"Image": models, code, and papers
TAAL: Test-time Augmentation for Active Learning in Medical Image Segmentation
Jan 16, 2023Deep learning methods typically depend on the availability of labeled data, which is expensive and time-consuming to obtain. Active learning addresses such effort by prioritizing which samples are best to annotate in order to maximize the performance of the task model. While frameworks for active learning have been widely explored in the context of classification of natural images, they have been only sparsely used in medical image segmentation. The challenge resides in obtaining an uncertainty measure that reveals the best candidate data for annotation. This paper proposes Test-time Augmentation for Active Learning (TAAL), a novel semi-supervised active learning approach for segmentation that exploits the uncertainty information offered by data transformations. Our method applies cross-augmentation consistency during training and inference to both improve model learning in a semi-supervised fashion and identify the most relevant unlabeled samples to annotate next. In addition, our consistency loss uses a modified version of the JSD to further improve model performance. By relying on data transformations rather than on external modules or simple heuristics typically used in uncertainty-based strategies, TAAL emerges as a simple, yet powerful task-agnostic semi-supervised active learning approach applicable to the medical domain. Our results on a publicly-available dataset of cardiac images show that TAAL outperforms existing baseline methods in both fully-supervised and semi-supervised settings. Our implementation is publicly available on https://github.com/melinphd/TAAL.
Fracture Detection in Pediatric Wrist Trauma X-ray Images Using YOLOv8 Algorithm
Apr 11, 2023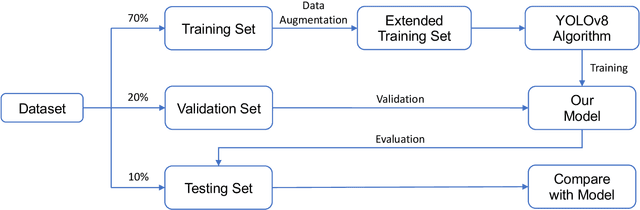
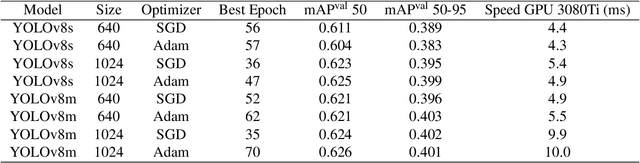
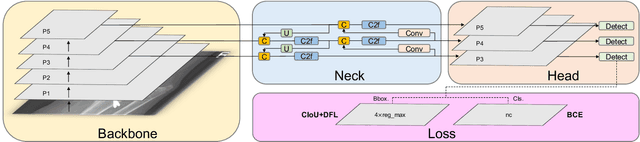
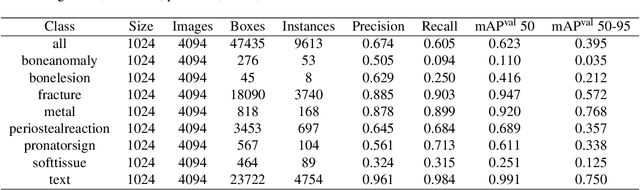
Hospital emergency departments frequently receive lots of bone fracture cases, with pediatric wrist trauma fracture accounting for the majority of them. Before pediatric surgeons perform surgery, they need to ask patients how the fracture occurred and analyze the fracture situation by interpreting X-ray images. The interpretation of X-ray images often requires a combination of techniques from radiologists and surgeons, which requires time-consuming specialized training. With the rise of deep learning in the field of computer vision, network models applying for fracture detection has become an important research topic. In this paper, YOLOv8 algorithm is used to train models on the GRAZPEDWRI-DX dataset, which includes X-ray images from 6,091 pediatric patients with wrist trauma. The experimental results show that YOLOv8 algorithm models have different advantages for different model sizes, with YOLOv8l model achieving the highest mean average precision (mAP 50) of 63.6\%, and YOLOv8n model achieving the inference time of 67.4ms per X-ray image on one single CPU with low computing power. In this way, we create "Fracture Detection Using YOLOv8 App" to assist surgeons in interpreting X-ray images without the help of radiologists. Our implementation code is released at https://github.com/RuiyangJu/Bone_Fracture_Detection_YOLOv8.
SPIRiT-Diffusion: Self-Consistency Driven Diffusion Model for Accelerated MRI
Apr 11, 2023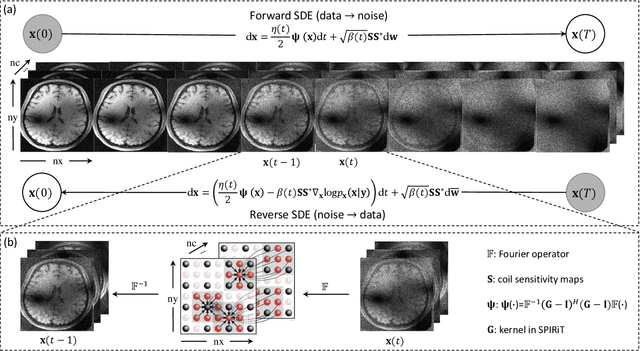
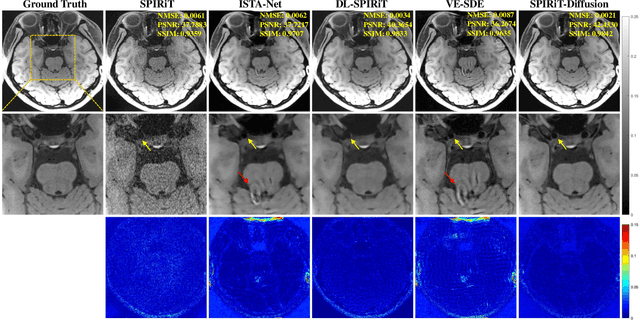
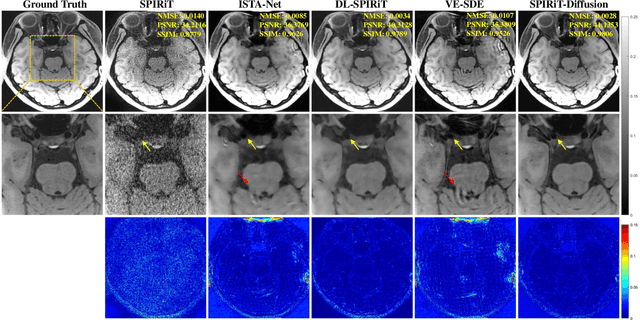
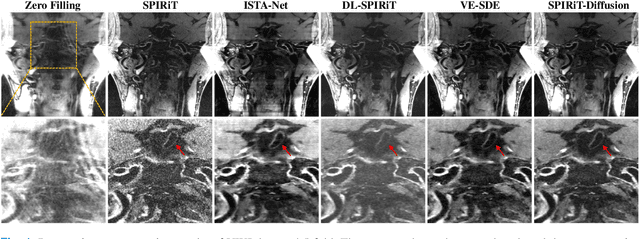
Diffusion models are a leading method for image generation and have been successfully applied in magnetic resonance imaging (MRI) reconstruction. Current diffusion-based reconstruction methods rely on coil sensitivity maps (CSM) to reconstruct multi-coil data. However, it is difficult to accurately estimate CSMs in practice use, resulting in degradation of the reconstruction quality. To address this issue, we propose a self-consistency-driven diffusion model inspired by the iterative self-consistent parallel imaging (SPIRiT), namely SPIRiT-Diffusion. Specifically, the iterative solver of the self-consistent term in SPIRiT is utilized to design a novel stochastic differential equation (SDE) for diffusion process. Then $\textit{k}$-space data can be interpolated directly during the reverse diffusion process, instead of using CSM to separate and combine individual coil images. This method indicates that the optimization model can be used to design SDE in diffusion models, driving the diffusion process strongly conforming with the physics involved in the optimization model, dubbed model-driven diffusion. The proposed SPIRiT-Diffusion method was evaluated on a 3D joint Intracranial and Carotid Vessel Wall imaging dataset. The results demonstrate that it outperforms the CSM-based reconstruction methods, and achieves high reconstruction quality at a high acceleration rate of 10.
Diffusion Models for Constrained Domains
Apr 11, 2023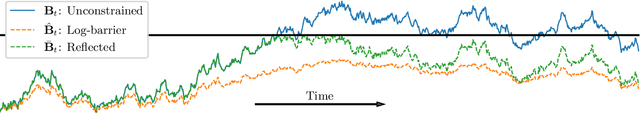

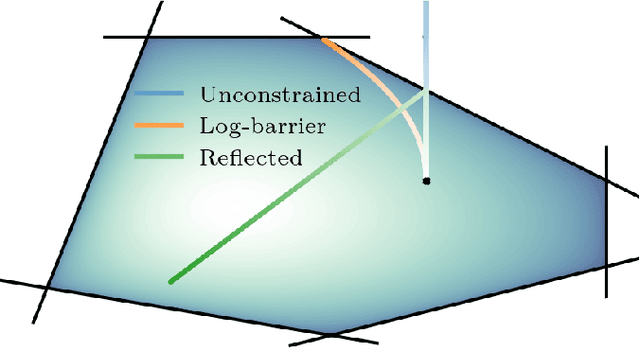
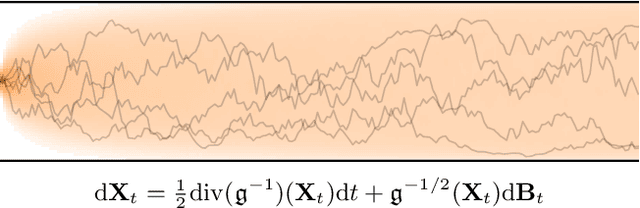
Denoising diffusion models are a recent class of generative models which achieve state-of-the-art results in many domains such as unconditional image generation and text-to-speech tasks. They consist of a noising process destroying the data and a backward stage defined as the time-reversal of the noising diffusion. Building on their success, diffusion models have recently been extended to the Riemannian manifold setting. Yet, these Riemannian diffusion models require geodesics to be defined for all times. While this setting encompasses many important applications, it does not include manifolds defined via a set of inequality constraints, which are ubiquitous in many scientific domains such as robotics and protein design. In this work, we introduce two methods to bridge this gap. First, we design a noising process based on the logarithmic barrier metric induced by the inequality constraints. Second, we introduce a noising process based on the reflected Brownian motion. As existing diffusion model techniques cannot be applied in this setting, we derive new tools to define such models in our framework. We empirically demonstrate the applicability of our methods to a number of synthetic and real-world tasks, including the constrained conformational modelling of protein backbones and robotic arms.
EVA-02: A Visual Representation for Neon Genesis
Mar 20, 2023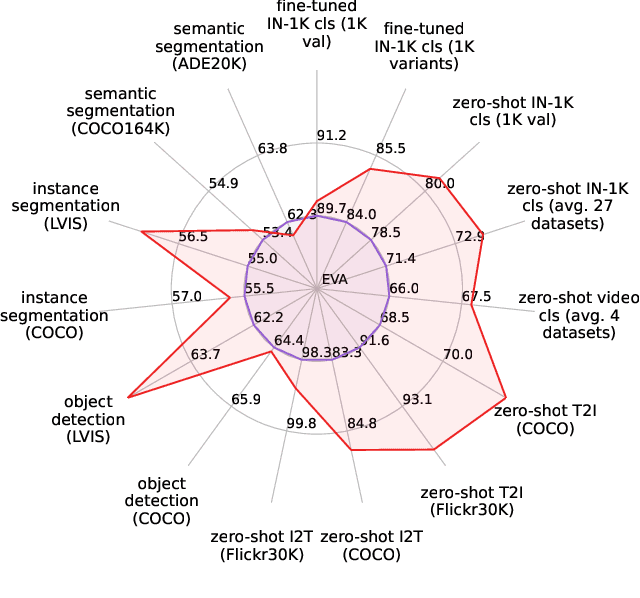
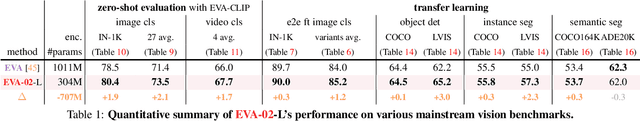
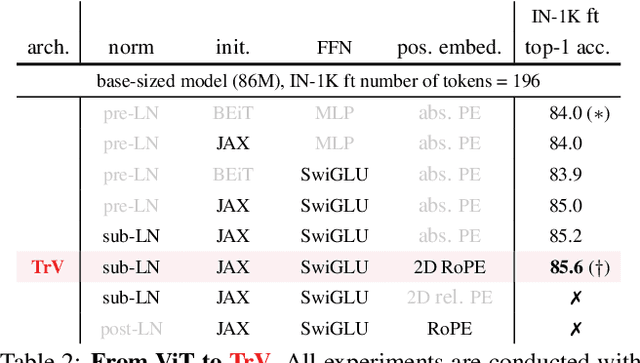
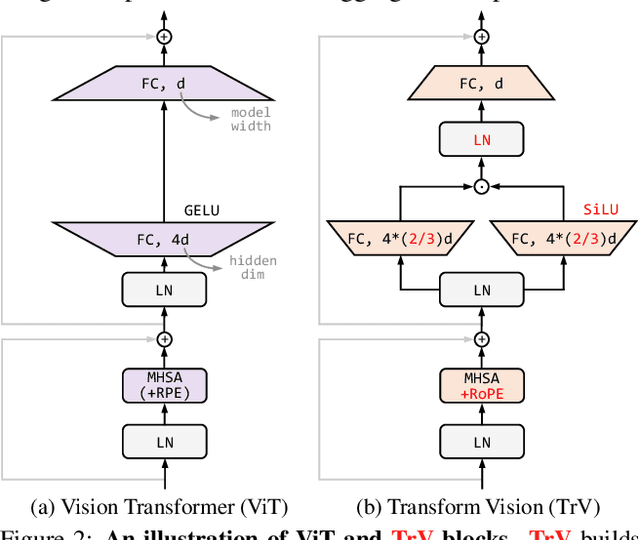
We launch EVA-02, a next-generation Transformer-based visual representation pre-trained to reconstruct strong and robust language-aligned vision features via masked image modeling. With an updated plain Transformer architecture as well as extensive pre-training from an open & accessible giant CLIP vision encoder, EVA-02 demonstrates superior performance compared to prior state-of-the-art approaches across various representative vision tasks, while utilizing significantly fewer parameters and compute budgets. Notably, using exclusively publicly accessible training data, EVA-02 with only 304M parameters achieves a phenomenal 90.0 fine-tuning top-1 accuracy on ImageNet-1K val set. Additionally, our EVA-02-CLIP can reach up to 80.4 zero-shot top-1 on ImageNet-1K, outperforming the previous largest & best open-sourced CLIP with only ~1/6 parameters and ~1/6 image-text training data. We offer four EVA-02 variants in various model sizes, ranging from 6M to 304M parameters, all with impressive performance. To facilitate open access and open research, we release the complete suite of EVA-02 to the community at https://github.com/baaivision/EVA/tree/master/EVA-02.
Contrastive Model Adaptation for Cross-Condition Robustness in Semantic Segmentation
Mar 09, 2023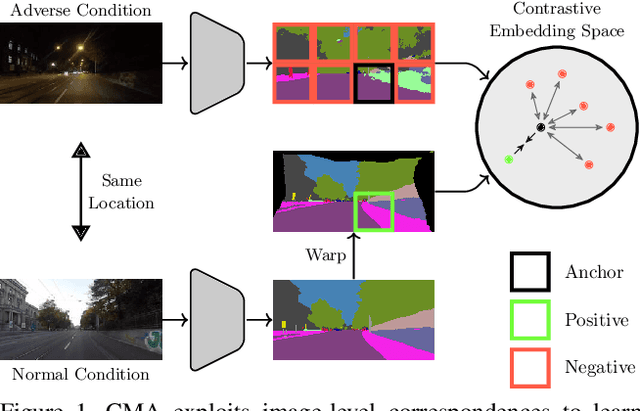
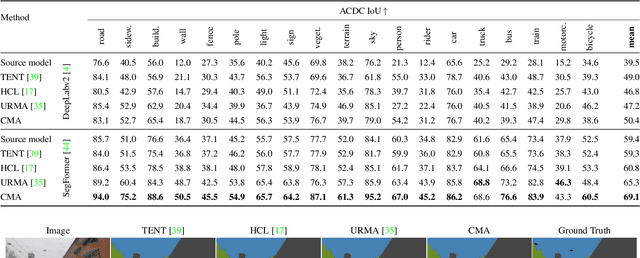
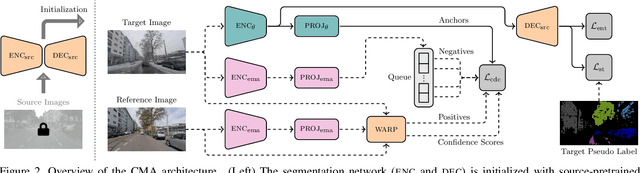
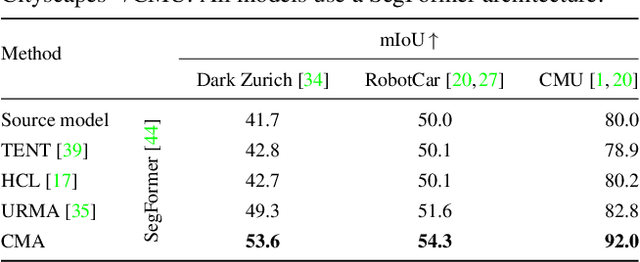
Standard unsupervised domain adaptation methods adapt models from a source to a target domain using labeled source data and unlabeled target data jointly. In model adaptation, on the other hand, access to the labeled source data is prohibited, i.e., only the source-trained model and unlabeled target data are available. We investigate normal-to-adverse condition model adaptation for semantic segmentation, whereby image-level correspondences are available in the target domain. The target set consists of unlabeled pairs of adverse- and normal-condition street images taken at GPS-matched locations. Our method -- CMA -- leverages such image pairs to learn condition-invariant features via contrastive learning. In particular, CMA encourages features in the embedding space to be grouped according to their condition-invariant semantic content and not according to the condition under which respective inputs are captured. To obtain accurate cross-domain semantic correspondences, we warp the normal image to the viewpoint of the adverse image and leverage warp-confidence scores to create robust, aggregated features. With this approach, we achieve state-of-the-art semantic segmentation performance for model adaptation on several normal-to-adverse adaptation benchmarks, such as ACDC and Dark Zurich. We also evaluate CMA on a newly procured adverse-condition generalization benchmark and report favorable results compared to standard unsupervised domain adaptation methods, despite the comparative handicap of CMA due to source data inaccessibility. Code is available at https://github.com/brdav/cma.
Improving 3D Imaging with Pre-Trained Perpendicular 2D Diffusion Models
Mar 15, 2023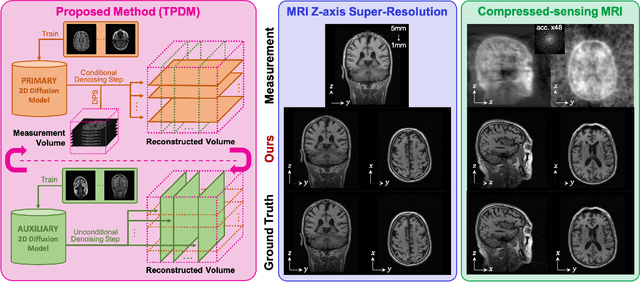
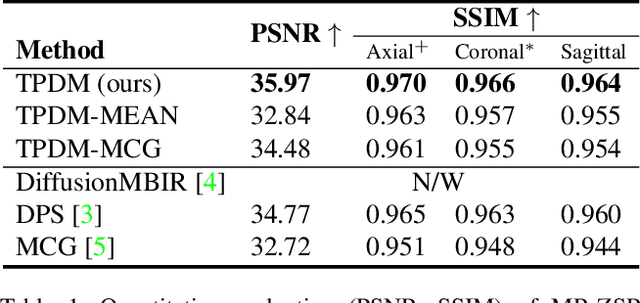
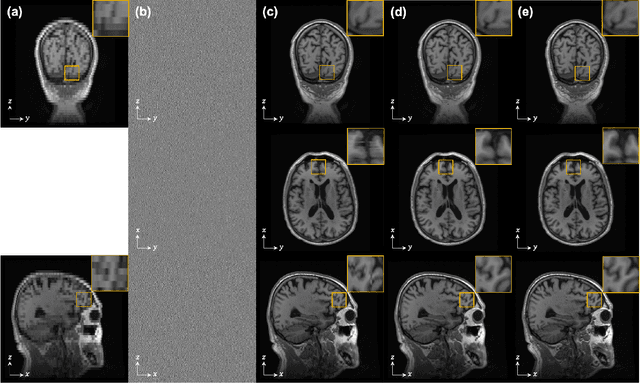
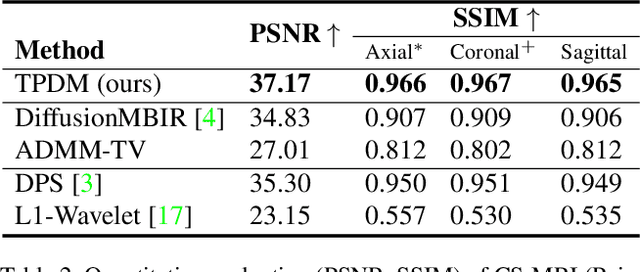
Diffusion models have become a popular approach for image generation and reconstruction due to their numerous advantages. However, most diffusion-based inverse problem-solving methods only deal with 2D images, and even recently published 3D methods do not fully exploit the 3D distribution prior. To address this, we propose a novel approach using two perpendicular pre-trained 2D diffusion models to solve the 3D inverse problem. By modeling the 3D data distribution as a product of 2D distributions sliced in different directions, our method effectively addresses the curse of dimensionality. Our experimental results demonstrate that our method is highly effective for 3D medical image reconstruction tasks, including MRI Z-axis super-resolution, compressed sensing MRI, and sparse-view CT. Our method can generate high-quality voxel volumes suitable for medical applications.
Direct Motif Extraction from High Resolution Crystalline STEM Images
Mar 13, 2023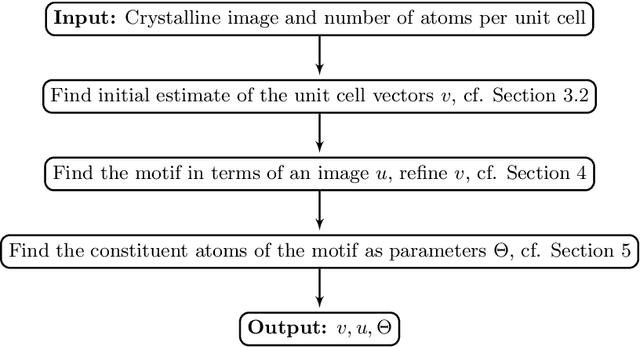
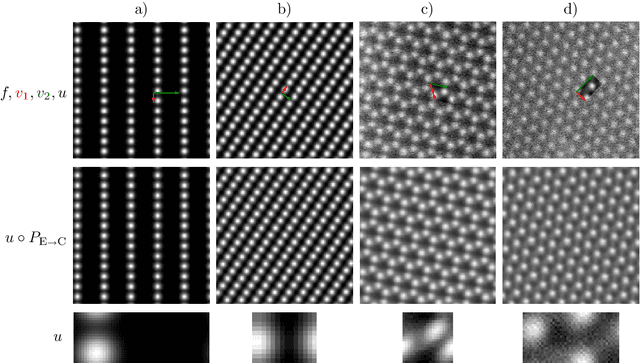
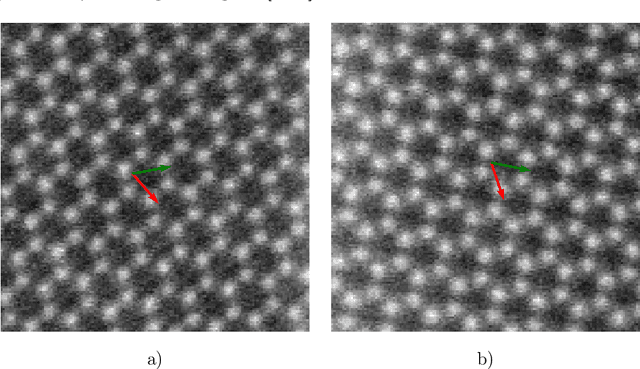
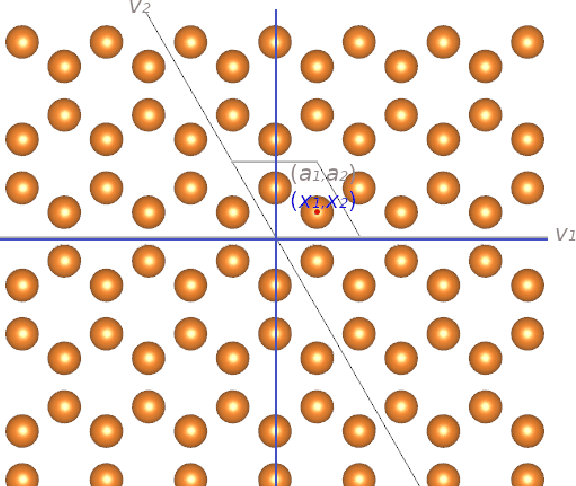

During the last decade, automatic data analysis methods concerning different aspects of crystal analysis have been developed, e.g., unsupervised primitive unit cell extraction and automated crystal distortion and defects detection. However, an automatic, unsupervised motif extraction method is still not widely available yet. Here, we propose and demonstrate a novel method for the automatic motif extraction in real space from crystalline images based on a variational approach involving the unit cell projection operator. Due to the non-convex nature of the resulting minimization problem, a multi-stage algorithm is used. First, we determine the primitive unit cell in form of two lattice vectors. Second, a motif image is estimated using the unit cell information. Finally, the motif is determined in terms of atom positions inside the unit cell. The method was tested on various synthetic and experimental HAADF STEM images. The results are a representation of the motif in form of an image, atomic positions, primitive unit cell vectors, and a denoised and a modeled reconstruction of the input image. The method was applied to extract the primitive cells of complex $\mu$-phase structures Nb$_\text{6.4}$Co$_\text{6.6}$ and Nb$_\text{7}$Co$_\text{6}$, where subtle differences between their interplanar spacings were determined.
On the De-duplication of LAION-2B
Mar 17, 2023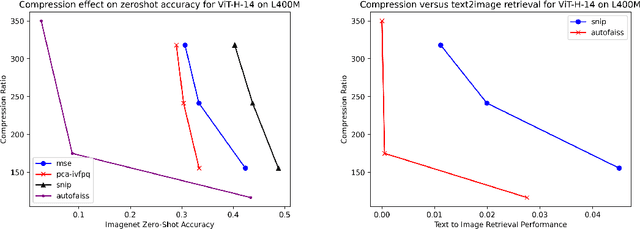
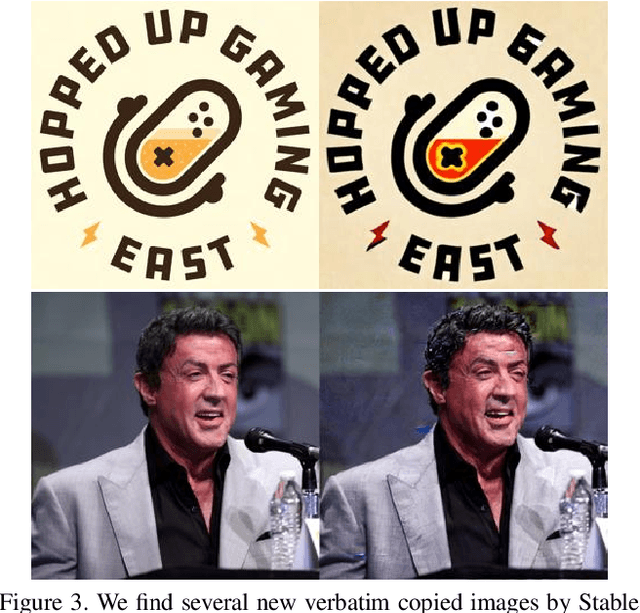
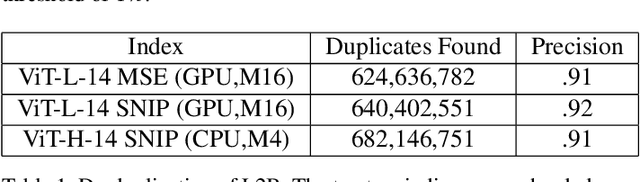
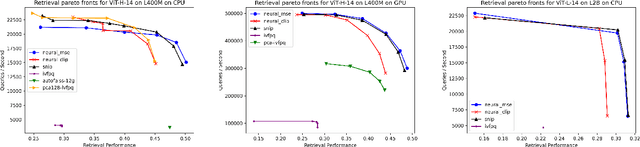

Generative models, such as DALL-E, Midjourney, and Stable Diffusion, have societal implications that extend beyond the field of computer science. These models require large image databases like LAION-2B, which contain two billion images. At this scale, manual inspection is difficult and automated analysis is challenging. In addition, recent studies show that duplicated images pose copyright problems for models trained on LAION2B, which hinders its usability. This paper proposes an algorithmic chain that runs with modest compute, that compresses CLIP features to enable efficient duplicate detection, even for vast image volumes. Our approach demonstrates that roughly 700 million images, or about 30\%, of LAION-2B's images are likely duplicated. Our method also provides the histograms of duplication on this dataset, which we use to reveal more examples of verbatim copies by Stable Diffusion and further justify the approach. The current version of the de-duplicated set will be distributed online.
Prototype Knowledge Distillation for Medical Segmentation with Missing Modality
Mar 17, 2023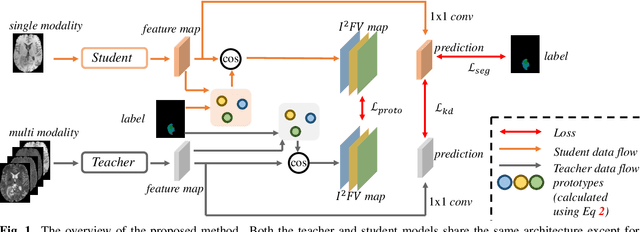
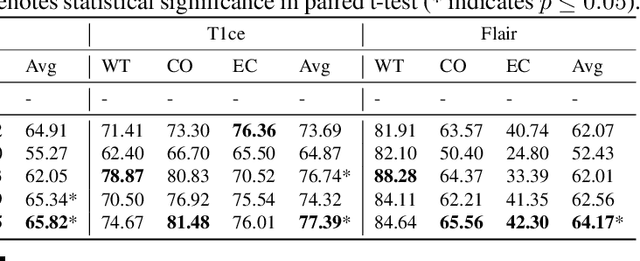
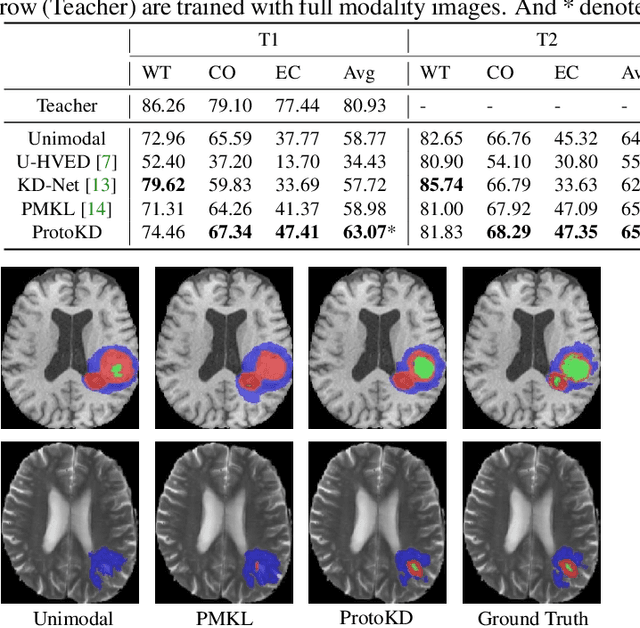
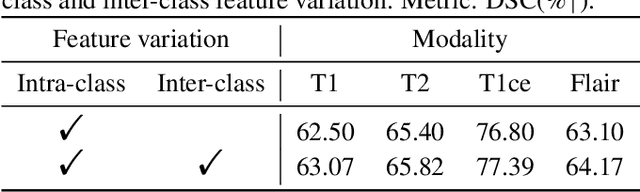
Multi-modality medical imaging is crucial in clinical treatment as it can provide complementary information for medical image segmentation. However, collecting multi-modal data in clinical is difficult due to the limitation of the scan time and other clinical situations. As such, it is clinically meaningful to develop an image segmentation paradigm to handle this missing modality problem. In this paper, we propose a prototype knowledge distillation (ProtoKD) method to tackle the challenging problem, especially for the toughest scenario when only single modal data can be accessed. Specifically, our ProtoKD can not only distillate the pixel-wise knowledge of multi-modality data to single-modality data but also transfer intra-class and inter-class feature variations, such that the student model could learn more robust feature representation from the teacher model and inference with only one single modality data. Our method achieves state-of-the-art performance on BraTS benchmark.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge