"Image": models, code, and papers
Spherical Image Inpainting with Frame Transformation and Data-driven Prior Deep Networks
Sep 29, 2022
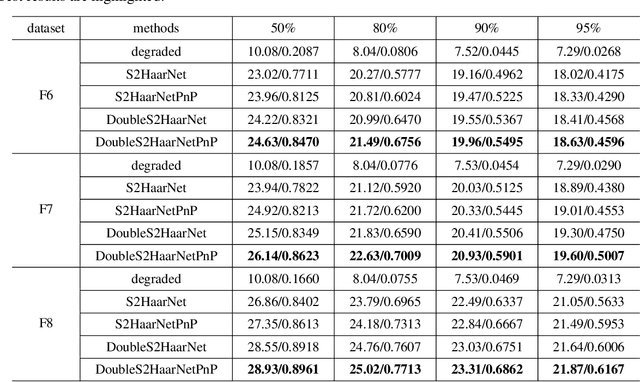
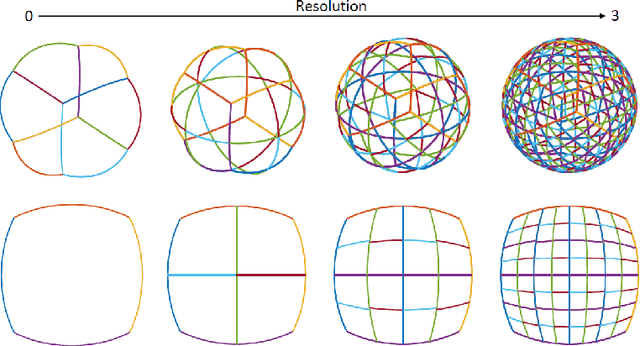
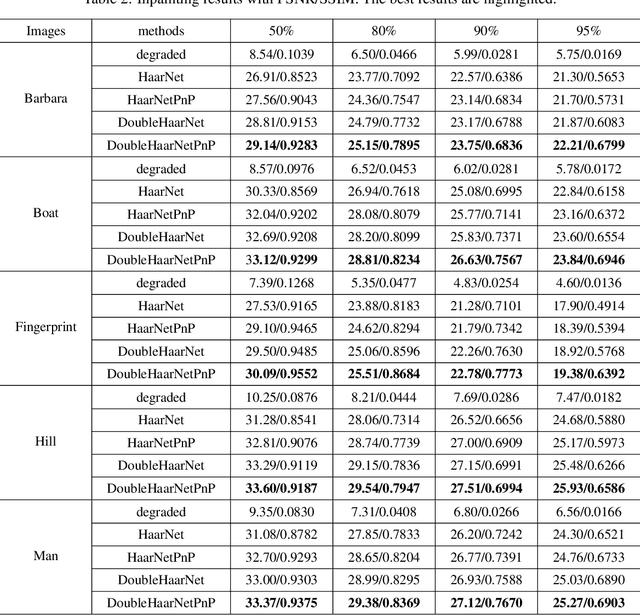
Spherical image processing has been widely applied in many important fields, such as omnidirectional vision for autonomous cars, global climate modelling, and medical imaging. It is non-trivial to extend an algorithm developed for flat images to the spherical ones. In this work, we focus on the challenging task of spherical image inpainting with deep learning-based regularizer. Instead of a naive application of existing models for planar images, we employ a fast directional spherical Haar framelet transform and develop a novel optimization framework based on a sparsity assumption of the framelet transform. Furthermore, by employing progressive encoder-decoder architecture, a new and better-performed deep CNN denoiser is carefully designed and works as an implicit regularizer. Finally, we use a plug-and-play method to handle the proposed optimization model, which can be implemented efficiently by training the CNN denoiser prior. Numerical experiments are conducted and show that the proposed algorithms can greatly recover damaged spherical images and achieve the best performance over purely using deep learning denoiser and plug-and-play model.
Automated Assessment of Transthoracic Echocardiogram Image Quality Using Deep Neural Networks
Sep 02, 2022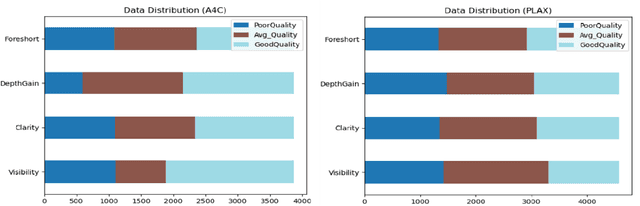
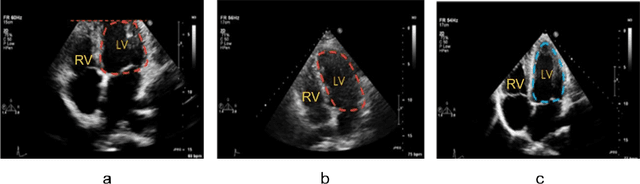
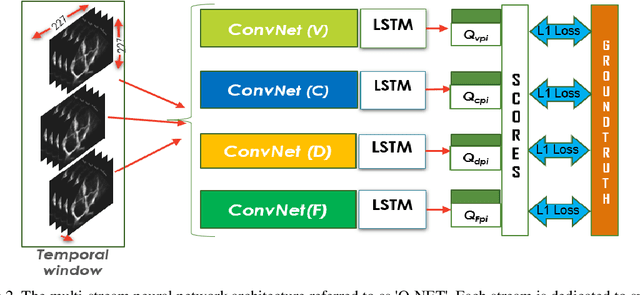
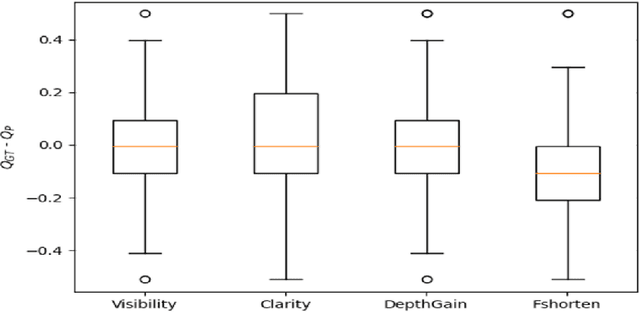
Standard views in two-dimensional echocardiography are well established but the quality of acquired images are highly dependent on operator skills and are assessed subjectively. This study is aimed at providing an objective assessment pipeline for echocardiogram image quality by defining a new set of domain-specific quality indicators. Consequently, image quality assessment can thus be automated to enhance clinical measurements, interpretation, and real-time optimization. We have developed deep neural networks for the automated assessment of echocardiographic frame which were randomly sampled from 11,262 adult patients. The private echocardiography dataset consists of 33,784 frames, previously acquired between 2010 and 2020. Deep learning approaches were used to extract the spatiotemporal features and the image quality indicators were evaluated against the mean absolute error. Our quality indicators encapsulate both anatomical and pathological elements to provide multivariate assessment scores for anatomical visibility, clarity, depth-gain and foreshortedness, respectively.
Deep Learning Computer Vision Algorithms for Real-time UAVs On-board Camera Image Processing
Nov 02, 2022
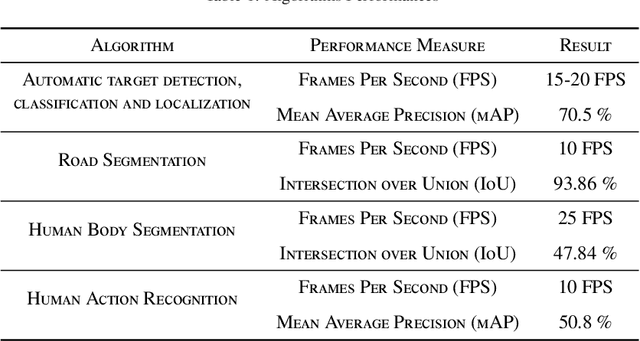



This paper describes how advanced deep learning based computer vision algorithms are applied to enable real-time on-board sensor processing for small UAVs. Four use cases are considered: target detection, classification and localization, road segmentation for autonomous navigation in GNSS-denied zones, human body segmentation, and human action recognition. All algorithms have been developed using state-of-the-art image processing methods based on deep neural networks. Acquisition campaigns have been carried out to collect custom datasets reflecting typical operational scenarios, where the peculiar point of view of a multi-rotor UAV is replicated. Algorithms architectures and trained models performances are reported, showing high levels of both accuracy and inference speed. Output examples and on-field videos are presented, demonstrating models operation when deployed on a GPU-powered commercial embedded device (NVIDIA Jetson Xavier) mounted on board of a custom quad-rotor, paving the way to enabling high level autonomy.
Image Reconstruction by Splitting Expectation Propagation Techniques from Iterative Inversion
Aug 25, 2022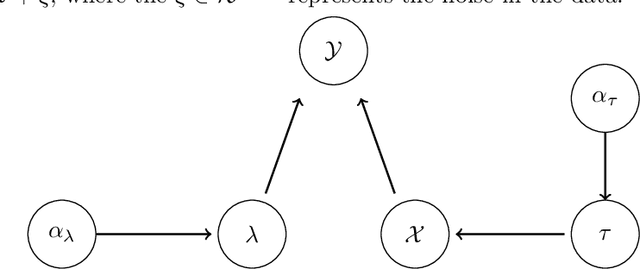
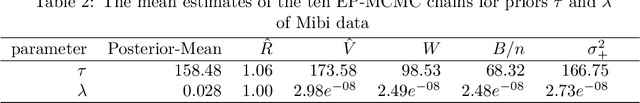

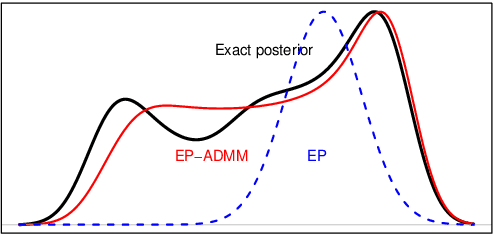

Reconstructing images from downsampled and noisy measurements, such as MRI and low dose Computed Tomography (CT), is a mathematically ill-posed inverse problem. We propose an easy-to-use reconstruction method based on Expectation Propagation (EP) techniques. We incorporate the Monte Carlo (MC) method, Markov Chain Monte Carlo (MCMC), and Alternating Direction Method of Multiplier (ADMM) algorithm into EP method to address the intractability issue encountered in EP. We demonstrate the approach on complex Bayesian models for image reconstruction. Our technique is applied to images from Gamma-camera scans. We compare EPMC, EP-MCMC, EP-ADMM methods with MCMC only. The metrics are the better image reconstruction, speed, and parameters estimation. Experiments with Gamma-camera imaging in real and simulated data show that our proposed method is convincingly less computationally expensive than MCMC and produces relatively a better image reconstruction.
Differentially Private Kernel Inducing Points (DP-KIP) for Privacy-preserving Data Distillation
Jan 31, 2023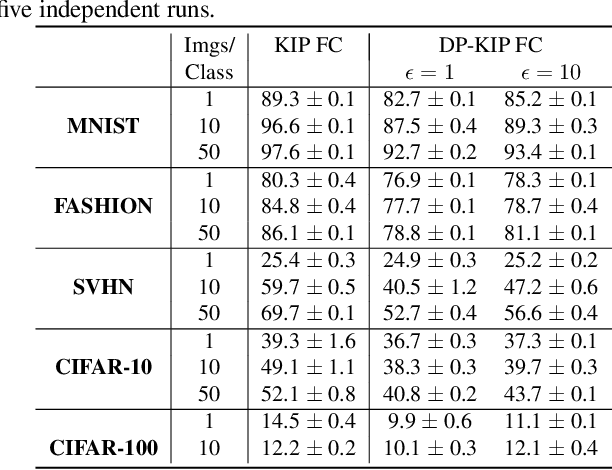
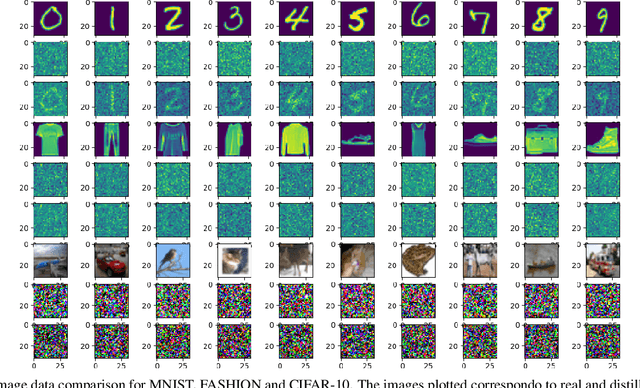

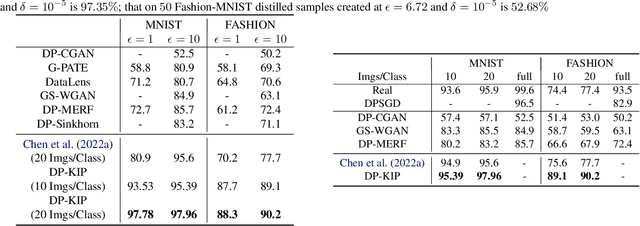
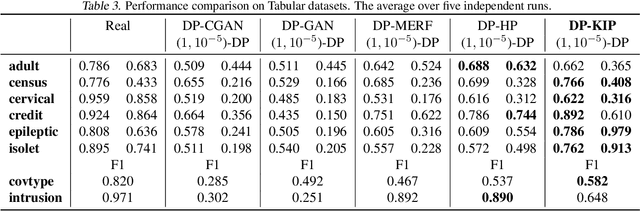
While it is tempting to believe that data distillation preserves privacy, distilled data's empirical robustness against known attacks does not imply a provable privacy guarantee. Here, we develop a provably privacy-preserving data distillation algorithm, called differentially private kernel inducing points (DP-KIP). DP-KIP is an instantiation of DP-SGD on kernel ridge regression (KRR). Following a recent work, we use neural tangent kernels and minimize the KRR loss to estimate the distilled datapoints (i.e., kernel inducing points). We provide a computationally efficient JAX implementation of DP-KIP, which we test on several popular image and tabular datasets to show its efficacy in data distillation with differential privacy guarantees.
Retrieval-Augmented Transformer for Image Captioning
Jul 26, 2022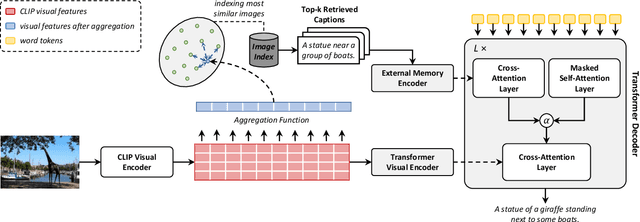

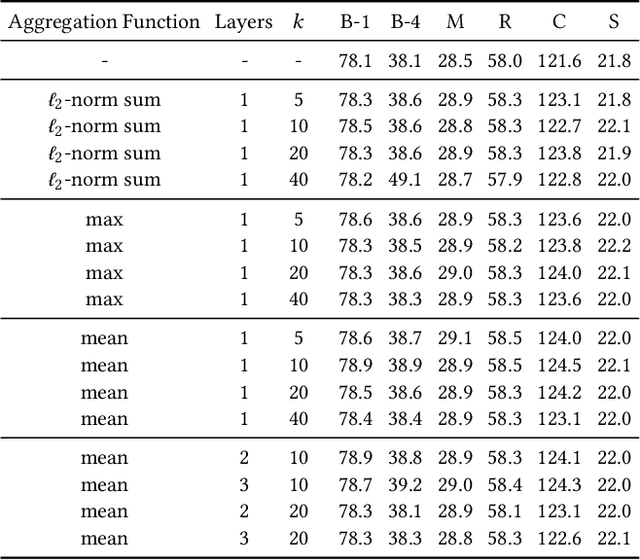
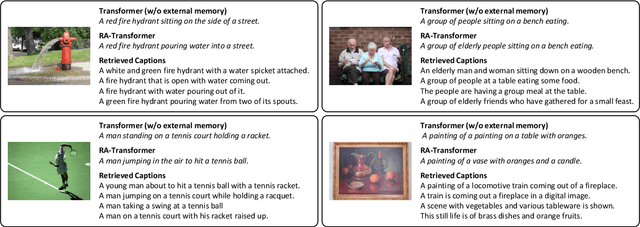
Image captioning models aim at connecting Vision and Language by providing natural language descriptions of input images. In the past few years, the task has been tackled by learning parametric models and proposing visual feature extraction advancements or by modeling better multi-modal connections. In this paper, we investigate the development of an image captioning approach with a kNN memory, with which knowledge can be retrieved from an external corpus to aid the generation process. Our architecture combines a knowledge retriever based on visual similarities, a differentiable encoder, and a kNN-augmented attention layer to predict tokens based on the past context and on text retrieved from the external memory. Experimental results, conducted on the COCO dataset, demonstrate that employing an explicit external memory can aid the generation process and increase caption quality. Our work opens up new avenues for improving image captioning models at larger scale.
Spiking Synaptic Penalty: Appropriate Penalty Term for Energy-Efficient Spiking Neural Networks
Feb 03, 2023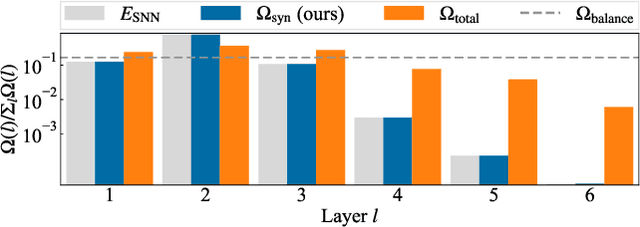
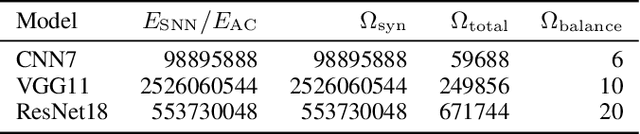

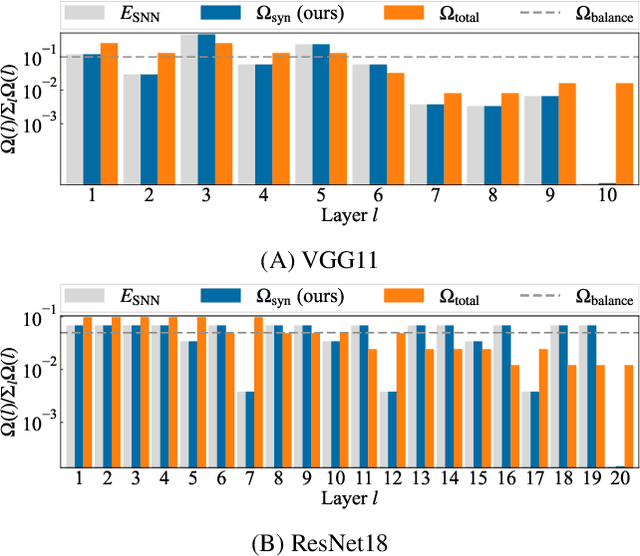
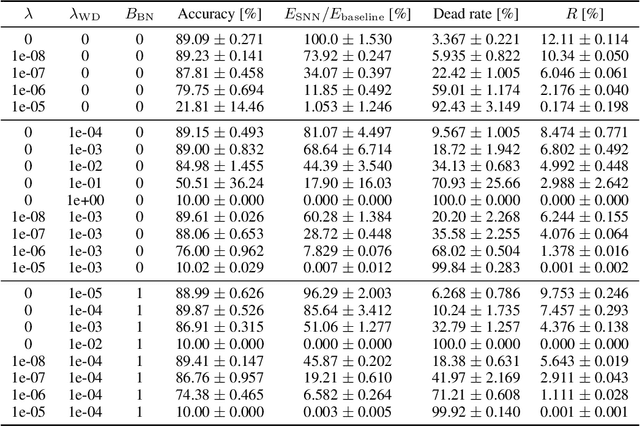
Spiking neural networks (SNNs) are energy-efficient neural networks because of their spiking nature. However, as the spike firing rate of SNNs increases, the energy consumption does as well, and thus, the advantage of SNNs diminishes. Here, we tackle this problem by introducing a novel penalty term for the spiking activity into the objective function in the training phase. Our method is designed so as to optimize the energy consumption metric directly without modifying the network architecture. Therefore, the proposed method can reduce the energy consumption more than other methods while maintaining the accuracy. We conducted experiments for image classification tasks, and the results indicate the effectiveness of the proposed method, which mitigates the dilemma of the energy--accuracy trade-off.
Quantifying U-Net Uncertainty in Multi-Parametric MRI-based Glioma Segmentation by Spherical Image Projection
Oct 12, 2022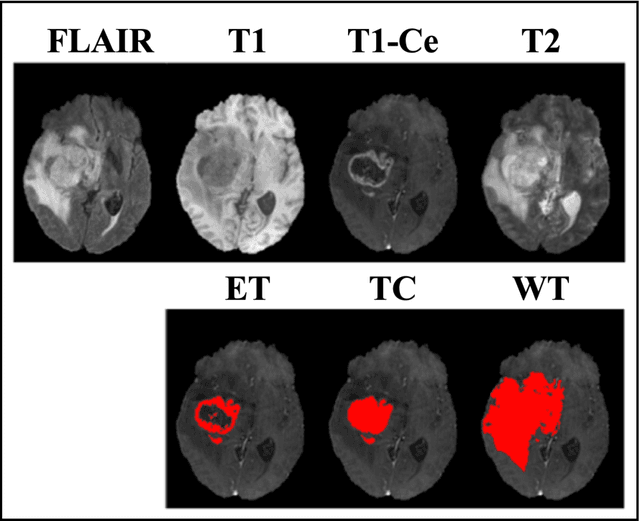

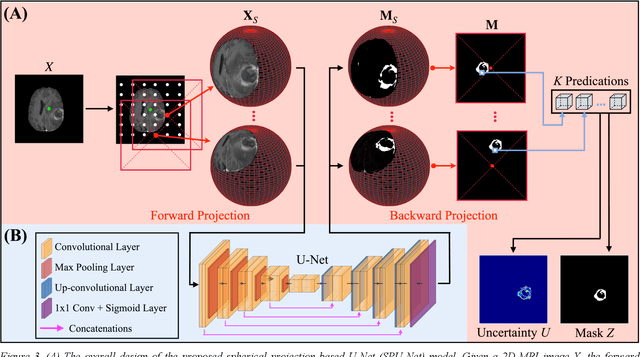
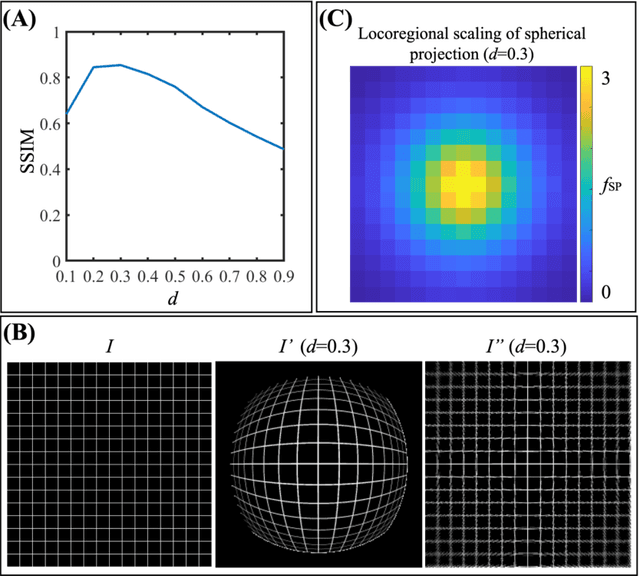
Purpose: To develop a U-Net segmentation uncertainty quantification method based on spherical image projection of multi-parametric MRI (MP-MRI) in glioma segmentation. Methods: The projection of planar MRI onto a spherical surface retains global anatomical information. By incorporating such image transformation in our proposed spherical projection-based U-Net (SPU-Net) segmentation model design, multiple segmentation predictions can be obtained for a single MRI. The final segmentation is the average of all predictions, and the variation can be shown as an uncertainty map. An uncertainty score was introduced to compare the uncertainty measurements' performance. The SPU-Net model was implemented on 369 glioma patients with MP-MRI scans. Three SPU-Nets were trained to segment enhancing tumor (ET), tumor core (TC), and whole tumor (WT), respectively. The SPU-Net was compared with (1) classic U-Net with test-time augmentation (TTA) and (2) linear scaling-based U-Net (LSU-Net) in both segmentation accuracy (Dice coefficient) and uncertainty (uncertainty map and uncertainty score). Results: The SPU-Net achieved low uncertainty for correct segmentation predictions (e.g., tumor interior or healthy tissue interior) and high uncertainty for incorrect results (e.g., tumor boundaries). This model could allow the identification of missed tumor targets or segmentation errors in U-Net. The SPU-Net achieved the highest uncertainty scores for 3 targets (ET/TC/WT): 0.826/0.848/0.936, compared to 0.784/0.643/0.872 for the U-Net with TTA and 0.743/0.702/0.876 for the LSU-Net. The SPU-Net also achieved statistically significantly higher Dice coefficients. Conclusion: The SPU-Net offers a powerful tool to quantify glioma segmentation uncertainty while improving segmentation accuracy. The proposed method can be generalized to other medical image-related deep-learning applications for uncertainty evaluation.
Information-Theoretic Diffusion
Feb 07, 2023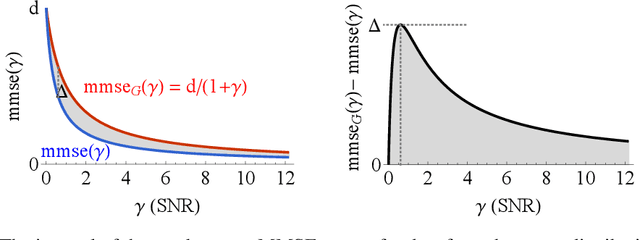
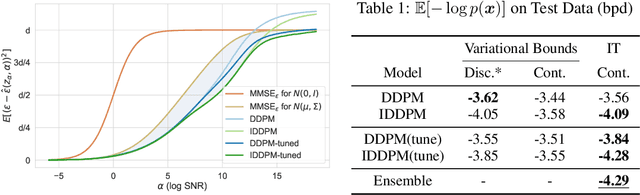
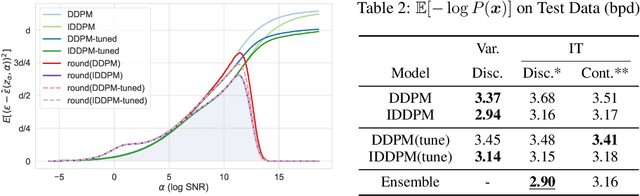
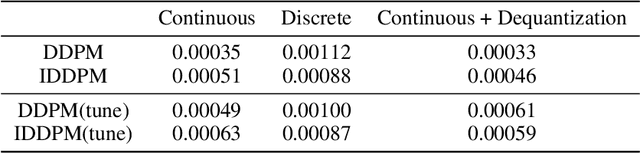
Denoising diffusion models have spurred significant gains in density modeling and image generation, precipitating an industrial revolution in text-guided AI art generation. We introduce a new mathematical foundation for diffusion models inspired by classic results in information theory that connect Information with Minimum Mean Square Error regression, the so-called I-MMSE relations. We generalize the I-MMSE relations to exactly relate the data distribution to an optimal denoising regression problem, leading to an elegant refinement of existing diffusion bounds. This new insight leads to several improvements for probability distribution estimation, including theoretical justification for diffusion model ensembling. Remarkably, our framework shows how continuous and discrete probabilities can be learned with the same regression objective, avoiding domain-specific generative models used in variational methods. Code to reproduce experiments is provided at http://github.com/kxh001/ITdiffusion and simplified demonstration code is at http://github.com/gregversteeg/InfoDiffusionSimple.
DrasCLR: A Self-supervised Framework of Learning Disease-related and Anatomy-specific Representation for 3D Medical Images
Feb 21, 2023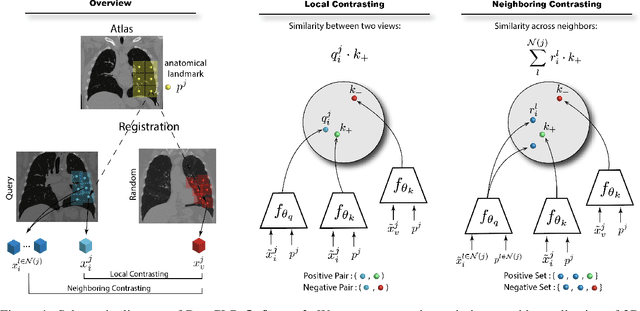
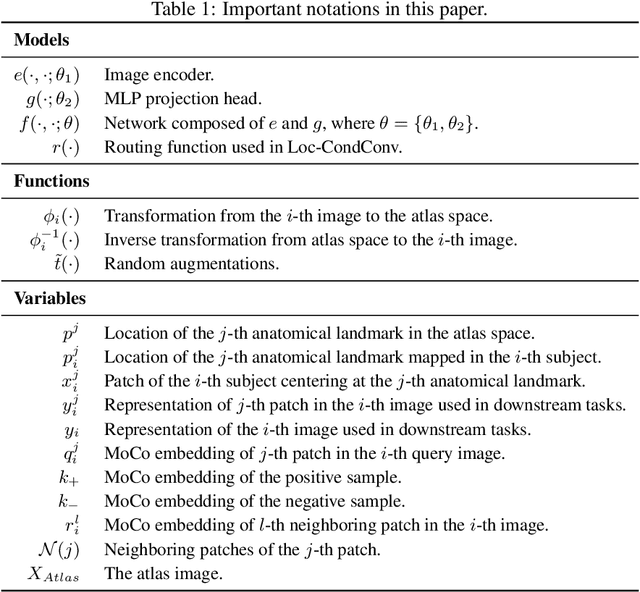
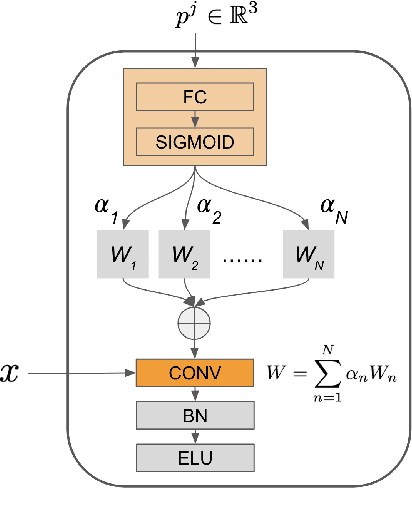
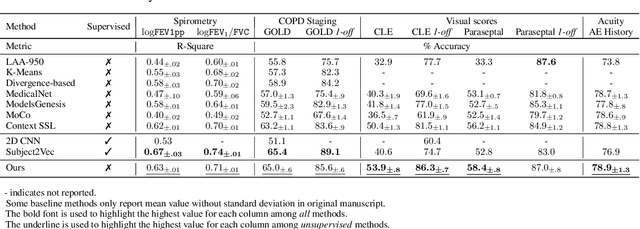
Large-scale volumetric medical images with annotation are rare, costly, and time prohibitive to acquire. Self-supervised learning (SSL) offers a promising pre-training and feature extraction solution for many downstream tasks, as it only uses unlabeled data. Recently, SSL methods based on instance discrimination have gained popularity in the medical imaging domain. However, SSL pre-trained encoders may use many clues in the image to discriminate an instance that are not necessarily disease-related. Moreover, pathological patterns are often subtle and heterogeneous, requiring the ability of the desired method to represent anatomy-specific features that are sensitive to abnormal changes in different body parts. In this work, we present a novel SSL framework, named DrasCLR, for 3D medical imaging to overcome these challenges. We propose two domain-specific contrastive learning strategies: one aims to capture subtle disease patterns inside a local anatomical region, and the other aims to represent severe disease patterns that span larger regions. We formulate the encoder using conditional hyper-parameterized network, in which the parameters are dependant on the anatomical location, to extract anatomically sensitive features. Extensive experiments on large-scale computer tomography (CT) datasets of lung images show that our method improves the performance of many downstream prediction and segmentation tasks. The patient-level representation improves the performance of the patient survival prediction task. We show how our method can detect emphysema subtypes via dense prediction. We demonstrate that fine-tuning the pre-trained model can significantly reduce annotation efforts without sacrificing emphysema detection accuracy. Our ablation study highlights the importance of incorporating anatomical context into the SSL framework.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge