"Image": models, code, and papers
Uncertainty-Aware Multiple-Instance Learning for Reliable Classification: Application to Optical Coherence Tomography
Feb 06, 2023
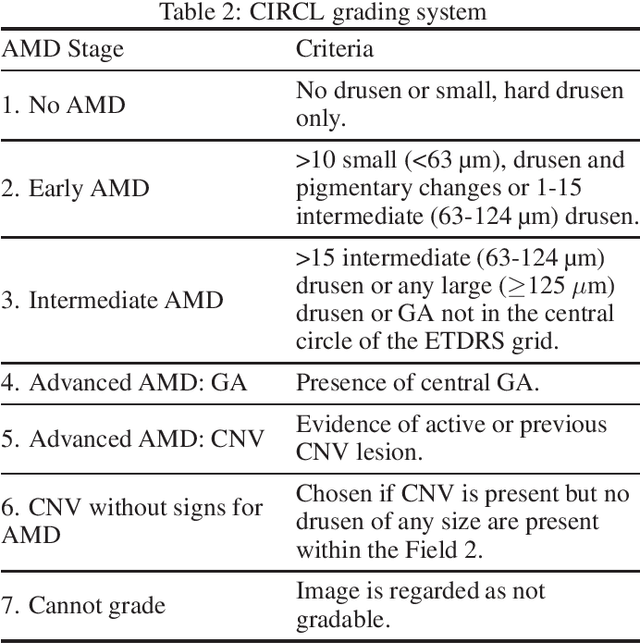
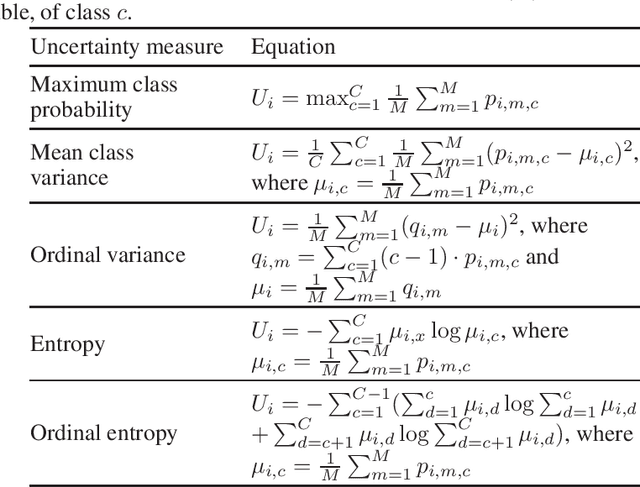
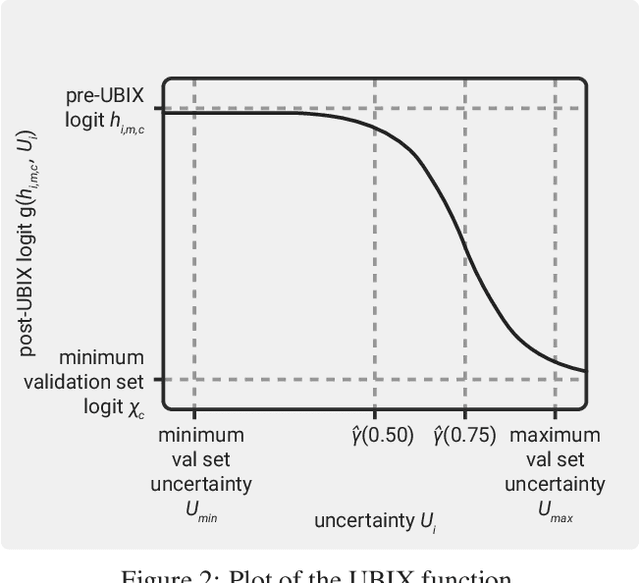

Deep learning classification models for medical image analysis often perform well on data from scanners that were used during training. However, when these models are applied to data from different vendors, their performance tends to drop substantially. Artifacts that only occur within scans from specific scanners are major causes of this poor generalizability. We aimed to improve the reliability of deep learning classification models by proposing Uncertainty-Based Instance eXclusion (UBIX). This technique, based on multiple-instance learning, reduces the effect of corrupted instances on the bag-classification by seamlessly integrating out-of-distribution (OOD) instance detection during inference. Although UBIX is generally applicable to different medical images and diverse classification tasks, we focused on staging of age-related macular degeneration in optical coherence tomography. After being trained using images from one vendor, UBIX showed a reliable behavior, with a slight decrease in performance (a decrease of the quadratic weighted kappa ($\kappa_w$) from 0.861 to 0.708), when applied to images from different vendors containing artifacts; while a state-of-the-art 3D neural network suffered from a significant detriment of performance ($\kappa_w$ from 0.852 to 0.084) on the same test set. We showed that instances with unseen artifacts can be identified with OOD detection and their contribution to the bag-level predictions can be reduced, improving reliability without the need for retraining on new data. This potentially increases the applicability of artificial intelligence models to data from other scanners than the ones for which they were developed.
CADA-GAN: Context-Aware GAN with Data Augmentation
Jan 21, 2023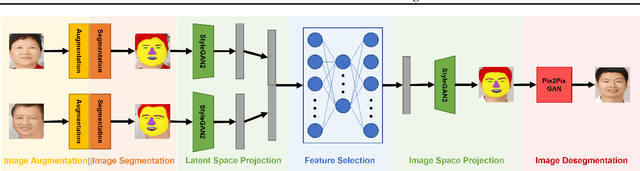


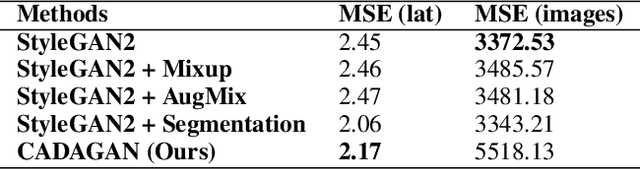
Current child face generators are restricted by the limited size of the available datasets. In addition, feature selection can prove to be a significant challenge, especially due to the large amount of features that need to be trained for. To manage these problems, we proposed CADA-GAN, a \textbf{C}ontext-\textbf{A}ware GAN that allows optimal feature extraction, with added robustness from additional \textbf{D}ata \textbf{A}ugmentation. CADA-GAN is adapted from the popular StyleGAN2-Ada model, with attention on augmentation and segmentation of the parent images. The model has the lowest \textit{Mean Squared Error Loss} (MSEloss) on latent feature representations and the generated child image is robust compared with the one that generated from baseline models.
FrustumFormer: Adaptive Instance-aware Resampling for Multi-view 3D Detection
Jan 10, 2023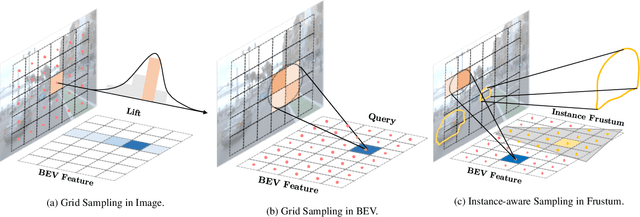
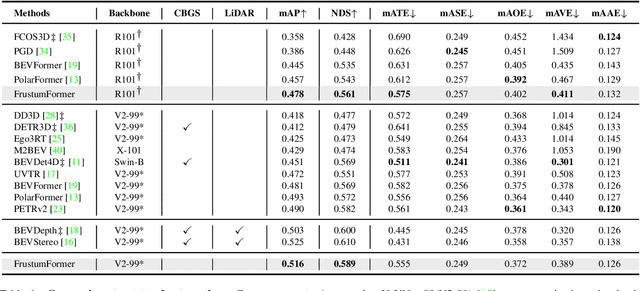
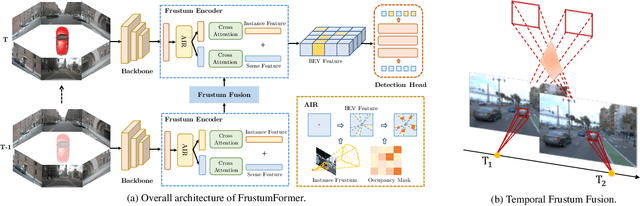
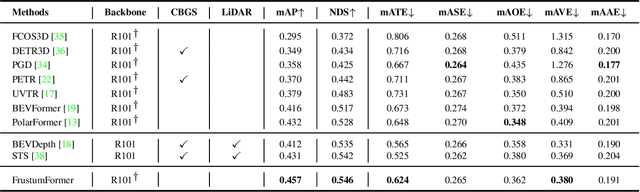
The transformation of features from 2D perspective space to 3D space is essential to multi-view 3D object detection. Recent approaches mainly focus on the design of view transformation, either pixel-wisely lifting perspective view features into 3D space with estimated depth or grid-wisely constructing BEV features via 3D projection, treating all pixels or grids equally. However, choosing what to transform is also important but has rarely been discussed before. The pixels of a moving car are more informative than the pixels of the sky. To fully utilize the information contained in images, the view transformation should be able to adapt to different image regions according to their contents. In this paper, we propose a novel framework named FrustumFormer, which pays more attention to the features in instance regions via adaptive instance-aware resampling. Specifically, the model obtains instance frustums on the bird's eye view by leveraging image view object proposals. An adaptive occupancy mask within the instance frustum is learned to refine the instance location. Moreover, the temporal frustum intersection could further reduce the localization uncertainty of objects. Comprehensive experiments on the nuScenes dataset demonstrate the effectiveness of FrustumFormer, and we achieve a new state-of-the-art performance on the benchmark. Codes will be released soon.
Self-Supervised Depth Estimation in Laparoscopic Image using 3D Geometric Consistency
Aug 17, 2022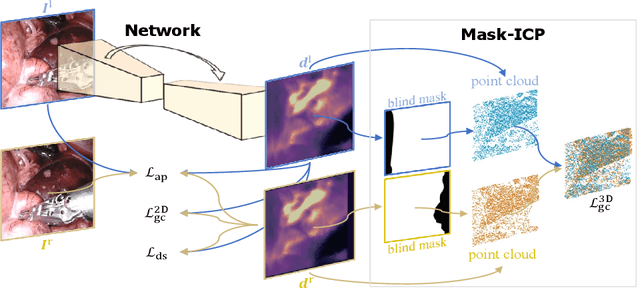
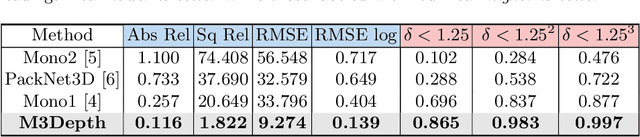
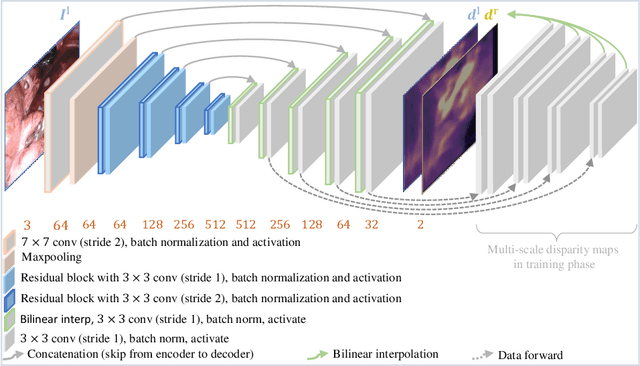
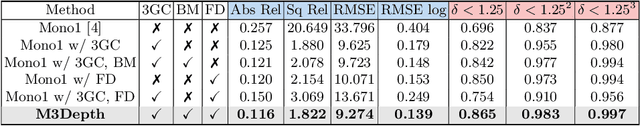
Depth estimation is a crucial step for image-guided intervention in robotic surgery and laparoscopic imaging system. Since per-pixel depth ground truth is difficult to acquire for laparoscopic image data, it is rarely possible to apply supervised depth estimation to surgical applications. As an alternative, self-supervised methods have been introduced to train depth estimators using only synchronized stereo image pairs. However, most recent work focused on the left-right consistency in 2D and ignored valuable inherent 3D information on the object in real world coordinates, meaning that the left-right 3D geometric structural consistency is not fully utilized. To overcome this limitation, we present M3Depth, a self-supervised depth estimator to leverage 3D geometric structural information hidden in stereo pairs while keeping monocular inference. The method also removes the influence of border regions unseen in at least one of the stereo images via masking, to enhance the correspondences between left and right images in overlapping areas. Intensive experiments show that our method outperforms previous self-supervised approaches on both a public dataset and a newly acquired dataset by a large margin, indicating a good generalization across different samples and laparoscopes.
Learning with Limited Annotations: A Survey on Deep Semi-Supervised Learning for Medical Image Segmentation
Aug 13, 2022
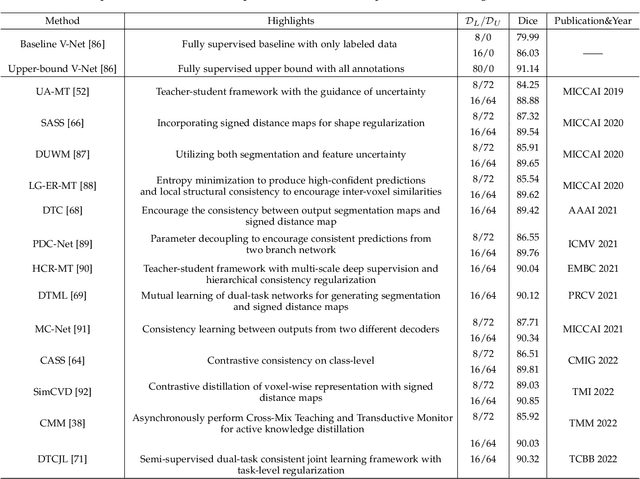
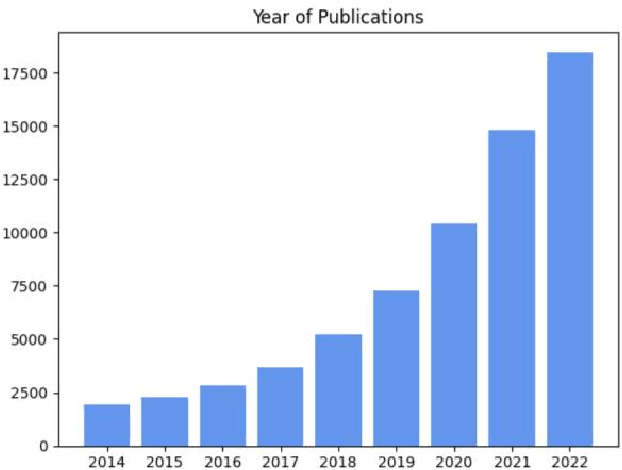
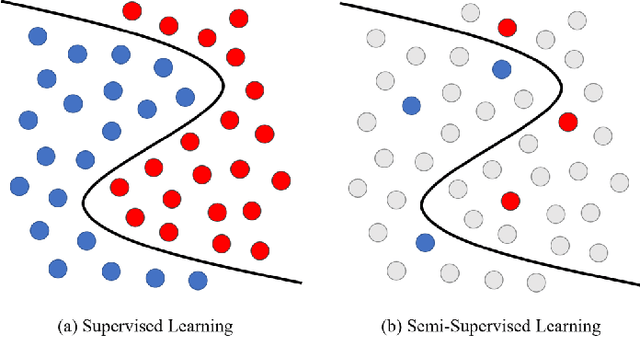
Medical image segmentation is a fundamental and critical step in many image-guided clinical approaches. Recent success of deep learning-based segmentation methods usually relies on a large amount of labeled data, which is particularly difficult and costly to obtain especially in the medical imaging domain where only experts can provide reliable and accurate annotations. Semi-supervised learning has emerged as an appealing strategy and been widely applied to medical image segmentation tasks to train deep models with limited annotations. In this paper, we present a comprehensive review of recently proposed semi-supervised learning methods for medical image segmentation and summarized both the technical novelties and empirical results. Furthermore, we analyze and discuss the limitations and several unsolved problems of existing approaches. We hope this review could inspire the research community to explore solutions for this challenge and further promote the developments in medical image segmentation field.
Basic Binary Convolution Unit for Binarized Image Restoration Network
Oct 02, 2022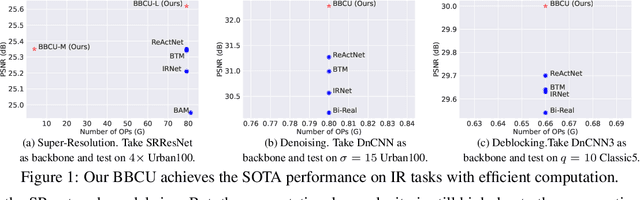
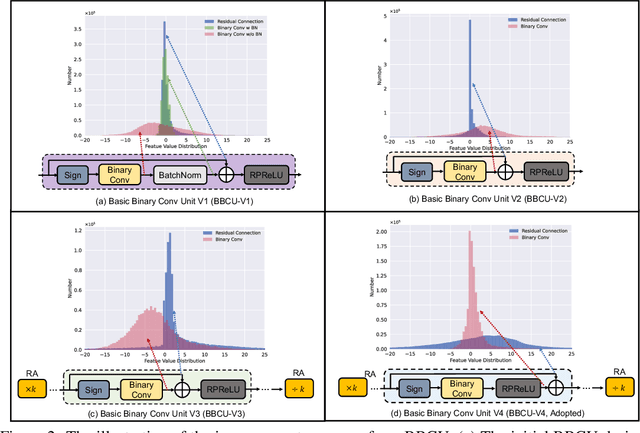
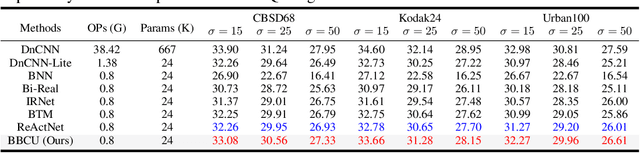
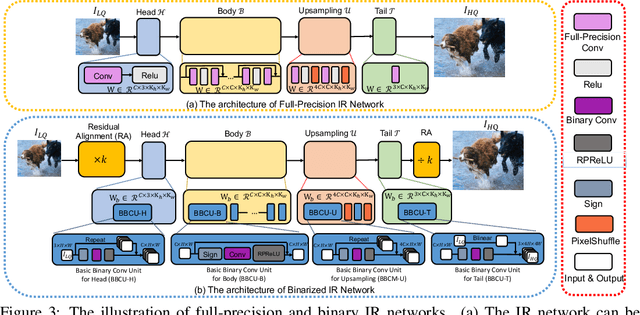
Lighter and faster image restoration (IR) models are crucial for the deployment on resource-limited devices. Binary neural network (BNN), one of the most promising model compression methods, can dramatically reduce the computations and parameters of full-precision convolutional neural networks (CNN). However, there are different properties between BNN and full-precision CNN, and we can hardly use the experience of designing CNN to develop BNN. In this study, we reconsider components in binary convolution, such as residual connection, BatchNorm, activation function, and structure, for IR tasks. We conduct systematic analyses to explain each component's role in binary convolution and discuss the pitfalls. Specifically, we find that residual connection can reduce the information loss caused by binarization; BatchNorm can solve the value range gap between residual connection and binary convolution; The position of the activation function dramatically affects the performance of BNN. Based on our findings and analyses, we design a simple yet efficient basic binary convolution unit (BBCU). Furthermore, we divide IR networks into four parts and specially design variants of BBCU for each part to explore the benefit of binarizing these parts. We conduct experiments on different IR tasks, and our BBCU significantly outperforms other BNNs and lightweight models, which shows that BBCU can serve as a basic unit for binarized IR networks. All codes and models will be released.
MOMA:Distill from Self-Supervised Teachers
Feb 04, 2023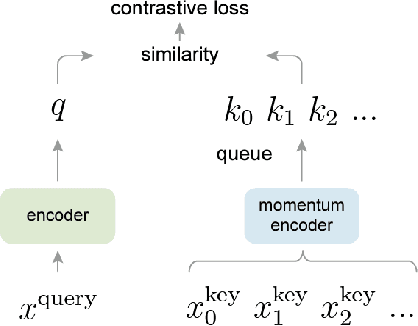
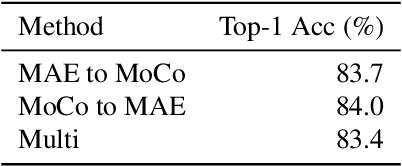
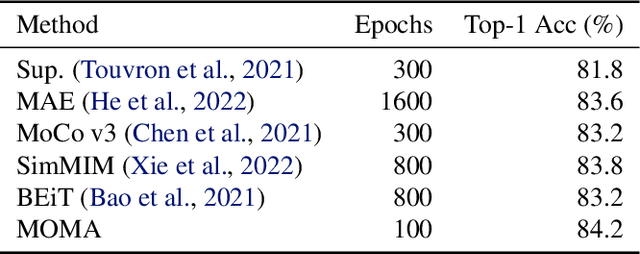
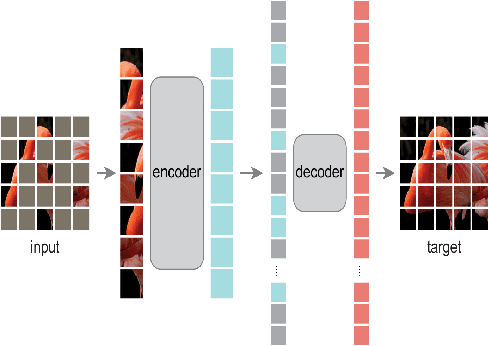

Contrastive Learning and Masked Image Modelling have demonstrated exceptional performance on self-supervised representation learning, where Momentum Contrast (i.e., MoCo) and Masked AutoEncoder (i.e., MAE) are the state-of-the-art, respectively. In this work, we propose MOMA to distill from pre-trained MoCo and MAE in a self-supervised manner to collaborate the knowledge from both paradigms. We introduce three different mechanisms of knowledge transfer in the propsoed MOMA framework. : (1) Distill pre-trained MoCo to MAE. (2) Distill pre-trained MAE to MoCo (3) Distill pre-trained MoCo and MAE to a random initialized student. During the distillation, the teacher and the student are fed with original inputs and masked inputs, respectively. The learning is enabled by aligning the normalized representations from the teacher and the projected representations from the student. This simple design leads to efficient computation with extremely high mask ratio and dramatically reduced training epochs, and does not require extra considerations on the distillation target. The experiments show MOMA delivers compact student models with comparable performance to existing state-of-the-art methods, combining the power of both self-supervised learning paradigms. It presents competitive results against different benchmarks in computer vision. We hope our method provides an insight on transferring and adapting the knowledge from large-scale pre-trained models in a computationally efficient way.
RevealED: Uncovering Pro-Eating Disorder Content on Twitter Using Deep Learning
Jan 02, 2023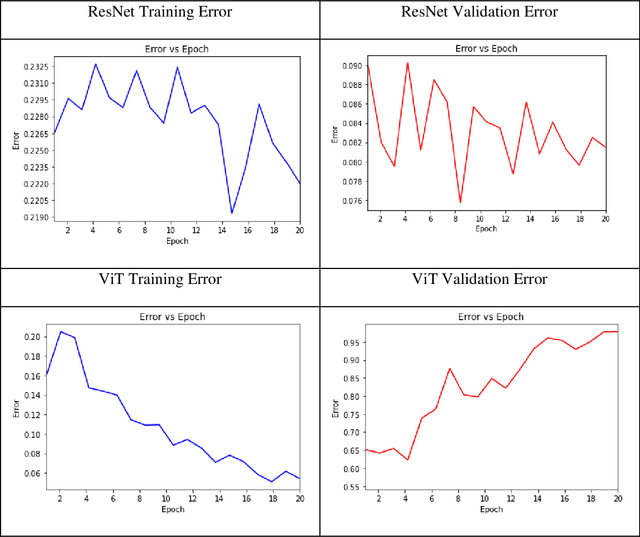
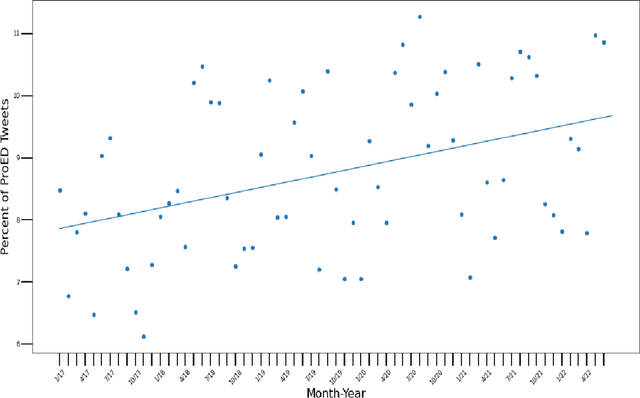
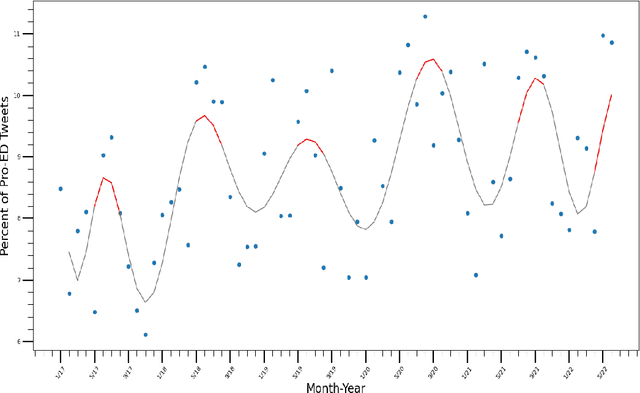
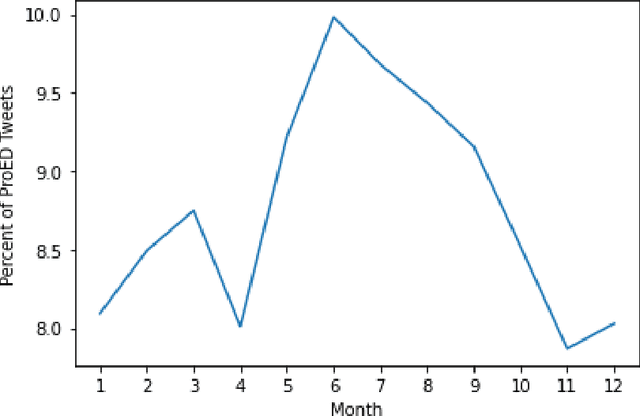
The Covid-19 pandemic induced a vast increase in adolescents diagnosed with eating disorders and hospitalized due to eating disorders. This immense growth stemmed partially from the stress of the pandemic but also from increased exposure to content that promotes eating disorders via social media, which, within the last decade, has become plagued by pro-eating disorder content. This study aimed to create a deep learning model capable of determining whether a given social media post promotes eating disorders based solely on image data. Tweets from hashtags that have been documented to promote eating disorders along with Tweets from unrelated hashtags were collected. After prepossessing, these images were labeled as either pro-eating disorder or not based on which Twitter hashtag they were scraped from. Several deep-learning models were trained on the scraped dataset and were evaluated based on their accuracy, F1 score, precision, and recall. Ultimately, the Vision Transformer model was determined to be the most accurate, attaining an F1 score of 0.877 and an accuracy of 86.7% on the test set. The model, which was applied to unlabeled Twitter image data scraped from "#selfie", uncovered seasonal fluctuations in the relative abundance of pro-eating disorder content, which reached its peak in the summertime. These fluctuations correspond not only to the seasons, but also to stressors, such as the Covid-19 pandemic. Moreover, the Twitter image data indicated that the relative amount of pro-eating disorder content has been steadily rising over the last five years and is likely to continue increasing in the future.
Rethinking Soft Label in Label Distribution Learning Perspective
Jan 31, 2023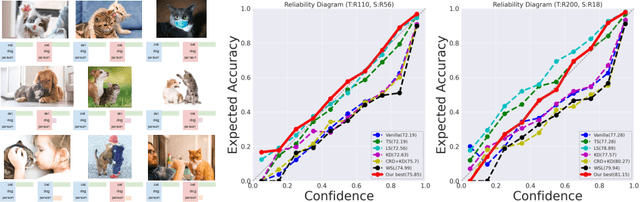
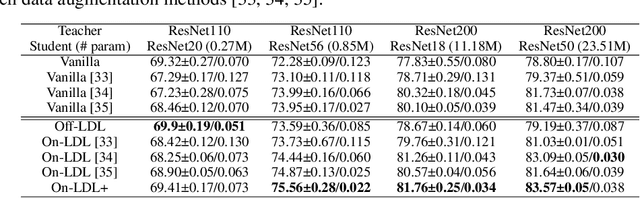
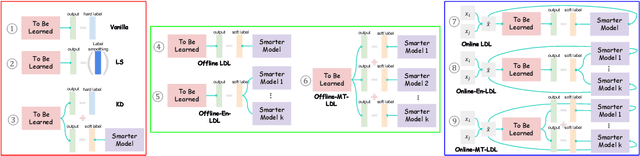
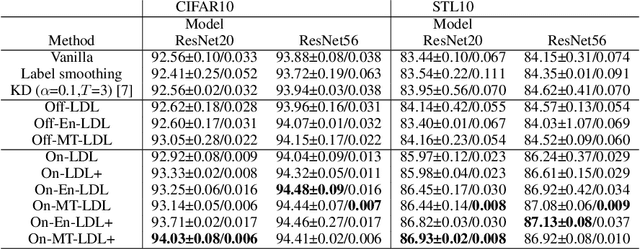
The primary goal of training in early convolutional neural networks (CNN) is the higher generalization performance of the model. However, as the expected calibration error (ECE), which quantifies the explanatory power of model inference, was recently introduced, research on training models that can be explained is in progress. We hypothesized that a gap in supervision criteria during training and inference leads to overconfidence, and investigated that performing label distribution learning (LDL) would enhance the model calibration in CNN training. To verify this assumption, we used a simple LDL setting with recent data augmentation techniques. Based on a series of experiments, the following results are obtained: 1) State-of-the-art KD methods significantly impede model calibration. 2) Training using LDL with recent data augmentation can have excellent effects on model calibration and even in generalization performance. 3) Online LDL brings additional improvements in model calibration and accuracy with long training, especially in large-size models. Using the proposed approach, we simultaneously achieved a lower ECE and higher generalization performance for the image classification datasets CIFAR10, 100, STL10, and ImageNet. We performed several visualizations and analyses and witnessed several interesting behaviors in CNN training with the LDL.
Biomedical image analysis competitions: The state of current participation practice
Dec 16, 2022The number of international benchmarking competitions is steadily increasing in various fields of machine learning (ML) research and practice. So far, however, little is known about the common practice as well as bottlenecks faced by the community in tackling the research questions posed. To shed light on the status quo of algorithm development in the specific field of biomedical imaging analysis, we designed an international survey that was issued to all participants of challenges conducted in conjunction with the IEEE ISBI 2021 and MICCAI 2021 conferences (80 competitions in total). The survey covered participants' expertise and working environments, their chosen strategies, as well as algorithm characteristics. A median of 72% challenge participants took part in the survey. According to our results, knowledge exchange was the primary incentive (70%) for participation, while the reception of prize money played only a minor role (16%). While a median of 80 working hours was spent on method development, a large portion of participants stated that they did not have enough time for method development (32%). 25% perceived the infrastructure to be a bottleneck. Overall, 94% of all solutions were deep learning-based. Of these, 84% were based on standard architectures. 43% of the respondents reported that the data samples (e.g., images) were too large to be processed at once. This was most commonly addressed by patch-based training (69%), downsampling (37%), and solving 3D analysis tasks as a series of 2D tasks. K-fold cross-validation on the training set was performed by only 37% of the participants and only 50% of the participants performed ensembling based on multiple identical models (61%) or heterogeneous models (39%). 48% of the respondents applied postprocessing steps.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge