"Image": models, code, and papers
LapGM: A Multisequence MR Bias Correction and Normalization Model
Sep 27, 2022
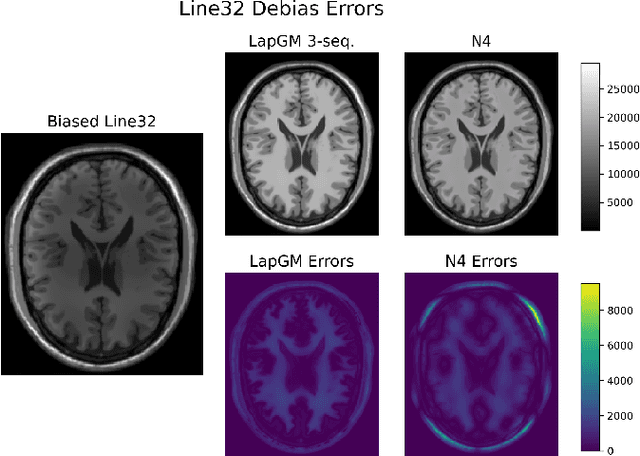
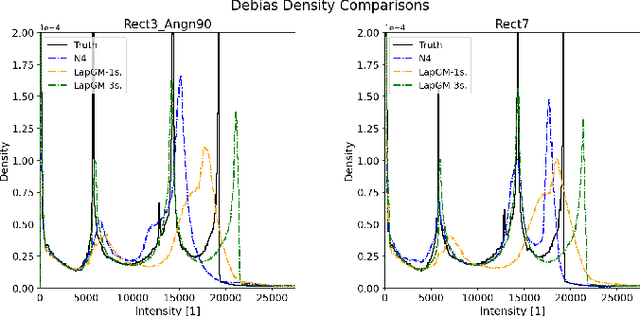
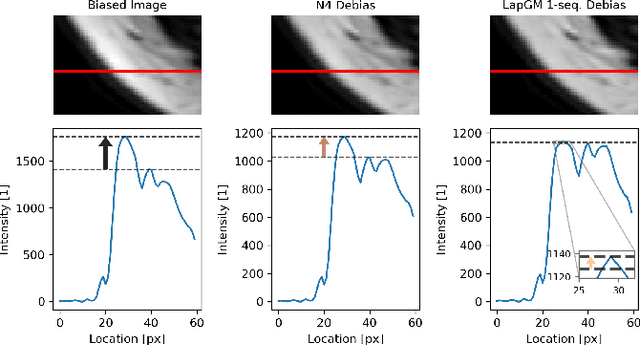
A spatially regularized Gaussian mixture model, LapGM, is proposed for the bias field correction and magnetic resonance normalization problem. The proposed spatial regularizer gives practitioners fine-tuned control between balancing bias field removal and preserving image contrast preservation for multi-sequence, magnetic resonance images. The fitted Gaussian parameters of LapGM serve as control values which can be used to normalize image intensities across different patient scans. LapGM is compared to well-known debiasing algorithm N4ITK in both the single and multi-sequence setting. As a normalization procedure, LapGM is compared to known techniques such as: max normalization, Z-score normalization, and a water-masked region-of-interest normalization. Lastly a CUDA-accelerated Python package $\texttt{lapgm}$ is provided from the authors for use.
Solving 3D Inverse Problems using Pre-trained 2D Diffusion Models
Nov 19, 2022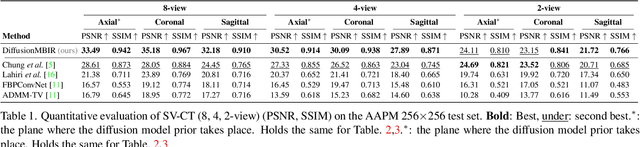
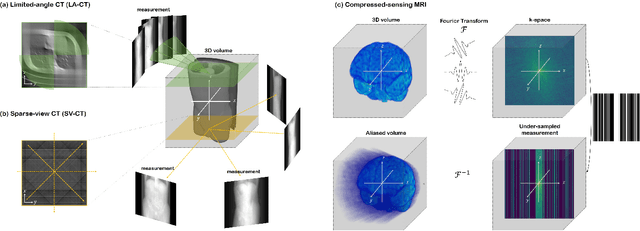
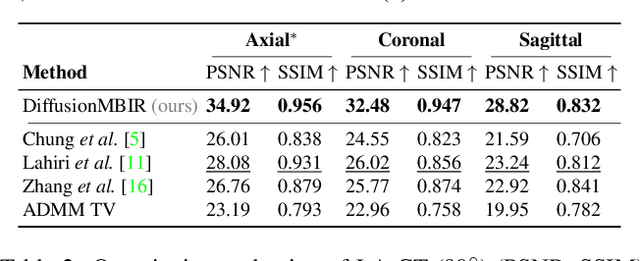
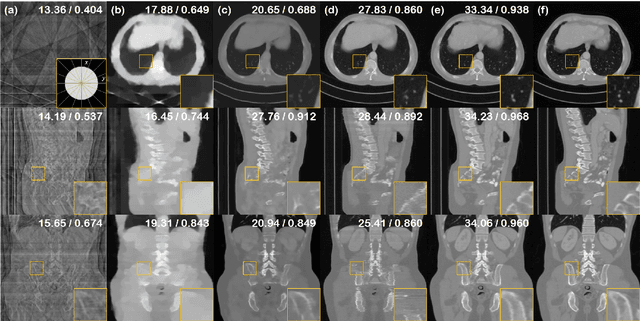
Diffusion models have emerged as the new state-of-the-art generative model with high quality samples, with intriguing properties such as mode coverage and high flexibility. They have also been shown to be effective inverse problem solvers, acting as the prior of the distribution, while the information of the forward model can be granted at the sampling stage. Nonetheless, as the generative process remains in the same high dimensional (i.e. identical to data dimension) space, the models have not been extended to 3D inverse problems due to the extremely high memory and computational cost. In this paper, we combine the ideas from the conventional model-based iterative reconstruction with the modern diffusion models, which leads to a highly effective method for solving 3D medical image reconstruction tasks such as sparse-view tomography, limited angle tomography, compressed sensing MRI from pre-trained 2D diffusion models. In essence, we propose to augment the 2D diffusion prior with a model-based prior in the remaining direction at test time, such that one can achieve coherent reconstructions across all dimensions. Our method can be run in a single commodity GPU, and establishes the new state-of-the-art, showing that the proposed method can perform reconstructions of high fidelity and accuracy even in the most extreme cases (e.g. 2-view 3D tomography). We further reveal that the generalization capacity of the proposed method is surprisingly high, and can be used to reconstruct volumes that are entirely different from the training dataset.
Discriminative Kernel Convolution Network for Multi-Label Ophthalmic Disease Detection on Imbalanced Fundus Image Dataset
Jul 16, 2022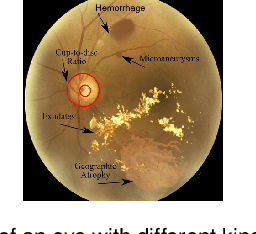
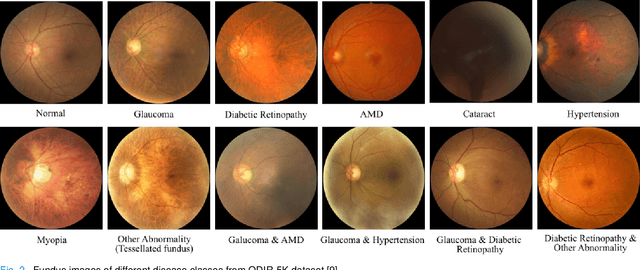
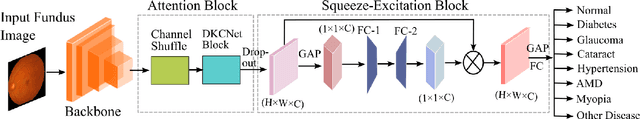
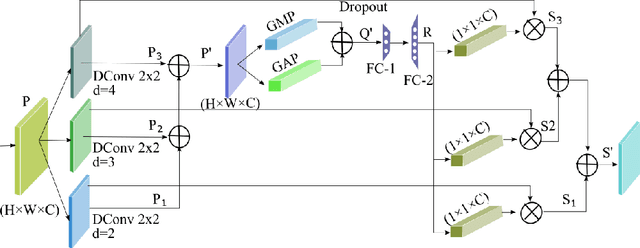
It is feasible to recognize the presence and seriousness of eye disease by investigating the progressions in retinal biological structure. Fundus examination is a diagnostic procedure to examine the biological structure and anomaly of the eye. Ophthalmic diseases like glaucoma, diabetic retinopathy, and cataract are the main reason for visual impairment around the world. Ocular Disease Intelligent Recognition (ODIR-5K) is a benchmark structured fundus image dataset utilized by researchers for multi-label multi-disease classification of fundus images. This work presents a discriminative kernel convolution network (DKCNet), which explores discriminative region-wise features without adding extra computational cost. DKCNet is composed of an attention block followed by a squeeze and excitation (SE) block. The attention block takes features from the backbone network and generates discriminative feature attention maps. The SE block takes the discriminative feature maps and improves channel interdependencies. Better performance of DKCNet is observed with InceptionResnet backbone network for multi-label classification of ODIR-5K fundus images with 96.08 AUC, 94.28 F1-score and 0.81 kappa score. The proposed method splits the common target label for an eye pair based on the diagnostic keyword. Based on these labels oversampling and undersampling is done to resolve class imbalance. To check the biasness of proposed model towards training data, the model trained on ODIR dataset is tested on three publicly available benchmark datasets. It is found to give good performance on completely unseen fundus images also.
A Guide to Image and Video based Small Object Detection using Deep Learning : Case Study of Maritime Surveillance
Jul 26, 2022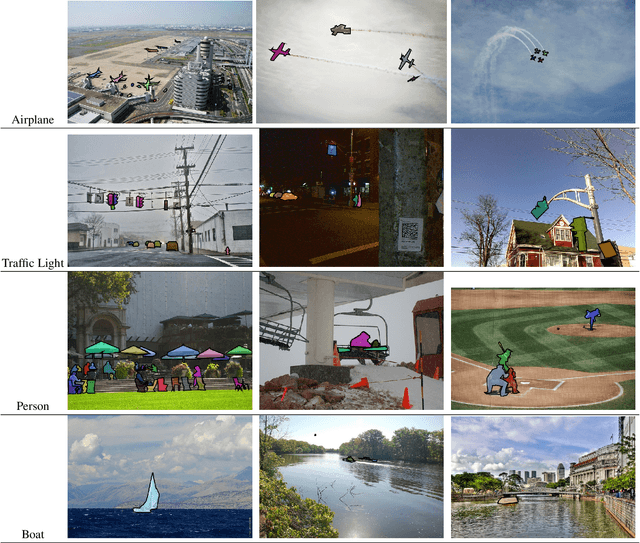
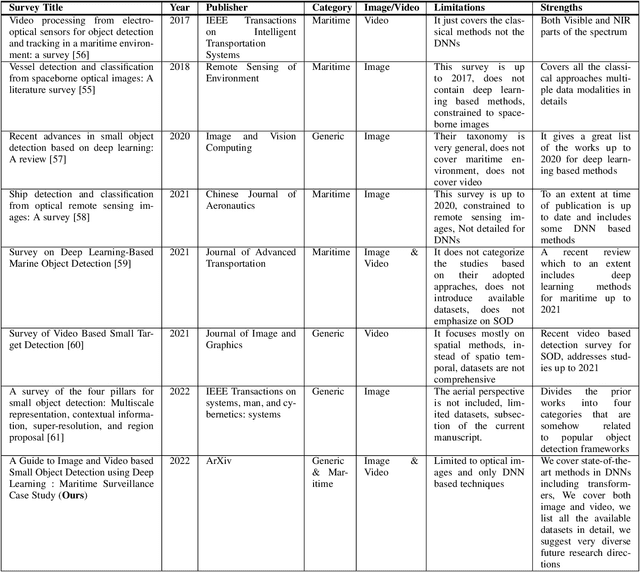
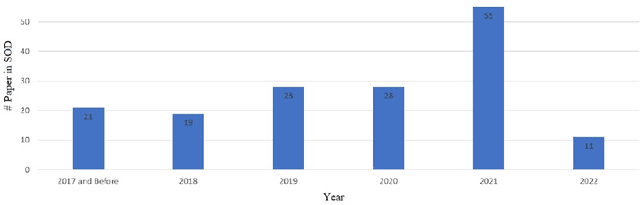
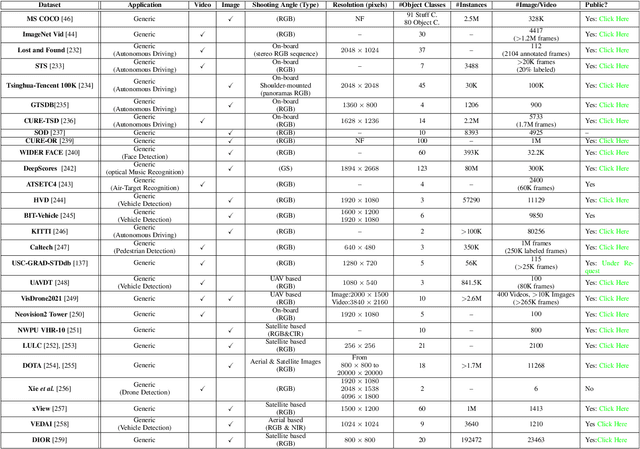
Small object detection (SOD) in optical images and videos is a challenging problem that even state-of-the-art generic object detection methods fail to accurately localize and identify such objects. Typically, small objects appear in real-world due to large camera-object distance. Because small objects occupy only a small area in the input image (e.g., less than 10%), the information extracted from such a small area is not always rich enough to support decision making. Multidisciplinary strategies are being developed by researchers working at the interface of deep learning and computer vision to enhance the performance of SOD deep learning based methods. In this paper, we provide a comprehensive review of over 160 research papers published between 2017 and 2022 in order to survey this growing subject. This paper summarizes the existing literature and provide a taxonomy that illustrates the broad picture of current research. We investigate how to improve the performance of small object detection in maritime environments, where increasing performance is critical. By establishing a connection between generic and maritime SOD research, future directions have been identified. In addition, the popular datasets that have been used for SOD for generic and maritime applications are discussed, and also well-known evaluation metrics for the state-of-the-art methods on some of the datasets are provided.
SUPRA: Superpixel Guided Loss for Improved Multi-modal Segmentation in Endoscopy
Nov 12, 2022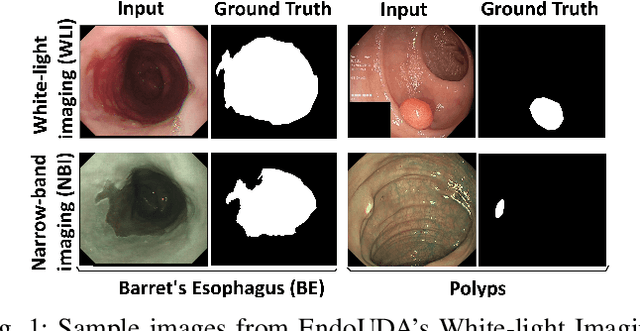
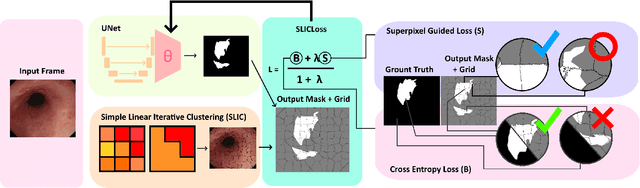
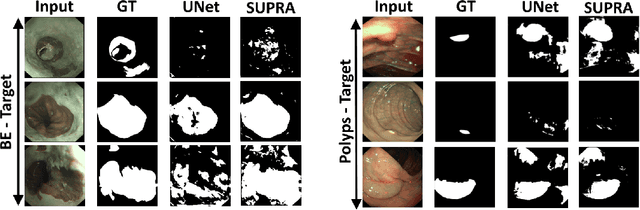
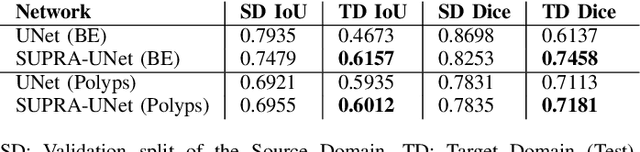
Domain shift is a well-known problem in the medical imaging community. In particular, for endoscopic image analysis where the data can have different modalities the performance of deep learning (DL) methods gets adversely affected. In other words, methods developed on one modality cannot be used for a different modality. However, in real clinical settings, endoscopists switch between modalities for better mucosal visualisation. In this paper, we explore the domain generalisation technique to enable DL methods to be used in such scenarios. To this extend, we propose to use super pixels generated with Simple Linear Iterative Clustering (SLIC) which we refer to as "SUPRA" for SUPeRpixel Augmented method. SUPRA first generates a preliminary segmentation mask making use of our new loss "SLICLoss" that encourages both an accurate and color-consistent segmentation. We demonstrate that SLICLoss when combined with Binary Cross Entropy loss (BCE) can improve the model's generalisability with data that presents significant domain shift. We validate this novel compound loss on a vanilla U-Net using the EndoUDA dataset, which contains images for Barret's Esophagus and polyps from two modalities. We show that our method yields an improvement of nearly 25% in the target domain set compared to the baseline.
Tables to LaTeX: structure and content extraction from scientific tables
Oct 31, 2022Scientific documents contain tables that list important information in a concise fashion. Structure and content extraction from tables embedded within PDF research documents is a very challenging task due to the existence of visual features like spanning cells and content features like mathematical symbols and equations. Most existing table structure identification methods tend to ignore these academic writing features. In this paper, we adapt the transformer-based language modeling paradigm for scientific table structure and content extraction. Specifically, the proposed model converts a tabular image to its corresponding LaTeX source code. Overall, we outperform the current state-of-the-art baselines and achieve an exact match accuracy of 70.35 and 49.69% on table structure and content extraction, respectively. Further analysis demonstrates that the proposed models efficiently identify the number of rows and columns, the alphanumeric characters, the LaTeX tokens, and symbols.
Language Models with Image Descriptors are Strong Few-Shot Video-Language Learners
May 24, 2022
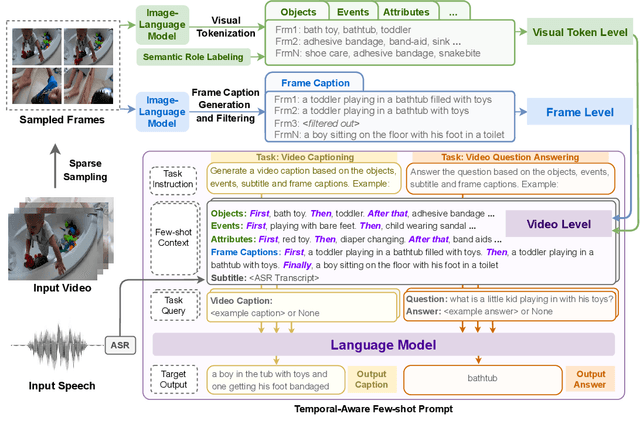
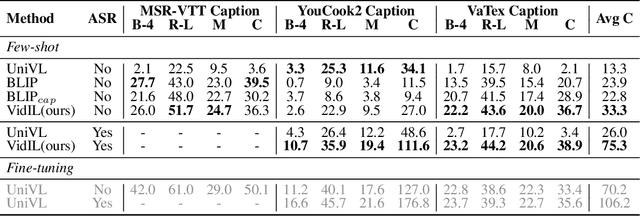

The goal of this work is to build flexible video-language models that can generalize to various video-to-text tasks from few examples, such as domain-specific captioning, question answering, and future event prediction. Existing few-shot video-language learners focus exclusively on the encoder, resulting in the absence of a video-to-text decoder to handle generative tasks. Video captioners have been pretrained on large-scale video-language datasets, but they rely heavily on finetuning and lack the ability to generate text for unseen tasks in a few-shot setting. We propose VidIL, a few-shot Video-language Learner via Image and Language models, which demonstrates strong performance on few-shot video-to-text tasks without the necessity of pretraining or finetuning on any video datasets. We use the image-language models to translate the video content into frame captions, object, attribute, and event phrases, and compose them into a temporal structure template. We then instruct a language model, with a prompt containing a few in-context examples, to generate a target output from the composed content. The flexibility of prompting allows the model to capture any form of text input, such as automatic speech recognition (ASR) transcripts. Our experiments demonstrate the power of language models in understanding videos on a wide variety of video-language tasks, including video captioning, video question answering, video caption retrieval, and video future event prediction. Especially, on video future event prediction, our few-shot model significantly outperforms state-of-the-art supervised models trained on large-scale video datasets. Code and resources are publicly available for research purposes at https://github.com/MikeWangWZHL/VidIL .
Transformer-based SAR Image Despeckling
Jan 23, 2022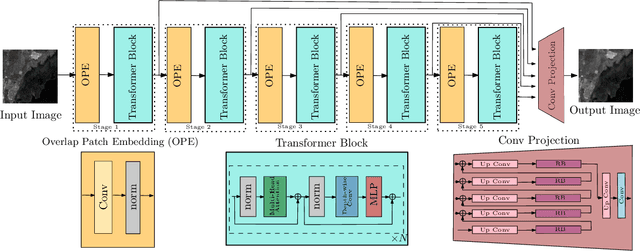
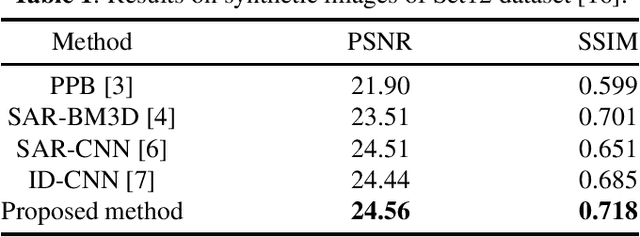
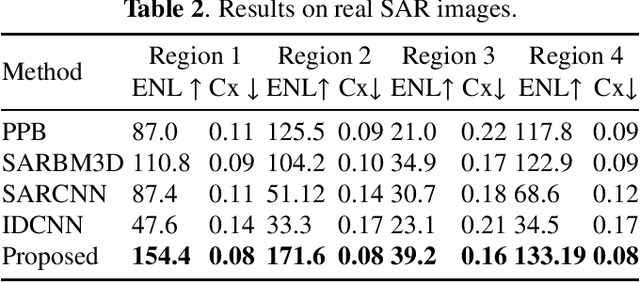
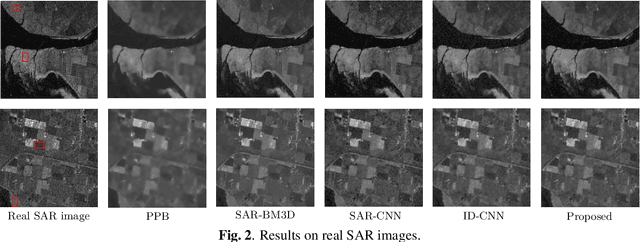
Synthetic Aperture Radar (SAR) images are usually degraded by a multiplicative noise known as speckle which makes processing and interpretation of SAR images difficult. In this paper, we introduce a transformer-based network for SAR image despeckling. The proposed despeckling network comprises of a transformer-based encoder which allows the network to learn global dependencies between different image regions - aiding in better despeckling. The network is trained end-to-end with synthetically generated speckled images using a composite loss function. Experiments show that the proposed method achieves significant improvements over traditional and convolutional neural network-based despeckling methods on both synthetic and real SAR images.
Deep Convolutional Pooling Transformer for Deepfake Detection
Sep 12, 2022



Recently, Deepfake has drawn considerable public attention due to security and privacy concerns in social media digital forensics. As the wildly spreading Deepfake videos on the Internet become more realistic, traditional detection techniques have failed in distinguishing between the real and fake. Most existing deep learning methods mainly focus on local features and relations within the face image using convolutional neural networks as a backbone. However, local features and relations are insufficient for model training to learn enough general information for Deepfake detection. Therefore, the existing Deepfake detection methods have reached a bottleneck to further improving the detection performance. To address this issue, we propose a deep convolutional Transformer to incorporate the decisive image features both locally and globally. Specifically, we apply convolutional pooling and re-attention to enrich the extracted features and enhance the efficacy. Moreover, we employ the barely discussed image keyframes in model training for performance improvement and visualize the feature quantity gap between the key and normal image frames caused by video compression. We finally illustrate the transferability with extensive experiments on several Deepfake benchmark datasets. The proposed solution consistently outperforms several state-of-the-art baselines on both within- and cross-dataset experiments.
Imrpoving Strain Estimation in Breast Ultrasound Images Using Novel 1.5D Approach (Simulation and In-vivo results
Oct 13, 2022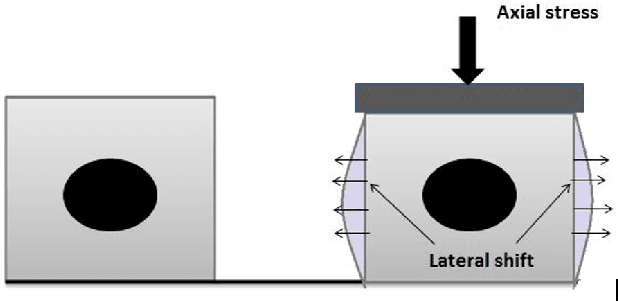
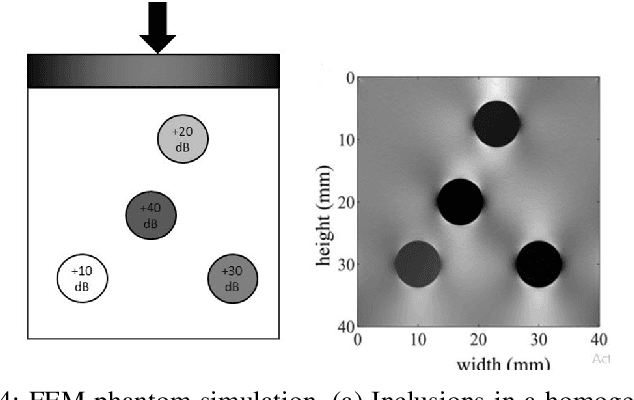
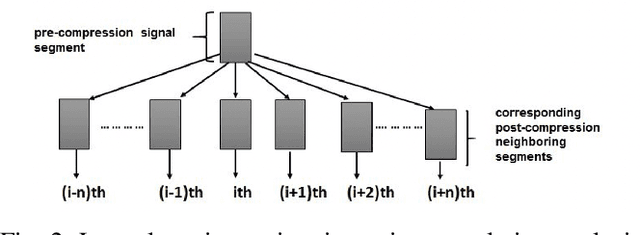
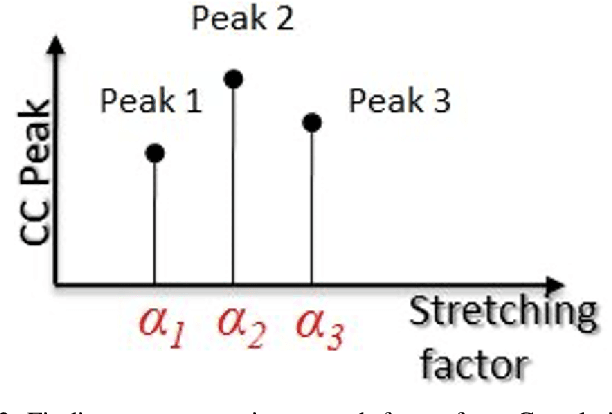

Ultrasound elastography is the method to image the elasticity of compliant tissues due to a mechanical compression applied to it. In elastography, the local strain of explored tissue is estimated by analyzing the echo signals. This is accomplished by a sonographer who uses ultrasound transducer to apply pressure on the tissue area causing displacement of the tissue. A set of data is obtained before and after the compression. This stress along the axial direction not only causes deformation in that direction but it can also cause lateral deformation in the tissue. This relative deformation is later calculated using tracking algorithms. Finally, estimating the elasticity, images are formed. In this paper, we introduce a novel strain estimator. The proposed 1.5D strain estimator uses 1D windows for swift computations and searches the lateral direction for taking the non-axial movement into account. We have investigated the performance of our estimator using simulated and in-vivo data of breast tissue. The performance of our method is verified by comparing with different methods up to 16\% applied strain. The proposed estimator has exhibited a superior performance compared to the conventional methods. The image quality significantly improves, especially in the presence of higher applied strain, when the non-axial motion is significant.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge