"Image": models, code, and papers
Histo-Genomic Knowledge Distillation For Cancer Prognosis From Histopathology Whole Slide Images
Mar 18, 2024
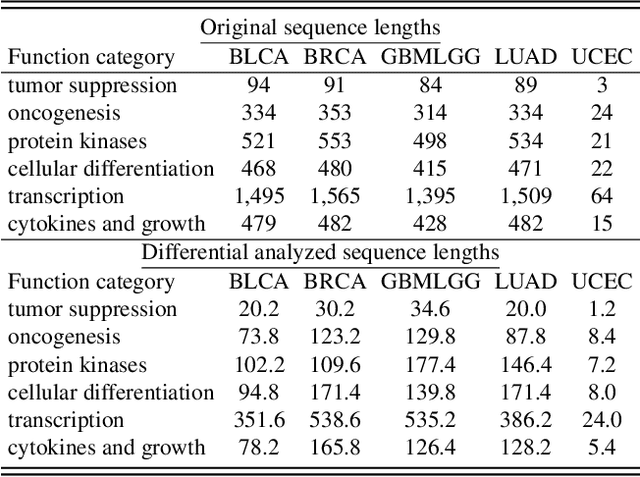
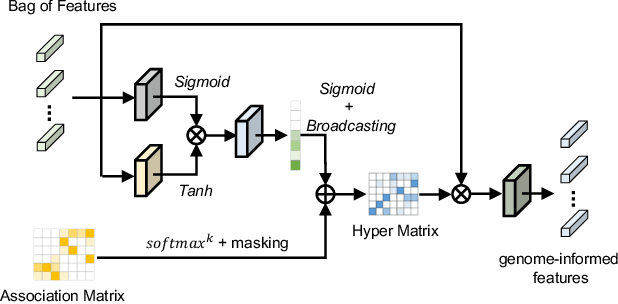
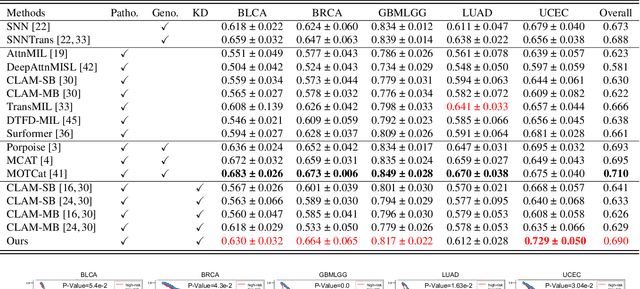
Histo-genomic multi-modal methods have recently emerged as a powerful paradigm, demonstrating significant potential for improving cancer prognosis. However, genome sequencing, unlike histopathology imaging, is still not widely accessible in underdeveloped regions, limiting the application of these multi-modal approaches in clinical settings. To address this, we propose a novel Genome-informed Hyper-Attention Network, termed G-HANet, which is capable of effectively distilling the histo-genomic knowledge during training to elevate uni-modal whole slide image (WSI)-based inference for the first time. Compared with traditional knowledge distillation methods (i.e., teacher-student architecture) in other tasks, our end-to-end model is superior in terms of training efficiency and learning cross-modal interactions. Specifically, the network comprises the cross-modal associating branch (CAB) and hyper-attention survival branch (HSB). Through the genomic data reconstruction from WSIs, CAB effectively distills the associations between functional genotypes and morphological phenotypes and offers insights into the gene expression profiles in the feature space. Subsequently, HSB leverages the distilled histo-genomic associations as well as the generated morphology-based weights to achieve the hyper-attention modeling of the patients from both histopathology and genomic perspectives to improve cancer prognosis. Extensive experiments are conducted on five TCGA benchmarking datasets and the results demonstrate that G-HANet significantly outperforms the state-of-the-art WSI-based methods and achieves competitive performance with genome-based and multi-modal methods. G-HANet is expected to be explored as a useful tool by the research community to address the current bottleneck of insufficient histo-genomic data pairing in the context of cancer prognosis and precision oncology.
VideoMV: Consistent Multi-View Generation Based on Large Video Generative Model
Mar 18, 2024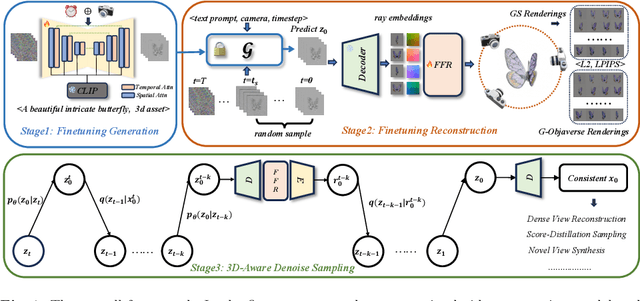
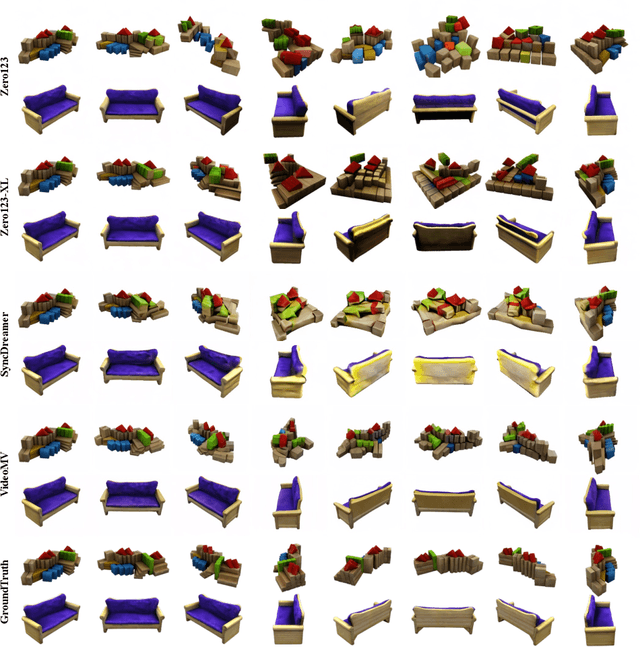
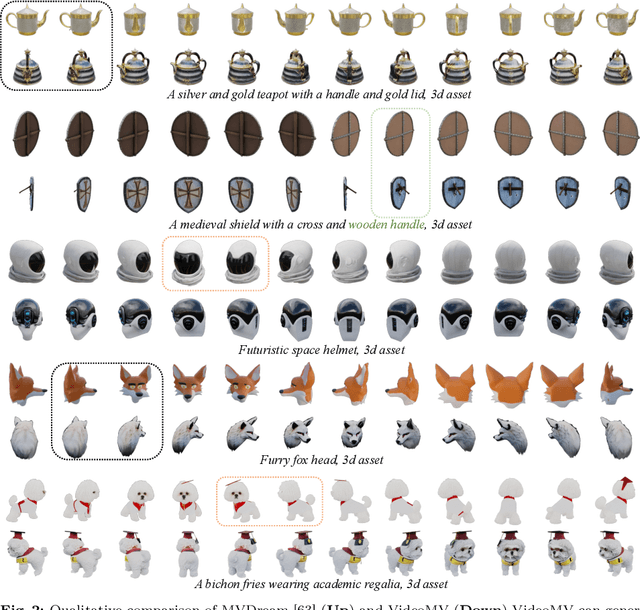
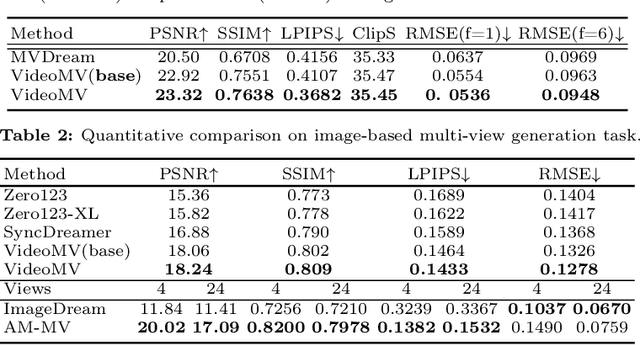


Generating multi-view images based on text or single-image prompts is a critical capability for the creation of 3D content. Two fundamental questions on this topic are what data we use for training and how to ensure multi-view consistency. This paper introduces a novel framework that makes fundamental contributions to both questions. Unlike leveraging images from 2D diffusion models for training, we propose a dense consistent multi-view generation model that is fine-tuned from off-the-shelf video generative models. Images from video generative models are more suitable for multi-view generation because the underlying network architecture that generates them employs a temporal module to enforce frame consistency. Moreover, the video data sets used to train these models are abundant and diverse, leading to a reduced train-finetuning domain gap. To enhance multi-view consistency, we introduce a 3D-Aware Denoising Sampling, which first employs a feed-forward reconstruction module to get an explicit global 3D model, and then adopts a sampling strategy that effectively involves images rendered from the global 3D model into the denoising sampling loop to improve the multi-view consistency of the final images. As a by-product, this module also provides a fast way to create 3D assets represented by 3D Gaussians within a few seconds. Our approach can generate 24 dense views and converges much faster in training than state-of-the-art approaches (4 GPU hours versus many thousand GPU hours) with comparable visual quality and consistency. By further fine-tuning, our approach outperforms existing state-of-the-art methods in both quantitative metrics and visual effects. Our project page is aigc3d.github.io/VideoMV.
Uncertainty-Calibrated Test-Time Model Adaptation without Forgetting
Mar 18, 2024
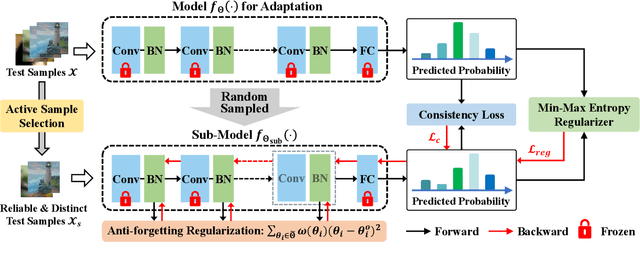
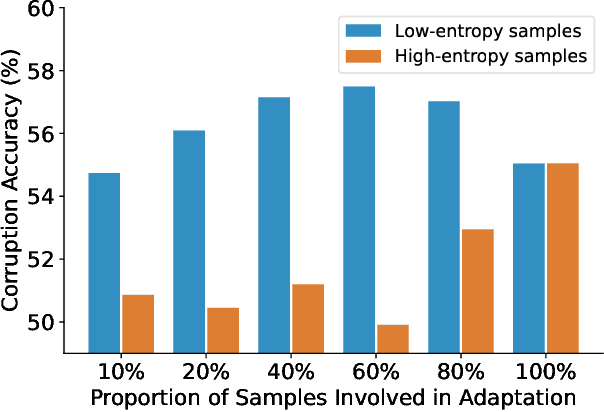
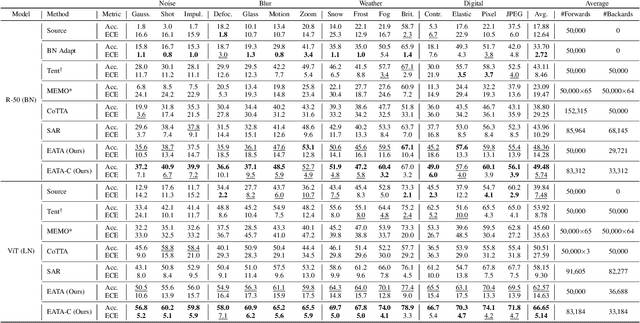
Test-time adaptation (TTA) seeks to tackle potential distribution shifts between training and test data by adapting a given model w.r.t. any test sample. Although recent TTA has shown promising performance, we still face two key challenges: 1) prior methods perform backpropagation for each test sample, resulting in unbearable optimization costs to many applications; 2) while existing TTA can significantly improve the test performance on out-of-distribution data, they often suffer from severe performance degradation on in-distribution data after TTA (known as forgetting). To this end, we have proposed an Efficient Anti-Forgetting Test-Time Adaptation (EATA) method which develops an active sample selection criterion to identify reliable and non-redundant samples for test-time entropy minimization. To alleviate forgetting, EATA introduces a Fisher regularizer estimated from test samples to constrain important model parameters from drastic changes. However, in EATA, the adopted entropy loss consistently assigns higher confidence to predictions even for samples that are underlying uncertain, leading to overconfident predictions. To tackle this, we further propose EATA with Calibration (EATA-C) to separately exploit the reducible model uncertainty and the inherent data uncertainty for calibrated TTA. Specifically, we measure the model uncertainty by the divergence between predictions from the full network and its sub-networks, on which we propose a divergence loss to encourage consistent predictions instead of overconfident ones. To further recalibrate prediction confidence, we utilize the disagreement among predicted labels as an indicator of the data uncertainty, and then devise a min-max entropy regularizer to selectively increase and decrease prediction confidence for different samples. Experiments on image classification and semantic segmentation verify the effectiveness of our methods.
Spatiotemporal Representation Learning for Short and Long Medical Image Time Series
Mar 12, 2024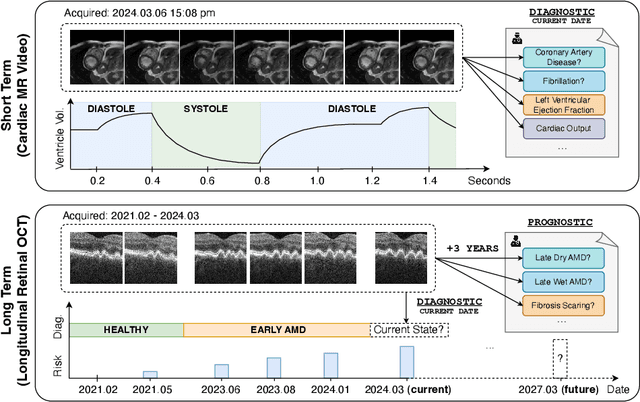
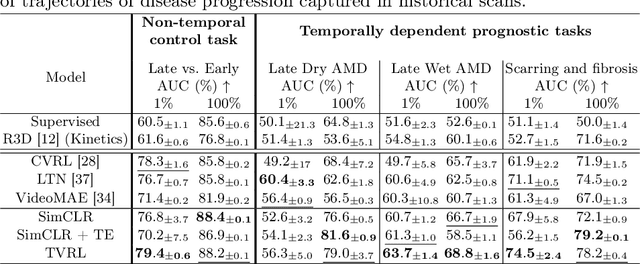
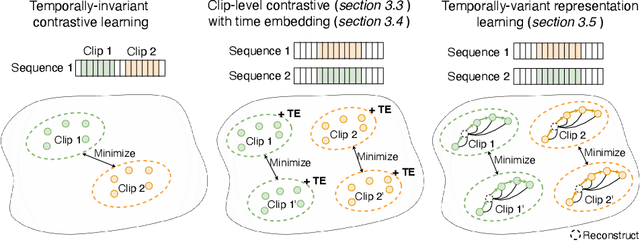
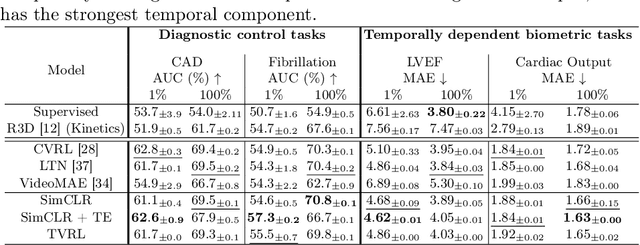
Analyzing temporal developments is crucial for the accurate prognosis of many medical conditions. Temporal changes that occur over short time scales are key to assessing the health of physiological functions, such as the cardiac cycle. Moreover, tracking longer term developments that occur over months or years in evolving processes, such as age-related macular degeneration (AMD), is essential for accurate prognosis. Despite the importance of both short and long term analysis to clinical decision making, they remain understudied in medical deep learning. State of the art methods for spatiotemporal representation learning, developed for short natural videos, prioritize the detection of temporal constants rather than temporal developments. Moreover, they do not account for varying time intervals between acquisitions, which are essential for contextualizing observed changes. To address these issues, we propose two approaches. First, we combine clip-level contrastive learning with a novel temporal embedding to adapt to irregular time series. Second, we propose masking and predicting latent frame representations of the temporal sequence. Our two approaches outperform all prior methods on temporally-dependent tasks including cardiac output estimation and three prognostic AMD tasks. Overall, this enables the automated analysis of temporal patterns which are typically overlooked in applications of deep learning to medicine.
Multi-LoRA Composition for Image Generation
Feb 26, 2024Low-Rank Adaptation (LoRA) is extensively utilized in text-to-image models for the accurate rendition of specific elements like distinct characters or unique styles in generated images. Nonetheless, existing methods face challenges in effectively composing multiple LoRAs, especially as the number of LoRAs to be integrated grows, thus hindering the creation of complex imagery. In this paper, we study multi-LoRA composition through a decoding-centric perspective. We present two training-free methods: LoRA Switch, which alternates between different LoRAs at each denoising step, and LoRA Composite, which simultaneously incorporates all LoRAs to guide more cohesive image synthesis. To evaluate the proposed approaches, we establish ComposLoRA, a new comprehensive testbed as part of this research. It features a diverse range of LoRA categories with 480 composition sets. Utilizing an evaluation framework based on GPT-4V, our findings demonstrate a clear improvement in performance with our methods over the prevalent baseline, particularly evident when increasing the number of LoRAs in a composition.
Two-stage Cytopathological Image Synthesis for Augmenting Cervical Abnormality Screening
Feb 25, 2024Automatic thin-prep cytologic test (TCT) screening can assist pathologists in finding cervical abnormality towards accurate and efficient cervical cancer diagnosis. Current automatic TCT screening systems mostly involve abnormal cervical cell detection, which generally requires large-scale and diverse training data with high-quality annotations to achieve promising performance. Pathological image synthesis is naturally raised to minimize the efforts in data collection and annotation. However, it is challenging to generate realistic large-size cytopathological images while simultaneously synthesizing visually plausible appearances for small-size abnormal cervical cells. In this paper, we propose a two-stage image synthesis framework to create synthetic data for augmenting cervical abnormality screening. In the first Global Image Generation stage, a Normal Image Generator is designed to generate cytopathological images full of normal cervical cells. In the second Local Cell Editing stage, normal cells are randomly selected from the generated images and then are converted to different types of abnormal cells using the proposed Abnormal Cell Synthesizer. Both Normal Image Generator and Abnormal Cell Synthesizer are built upon Stable Diffusion, a pre-trained foundation model for image synthesis, via parameter-efficient fine-tuning methods for customizing cytopathological image contents and extending spatial layout controllability, respectively. Our experiments demonstrate the synthetic image quality, diversity, and controllability of the proposed synthesis framework, and validate its data augmentation effectiveness in enhancing the performance of abnormal cervical cell detection.
Matching Non-Identical Objects
Mar 13, 2024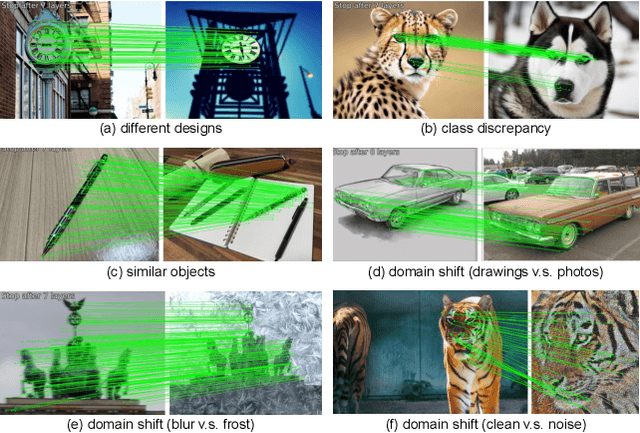
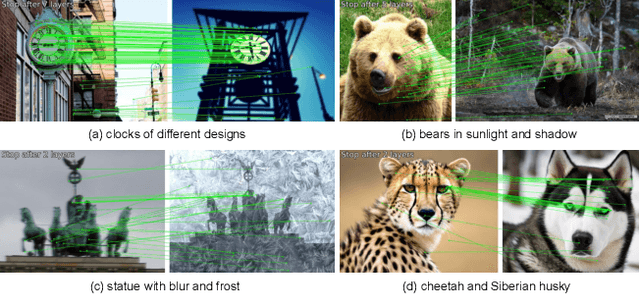
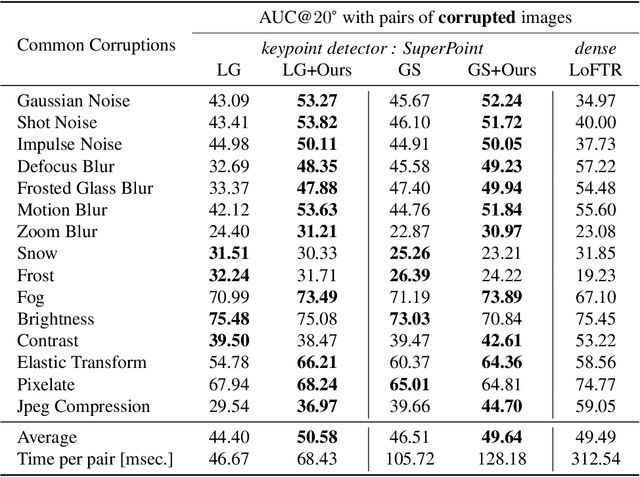

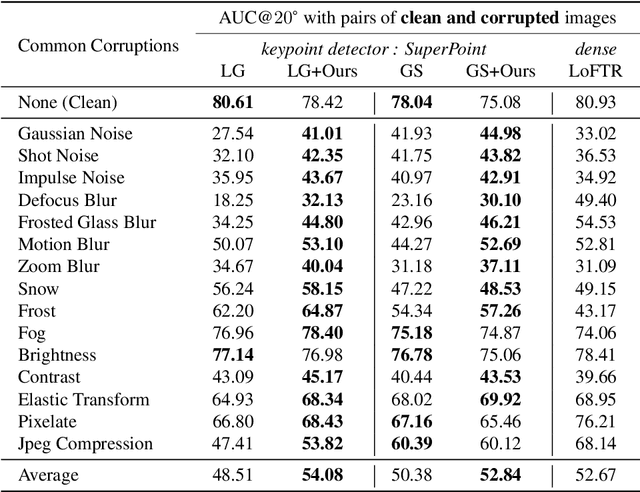
Not identical but similar objects are everywhere in the world. Examples include four-legged animals such as dogs and cats, cars of different models, akin flowers in various colors, and countless others. In this study, we address a novel task of matching such non-identical objects. We propose a simple weighting scheme of descriptors that enhance various sparse image matching methods, which are originally designed for matching identical objects captured from different perspectives, and achieve semantically robust matching. The experiments show successful matching between non-identical objects in various cases including domain shift. Further, we present a first evaluation of the robustness of the image matching methods under common corruptions, which is a sort of domain shift, and the proposed method improves the matching in this case as well.
TTA-Nav: Test-time Adaptive Reconstruction for Point-Goal Navigation under Visual Corruptions
Mar 14, 2024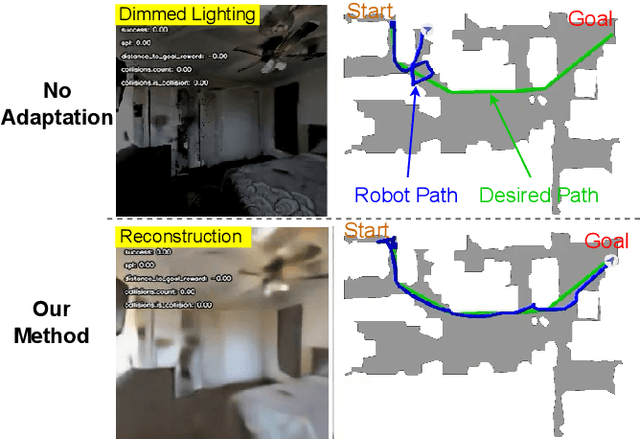
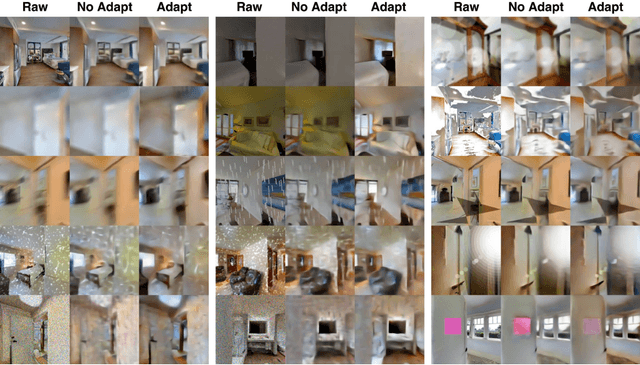
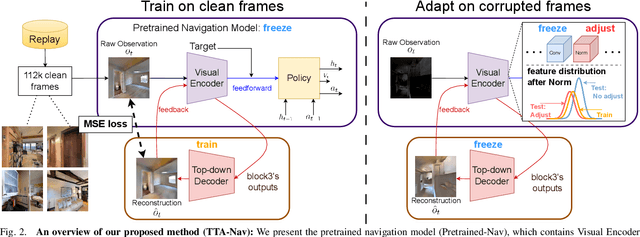


Robot navigation under visual corruption presents a formidable challenge. To address this, we propose a Test-time Adaptation (TTA) method, named as TTA-Nav, for point-goal navigation under visual corruptions. Our "plug-and-play" method incorporates a top-down decoder to a pre-trained navigation model. Firstly, the pre-trained navigation model gets a corrupted image and extracts features. Secondly, the top-down decoder produces the reconstruction given the high-level features extracted by the pre-trained model. Then, it feeds the reconstruction of a corrupted image back to the pre-trained model. Finally, the pre-trained model does forward pass again to output action. Despite being trained solely on clean images, the top-down decoder can reconstruct cleaner images from corrupted ones without the need for gradient-based adaptation. The pre-trained navigation model with our top-down decoder significantly enhances navigation performance across almost all visual corruptions in our benchmarks. Our method improves the success rate of point-goal navigation from the state-of-the-art result of 46% to 94% on the most severe corruption. This suggests its potential for broader application in robotic visual navigation. Project page: https://sites.google.com/view/tta-nav
ConDiSR: Contrastive Disentanglement and Style Regularization for Single Domain Generalization
Mar 14, 2024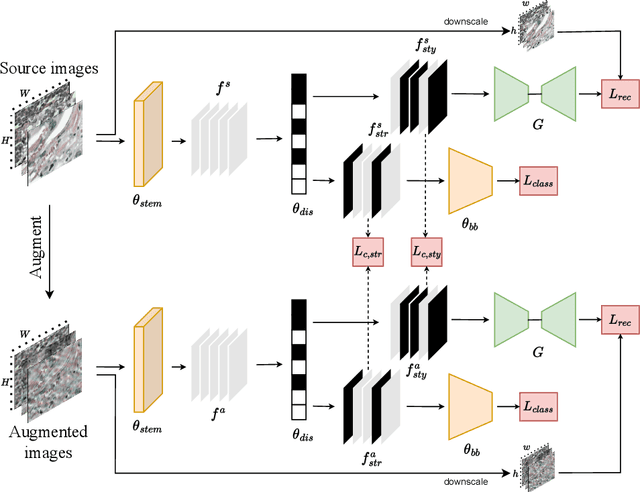


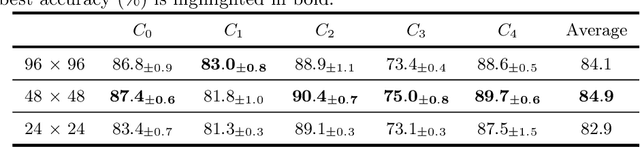
Medical data often exhibits distribution shifts, which cause test-time performance degradation for deep learning models trained using standard supervised learning pipelines. This challenge is addressed in the field of Domain Generalization (DG) with the sub-field of Single Domain Generalization (SDG) being specifically interesting due to the privacy- or logistics-related issues often associated with medical data. Existing disentanglement-based SDG methods heavily rely on structural information embedded in segmentation masks, however classification labels do not provide such dense information. This work introduces a novel SDG method aimed at medical image classification that leverages channel-wise contrastive disentanglement. It is further enhanced with reconstruction-based style regularization to ensure extraction of distinct style and structure feature representations. We evaluate our method on the complex task of multicenter histopathology image classification, comparing it against state-of-the-art (SOTA) SDG baselines. Results demonstrate that our method surpasses the SOTA by a margin of 1% in average accuracy while also showing more stable performance. This study highlights the importance and challenges of exploring SDG frameworks in the context of the classification task. The code is publicly available at https://github.com/BioMedIA-MBZUAI/ConDiSR
CLIP-EBC: CLIP Can Count Accurately through Enhanced Blockwise Classification
Mar 14, 2024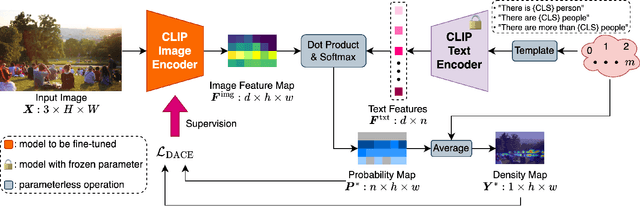
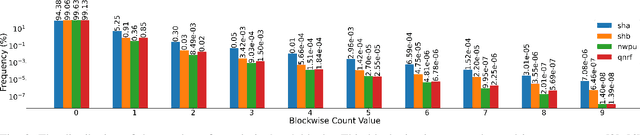
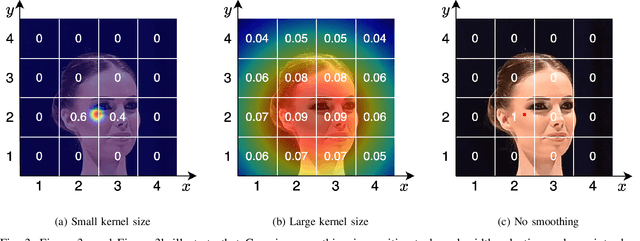
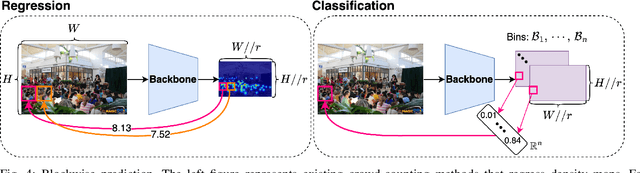
The CLIP (Contrastive Language-Image Pretraining) model has exhibited outstanding performance in recognition problems, such as zero-shot image classification and object detection. However, its ability to count remains understudied due to the inherent challenges of transforming counting--a regression task--into a recognition task. In this paper, we investigate CLIP's potential in counting, focusing specifically on estimating crowd sizes. Existing classification-based crowd-counting methods have encountered issues, including inappropriate discretization strategies, which impede the application of CLIP and result in suboptimal performance. To address these challenges, we propose the Enhanced Blockwise Classification (EBC) framework. In contrast to previous methods, EBC relies on integer-valued bins that facilitate the learning of robust decision boundaries. Within our model-agnostic EBC framework, we introduce CLIP-EBC, the first fully CLIP-based crowd-counting model capable of generating density maps. Comprehensive evaluations across diverse crowd-counting datasets demonstrate the state-of-the-art performance of our methods. Particularly, EBC can improve existing models by up to 76.9%. Moreover, our CLIP-EBC model surpasses current crowd-counting methods, achieving mean absolute errors of 55.0 and 6.3 on ShanghaiTech part A and part B datasets, respectively. The code will be made publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge