"Image": models, code, and papers
Strong Baseline and Bag of Tricks for COVID-19 Detection of CT Scans
Mar 15, 2023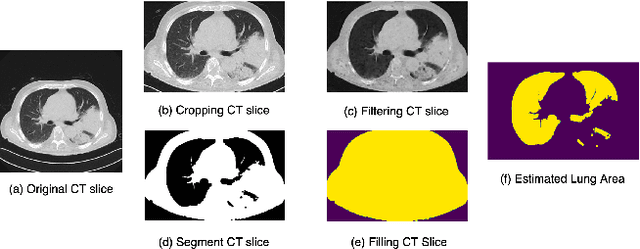

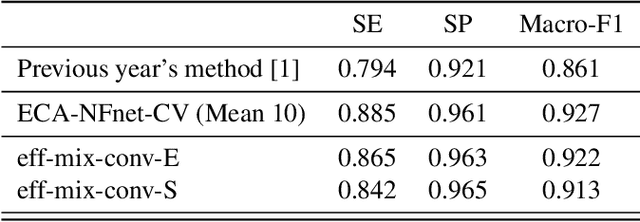
This paper investigates the application of deep learning models for lung Computed Tomography (CT) image analysis. Traditional deep learning frameworks encounter compatibility issues due to variations in slice numbers and resolutions in CT images, which stem from the use of different machines. Commonly, individual slices are predicted and subsequently merged to obtain the final result; however, this approach lacks slice-wise feature learning and consequently results in decreased performance. We propose a novel slice selection method for each CT dataset to address this limitation, effectively filtering out uncertain slices and enhancing the model's performance. Furthermore, we introduce a spatial-slice feature learning (SSFL) technique\cite{hsu2022} that employs a conventional and efficient backbone model for slice feature training, followed by extracting one-dimensional data from the trained model for COVID and non-COVID classification using a dedicated classification model. Leveraging these experimental steps, we integrate one-dimensional features with multiple slices for channel merging and employ a 2D convolutional neural network (CNN) model for classification. In addition to the aforementioned methods, we explore various high-performance classification models, ultimately achieving promising results.
SeqCo-DETR: Sequence Consistency Training for Self-Supervised Object Detection with Transformers
Mar 15, 2023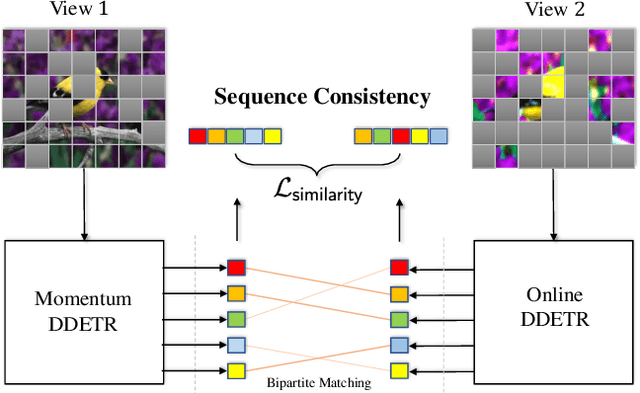
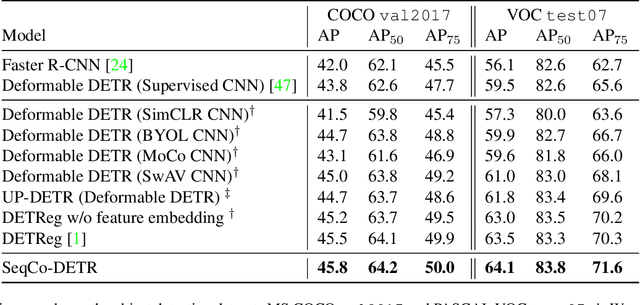
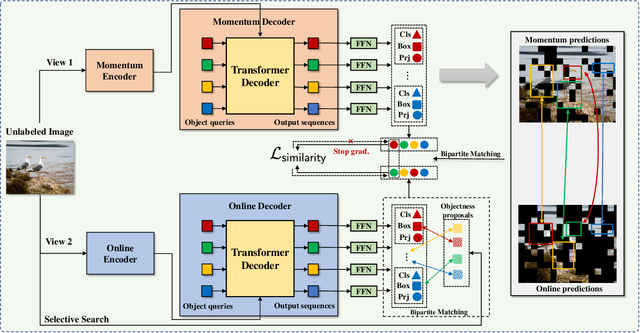
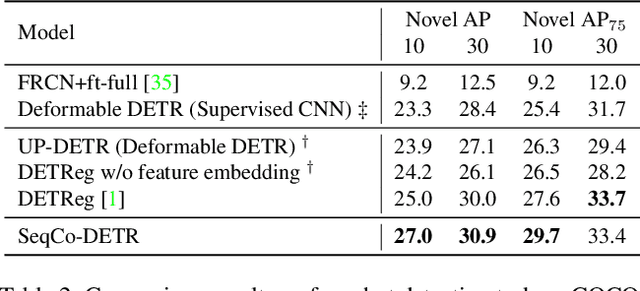
Self-supervised pre-training and transformer-based networks have significantly improved the performance of object detection. However, most of the current self-supervised object detection methods are built on convolutional-based architectures. We believe that the transformers' sequence characteristics should be considered when designing a transformer-based self-supervised method for the object detection task. To this end, we propose SeqCo-DETR, a novel Sequence Consistency-based self-supervised method for object DEtection with TRansformers. SeqCo-DETR defines a simple but effective pretext by minimizes the discrepancy of the output sequences of transformers with different image views as input and leverages bipartite matching to find the most relevant sequence pairs to improve the sequence-level self-supervised representation learning performance. Furthermore, we provide a mask-based augmentation strategy incorporated with the sequence consistency strategy to extract more representative contextual information about the object for the object detection task. Our method achieves state-of-the-art results on MS COCO (45.8 AP) and PASCAL VOC (64.1 AP), demonstrating the effectiveness of our approach.
SpiderMesh: Spatial-aware Demand-guided Recursive Meshing for RGB-T Semantic Segmentation
Mar 15, 2023
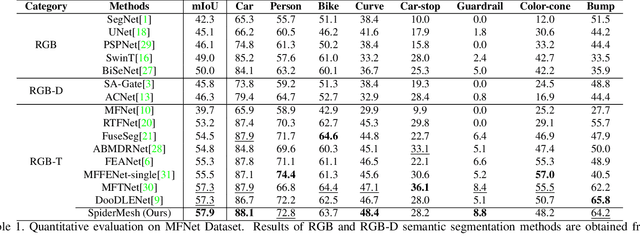
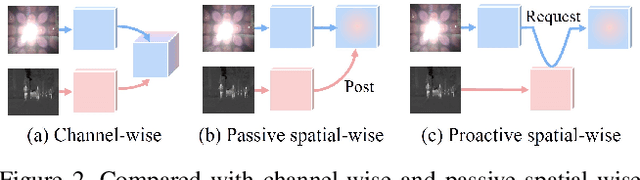
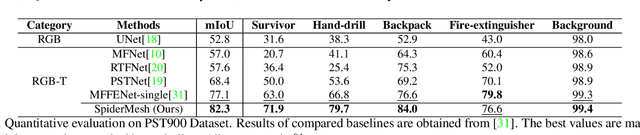
For semantic segmentation in urban scene understanding, RGB cameras alone often fail to capture a clear holistic topology, especially in challenging lighting conditions. Thermal signal is an informative additional channel that can bring to light the contour and fine-grained texture of blurred regions in low-quality RGB image. Aiming at RGB-T (thermal) segmentation, existing methods either use simple passive channel/spatial-wise fusion for cross-modal interaction, or rely on heavy labeling of ambiguous boundaries for fine-grained supervision. We propose a Spatial-aware Demand-guided Recursive Meshing (SpiderMesh) framework that: 1) proactively compensates inadequate contextual semantics in optically-impaired regions via a demand-guided target masking algorithm; 2) refines multimodal semantic features with recursive meshing to improve pixel-level semantic analysis performance. We further introduce an asymmetric data augmentation technique M-CutOut, and enable semi-supervised learning to fully utilize RGB-T labels only sparsely available in practical use. Extensive experiments on MFNet and PST900 datasets demonstrate that SpiderMesh achieves new state-of-the-art performance on standard RGB-T segmentation benchmarks.
Reversing the Abnormal: Pseudo-Healthy Generative Networks for Anomaly Detection
Mar 15, 2023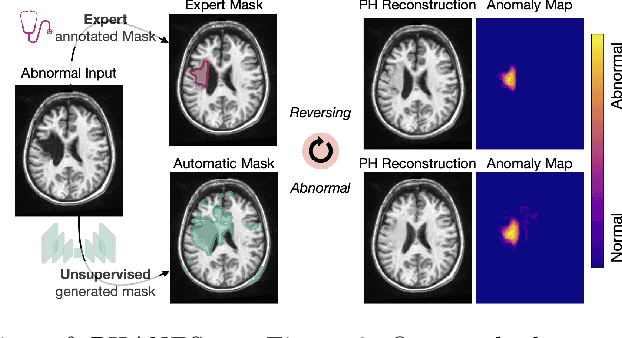
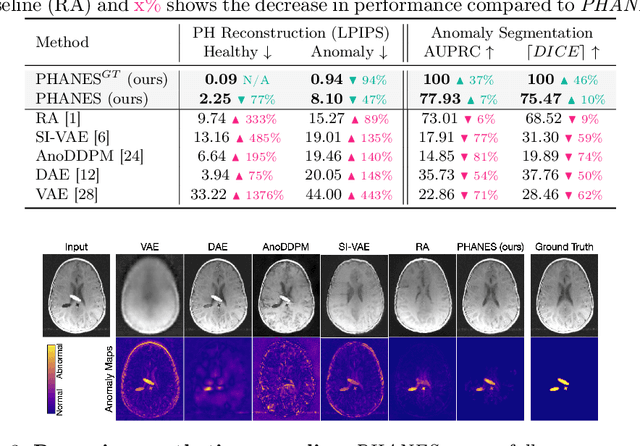
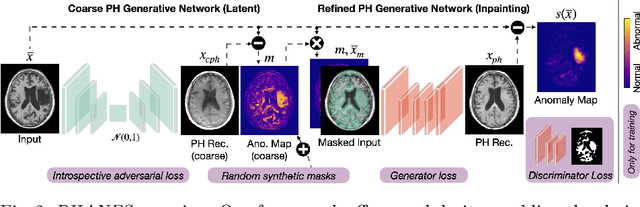
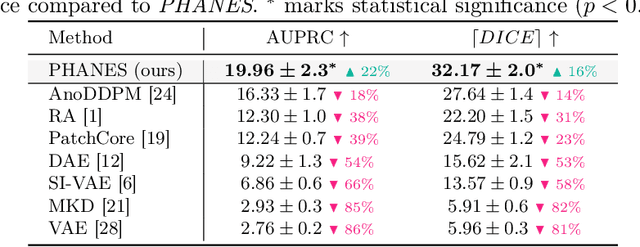
Early and accurate disease detection is crucial for patient management and successful treatment outcomes. However, the automatic identification of anomalies in medical images can be challenging. Conventional methods rely on large labeled datasets which are difficult to obtain. To overcome these limitations, we introduce a novel unsupervised approach, called PHANES (Pseudo Healthy generative networks for ANomaly Segmentation). Our method has the capability of reversing anomalies, i.e., preserving healthy tissue and replacing anomalous regions with pseudo-healthy (PH) reconstructions. Unlike recent diffusion models, our method does not rely on a learned noise distribution nor does it introduce random alterations to the entire image. Instead, we use latent generative networks to create masks around possible anomalies, which are refined using inpainting generative networks. We demonstrate the effectiveness of PHANES in detecting stroke lesions in T1w brain MRI datasets and show significant improvements over state-of-the-art (SOTA) methods. We believe that our proposed framework will open new avenues for interpretable, fast, and accurate anomaly segmentation with the potential to support various clinical-oriented downstream tasks.
SegPrompt: Using Segmentation Map as a Better Prompt to Finetune Deep Models for Kidney Stone Classification
Mar 15, 2023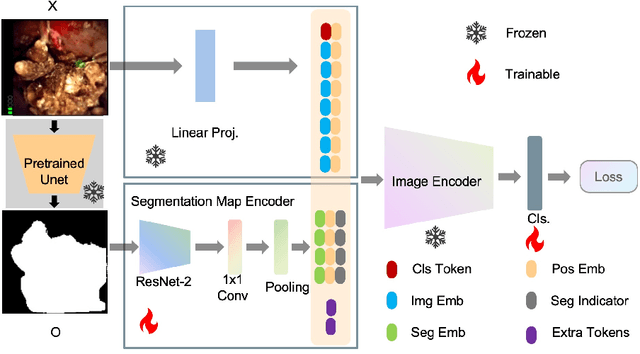
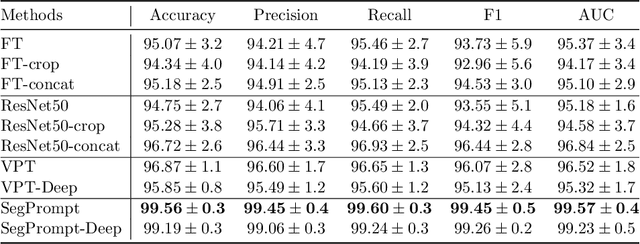
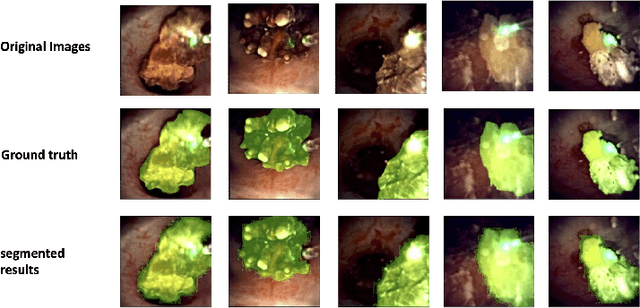
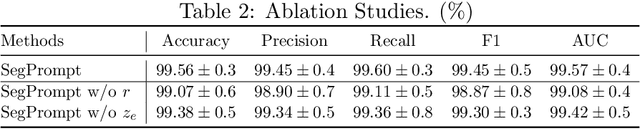
Recently, deep learning has produced encouraging results for kidney stone classification using endoscope images. However, the shortage of annotated training data poses a severe problem in improving the performance and generalization ability of the trained model. It is thus crucial to fully exploit the limited data at hand. In this paper, we propose SegPrompt to alleviate the data shortage problems by exploiting segmentation maps from two aspects. First, SegPrompt integrates segmentation maps to facilitate classification training so that the classification model is aware of the regions of interest. The proposed method allows the image and segmentation tokens to interact with each other to fully utilize the segmentation map information. Second, we use the segmentation maps as prompts to tune the pretrained deep model, resulting in much fewer trainable parameters than vanilla finetuning. We perform extensive experiments on the collected kidney stone dataset. The results show that SegPrompt can achieve an advantageous balance between the model fitting ability and the generalization ability, eventually leading to an effective model with limited training data.
Data fusion of satellite imagery for generation of daily cloud free images at high resolution level
Feb 24, 2023
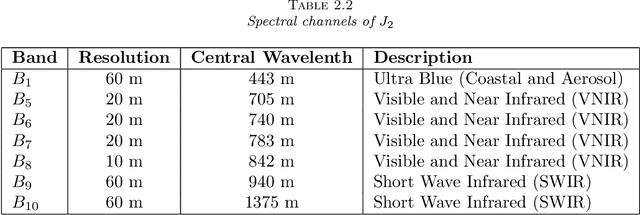



In this paper we discuss a new variational approach to the Date Fusion problem of multi-spectral satellite images from Sentinel-2 and MODIS that have been captured at different resolution level and, arguably, on different days. The crucial point of our approach that the MODIS image is cloud-free whereas the images from Sentinel-2 can be corrupted by clouds or noise.
Hierarchical discriminative learning improves visual representations of biomedical microscopy
Mar 02, 2023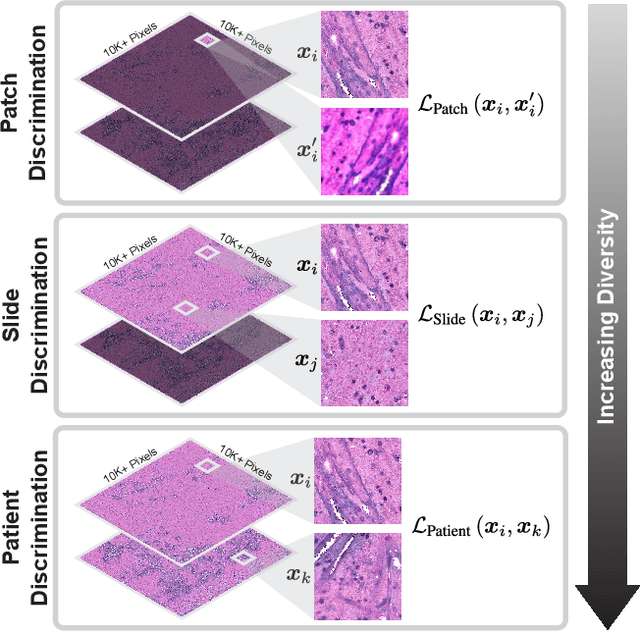
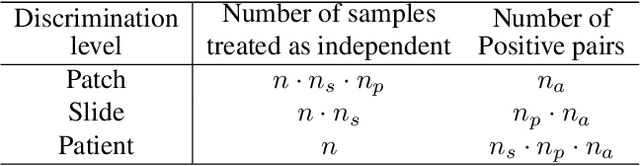

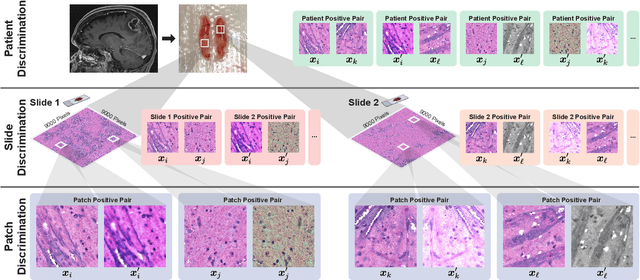
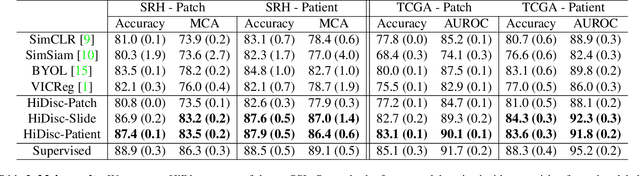
Learning high-quality, self-supervised, visual representations is essential to advance the role of computer vision in biomedical microscopy and clinical medicine. Previous work has focused on self-supervised representation learning (SSL) methods developed for instance discrimination and applied them directly to image patches, or fields-of-view, sampled from gigapixel whole-slide images (WSIs) used for cancer diagnosis. However, this strategy is limited because it (1) assumes patches from the same patient are independent, (2) neglects the patient-slide-patch hierarchy of clinical biomedical microscopy, and (3) requires strong data augmentations that can degrade downstream performance. Importantly, sampled patches from WSIs of a patient's tumor are a diverse set of image examples that capture the same underlying cancer diagnosis. This motivated HiDisc, a data-driven method that leverages the inherent patient-slide-patch hierarchy of clinical biomedical microscopy to define a hierarchical discriminative learning task that implicitly learns features of the underlying diagnosis. HiDisc uses a self-supervised contrastive learning framework in which positive patch pairs are defined based on a common ancestry in the data hierarchy, and a unified patch, slide, and patient discriminative learning objective is used for visual SSL. We benchmark HiDisc visual representations on two vision tasks using two biomedical microscopy datasets, and demonstrate that (1) HiDisc pretraining outperforms current state-of-the-art self-supervised pretraining methods for cancer diagnosis and genetic mutation prediction, and (2) HiDisc learns high-quality visual representations using natural patch diversity without strong data augmentations.
The Carbon Emissions of Writing and Illustrating Are Lower for AI than for Humans
Mar 08, 2023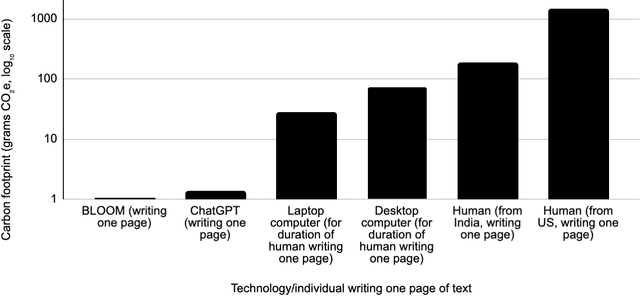
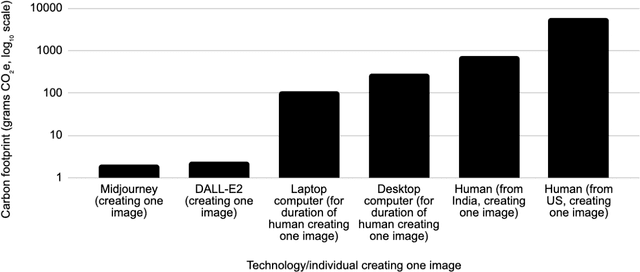
As AI systems proliferate, their greenhouse gas emissions are an increasingly important concern for human societies. We analyze the emissions of several AI systems (ChatGPT, BLOOM, DALL-E2, Midjourney) relative to those of humans completing the same tasks. We find that an AI writing a page of text emits 130 to 1500 times less CO2e than a human doing so. Similarly, an AI creating an image emits 310 to 2900 times less. Emissions analysis do not account for social impacts such as professional displacement, legality, and rebound effects. In addition, AI is not a substitute for all human tasks. Nevertheless, at present, the use of AI holds the potential to carry out several major activities at much lower emission levels than can humans.
Fast Cauchy-Rician Modelling of SAR Images with Method of Algebraic Moments Estimator
Mar 08, 2023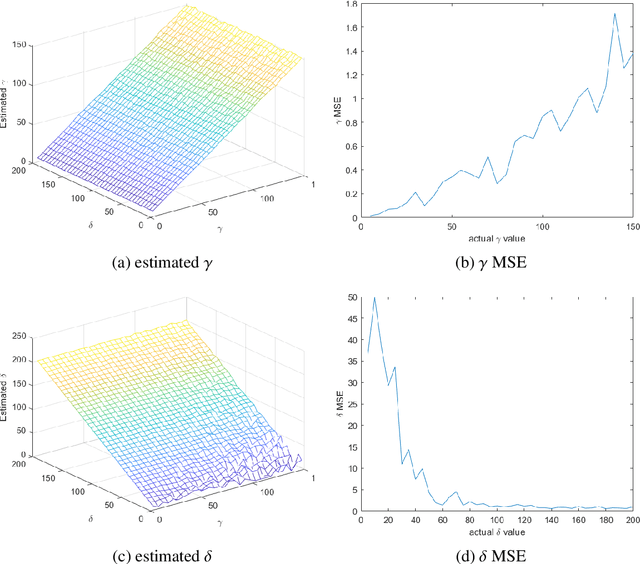
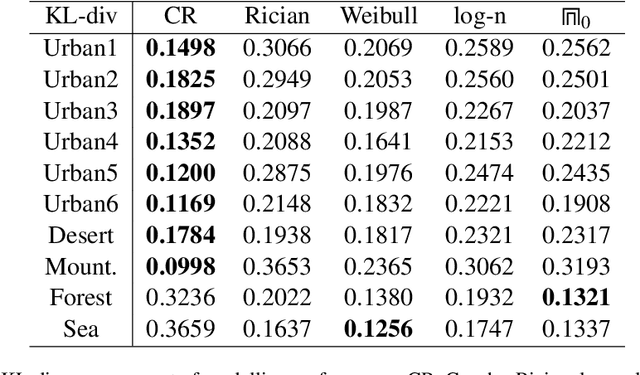

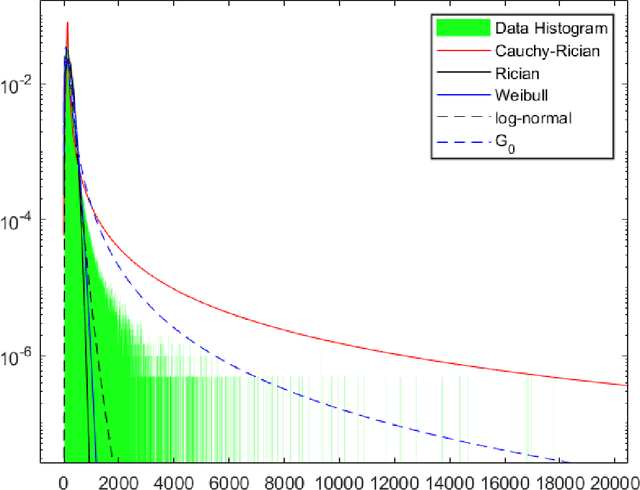
SAR technology has been intensively implemented for geo-sensing and mapping purposes due to its advantages of high azimuthal resolution and weather-independent operation compared to other remote sensing technologies. Modelling SAR image data consequently becomes a prominent topic of interest, especially for data populations with impulsive signal features, which are common in SAR images of urban areas. A recently proposed model named Cauchy-Rician has manifested great potential in modelling extremely heterogeneous SAR images, yet the work only provided a MCMC-based parameter estimator that demands considerable computational power. In this work, a novel analytical parameter estimation method based on algebraic moments is proposed to provide stable and accurate estimation of the parameters of the Cauchy-Rician model with significant improvement on computation speed.
M-EBM: Towards Understanding the Manifolds of Energy-Based Models
Mar 08, 2023
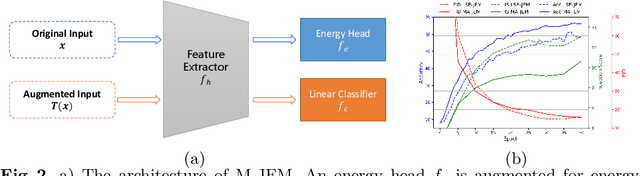
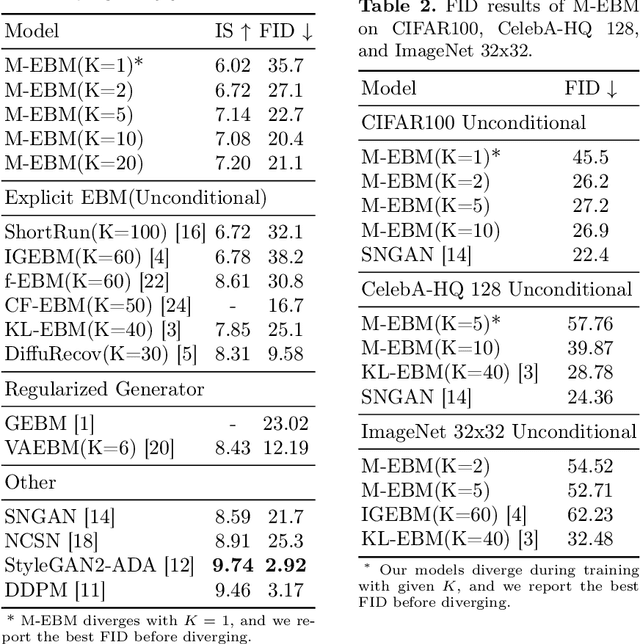

Energy-based models (EBMs) exhibit a variety of desirable properties in predictive tasks, such as generality, simplicity and compositionality. However, training EBMs on high-dimensional datasets remains unstable and expensive. In this paper, we present a Manifold EBM (M-EBM) to boost the overall performance of unconditional EBM and Joint Energy-based Model (JEM). Despite its simplicity, M-EBM significantly improves unconditional EBMs in training stability and speed on a host of benchmark datasets, such as CIFAR10, CIFAR100, CelebA-HQ, and ImageNet 32x32. Once class labels are available, label-incorporated M-EBM (M-JEM) further surpasses M-EBM in image generation quality with an over 40% FID improvement, while enjoying improved accuracy. The code can be found at https://github.com/sndnyang/mebm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge