"Image": models, code, and papers
Cross-Layer Retrospective Retrieving via Layer Attention
Feb 28, 2023
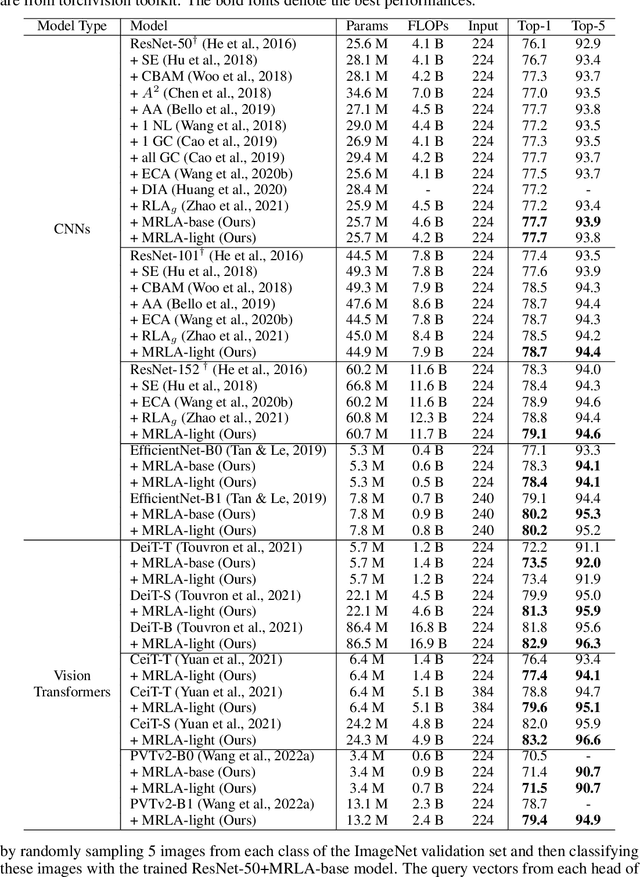
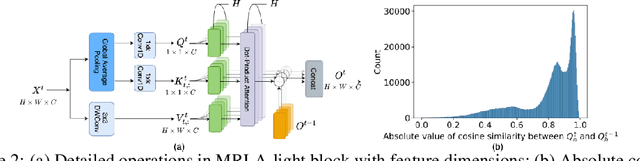
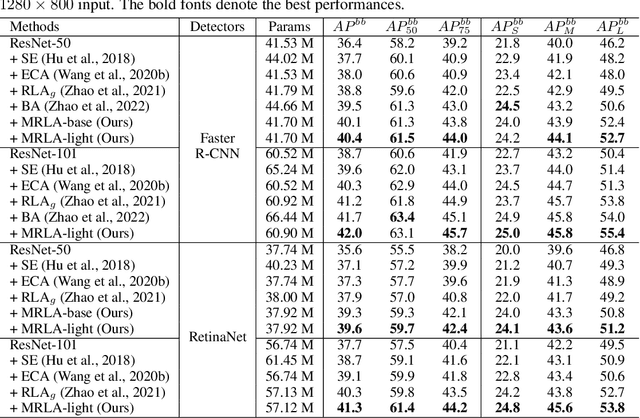
More and more evidence has shown that strengthening layer interactions can enhance the representation power of a deep neural network, while self-attention excels at learning interdependencies by retrieving query-activated information. Motivated by this, we devise a cross-layer attention mechanism, called multi-head recurrent layer attention (MRLA), that sends a query representation of the current layer to all previous layers to retrieve query-related information from different levels of receptive fields. A light-weighted version of MRLA is also proposed to reduce the quadratic computation cost. The proposed layer attention mechanism can enrich the representation power of many state-of-the-art vision networks, including CNNs and vision transformers. Its effectiveness has been extensively evaluated in image classification, object detection and instance segmentation tasks, where improvements can be consistently observed. For example, our MRLA can improve 1.6% Top-1 accuracy on ResNet-50, while only introducing 0.16M parameters and 0.07B FLOPs. Surprisingly, it can boost the performances by a large margin of 3-4% box AP and mask AP in dense prediction tasks. Our code is available at https://github.com/joyfang1106/MRLA.
Revisiting Self-Training with Regularized Pseudo-Labeling for Tabular Data
Feb 28, 2023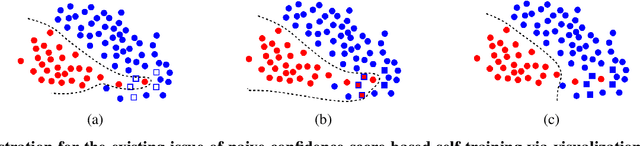
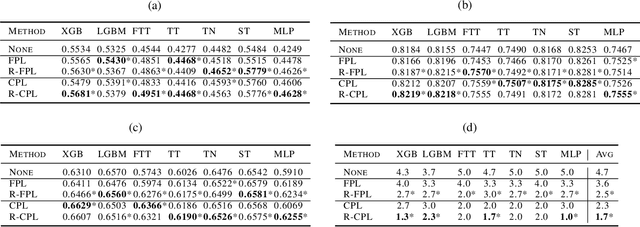
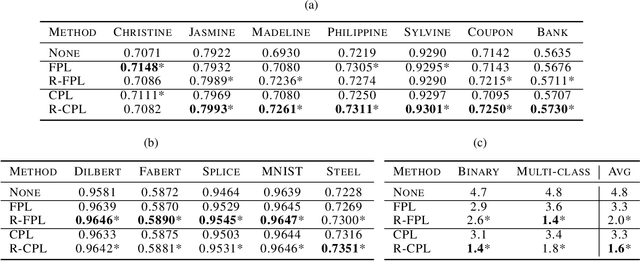
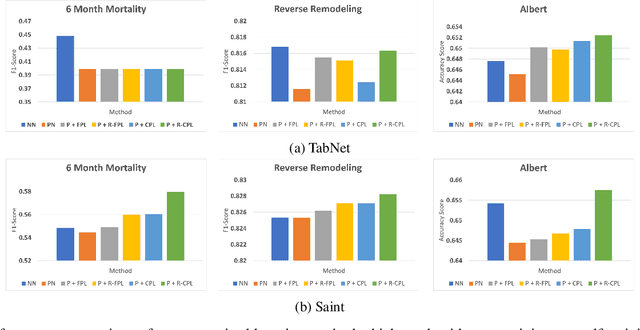
Recent progress in semi- and self-supervised learning has caused a rift in the long-held belief about the need for an enormous amount of labeled data for machine learning and the irrelevancy of unlabeled data. Although it has been successful in various data, there is no dominant semi- and self-supervised learning method that can be generalized for tabular data (i.e. most of the existing methods require appropriate tabular datasets and architectures). In this paper, we revisit self-training which can be applied to any kind of algorithm including the most widely used architecture, gradient boosting decision tree, and introduce curriculum pseudo-labeling (a state-of-the-art pseudo-labeling technique in image) for a tabular domain. Furthermore, existing pseudo-labeling techniques do not assure the cluster assumption when computing confidence scores of pseudo-labels generated from unlabeled data. To overcome this issue, we propose a novel pseudo-labeling approach that regularizes the confidence scores based on the likelihoods of the pseudo-labels so that more reliable pseudo-labels which lie in high density regions can be obtained. We exhaustively validate the superiority of our approaches using various models and tabular datasets.
A Hierarchical Representation Network for Accurate and Detailed Face Reconstruction from In-The-Wild Images
Feb 28, 2023
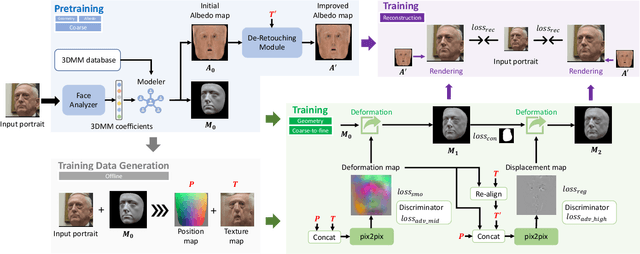
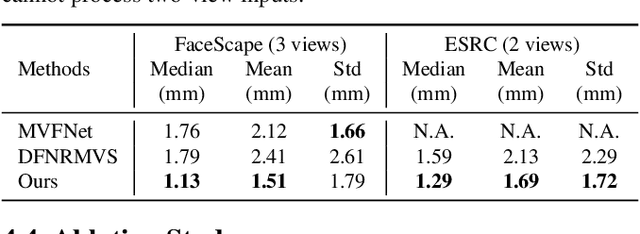
Limited by the nature of the low-dimensional representational capacity of 3DMM, most of the 3DMM-based face reconstruction (FR) methods fail to recover high-frequency facial details, such as wrinkles, dimples, etc. Some attempt to solve the problem by introducing detail maps or non-linear operations, however, the results are still not vivid. To this end, we in this paper present a novel hierarchical representation network (HRN) to achieve accurate and detailed face reconstruction from a single image. Specifically, we implement the geometry disentanglement and introduce the hierarchical representation to fulfill detailed face modeling. Meanwhile, 3D priors of facial details are incorporated to enhance the accuracy and authenticity of the reconstruction results. We also propose a de-retouching module to achieve better decoupling of the geometry and appearance. It is noteworthy that our framework can be extended to a multi-view fashion by considering detail consistency of different views. Extensive experiments on two single-view and two multi-view FR benchmarks demonstrate that our method outperforms the existing methods in both reconstruction accuracy and visual effects. Finally, we introduce a high-quality 3D face dataset FaceHD-100 to boost the research of high-fidelity face reconstruction.
A Self-attention Guided Multi-scale Gradient GAN for Diversified X-ray Image Synthesis
Oct 09, 2022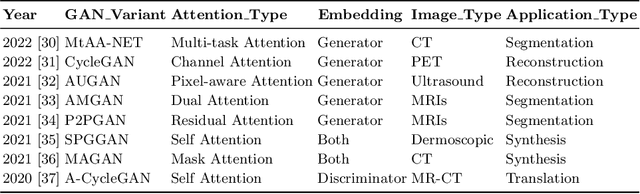
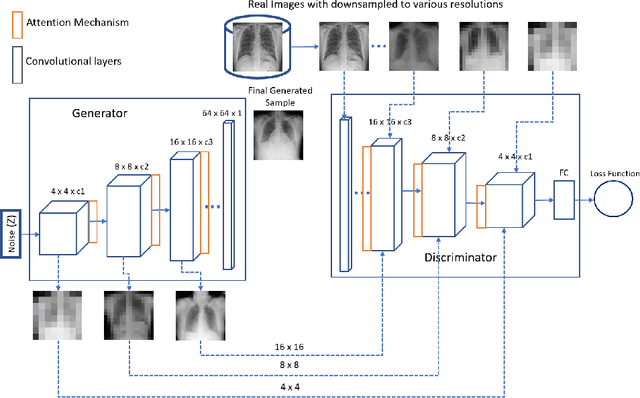
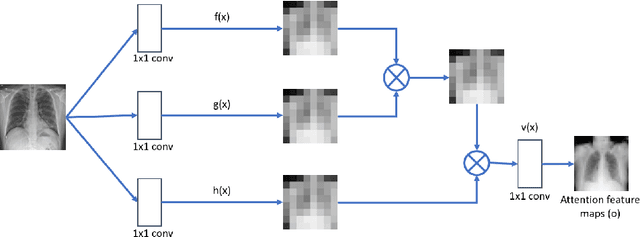
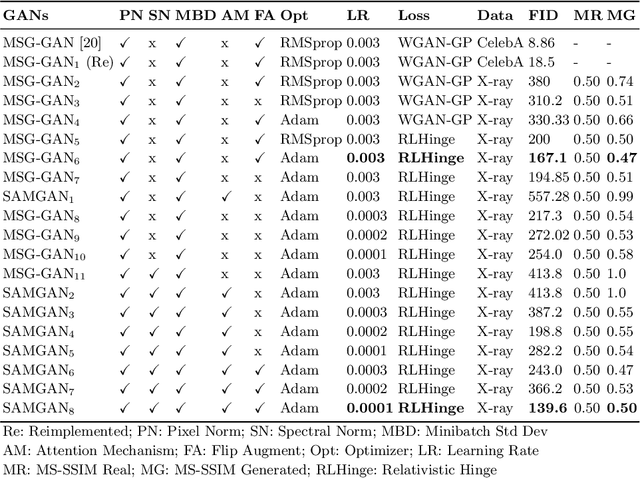
Imbalanced image datasets are commonly available in the domain of biomedical image analysis. Biomedical images contain diversified features that are significant in predicting targeted diseases. Generative Adversarial Networks (GANs) are utilized to address the data limitation problem via the generation of synthetic images. Training challenges such as mode collapse, non-convergence, and instability degrade a GAN's performance in synthesizing diversified and high-quality images. In this work, SAMGAN, an attention-guided multi-scale gradient GAN architecture is proposed to model the relationship between long-range dependencies of biomedical image features and improves the training performance using a flow of multi-scale gradients at multiple resolutions in the layers of generator and discriminator models. The intent is to reduce the impact of mode collapse and stabilize the training of GAN using an attention mechanism with multi-scale gradient learning for diversified X-ray image synthesis. Multi-scale Structural Similarity Index Measure (MS-SSIM) and Frechet Inception Distance (FID) are used to identify the occurrence of mode collapse and evaluate the diversity of synthetic images generated. The proposed architecture is compared with the multi-scale gradient GAN (MSG-GAN) to assess the diversity of generated synthetic images. Results indicate that the SAMGAN outperforms MSG-GAN in synthesizing diversified images as evidenced by the MS-SSIM and FID scores.
Classification of Lung Pathologies in Neonates using Dual Tree Complex Wavelet Transform
Feb 17, 2023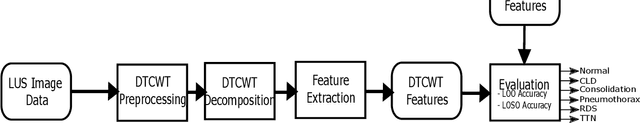
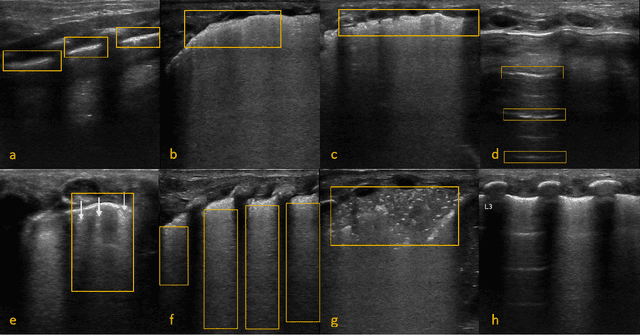
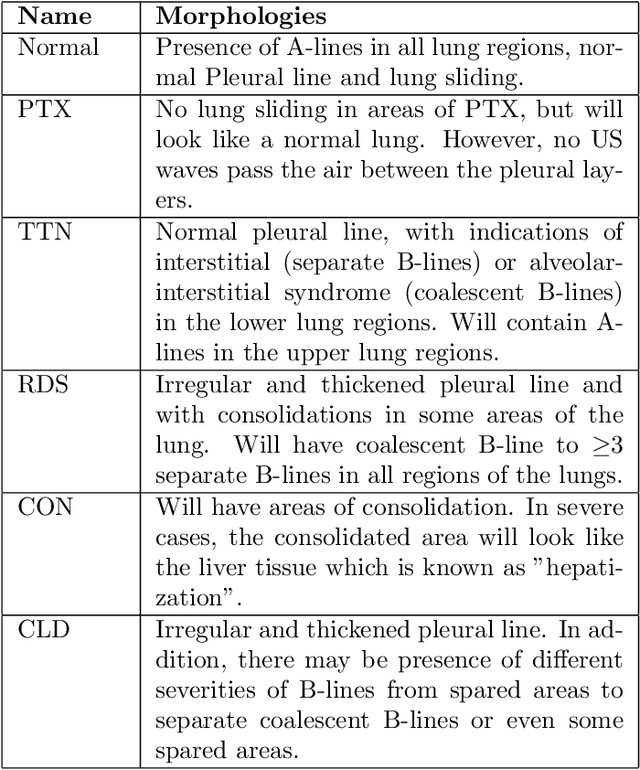

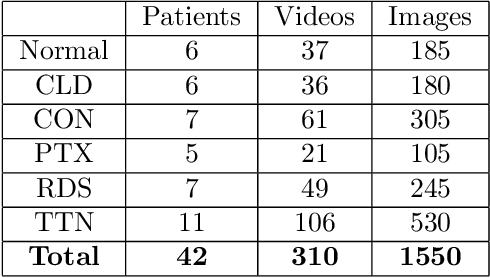
Annually 8500 neonatal deaths are reported in the US due to respiratory failure. Recently, Lung Ultrasound (LUS), due to its radiation free nature, portability, and being cheaper is gaining wide acceptability as a diagnostic tool for lung conditions. However, lack of highly trained medical professionals has limited its use especially in remote areas. To address this, an automated screening system that captures characteristics of the LUS patterns can be of significant assistance to clinicians who are not experts in lung ultrasound (LUS) images. In this paper, we propose a feature extraction method designed to quantify the spatially-localized line patterns and texture patterns found in LUS images. Using the dual-tree complex wavelet transform (DTCWT) and four types of common image features we propose a method to classify the LUS images into 6 common neonatal lung conditions. These conditions are normal lung, pneumothorax (PTX), transient tachypnea of the newborn (TTN), respiratory distress syndrome (RDS), chronic lung disease (CLD) and consolidation (CON) that could be pneumonia or atelectasis. The proposed method using DTCWT decomposition extracted global statistical, grey-level co-occurrence matrix (GLCM), grey-level run length matrix (GLRLM) and linear binary pattern (LBP) features to be fed to a linear discriminative analysis (LDA) based classifier. Using 15 best DTCWT features along with 3 clinical features the proposed approach achieved a per-image classification accuracy of 92.78% with a balanced dataset containing 720 images from 24 patients and 74.39% with the larger unbalanced dataset containing 1550 images from 42 patients. Likewise, the proposed method achieved a maximum per-subject classification accuracy of 81.53% with 43 DTCWT features and 3 clinical features using the balanced dataset and 64.97% with 13 DTCWT features and 3 clinical features using the unbalanced dataset.
Learned Distributed Image Compression with Multi-Scale Patch Matching in Feature Domai
Sep 06, 2022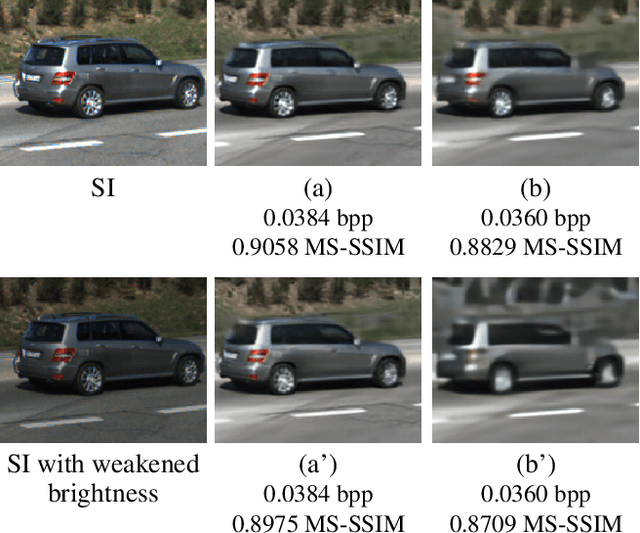
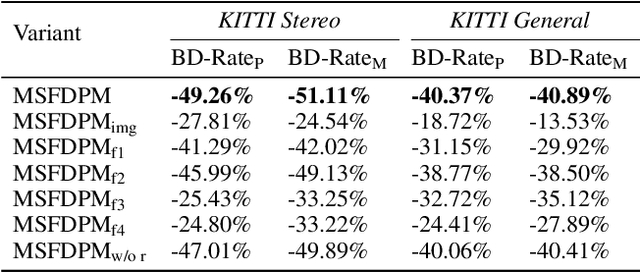
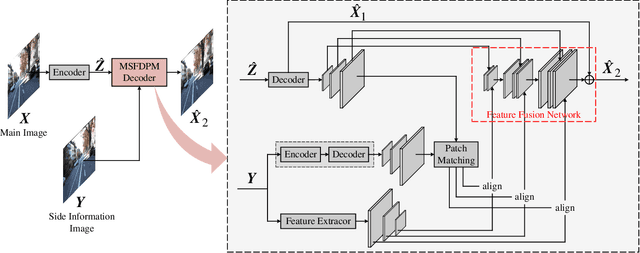

Beyond achieving higher compression efficiency over classical image compression codecs, deep image compression is expected to be improved with additional side information, e.g., another image from a different perspective of the same scene. To better utilize the side information under the distributed compression scenario, the existing method (Ayzik and Avidan 2020) only implements patch matching at the image domain to solve the parallax problem caused by the difference in viewing points. However, the patch matching at the image domain is not robust to the variance of scale, shape, and illumination caused by the different viewing angles, and can not make full use of the rich texture information of the side information image. To resolve this issue, we propose Multi-Scale Feature Domain Patch Matching (MSFDPM) to fully utilizes side information at the decoder of the distributed image compression model. Specifically, MSFDPM consists of a side information feature extractor, a multi-scale feature domain patch matching module, and a multi-scale feature fusion network. Furthermore, we reuse inter-patch correlation from the shallow layer to accelerate the patch matching of the deep layer. Finally, we nd that our patch matching in a multi-scale feature domain further improves compression rate by about 20% compared with the patch matching method at image domain (Ayzik and Avidan 2020).
On the Feasibility of Machine Learning Augmented Magnetic Resonance for Point-of-Care Identification of Disease
Jan 27, 2023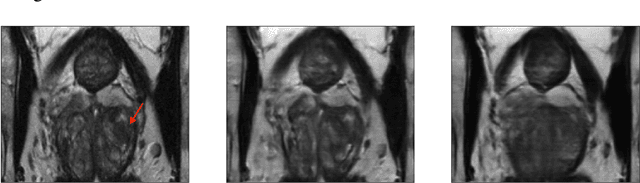
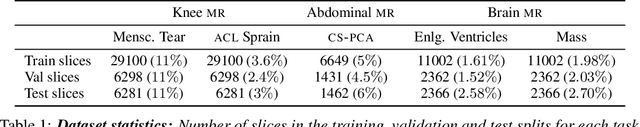
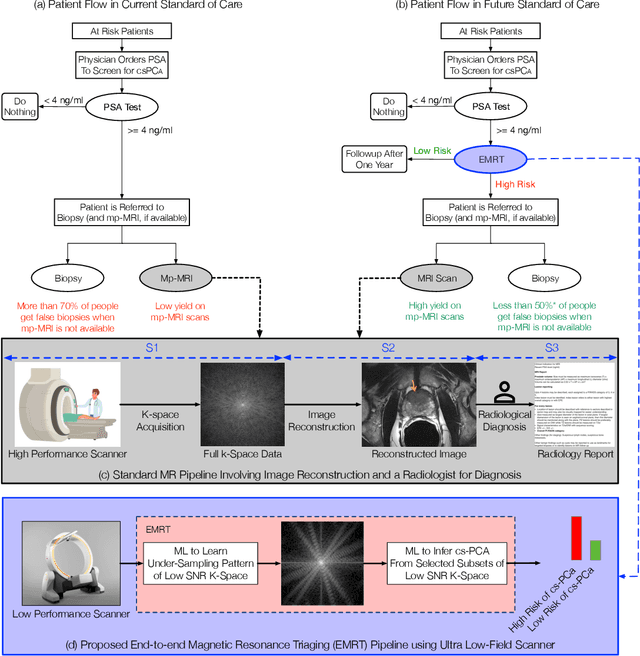
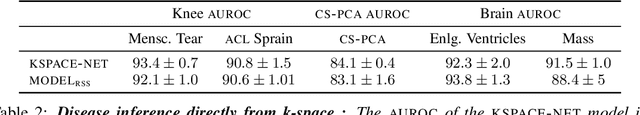
Early detection of many life-threatening diseases (e.g., prostate and breast cancer) within at-risk population can improve clinical outcomes and reduce cost of care. While numerous disease-specific "screening" tests that are closer to Point-of-Care (POC) are in use for this task, their low specificity results in unnecessary biopsies, leading to avoidable patient trauma and wasteful healthcare spending. On the other hand, despite the high accuracy of Magnetic Resonance (MR) imaging in disease diagnosis, it is not used as a POC disease identification tool because of poor accessibility. The root cause of poor accessibility of MR stems from the requirement to reconstruct high-fidelity images, as it necessitates a lengthy and complex process of acquiring large quantities of high-quality k-space measurements. In this study we explore the feasibility of an ML-augmented MR pipeline that directly infers the disease sidestepping the image reconstruction process. We hypothesise that the disease classification task can be solved using a very small tailored subset of k-space data, compared to image reconstruction. Towards that end, we propose a method that performs two tasks: 1) identifies a subset of the k-space that maximizes disease identification accuracy, and 2) infers the disease directly using the identified k-space subset, bypassing the image reconstruction step. We validate our hypothesis by measuring the performance of the proposed system across multiple diseases and anatomies. We show that comparable performance to image-based classifiers, trained on images reconstructed with full k-space data, can be achieved using small quantities of data: 8% of the data for detecting multiple abnormalities in prostate and brain scans, and 5% of the data for knee abnormalities. To better understand the proposed approach and instigate future research, we provide an extensive analysis and release code.
HDRfeat: A Feature-Rich Network for High Dynamic Range Image Reconstruction
Nov 08, 2022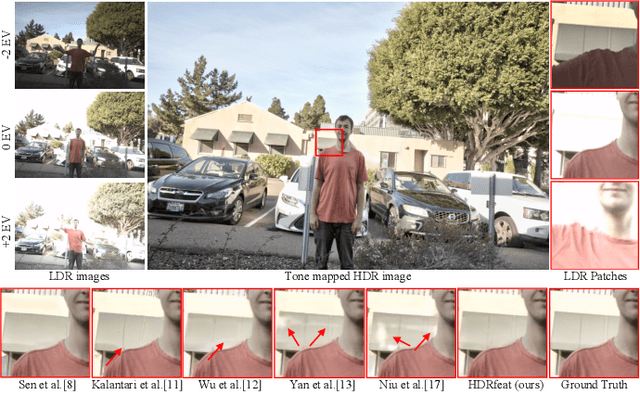
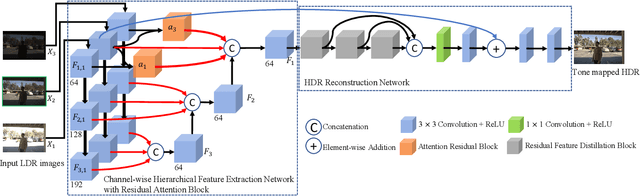
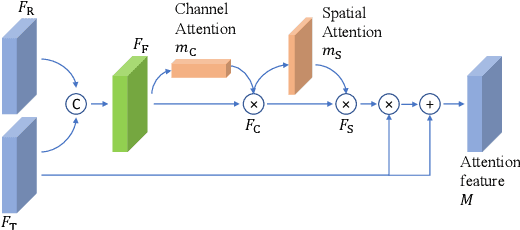

A major challenge for high dynamic range (HDR) image reconstruction from multi-exposed low dynamic range (LDR) images, especially with dynamic scenes, is the extraction and merging of relevant contextual features in order to suppress any ghosting and blurring artifacts from moving objects. To tackle this, in this work we propose a novel network for HDR reconstruction with deep and rich feature extraction layers, including residual attention blocks with sequential channel and spatial attention. For the compression of the rich-features to the HDR domain, a residual feature distillation block (RFDB) based architecture is adopted. In contrast to earlier deep-learning methods for HDR, the above contributions shift focus from merging/compression to feature extraction, the added value of which we demonstrate with ablation experiments. We present qualitative and quantitative comparisons on a public benchmark dataset, showing that our proposed method outperforms the state-of-the-art.
Vision, Deduction and Alignment: An Empirical Study on Multi-modal Knowledge Graph Alignment
Feb 17, 2023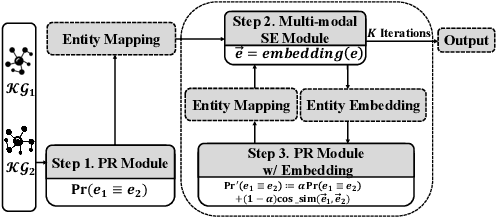
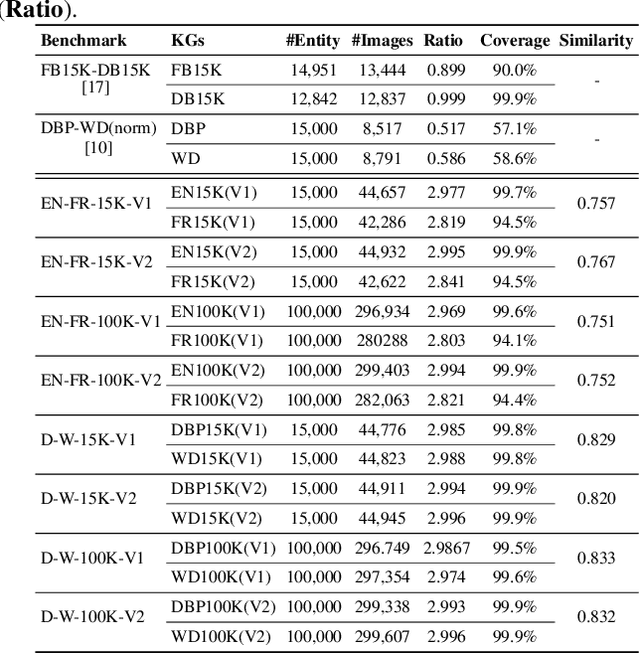
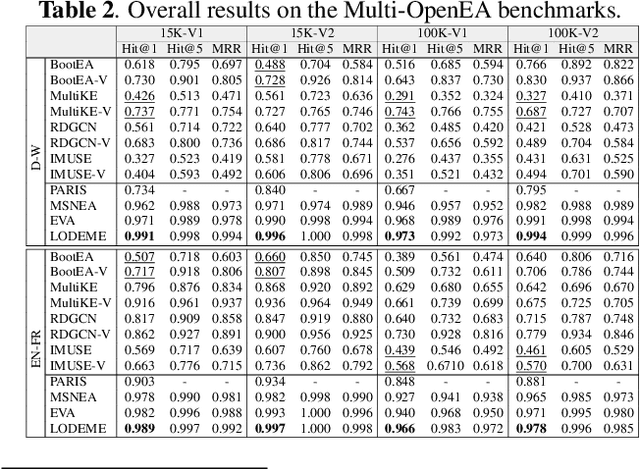
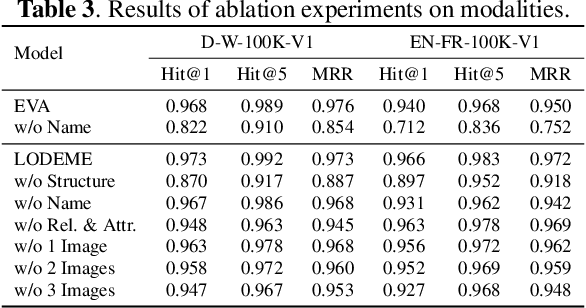
Entity alignment (EA) for knowledge graphs (KGs) plays a critical role in knowledge engineering. Existing EA methods mostly focus on utilizing the graph structures and entity attributes (including literals), but ignore images that are common in modern multi-modal KGs. In this study we first constructed Multi-OpenEA -- eight large-scale, image-equipped EA benchmarks, and then evaluated some existing embedding-based methods for utilizing images. In view of the complementary nature of visual modal information and logical deduction, we further developed a new multi-modal EA method named LODEME using logical deduction and multi-modal KG embedding, with state-of-the-art performance achieved on Multi-OpenEA and other existing multi-modal EA benchmarks.
Generalizable Medical Image Segmentation via Random Amplitude Mixup and Domain-Specific Image Restoration
Aug 08, 2022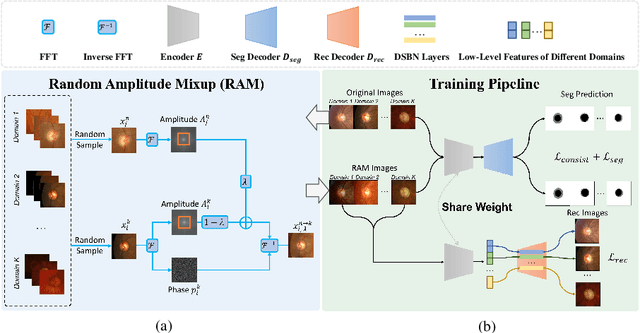
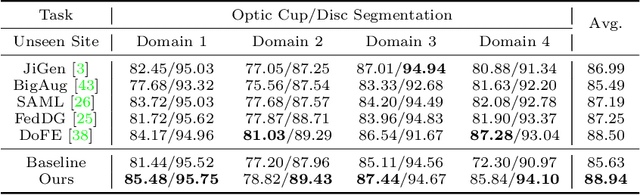
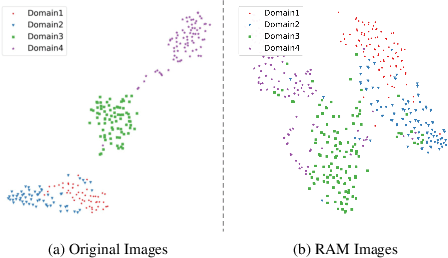
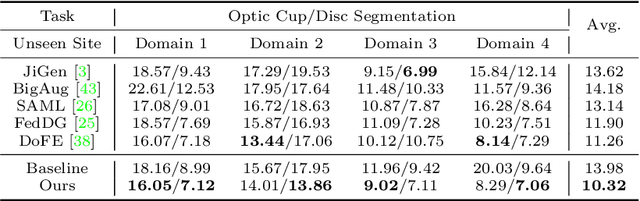
For medical image analysis, segmentation models trained on one or several domains lack generalization ability to unseen domains due to discrepancies between different data acquisition policies. We argue that the degeneration in segmentation performance is mainly attributed to overfitting to source domains and domain shift. To this end, we present a novel generalizable medical image segmentation method. To be specific, we design our approach as a multi-task paradigm by combining the segmentation model with a self-supervision domain-specific image restoration (DSIR) module for model regularization. We also design a random amplitude mixup (RAM) module, which incorporates low-level frequency information of different domain images to synthesize new images. To guide our model be resistant to domain shift, we introduce a semantic consistency loss. We demonstrate the performance of our method on two public generalizable segmentation benchmarks in medical images, which validates our method could achieve the state-of-the-art performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge