"Image": models, code, and papers
EndoBoost: a plug-and-play module for false positive suppression during computer-aided polyp detection in real-world colonoscopy (with dataset)
Dec 23, 2022
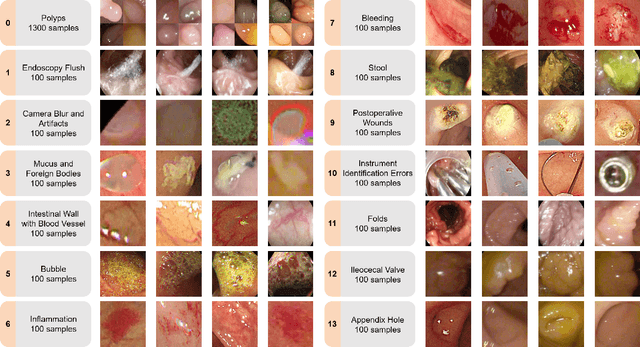
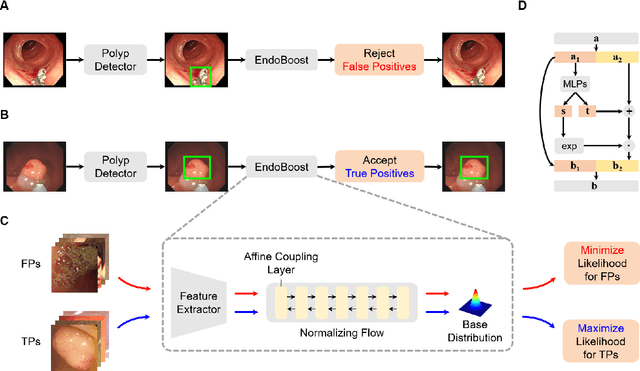
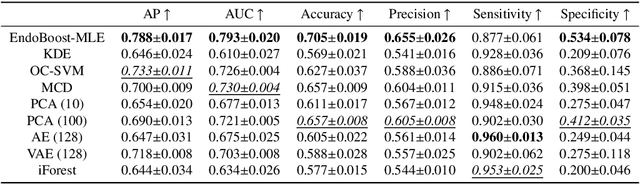
The advance of computer-aided detection systems using deep learning opened a new scope in endoscopic image analysis. However, the learning-based models developed on closed datasets are susceptible to unknown anomalies in complex clinical environments. In particular, the high false positive rate of polyp detection remains a major challenge in clinical practice. In this work, we release the FPPD-13 dataset, which provides a taxonomy and real-world cases of typical false positives during computer-aided polyp detection in real-world colonoscopy. We further propose a post-hoc module EndoBoost, which can be plugged into generic polyp detection models to filter out false positive predictions. This is realized by generative learning of the polyp manifold with normalizing flows and rejecting false positives through density estimation. Compared to supervised classification, this anomaly detection paradigm achieves better data efficiency and robustness in open-world settings. Extensive experiments demonstrate a promising false positive suppression in both retrospective and prospective validation. In addition, the released dataset can be used to perform 'stress' tests on established detection systems and encourages further research toward robust and reliable computer-aided endoscopic image analysis. The dataset and code will be publicly available at http://endoboost.miccai.cloud.
Self-Supervised Learning of Image Scale and Orientation
Jun 15, 2022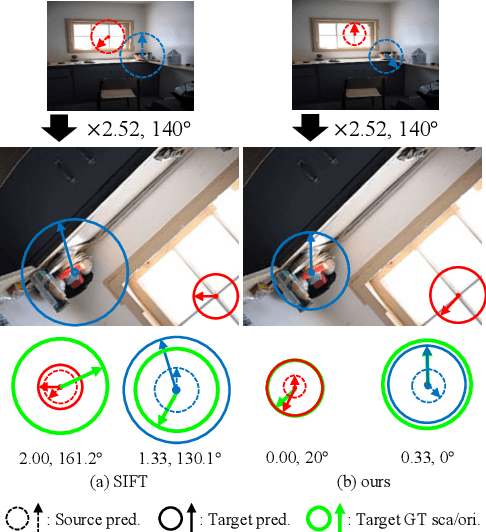
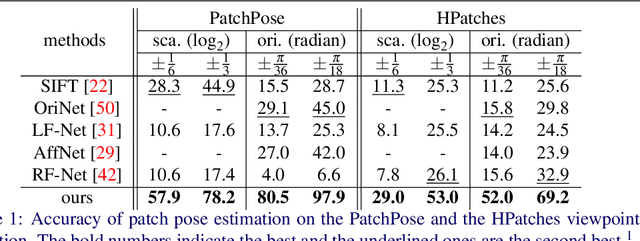
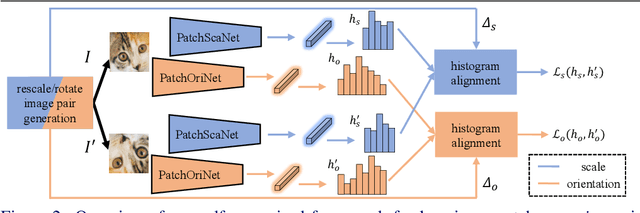
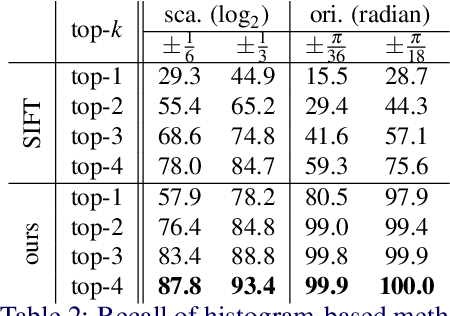
We study the problem of learning to assign a characteristic pose, i.e., scale and orientation, for an image region of interest. Despite its apparent simplicity, the problem is non-trivial; it is hard to obtain a large-scale set of image regions with explicit pose annotations that a model directly learns from. To tackle the issue, we propose a self-supervised learning framework with a histogram alignment technique. It generates pairs of image patches by random rescaling/rotating and then train an estimator to predict their scale/orientation values so that their relative difference is consistent with the rescaling/rotating used. The estimator learns to predict a non-parametric histogram distribution of scale/orientation without any supervision. Experiments show that it significantly outperforms previous methods in scale/orientation estimation and also improves image matching and 6 DoF camera pose estimation by incorporating our patch poses into a matching process.
Distributed Parallel Image Signal Extrapolation Framework using Message Passing Interface
Jul 01, 2022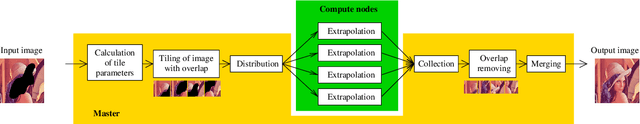
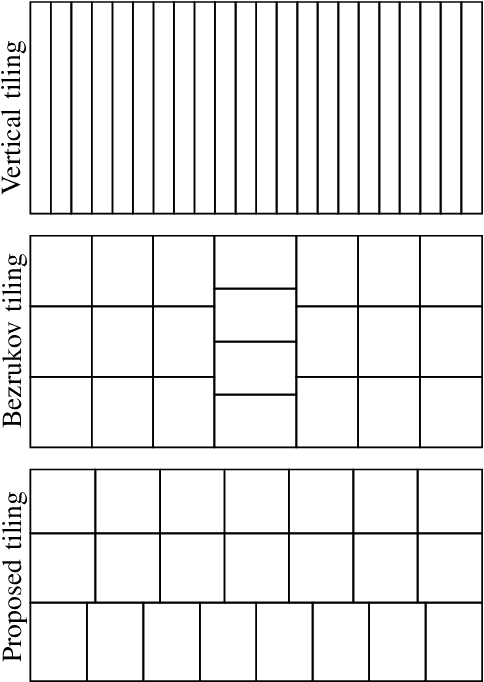

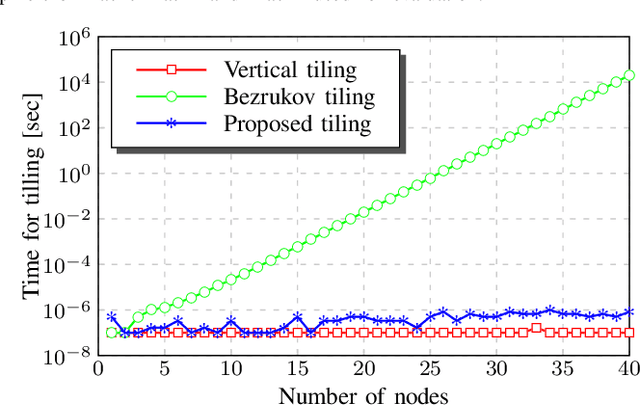
This paper introduces a framework for distributed parallel image signal extrapolation. Since high-quality image signal processing often comes along with a high computational complexity, a parallel execution is desirable. The proposed framework allows for the application of existing image signal extrapolation algorithms without the need to modify them for a parallel processing. The unaltered application of existing algorithms is achieved by dividing input images into overlapping tiles which are distributed to compute nodes via Message Passing Interface. In order to keep the computational overhead low, a novel image tiling algorithm is proposed. Using this algorithm, a nearly optimum tiling is possible at a very small processing time. For showing the efficacy of the framework, it is used for parallelizing a high-complexity extrapolation algorithm. Simulation results show that the proposed framework has no negative impact on extrapolation quality while at the same time offering good scaling behavior on compute clusters.
Unsupervised Superpixel Generation using Edge-Sparse Embedding
Nov 29, 2022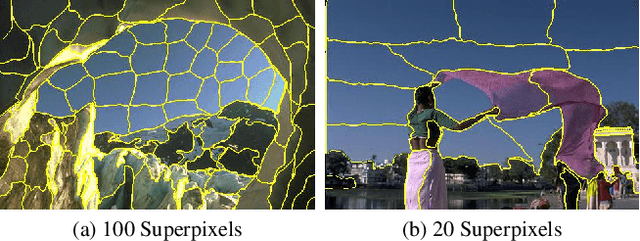
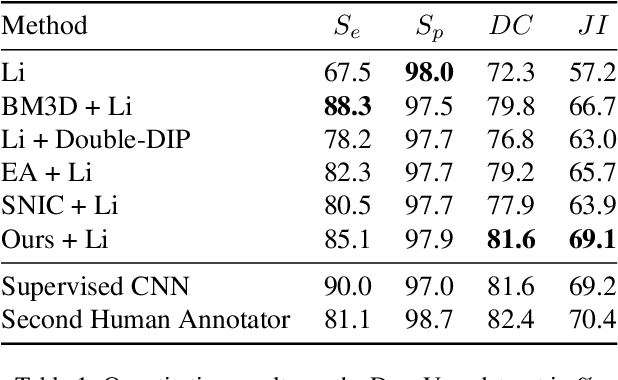
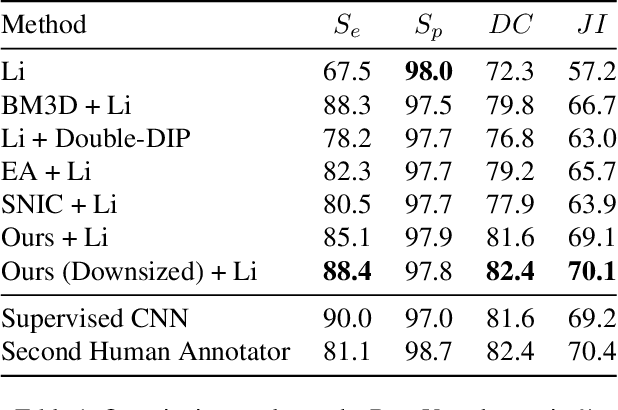
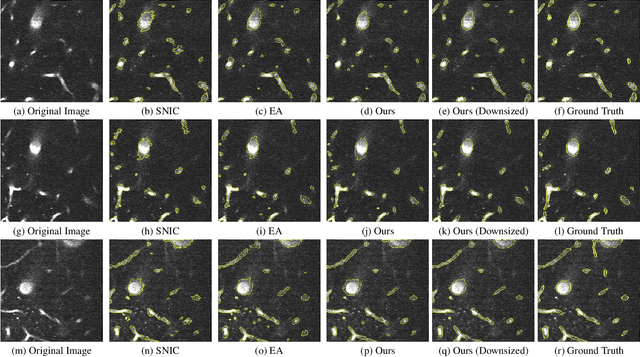
Partitioning an image into superpixels based on the similarity of pixels with respect to features such as colour or spatial location can significantly reduce data complexity and improve subsequent image processing tasks. Initial algorithms for unsupervised superpixel generation solely relied on local cues without prioritizing significant edges over arbitrary ones. On the other hand, more recent methods based on unsupervised deep learning either fail to properly address the trade-off between superpixel edge adherence and compactness or lack control over the generated number of superpixels. By using random images with strong spatial correlation as input, \ie, blurred noise images, in a non-convolutional image decoder we can reduce the expected number of contrasts and enforce smooth, connected edges in the reconstructed image. We generate edge-sparse pixel embeddings by encoding additional spatial information into the piece-wise smooth activation maps from the decoder's last hidden layer and use a standard clustering algorithm to extract high quality superpixels. Our proposed method reaches state-of-the-art performance on the BSDS500, PASCAL-Context and a microscopy dataset.
Taming a Generative Model
Nov 29, 2022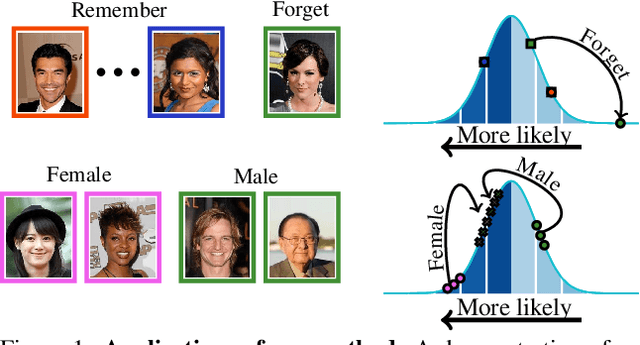
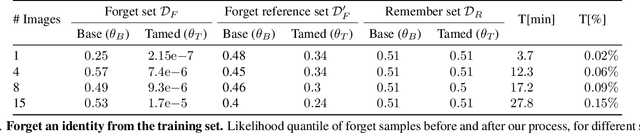
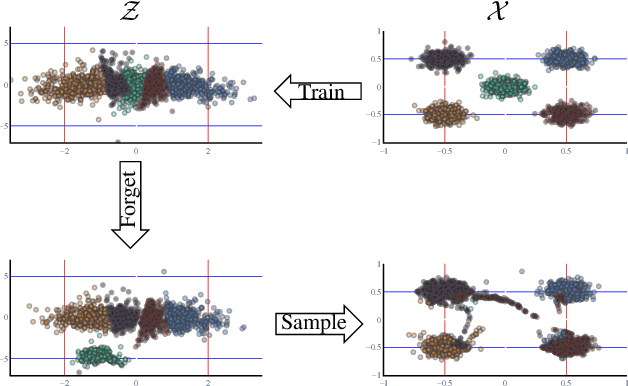
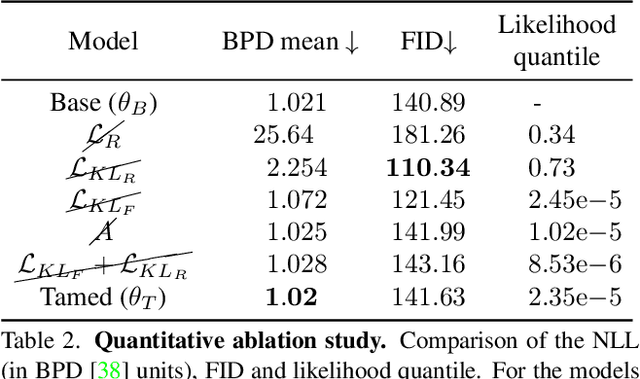
Generative models are becoming ever more powerful, being able to synthesize highly realistic images. We propose an algorithm for taming these models - changing the probability that the model will produce a specific image or image category. We consider generative models that are powered by normalizing flows, which allows us to reason about the exact generation probability likelihood for a given image. Our method is general purpose, and we exemplify it using models that generate human faces, a subdomain with many interesting privacy and bias considerations. Our method can be used in the context of privacy, e.g., removing a specific person from the output of a model, and also in the context of de-biasing by forcing a model to output specific image categories according to a given target distribution. Our method uses a fast fine-tuning process without retraining the model from scratch, achieving the goal in less than 1% of the time taken to initially train the generative model. We evaluate qualitatively and quantitatively, to examine the success of the taming process and output quality.
Image Keypoint Matching using Graph Neural Networks
May 27, 2022Image matching is a key component of many tasks in computer vision and its main objective is to find correspondences between features extracted from different natural images. When images are represented as graphs, image matching boils down to the problem of graph matching which has been studied intensively in the past. In recent years, graph neural networks have shown great potential in the graph matching task, and have also been applied to image matching. In this paper, we propose a graph neural network for the problem of image matching. The proposed method first generates initial soft correspondences between keypoints using localized node embeddings and then iteratively refines the initial correspondences using a series of graph neural network layers. We evaluate our method on natural image datasets with keypoint annotations and show that, in comparison to a state-of-the-art model, our method speeds up inference times without sacrificing prediction accuracy.
Multimodality Helps Unimodality: Cross-Modal Few-Shot Learning with Multimodal Models
Jan 18, 2023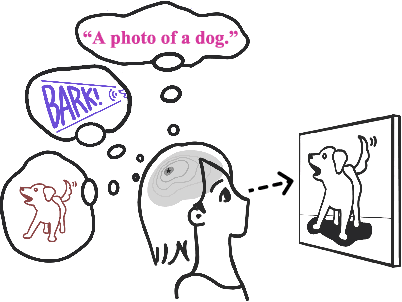

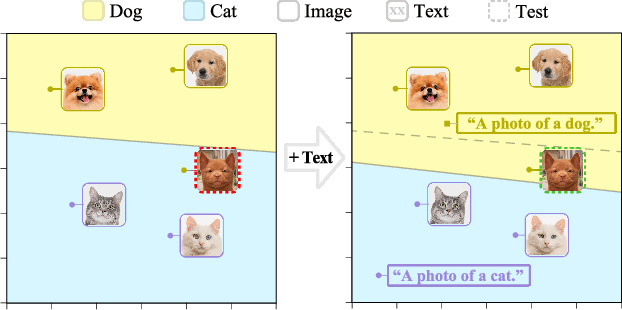
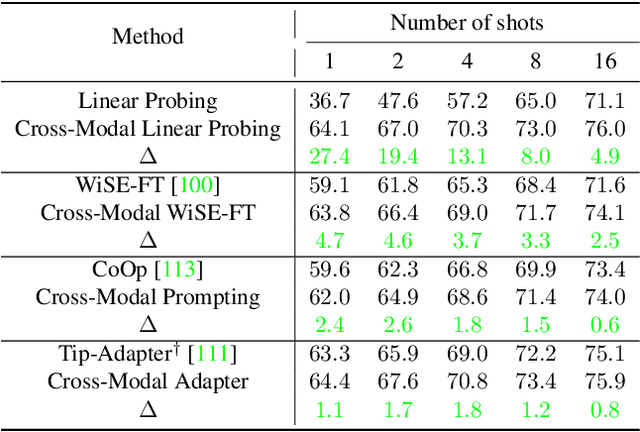
The ability to quickly learn a new task with minimal instruction - known as few-shot learning - is a central aspect of intelligent agents. Classical few-shot benchmarks make use of few-shot samples from a single modality, but such samples may not be sufficient to characterize an entire concept class. In contrast, humans use cross-modal information to learn new concepts efficiently. In this work, we demonstrate that one can indeed build a better ${\bf visual}$ dog classifier by ${\bf read}$ing about dogs and ${\bf listen}$ing to them bark. To do so, we exploit the fact that recent multimodal foundation models such as CLIP are inherently cross-modal, mapping different modalities to the same representation space. Specifically, we propose a simple cross-modal adaptation approach that learns from few-shot examples spanning different modalities. By repurposing class names as additional one-shot training samples, we achieve SOTA results with an embarrassingly simple linear classifier for vision-language adaptation. Furthermore, we show that our approach can benefit existing methods such as prefix tuning, adapters, and classifier ensembling. Finally, to explore other modalities beyond vision and language, we construct the first (to our knowledge) audiovisual few-shot benchmark and use cross-modal training to improve the performance of both image and audio classification.
pyssam -- a Python library for statistical modelling of biomedical shape and appearance
Jan 11, 2023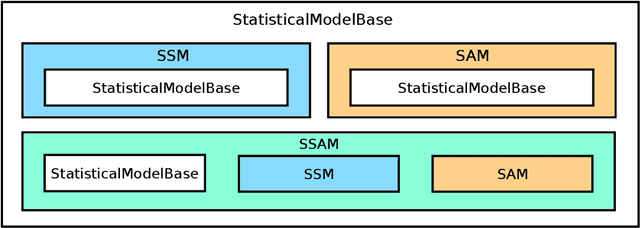
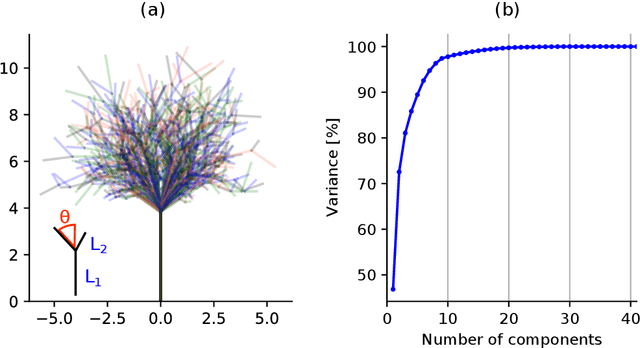
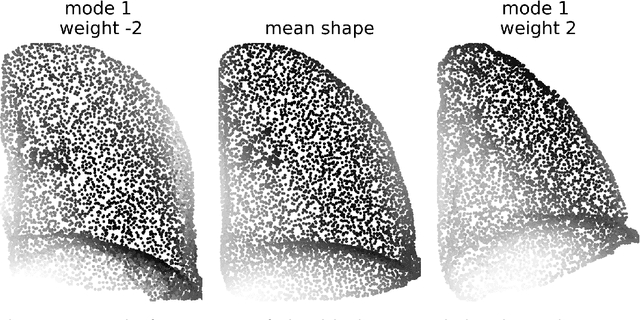
pyssam is a Python library for creating statistical shape and appearance models (SSAMs) for biological (and other) shapes such as bones, lungs or other organs. A point cloud best describing the anatomical 'landmarks' of the organ are required from each sample in a small population as an input. Additional information such as landmark gray-value can be included to incorporate joint correlations of shape and 'appearance' into the model. Our library performs alignment and scaling of the input data and creates a SSAM based on covariance across the population. The output SSAM can be used to parameterise and quantify shape change across a population. pyssam is a small and low dependency codebase with examples included as Jupyter notebooks for several common SSAM computations. The given examples can easily be extended to alternative datasets, and also alternative tasks such as medical image segmentation by incorporating a SSAM as a constraint for segmented organs.
SynMotor: A Benchmark Suite for Object Attribute Regression and Multi-task Learning
Jan 11, 2023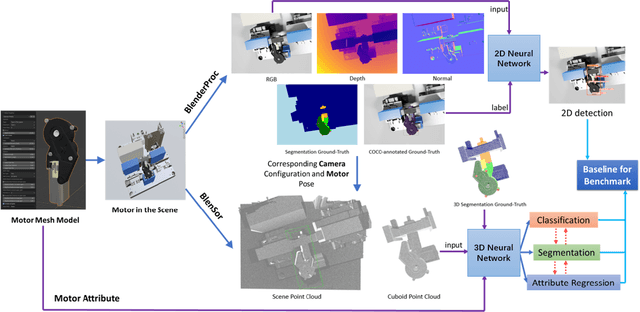
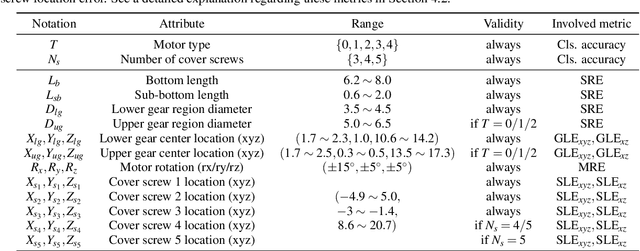
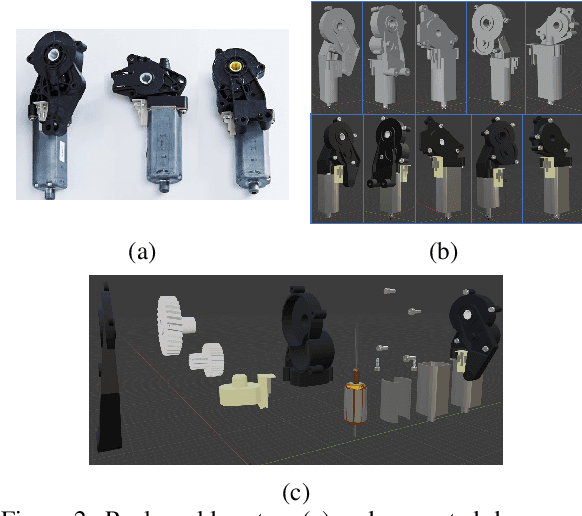
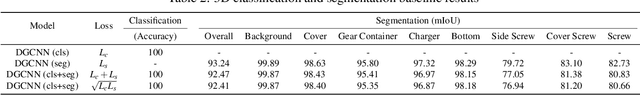
In this paper, we develop a novel benchmark suite including both a 2D synthetic image dataset and a 3D synthetic point cloud dataset. Our work is a sub-task in the framework of a remanufacturing project, in which small electric motors are used as fundamental objects. Apart from the given detection, classification, and segmentation annotations, the key objects also have multiple learnable attributes with ground truth provided. This benchmark can be used for computer vision tasks including 2D/3D detection, classification, segmentation, and multi-attribute learning. It is worth mentioning that most attributes of the motors are quantified as continuously variable rather than binary, which makes our benchmark well-suited for the less explored regression tasks. In addition, appropriate evaluation metrics are adopted or developed for each task and promising baseline results are provided. We hope this benchmark can stimulate more research efforts on the sub-domain of object attribute learning and multi-task learning in the future.
Towards Microstructural State Variables in Materials Systems
Jan 11, 2023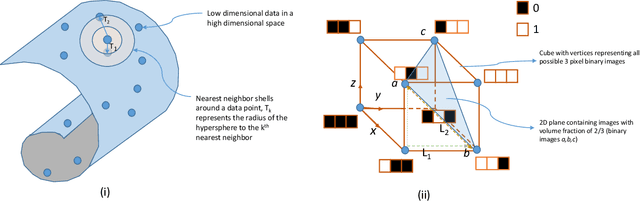
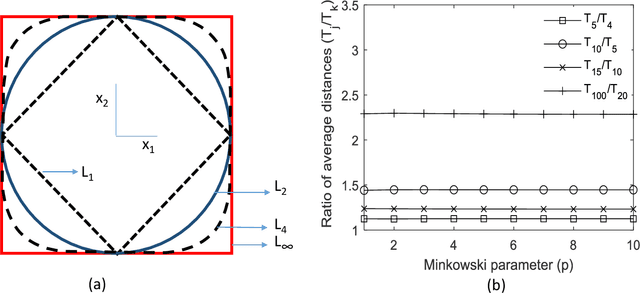
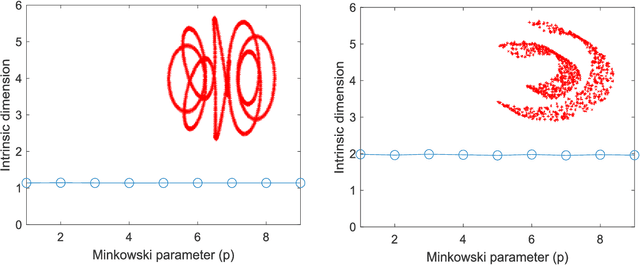
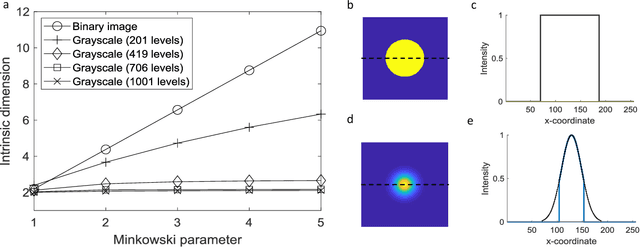
The vast combination of material properties seen in nature are achieved by the complexity of the material microstructure. Advanced characterization and physics based simulation techniques have led to generation of extremely large microstructural datasets. There is a need for machine learning techniques that can manage data complexity by capturing the maximal amount of information about the microstructure using the least number of variables. This paper aims to formulate dimensionality and state variable estimation techniques focused on reducing microstructural image data. It is shown that local dimensionality estimation based on nearest neighbors tend to give consistent dimension estimates for natural images for all p-Minkowski distances. However, it is found that dimensionality estimates have a systematic error for low-bit depth microstructural images. The use of Manhattan distance to alleviate this issue is demonstrated. It is also shown that stacked autoencoders can reconstruct the generator space of high dimensional microstructural data and provide a sparse set of state variables to fully describe the variability in material microstructures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge