"Image": models, code, and papers
Causal Triplet: An Open Challenge for Intervention-centric Causal Representation Learning
Jan 12, 2023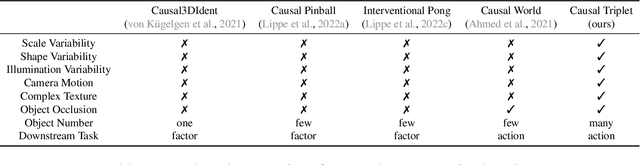
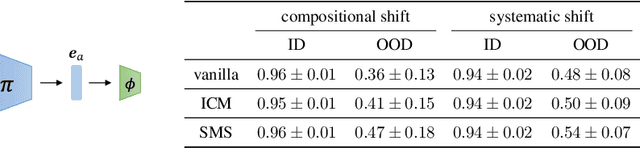
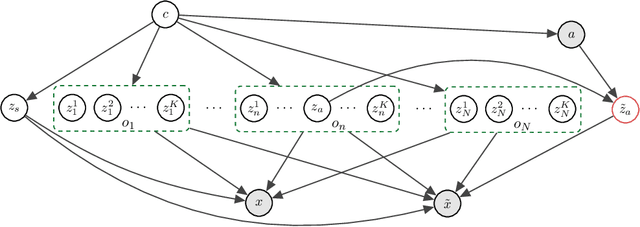


Recent years have seen a surge of interest in learning high-level causal representations from low-level image pairs under interventions. Yet, existing efforts are largely limited to simple synthetic settings that are far away from real-world problems. In this paper, we present Causal Triplet, a causal representation learning benchmark featuring not only visually more complex scenes, but also two crucial desiderata commonly overlooked in previous works: (i) an actionable counterfactual setting, where only certain object-level variables allow for counterfactual observations whereas others do not; (ii) an interventional downstream task with an emphasis on out-of-distribution robustness from the independent causal mechanisms principle. Through extensive experiments, we find that models built with the knowledge of disentangled or object-centric representations significantly outperform their distributed counterparts. However, recent causal representation learning methods still struggle to identify such latent structures, indicating substantial challenges and opportunities for future work. Our code and datasets will be available at https://sites.google.com/view/causaltriplet.
Fine-Grained Entity Segmentation
Nov 12, 2022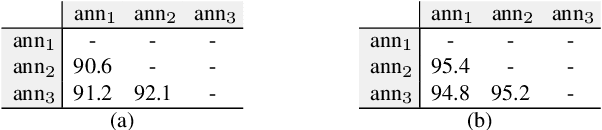
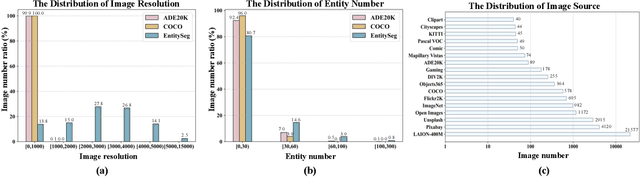
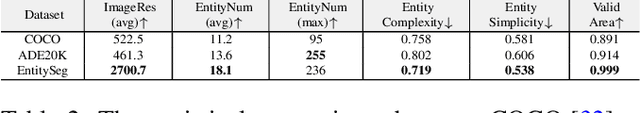
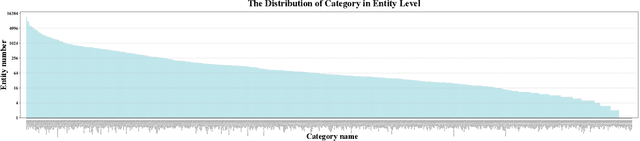
In dense image segmentation tasks (e.g., semantic, panoptic), existing methods can hardly generalize well to unseen image domains, predefined classes, and image resolution & quality variations. Motivated by these observations, we construct a large-scale entity segmentation dataset to explore fine-grained entity segmentation, with a strong focus on open-world and high-quality dense segmentation. The dataset contains images spanning diverse image domains and resolutions, along with high-quality mask annotations for training and testing. Given the high-quality and -resolution nature of the dataset, we propose CropFormer for high-quality segmentation, which can improve mask prediction using high-res image crops that provide more fine-grained image details than the full image. CropFormer is the first query-based Transformer architecture that can effectively ensemble mask predictions from multiple image crops, by learning queries that can associate the same entities across the full image and its crop. With CropFormer, we achieve a significant AP gain of $1.9$ on the challenging fine-grained entity segmentation task. The dataset and code will be released at http://luqi.info/entityv2.github.io/.
Saliency-Augmented Memory Completion for Continual Learning
Dec 26, 2022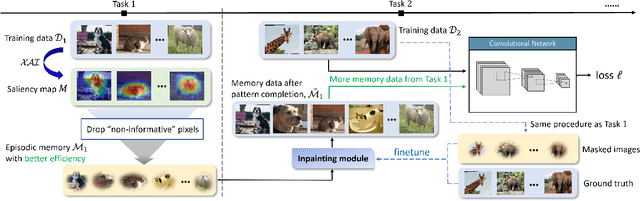
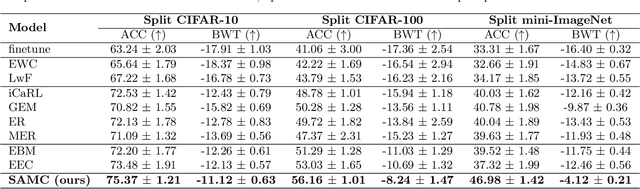
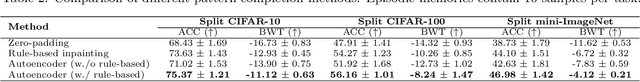
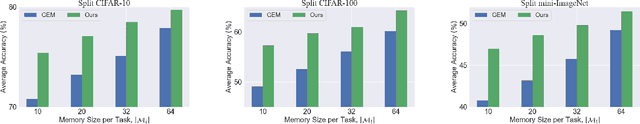
Continual Learning is considered a key step toward next-generation Artificial Intelligence. Among various methods, replay-based approaches that maintain and replay a small episodic memory of previous samples are one of the most successful strategies against catastrophic forgetting. However, since forgetting is inevitable given bounded memory and unbounded tasks, how to forget is a problem continual learning must address. Therefore, beyond simply avoiding catastrophic forgetting, an under-explored issue is how to reasonably forget while ensuring the merits of human memory, including 1. storage efficiency, 2. generalizability, and 3. some interpretability. To achieve these simultaneously, our paper proposes a new saliency-augmented memory completion framework for continual learning, inspired by recent discoveries in memory completion separation in cognitive neuroscience. Specifically, we innovatively propose to store the part of the image most important to the tasks in episodic memory by saliency map extraction and memory encoding. When learning new tasks, previous data from memory are inpainted by an adaptive data generation module, which is inspired by how humans complete episodic memory. The module's parameters are shared across all tasks and it can be jointly trained with a continual learning classifier as bilevel optimization. Extensive experiments on several continual learning and image classification benchmarks demonstrate the proposed method's effectiveness and efficiency.
SimpleMind adds thinking to deep neural networks
Dec 02, 2022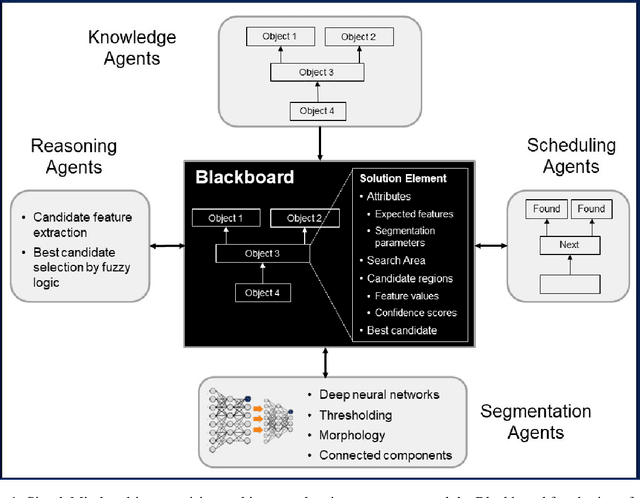
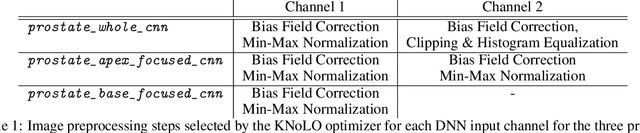

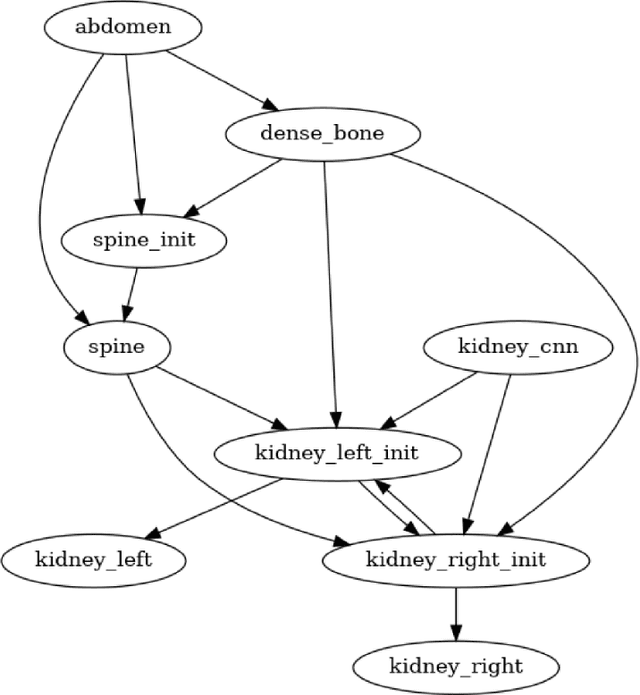
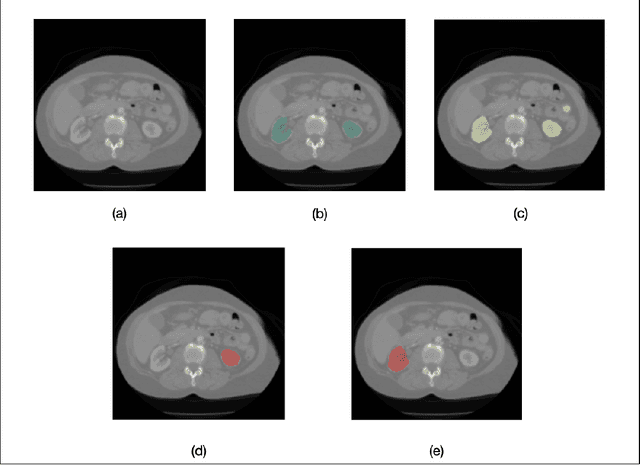
Deep neural networks (DNNs) detect patterns in data and have shown versatility and strong performance in many computer vision applications. However, DNNs alone are susceptible to obvious mistakes that violate simple, common sense concepts and are limited in their ability to use explicit knowledge to guide their search and decision making. While overall DNN performance metrics may be good, these obvious errors, coupled with a lack of explainability, have prevented widespread adoption for crucial tasks such as medical image analysis. The purpose of this paper is to introduce SimpleMind, an open-source software framework for Cognitive AI focused on medical image understanding. It allows creation of a knowledge base that describes expected characteristics and relationships between image objects in an intuitive human-readable form. The SimpleMind framework brings thinking to DNNs by: (1) providing methods for reasoning with the knowledge base about image content, such as spatial inferencing and conditional reasoning to check DNN outputs; (2) applying process knowledge, in the form of general-purpose software agents, that are chained together to accomplish image preprocessing, DNN prediction, and result post-processing, and (3) performing automatic co-optimization of all knowledge base parameters to adapt agents to specific problems. SimpleMind enables reasoning on multiple detected objects to ensure consistency, providing cross checking between DNN outputs. This machine reasoning improves the reliability and trustworthiness of DNNs through an interpretable model and explainable decisions. Example applications are provided that demonstrate how SimpleMind supports and improves deep neural networks by embedding them within a Cognitive AI framework.
A Segmentation Method for fluorescence images without a machine learning approach
Dec 28, 2022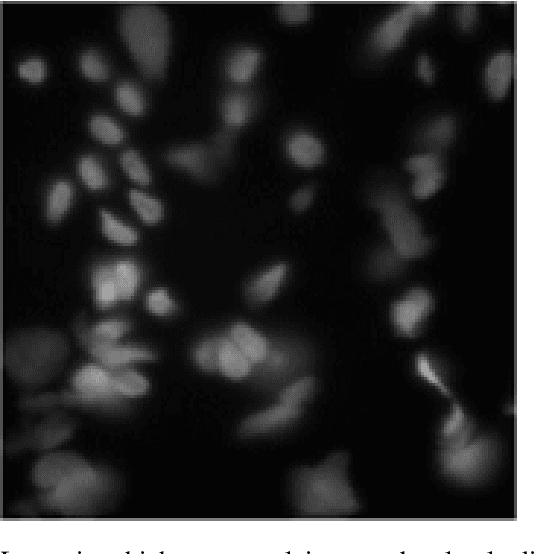
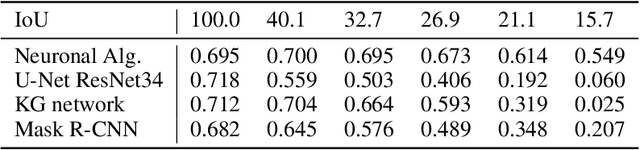
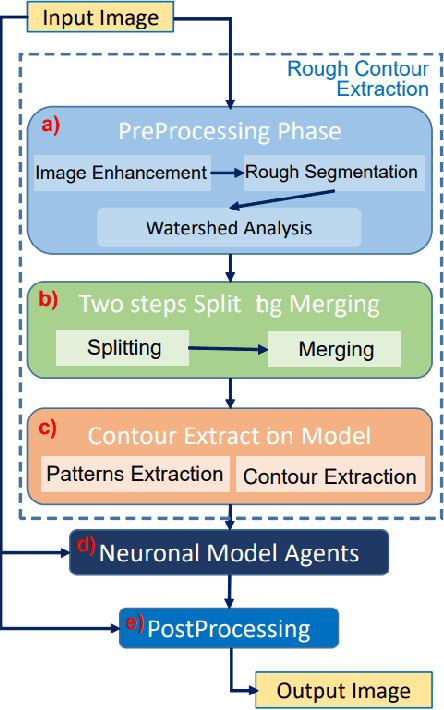
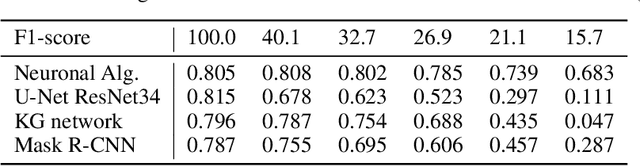
Background: Image analysis applications in digital pathology include various methods for segmenting regions of interest. Their identification is one of the most complex steps, and therefore of great interest for the study of robust methods that do not necessarily rely on a machine learning (ML) approach. Method: A fully automatic and optimized segmentation process for different datasets is a prerequisite for classifying and diagnosing Indirect ImmunoFluorescence (IIF) raw data. This study describes a deterministic computational neuroscience approach for identifying cells and nuclei. It is far from the conventional neural network approach, but it is equivalent to their quantitative and qualitative performance, and it is also solid to adversative noise. The method is robust, based on formally correct functions, and does not suffer from tuning on specific data sets. Results: This work demonstrates the robustness of the method against the variability of parameters, such as image size, mode, and signal-to-noise ratio. We validated the method on two datasets (Neuroblastoma and NucleusSegData) using images annotated by independent medical doctors. Conclusions: The definition of deterministic and formally correct methods, from a functional to a structural point of view, guarantees the achievement of optimized and functionally correct results. The excellent performance of our deterministic method (NeuronalAlg) to segment cells and nuclei from fluorescence images was measured with quantitative indicators and compared with those achieved by three published ML approaches.
3D-LDM: Neural Implicit 3D Shape Generation with Latent Diffusion Models
Dec 01, 2022
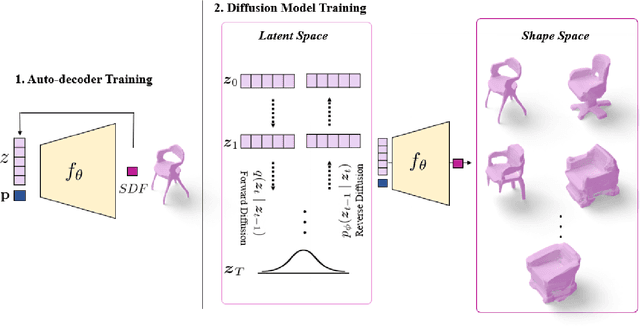
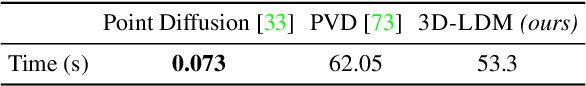
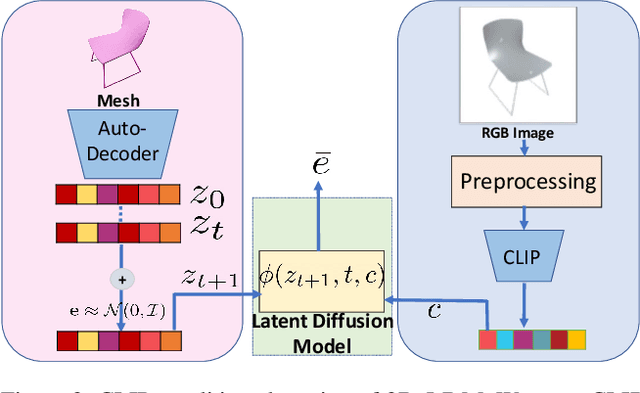
Diffusion models have shown great promise for image generation, beating GANs in terms of generation diversity, with comparable image quality. However, their application to 3D shapes has been limited to point or voxel representations that can in practice not accurately represent a 3D surface. We propose a diffusion model for neural implicit representations of 3D shapes that operates in the latent space of an auto-decoder. This allows us to generate diverse and high quality 3D surfaces. We additionally show that we can condition our model on images or text to enable image-to-3D generation and text-to-3D generation using CLIP embeddings. Furthermore, adding noise to the latent codes of existing shapes allows us to explore shape variations.
TFormer: A throughout fusion transformer for multi-modal skin lesion diagnosis
Nov 21, 2022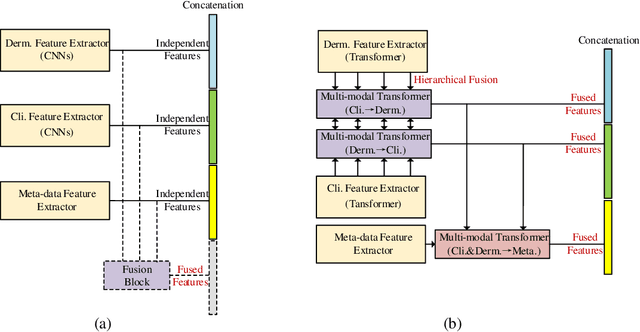
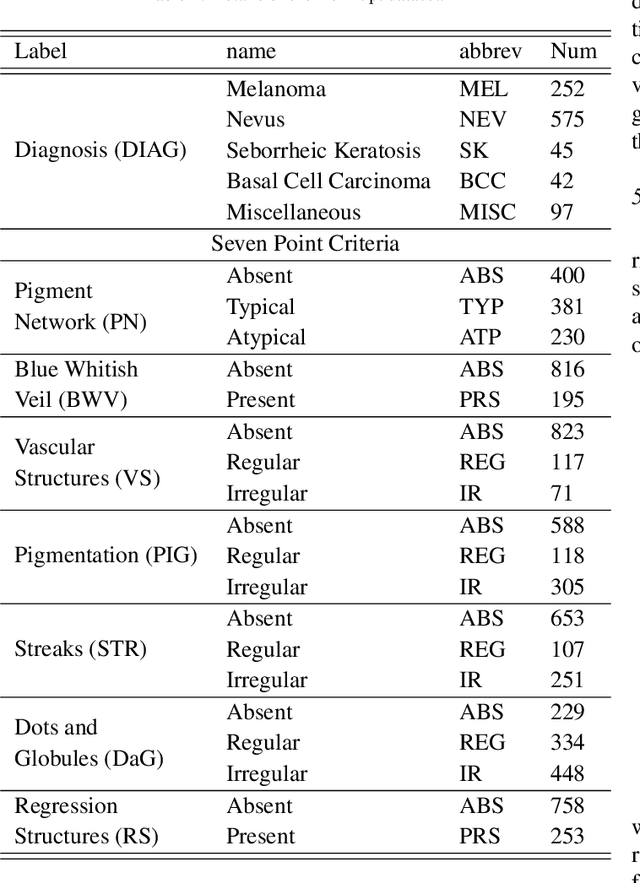
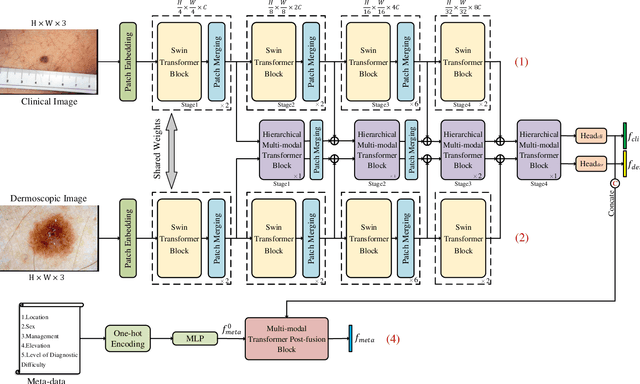

Multi-modal skin lesion diagnosis (MSLD) has achieved remarkable success by modern computer-aided diagnosis technology based on deep convolutions. However, the information aggregation across modalities in MSLD remains challenging due to severity unaligned spatial resolution (dermoscopic image and clinical image) and heterogeneous data (dermoscopic image and patients' meta-data). Limited by the intrinsic local attention, most recent MSLD pipelines using pure convolutions struggle to capture representative features in shallow layers, thus the fusion across different modalities is usually done at the end of the pipelines, even at the last layer, leading to an insufficient information aggregation. To tackle the issue, we introduce a pure transformer-based method, which we refer to as ``Throughout Fusion Transformer (TFormer)", for sufficient information intergration in MSLD. Different from the existing approaches with convolutions, the proposed network leverages transformer as feature extraction backbone, bringing more representative shallow features. We then carefully design a stack of dual-branch hierarchical multi-modal transformer (HMT) blocks to fuse information across different image modalities in a stage-by-stage way. With the aggregated information of image modalities, a multi-modal transformer post-fusion (MTP) block is designed to integrate features across image and non-image data. Such a strategy that information of the image modalities is firstly fused then the heterogeneous ones enables us to better divide and conquer the two major challenges while ensuring inter-modality dynamics are effectively modeled. Experiments conducted on the public Derm7pt dataset validate the superiority of the proposed method. Our TFormer outperforms other state-of-the-art methods. Ablation experiments also suggest the effectiveness of our designs.
RE-Tagger: A light-weight Real-Estate Image Classifier
Jul 12, 2022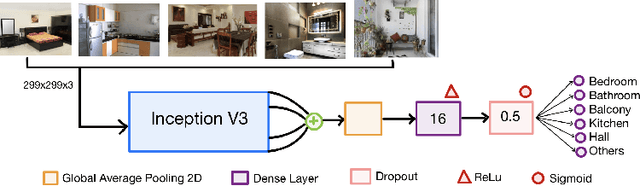
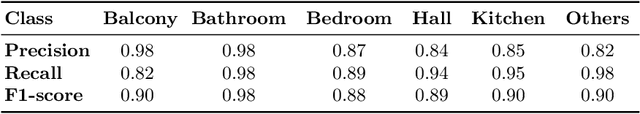
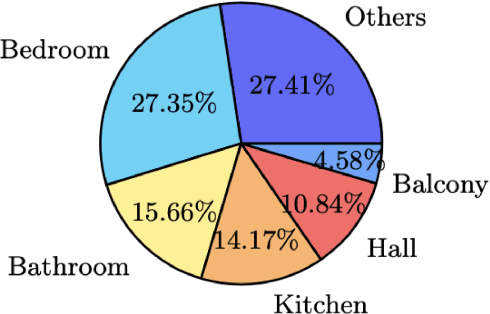

Real-estate image tagging is one of the essential use-cases to save efforts involved in manual annotation and enhance the user experience. This paper proposes an end-to-end pipeline (referred to as RE-Tagger) for the real-estate image classification problem. We present a two-stage transfer learning approach using custom InceptionV3 architecture to classify images into different categories (i.e., bedroom, bathroom, kitchen, balcony, hall, and others). Finally, we released the application as REST API hosted as a web application running on 2 cores machine with 2 GB RAM. The demo video is available here.
HumanGen: Generating Human Radiance Fields with Explicit Priors
Dec 10, 2022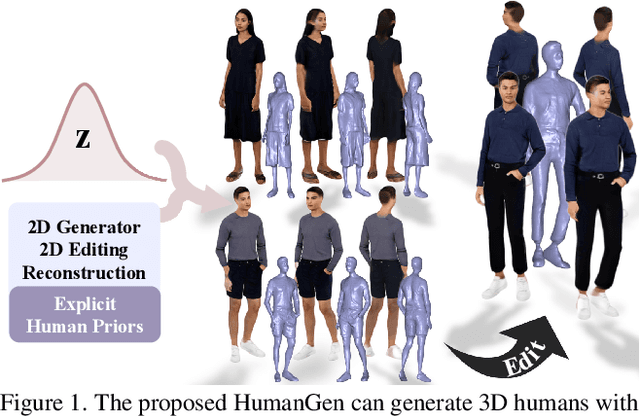
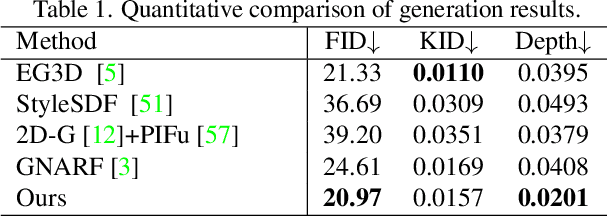
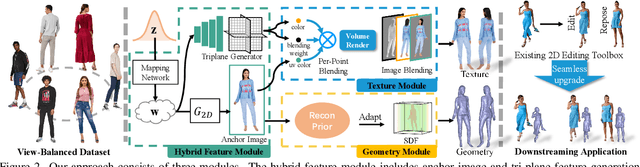
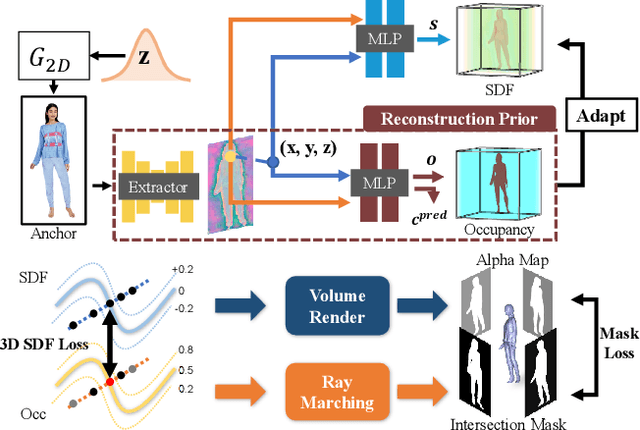
Recent years have witnessed the tremendous progress of 3D GANs for generating view-consistent radiance fields with photo-realism. Yet, high-quality generation of human radiance fields remains challenging, partially due to the limited human-related priors adopted in existing methods. We present HumanGen, a novel 3D human generation scheme with detailed geometry and $\text{360}^{\circ}$ realistic free-view rendering. It explicitly marries the 3D human generation with various priors from the 2D generator and 3D reconstructor of humans through the design of "anchor image". We introduce a hybrid feature representation using the anchor image to bridge the latent space of HumanGen with the existing 2D generator. We then adopt a pronged design to disentangle the generation of geometry and appearance. With the aid of the anchor image, we adapt a 3D reconstructor for fine-grained details synthesis and propose a two-stage blending scheme to boost appearance generation. Extensive experiments demonstrate our effectiveness for state-of-the-art 3D human generation regarding geometry details, texture quality, and free-view performance. Notably, HumanGen can also incorporate various off-the-shelf 2D latent editing methods, seamlessly lifting them into 3D.
Evaluation of Medical Image Segmentation Models for Uncertain, Small or Empty Reference Annotations
Sep 30, 2022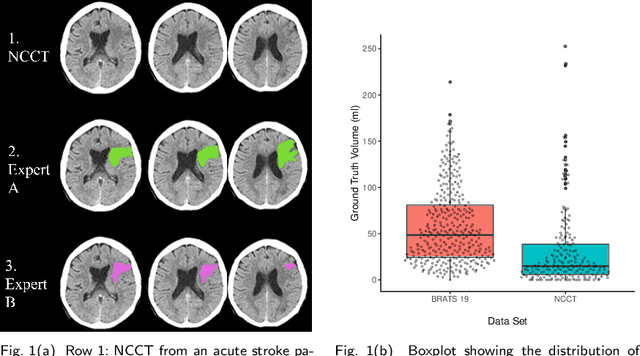
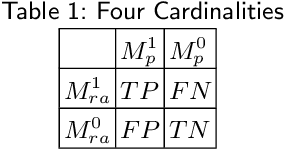


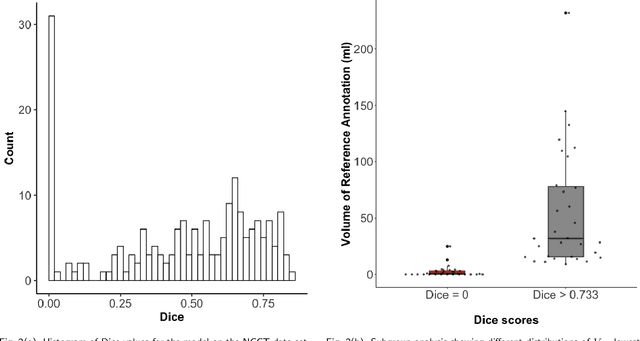
Performance metrics for medical image segmentation models are used to measure agreement between the reference annotation and the prediction. A common set of metrics is used in the development of such models to make results more comparable. However, there is a mismatch between the distributions in public data sets and cases encountered in clinical practice. Many common metrics fail to measure the impact of this mismatch, especially for clinical data sets containing uncertain, small or empty reference annotation. Thus, models may not be validated for clinically meaningful agreement by such metrics. Dimensions of evaluating clinical value include independence from reference annotation volume size, consideration of uncertainty of reference annotations, reward of volumetric and/or location agreement and reward of correct classification of empty reference annotations. Unlike common public data sets, our in-house data set is more representative. It contains uncertain, small or empty reference annotations. We examine publicly available metrics on the predictions of a deep learning framework in order to identify for which settings common metrics provide clinical meaningful results. We compare to a public benchmark data set without uncertain, small or empty reference annotations. https://github.com/SophieOstmeier/UncertainSmallEmpty
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge