTs, the requirement of pre-training them leads to a massive amount of computational cost, thereby incurring both algorithmic and computational complexity. In this paper, we address this problem by proposing a novel idea of disentangling the hierarchical architecture design from the self-supervised pre-training. We transform the plain ViT into a hierarchical one with minimal changes. Technically, we change the stride of linear embedding layer from 16 to 4 and add convolution (or simple average) pooling layers between the transformer blocks, thereby reducing the feature size from 1/4 to 1/32 sequentially. Despite its simplicity, it outperforms the plain ViT baseline in classification, detection, and segmentation tasks on ImageNet, MS COCO, Cityscapes, and ADE20K benchmarks, respectively. We hope this preliminary study could draw more attention from the community on developing effective (hierarchical) ViTs while avoiding the pre-training cost by leveraging the off-the-shelf checkpoints. The code and models will be released at https://github.com/ViTAE-Transformer/HPViT.
Privacy Meets Explainability: A Comprehensive Impact Benchmark
Nov 08, 2022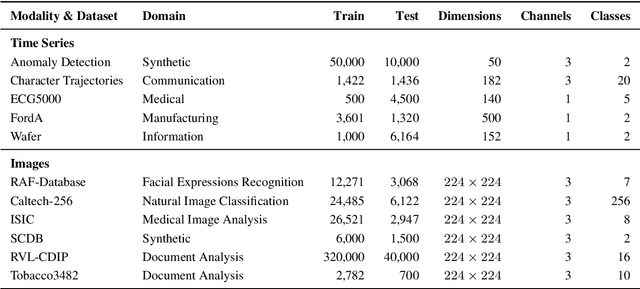
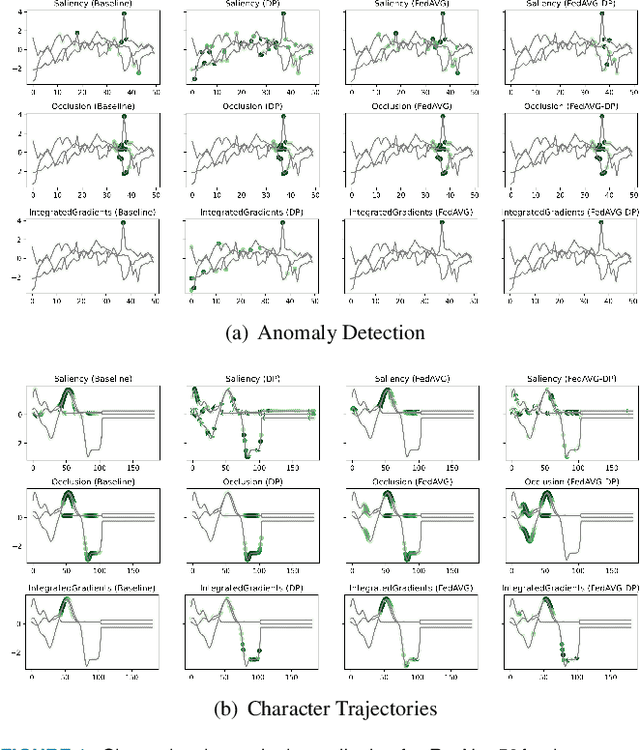
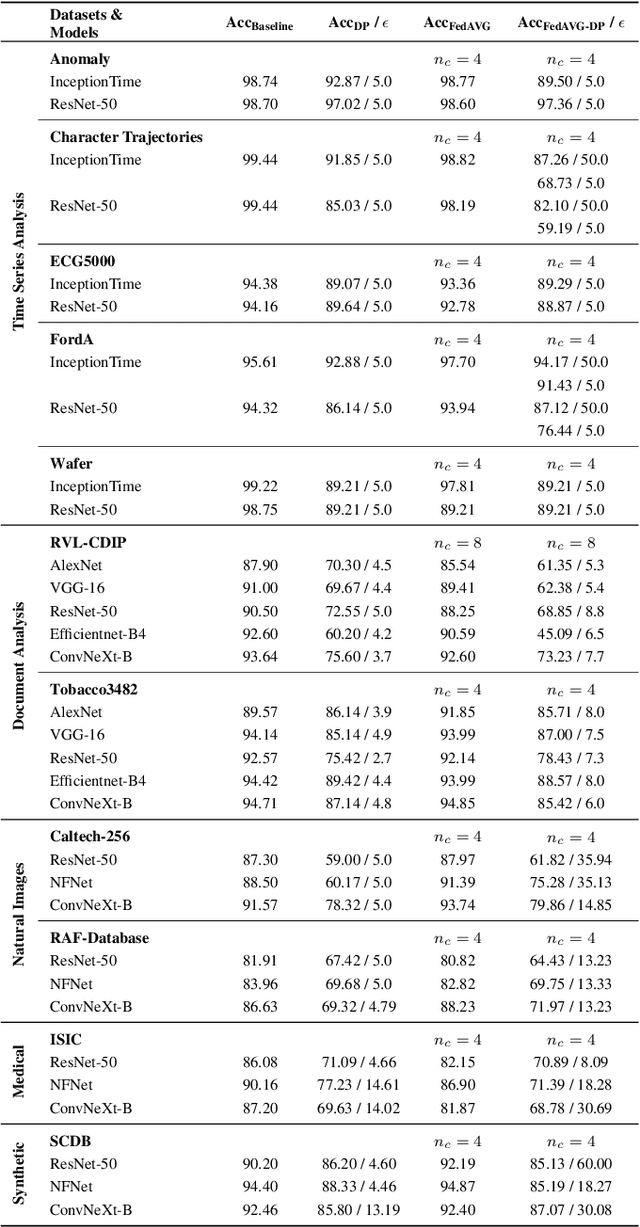

Since the mid-10s, the era of Deep Learning (DL) has continued to this day, bringing forth new superlatives and innovations each year. Nevertheless, the speed with which these innovations translate into real applications lags behind this fast pace. Safety-critical applications, in particular, underlie strict regulatory and ethical requirements which need to be taken care of and are still active areas of debate. eXplainable AI (XAI) and privacy-preserving machine learning (PPML) are both crucial research fields, aiming at mitigating some of the drawbacks of prevailing data-hungry black-box models in DL. Despite brisk research activity in the respective fields, no attention has yet been paid to their interaction. This work is the first to investigate the impact of private learning techniques on generated explanations for DL-based models. In an extensive experimental analysis covering various image and time series datasets from multiple domains, as well as varying privacy techniques, XAI methods, and model architectures, the effects of private training on generated explanations are studied. The findings suggest non-negligible changes in explanations through the introduction of privacy. Apart from reporting individual effects of PPML on XAI, the paper gives clear recommendations for the choice of techniques in real applications. By unveiling the interdependencies of these pivotal technologies, this work is a first step towards overcoming the remaining hurdles for practically applicable AI in safety-critical domains.
CDNet: Contrastive Disentangled Network for Fine-Grained Image Categorization of Ocular B-Scan Ultrasound
Jun 17, 2022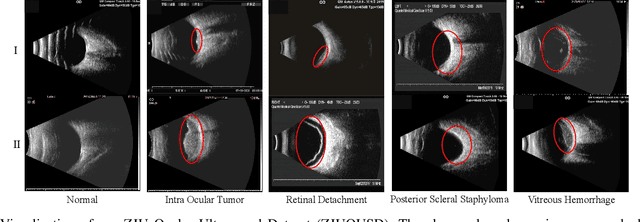
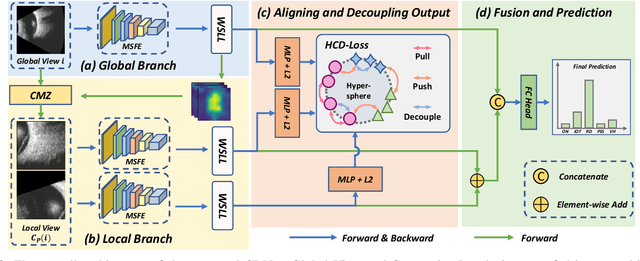

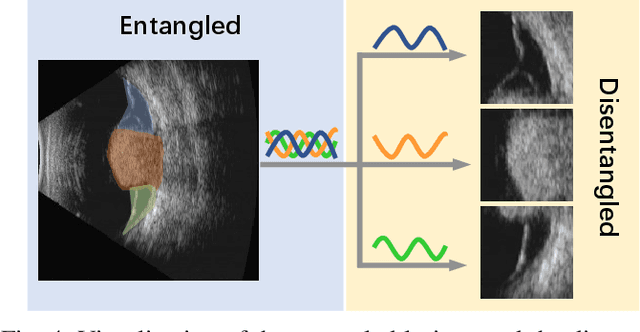
Precise and rapid categorization of images in the B-scan ultrasound modality is vital for diagnosing ocular diseases. Nevertheless, distinguishing various diseases in ultrasound still challenges experienced ophthalmologists. Thus a novel contrastive disentangled network (CDNet) is developed in this work, aiming to tackle the fine-grained image categorization (FGIC) challenges of ocular abnormalities in ultrasound images, including intraocular tumor (IOT), retinal detachment (RD), posterior scleral staphyloma (PSS), and vitreous hemorrhage (VH). Three essential components of CDNet are the weakly-supervised lesion localization module (WSLL), contrastive multi-zoom (CMZ) strategy, and hyperspherical contrastive disentangled loss (HCD-Loss), respectively. These components facilitate feature disentanglement for fine-grained recognition in both the input and output aspects. The proposed CDNet is validated on our ZJU Ocular Ultrasound Dataset (ZJUOUSD), consisting of 5213 samples. Furthermore, the generalization ability of CDNet is validated on two public and widely-used chest X-ray FGIC benchmarks. Quantitative and qualitative results demonstrate the efficacy of our proposed CDNet, which achieves state-of-the-art performance in the FGIC task. Code is available at: https://github.com/ZeroOneGame/CDNet-for-OUS-FGIC .
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge