"Image": models, code, and papers
Inpainting borehole images using Generative Adversarial Networks
Jan 15, 2023
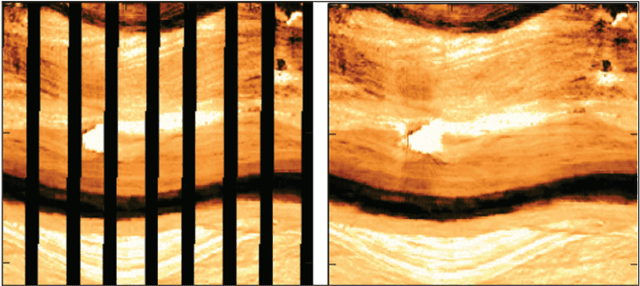
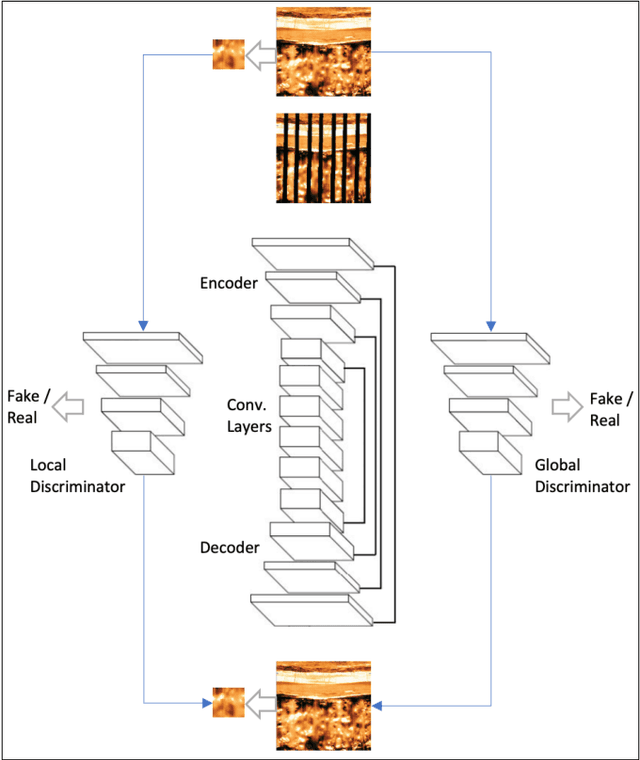

In this paper, we propose a GAN-based approach for gap filling in borehole images created by wireline microresistivity imaging tools. The proposed method utilizes a generator, global discriminator, and local discriminator to inpaint the missing regions of the image. The generator is based on an auto-encoder architecture with skip-connections, and the loss function used is the Wasserstein GAN loss. Our experiments on a dataset of borehole images demonstrate that the proposed model can effectively deal with large-scale missing pixels and generate realistic completion results. This approach can improve the quantitative evaluation of reservoirs and provide an essential basis for interpreting geological phenomena and reservoir parameters.
simple diffusion: End-to-end diffusion for high resolution images
Jan 26, 2023
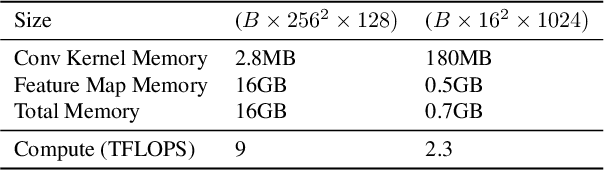

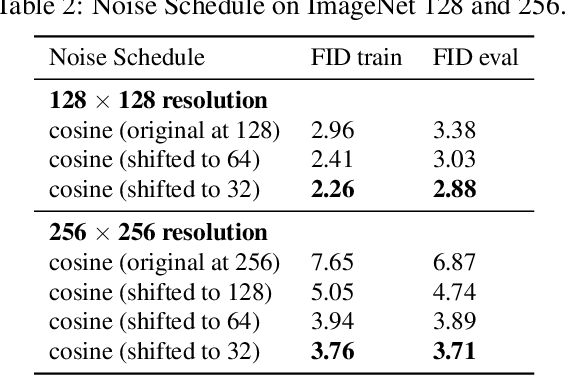
Currently, applying diffusion models in pixel space of high resolution images is difficult. Instead, existing approaches focus on diffusion in lower dimensional spaces (latent diffusion), or have multiple super-resolution levels of generation referred to as cascades. The downside is that these approaches add additional complexity to the diffusion framework. This paper aims to improve denoising diffusion for high resolution images while keeping the model as simple as possible. The paper is centered around the research question: How can one train a standard denoising diffusion models on high resolution images, and still obtain performance comparable to these alternate approaches? The four main findings are: 1) the noise schedule should be adjusted for high resolution images, 2) It is sufficient to scale only a particular part of the architecture, 3) dropout should be added at specific locations in the architecture, and 4) downsampling is an effective strategy to avoid high resolution feature maps. Combining these simple yet effective techniques, we achieve state-of-the-art on image generation among diffusion models without sampling modifiers on ImageNet.
Studying Therapy Effects and Disease Outcomes in Silico using Artificial Counterfactual Tissue Samples
Feb 06, 2023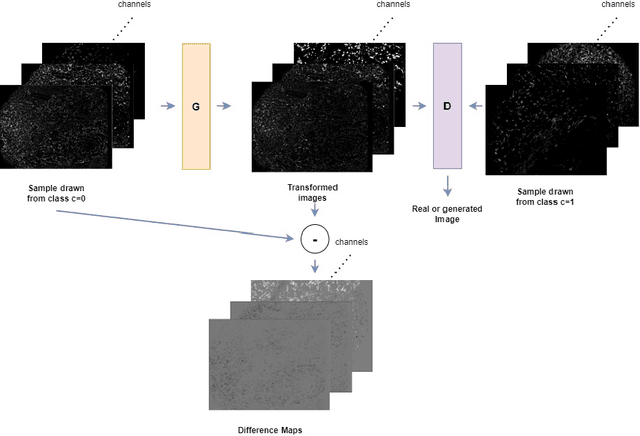
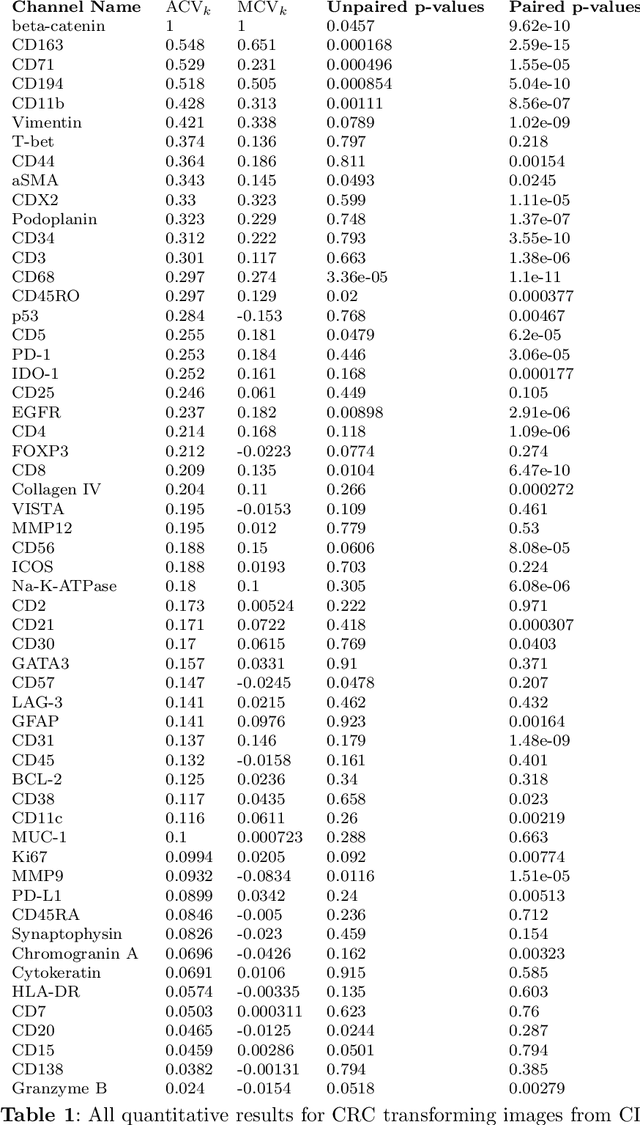
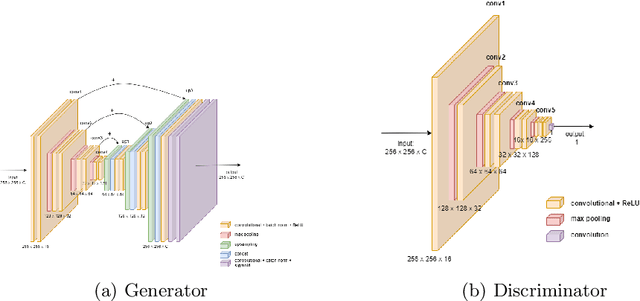
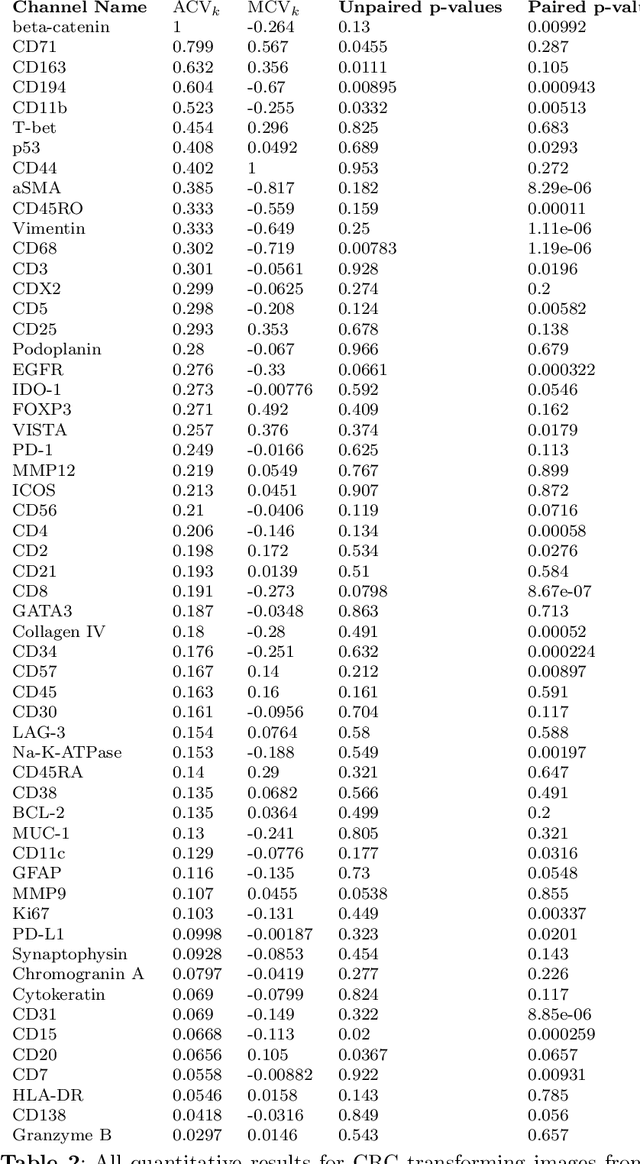
Understanding the interactions of different cell types inside the immune tumor microenvironment (iTME) is crucial for the development of immunotherapy treatments as well as for predicting their outcomes. Highly multiplexed tissue imaging (HMTI) technologies offer a tool which can capture cell properties of tissue samples by measuring expression of various proteins and storing them in separate image channels. HMTI technologies can be used to gain insights into the iTME and in particular how the iTME differs for different patient outcome groups of interest (e.g., treatment responders vs. non-responders). Understanding the systematic differences in the iTME of different patient outcome groups is crucial for developing better treatments and personalising existing treatments. However, such analyses are inherently limited by the fact that any two tissue samples vary due to a large number of factors unrelated to the outcome. Here, we present CF-HistoGAN, a machine learning framework that employs generative adversarial networks (GANs) to create artificial counterfactual tissue samples that resemble the original tissue samples as closely as possible but capture the characteristics of a different patient outcome group. Specifically, we learn to "translate" HMTI samples from one patient group to create artificial paired samples. We show that this approach allows to directly study the effects of different patient outcomes on the iTMEs of individual tissue samples. We demonstrate that CF-HistoGAN can be employed as an explorative tool for understanding iTME effects on the pixel level. Moreover, we show that our method can be used to identify statistically significant differences in the expression of different proteins between patient groups with greater sensitivity compared to conventional approaches.
Beyond Statistical Similarity: Rethinking Metrics for Deep Generative Models in Engineering Design
Feb 06, 2023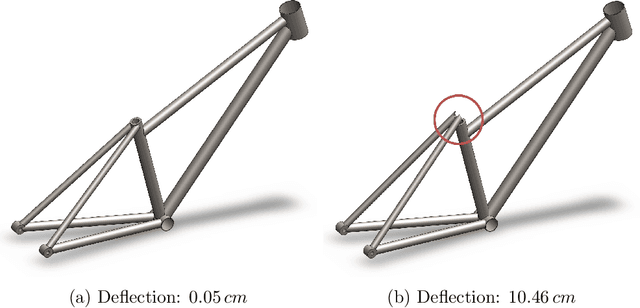
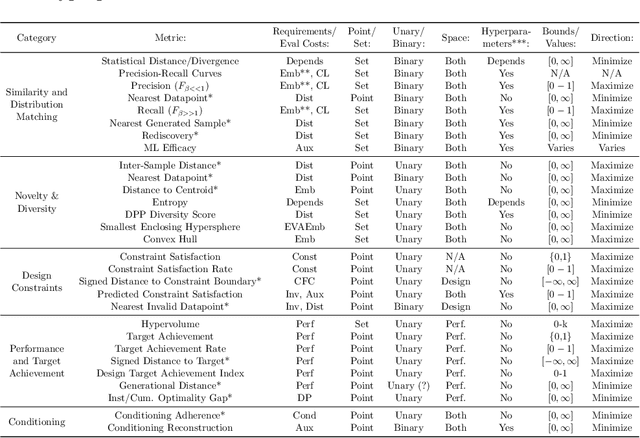
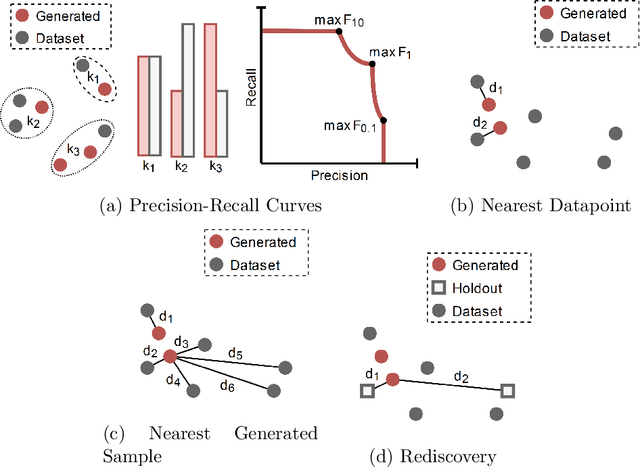
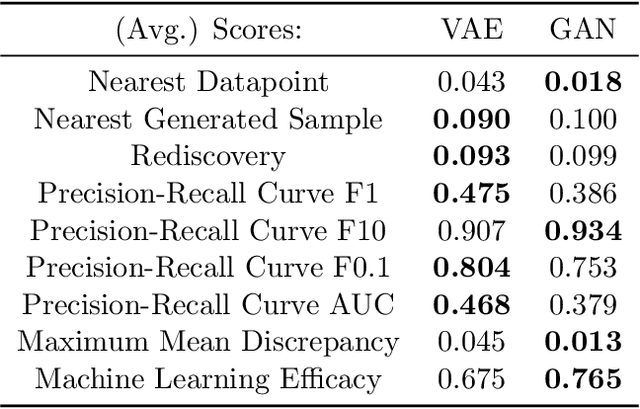
Deep generative models, such as Variational Autoencoders (VAEs), Generative Adversarial Networks (GANs), Diffusion Models, and Transformers, have shown great promise in a variety of applications, including image and speech synthesis, natural language processing, and drug discovery. However, when applied to engineering design problems, evaluating the performance of these models can be challenging, as traditional statistical metrics based on likelihood may not fully capture the requirements of engineering applications. This paper doubles as a review and a practical guide to evaluation metrics for deep generative models (DGMs) in engineering design. We first summarize well-accepted `classic' evaluation metrics for deep generative models grounded in machine learning theory and typical computer science applications. Using case studies, we then highlight why these metrics seldom translate well to design problems but see frequent use due to the lack of established alternatives. Next, we curate a set of design-specific metrics which have been proposed across different research communities and can be used for evaluating deep generative models. These metrics focus on unique requirements in design and engineering, such as constraint satisfaction, functional performance, novelty, and conditioning. We structure our review and discussion as a set of practical selection criteria and usage guidelines. Throughout our discussion, we apply the metrics to models trained on simple 2-dimensional example problems. Finally, to illustrate the selection process and classic usage of the presented metrics, we evaluate three deep generative models on a multifaceted bicycle frame design problem considering performance target achievement, design novelty, and geometric constraints. We publicly release the code for the datasets, models, and metrics used throughout the paper at decode.mit.edu/projects/metrics/.
Intra-operative Brain Tumor Detection with Deep Learning-Optimized Hyperspectral Imaging
Feb 06, 2023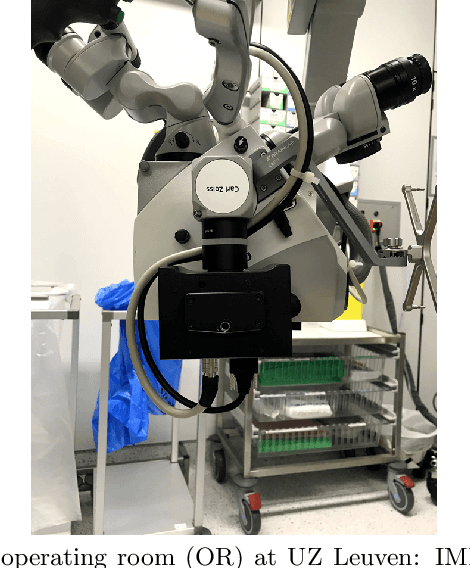
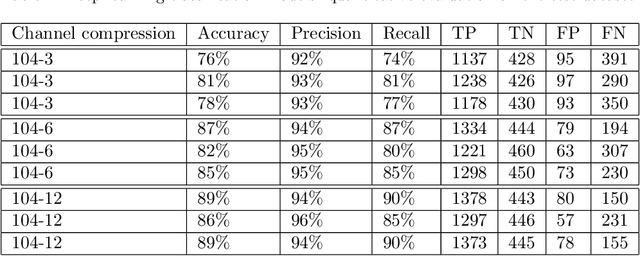
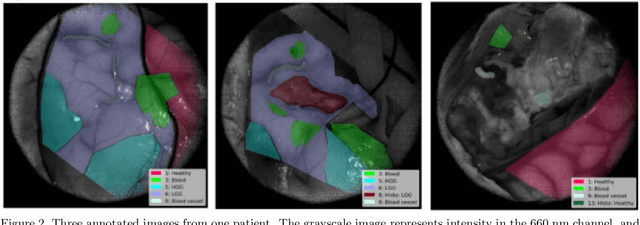
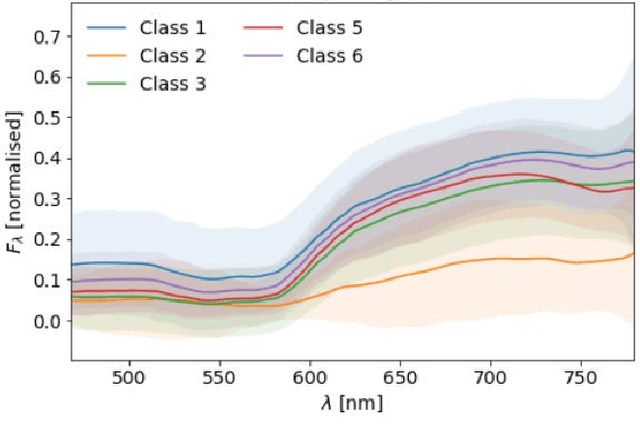
Surgery for gliomas (intrinsic brain tumors), especially when low-grade, is challenging due to the infiltrative nature of the lesion. Currently, no real-time, intra-operative, label-free and wide-field tool is available to assist and guide the surgeon to find the relevant demarcations for these tumors. While marker-based methods exist for the high-grade glioma case, there is no convenient solution available for the low-grade case; thus, marker-free optical techniques represent an attractive option. Although RGB imaging is a standard tool in surgical microscopes, it does not contain sufficient information for tissue differentiation. We leverage the richer information from hyperspectral imaging (HSI), acquired with a snapscan camera in the 468-787 nm range, coupled to a surgical microscope, to build a deep-learning-based diagnostic tool for cancer resection with potential for intra-operative guidance. However, the main limitation of the HSI snapscan camera is the image acquisition time, limiting its widespread deployment in the operation theater. Here, we investigate the effect of HSI channel reduction and pre-selection to scope the design space for the development of cheaper and faster sensors. Neural networks are used to identify the most important spectral channels for tumor tissue differentiation, optimizing the trade-off between the number of channels and precision to enable real-time intra-surgical application. We evaluate the performance of our method on a clinical dataset that was acquired during surgery on five patients. By demonstrating the possibility to efficiently detect low-grade glioma, these results can lead to better cancer resection demarcations, potentially improving treatment effectiveness and patient outcome.
Fine-Grained Hard Negative Mining: Generalizing Mitosis Detection with a Fifth of the MIDOG 2022 Dataset
Jan 03, 2023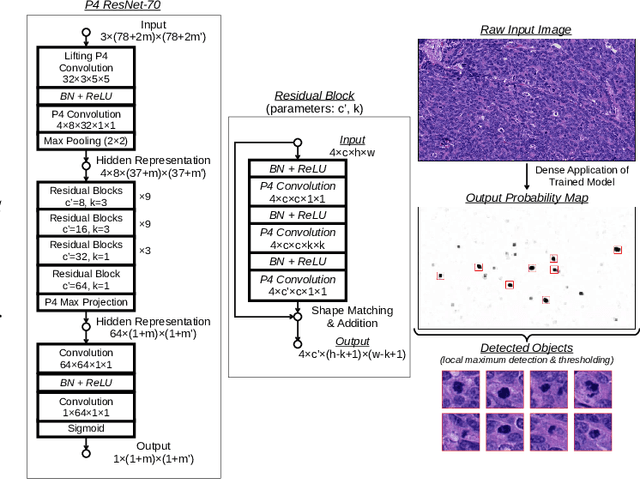
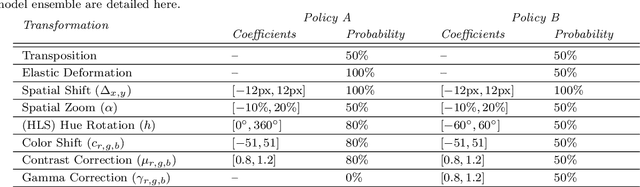
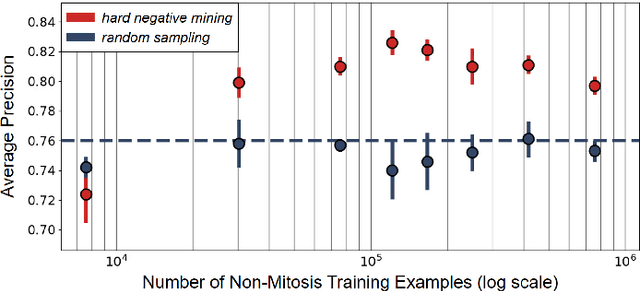
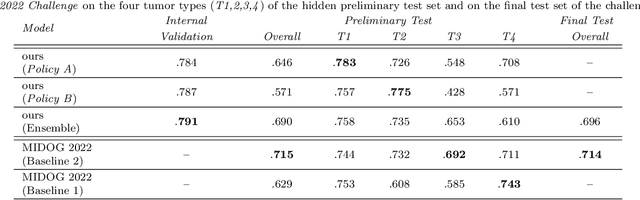
Making histopathology image classifiers robust to a wide range of real-world variability is a challenging task. Here, we describe a candidate deep learning solution for the Mitosis Domain Generalization Challenge 2022 (MIDOG) to address the problem of generalization for mitosis detection in images of hematoxylin-eosin-stained histology slides under high variability (scanner, tissue type and species variability). Our approach consists in training a rotation-invariant deep learning model using aggressive data augmentation with a training set enriched with hard negative examples and automatically selected negative examples from the unlabeled part of the challenge dataset. To optimize the performance of our models, we investigated a hard negative mining regime search procedure that lead us to train our best model using a subset of image patches representing 19.6% of our training partition of the challenge dataset. Our candidate model ensemble achieved a F1-score of .697 on the final test set after automated evaluation on the challenge platform, achieving the third best overall score in the MIDOG 2022 Challenge.
Bio-inspired Autonomous Exploration Policies with CNN-based Object Detection on Nano-drones
Feb 01, 2023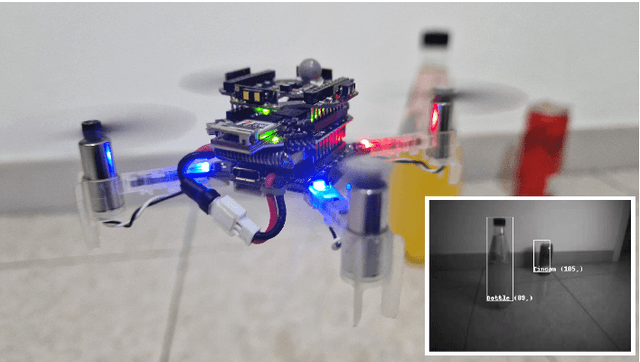
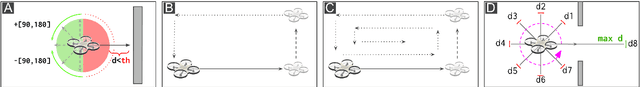
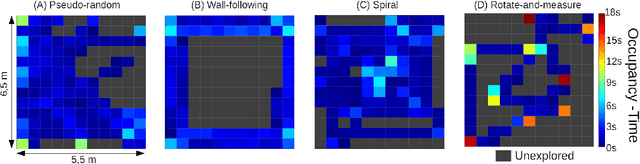

Nano-sized drones, with palm-sized form factor, are gaining relevance in the Internet-of-Things ecosystem. Achieving a high degree of autonomy for complex multi-objective missions (e.g., safe flight, exploration, object detection) is extremely challenging for the onboard chip-set due to tight size, payload (<10g), and power envelope constraints, which strictly limit both memory and computation. Our work addresses this complex problem by combining bio-inspired navigation policies, which rely on time-of-flight distance sensor data, with a vision-based convolutional neural network (CNN) for object detection. Our field-proven nano-drone is equipped with two microcontroller units (MCUs), a single-core ARM Cortex-M4 (STM32) for safe navigation and exploration policies, and a parallel ultra-low power octa-core RISC-V (GAP8) for onboard CNN inference, with a power envelope of just 134mW, including image sensors and external memories. The object detection task achieves a mean average precision of 50% (at 1.6 frame/s) on an in-field collected dataset. We compare four bio-inspired exploration policies and identify a pseudo-random policy to achieve the highest coverage area of 83% in a ~36m^2 unknown room in a 3 minutes flight. By combining the detection CNN and the exploration policy, we show an average detection rate of 90% on six target objects in a never-seen-before environment.
Synthesis-based Imaging-Differentiation Representation Learning for Multi-Sequence 3D/4D MRI
Feb 01, 2023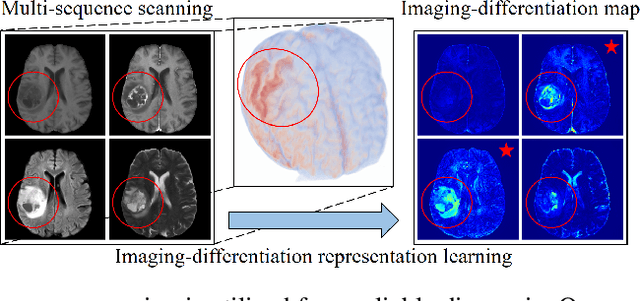
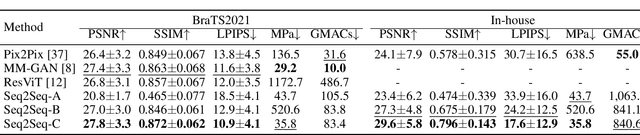
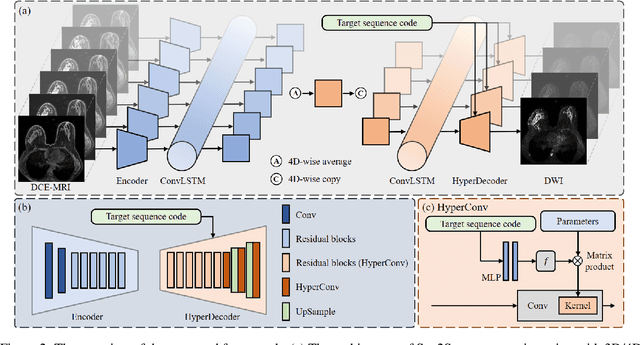
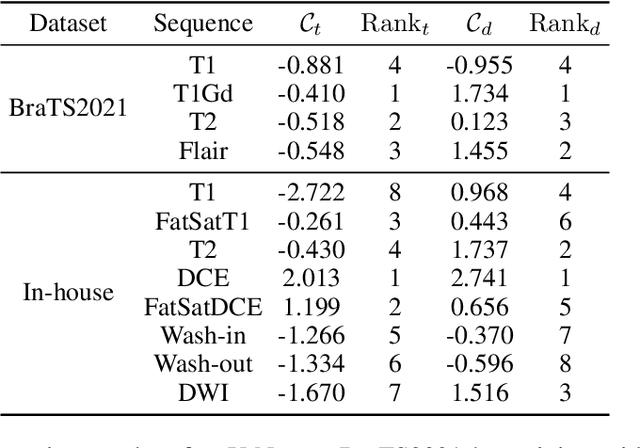
Multi-sequence MRIs can be necessary for reliable diagnosis in clinical practice due to the complimentary information within sequences. However, redundant information exists across sequences, which interferes with mining efficient representations by modern machine learning or deep learning models. To handle various clinical scenarios, we propose a sequence-to-sequence generation framework (Seq2Seq) for imaging-differentiation representation learning. In this study, not only do we propose arbitrary 3D/4D sequence generation within one model to generate any specified target sequence, but also we are able to rank the importance of each sequence based on a new metric estimating the difficulty of a sequence being generated. Furthermore, we also exploit the generation inability of the model to extract regions that contain unique information for each sequence. We conduct extensive experiments using three datasets including a toy dataset of 20,000 simulated subjects, a brain MRI dataset of 1,251 subjects, and a breast MRI dataset of 2,101 subjects, to demonstrate that (1) our proposed Seq2Seq is efficient and lightweight for complex clinical datasets and can achieve excellent image quality; (2) top-ranking sequences can be used to replace complete sequences with non-inferior performance; (3) combining MRI with our imaging-differentiation map leads to better performance in clinical tasks such as glioblastoma MGMT promoter methylation status prediction and breast cancer pathological complete response status prediction. Our code is available at https://github.com/fiy2W/mri_seq2seq.
Offline Clustering Approach to Self-supervised Learning for Class-imbalanced Image Data
Dec 22, 2022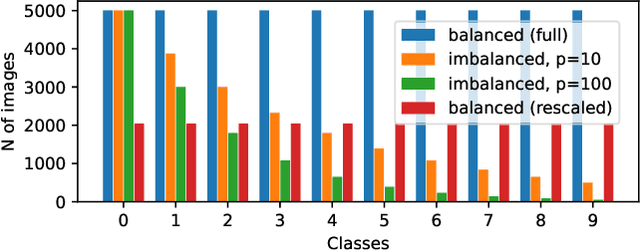
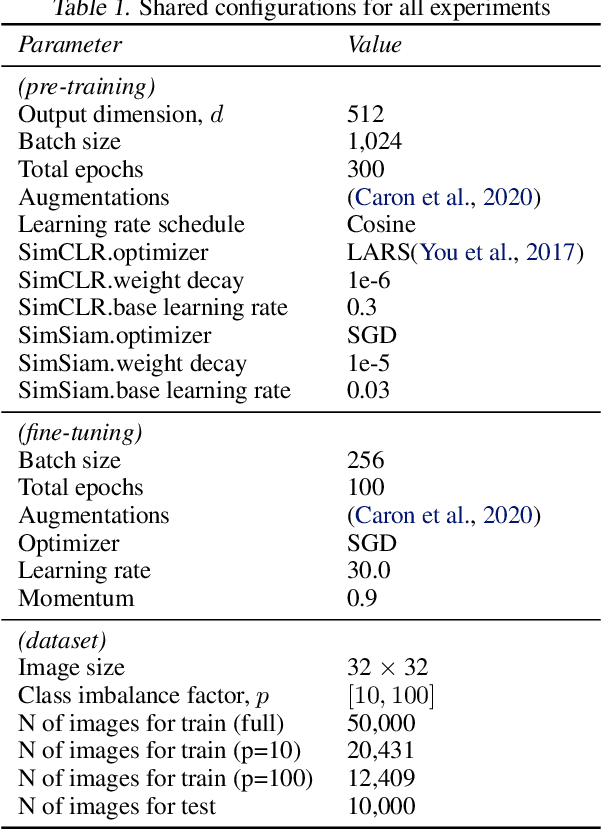
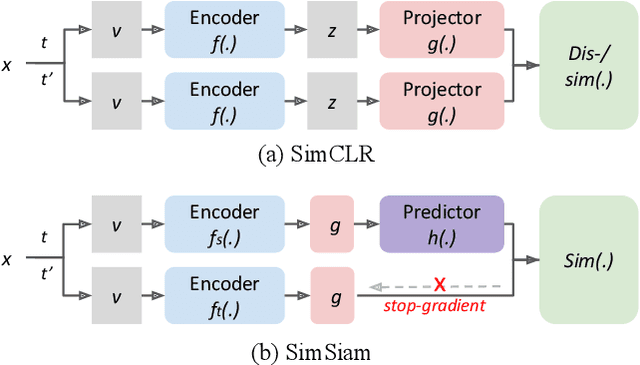
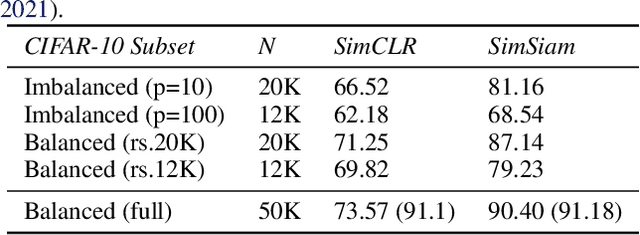
Class-imbalanced datasets are known to cause the problem of model being biased towards the majority classes. In this project, we set up two research questions: 1) when is the class-imbalance problem more prevalent in self-supervised pre-training? and 2) can offline clustering of feature representations help pre-training on class-imbalanced data? Our experiments investigate the former question by adjusting the degree of {\it class-imbalance} when training the baseline models, namely SimCLR and SimSiam on CIFAR-10 database. To answer the latter question, we train each expert model on each subset of the feature clusters. We then distill the knowledge of expert models into a single model, so that we will be able to compare the performance of this model to our baselines.
Artificial Image Tampering Distorts Spatial Distribution of Texture Landmarks and Quality Characteristics
Aug 04, 2022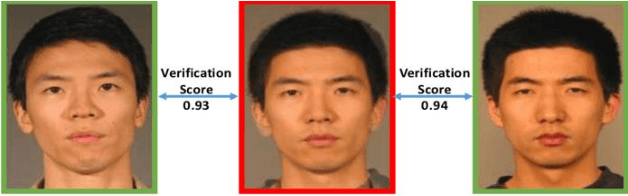
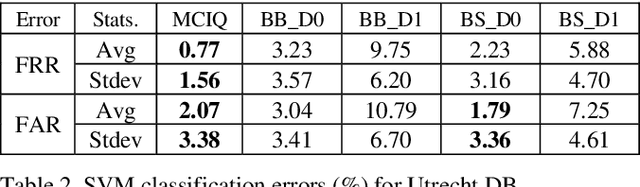
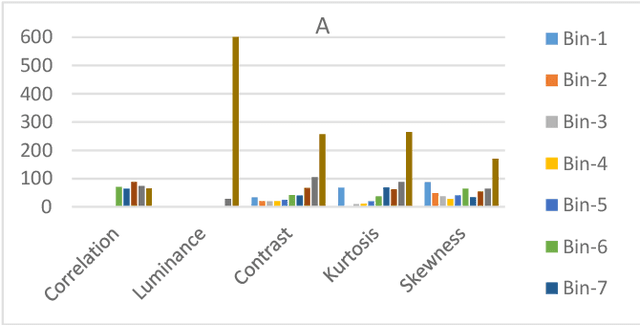
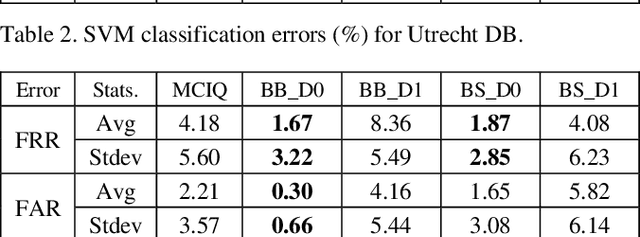
Advances in AI based computer vision has led to a significant growth in synthetic image generation and artificial image tampering with serious implications for unethical exploitations that undermine person identification and could make render AI predictions less explainable.Morphing, Deepfake and other artificial generation of face photographs undermine the reliability of face biometrics authentication using different electronic ID documents.Morphed face photographs on e-passports can fool automated border control systems and human guards.This paper extends our previous work on using the persistent homology (PH) of texture landmarks to detect morphing attacks.We demonstrate that artificial image tampering distorts the spatial distribution of texture landmarks (i.e. their PH) as well as that of a set of image quality characteristics.We shall demonstrate that the tamper caused distortion of these two slim feature vectors provide significant potentials for building explainable (Handcrafted) tamper detectors with low error rates and suitable for implementation on constrained devices.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge