"Image": models, code, and papers
Generalist Vision Foundation Models for Medical Imaging: A Case Study of Segment Anything Model on Zero-Shot Medical Segmentation
Apr 25, 2023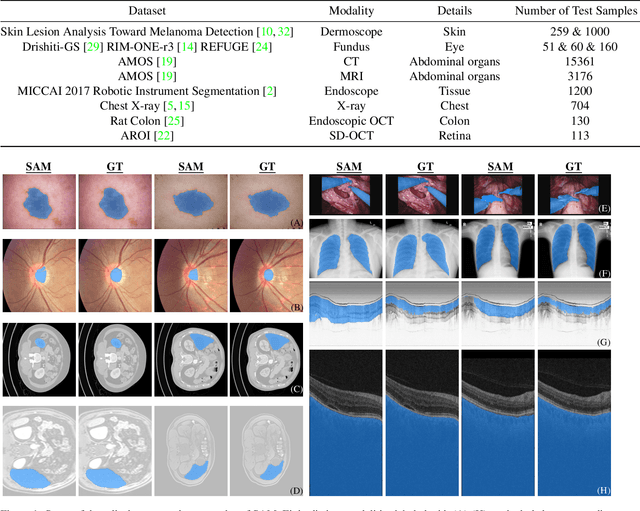
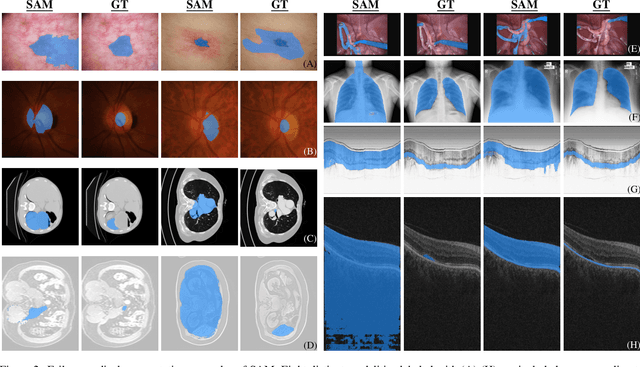
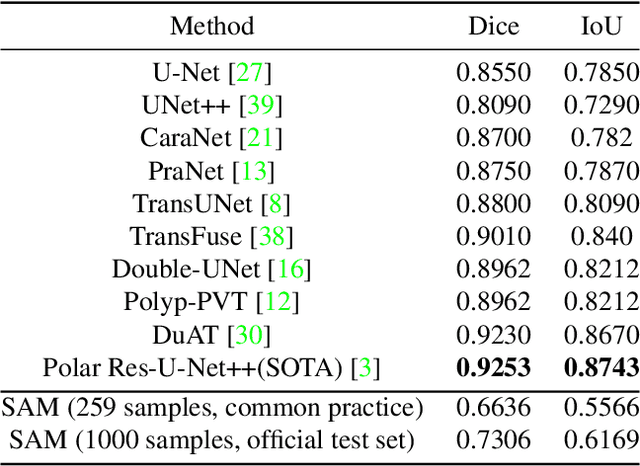
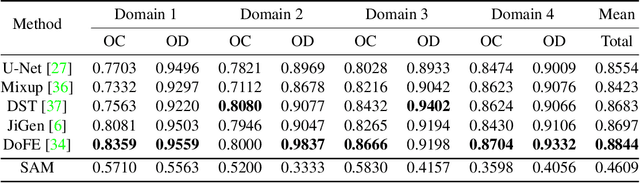
We examine the recent Segment Anything Model (SAM) on medical images, and report both quantitative and qualitative zero-shot segmentation results on nine medical image segmentation benchmarks, covering various imaging modalities, such as optical coherence tomography (OCT), magnetic resonance imaging (MRI), and computed tomography (CT), as well as different applications including dermatology, ophthalmology, and radiology. Our experiments reveal that while SAM demonstrates stunning segmentation performance on images from the general domain, for those out-of-distribution images, e.g., medical images, its zero-shot segmentation performance is still limited. Furthermore, SAM demonstrated varying zero-shot segmentation performance across different unseen medical domains. For example, it had a 0.8704 mean Dice score on segmenting under-bruch's membrane layer of retinal OCT, whereas the segmentation accuracy drops to 0.0688 when segmenting retinal pigment epithelium. For certain structured targets, e.g., blood vessels, the zero-shot segmentation of SAM completely failed, whereas a simple fine-tuning of it with small amount of data could lead to remarkable improvements of the segmentation quality. Our study indicates the versatility of generalist vision foundation models on solving specific tasks in medical imaging, and their great potential to achieve desired performance through fine-turning and eventually tackle the challenges of accessing large diverse medical datasets and the complexity of medical domains.
Segment anything, from space?
Apr 25, 2023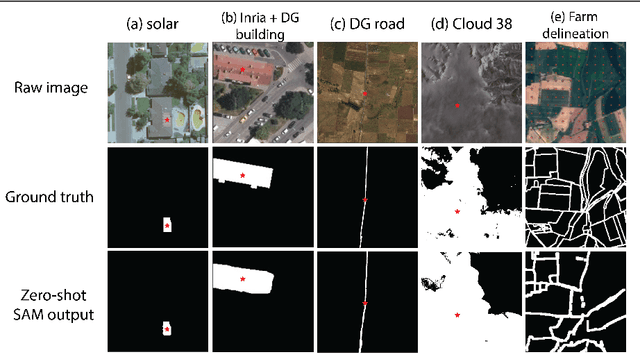
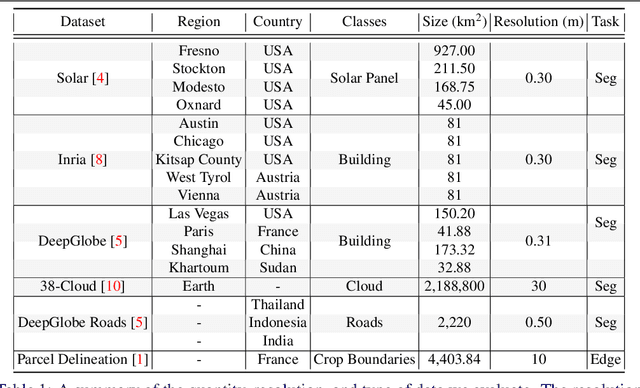
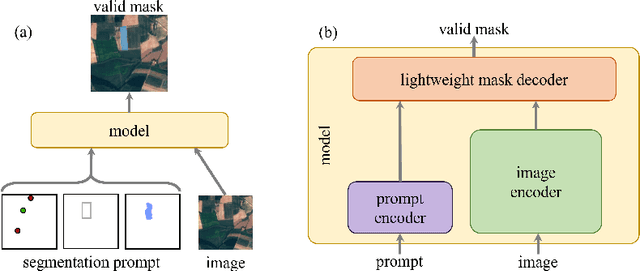
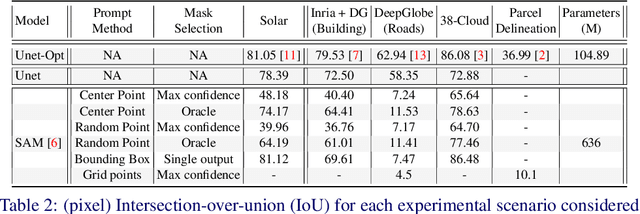
Recently, the first foundation model developed specifically for vision tasks was developed, termed the "Segment Anything Model" (SAM). SAM can segment objects in input imagery based upon cheap input prompts, such as one (or more) points, a bounding box, or a mask. The authors examined the zero-shot image segmentation accuracy of SAM on a large number of vision benchmark tasks and found that SAM usually achieved recognition accuracy similar to, or sometimes exceeding, vision models that had been trained on the target tasks. The impressive generalization of SAM for segmentation has major implications for vision researchers working on natural imagery. In this work, we examine whether SAM's impressive performance extends to overhead imagery problems, and help guide the community's response to its development. We examine SAM's performance on a set of diverse and widely-studied benchmark tasks. We find that SAM does often generalize well to overhead imagery, although it fails in some cases due to the unique characteristics of overhead imagery and the target objects. We report on these unique systematic failure cases for remote sensing imagery that may comprise useful future research for the community. Note that this is a working paper, and it will be updated as additional analysis and results are completed.
Feature Tracks are not Zero-Mean Gaussian
Mar 25, 2023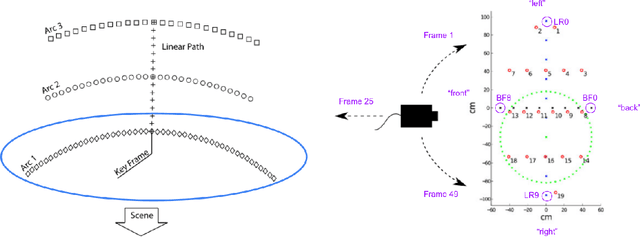

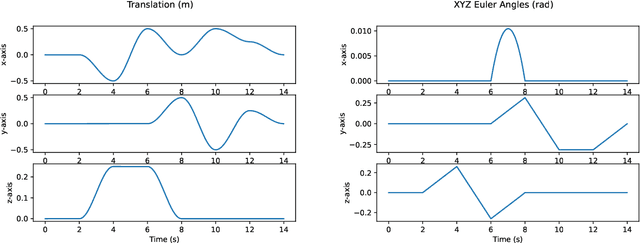

In state estimation algorithms that use feature tracks as input, it is customary to assume that the errors in feature track positions are zero-mean Gaussian. Using a combination of calibrated camera intrinsics, ground-truth camera pose, and depth images, it is possible to compute ground-truth positions for feature tracks extracted using an image processing algorithm. We find that feature track errors are not zero-mean Gaussian and that the distribution of errors is conditional on the type of motion, the speed of motion, and the image processing algorithm used to extract the tracks.
TruFor: Leveraging all-round clues for trustworthy image forgery detection and localization
Dec 21, 2022
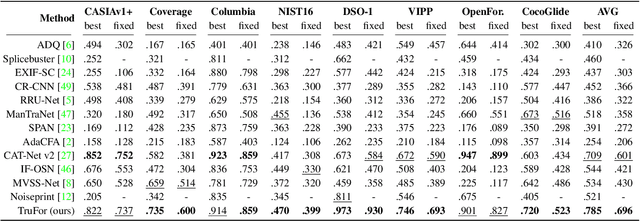
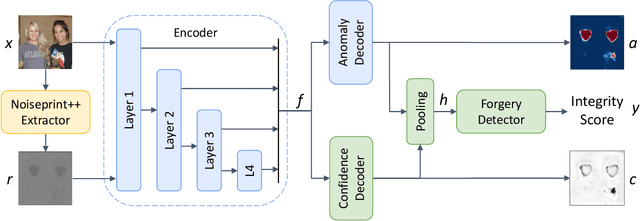
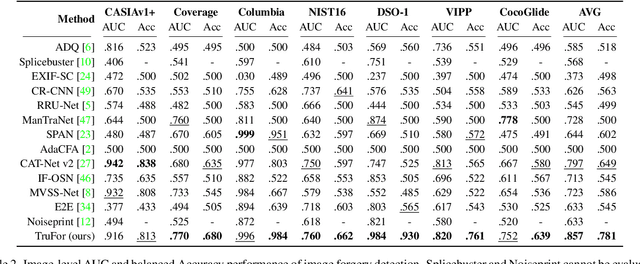
In this paper we present TruFor, a forensic framework that can be applied to a large variety of image manipulation methods, from classic cheapfakes to more recent manipulations based on deep learning. We rely on the extraction of both high-level and low-level traces through a transformer-based fusion architecture that combines the RGB image and a learned noise-sensitive fingerprint. The latter learns to embed the artifacts related to the camera internal and external processing by training only on real data in a self-supervised manner. Forgeries are detected as deviations from the expected regular pattern that characterizes each pristine image. Looking for anomalies makes the approach able to robustly detect a variety of local manipulations, ensuring generalization. In addition to a pixel-level localization map and a whole-image integrity score, our approach outputs a reliability map that highlights areas where localization predictions may be error-prone. This is particularly important in forensic applications in order to reduce false alarms and allow for a large scale analysis. Extensive experiments on several datasets show that our method is able to reliably detect and localize both cheapfakes and deepfakes manipulations outperforming state-of-the-art works. Code will be publicly available at https://grip-unina.github.io/TruFor/
Ensuring Trustworthy Medical Artificial Intelligence through Ethical and Philosophical Principles
Apr 29, 2023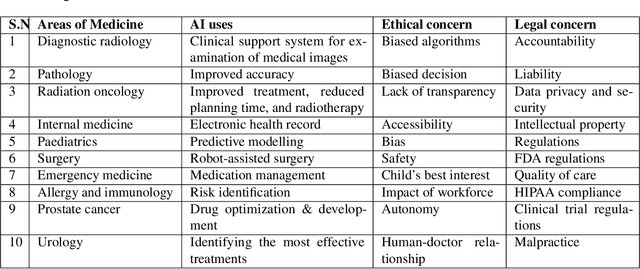
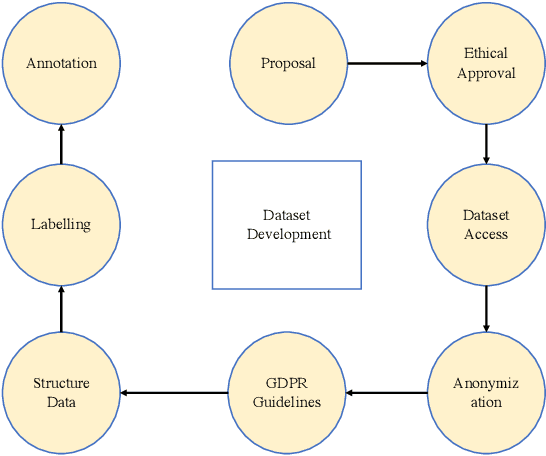

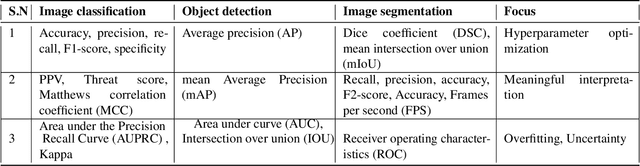
Artificial intelligence (AI) methods have great potential to revolutionize numerous medical care by enhancing the experience of medical experts and patients. AI based computer-assisted diagnosis tools can have a tremendous benefit if they can outperform or perform similarly to the level of a clinical expert. As a result, advanced healthcare services can be affordable in developing nations, and the problem of a lack of expert medical practitioners can be addressed. AI based tools can save time, resources, and overall cost for patient treatment. Furthermore, in contrast to humans, AI can uncover complex relations in the data from a large set of inputs and even lead to new evidence-based knowledge in medicine. However, integrating AI in healthcare raises several ethical and philosophical concerns, such as bias, transparency, autonomy, responsibility and accountability, which must be addressed before integrating such tools into clinical settings. In this article, we emphasize recent advances in AI-assisted medical image analysis, existing standards, and the significance of comprehending ethical issues and best practices for the applications of AI in clinical settings. We cover the technical and ethical challenges of AI and the implications of deploying AI in hospitals and public organizations. We also discuss promising key measures and techniques to address the ethical challenges, data scarcity, racial bias, lack of transparency, and algorithmic bias. Finally, we provide our recommendation and future directions for addressing the ethical challenges associated with AI in healthcare applications, with the goal of deploying AI into the clinical settings to make the workflow more efficient, accurate, accessible, transparent, and reliable for the patient worldwide.
Real-Time Superficial Vein Imaging System for Observing Abnormalities on Vascular Structures
Apr 29, 2023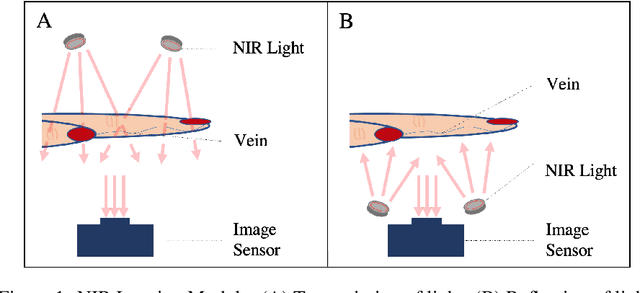
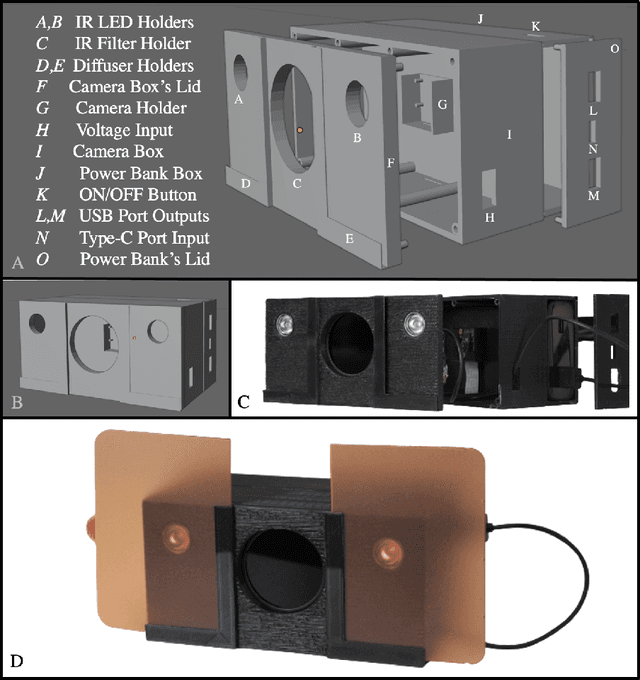
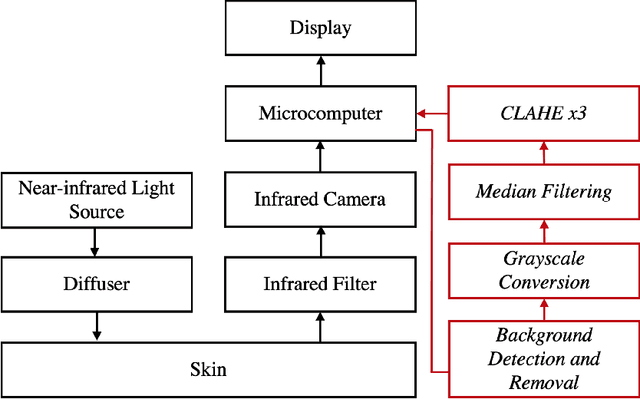

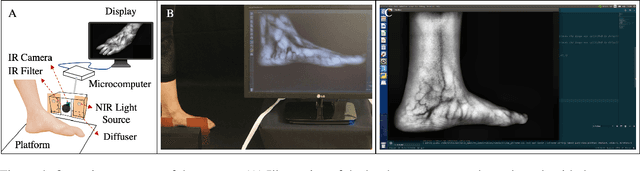
Circulatory system abnormalities might be an indicator of diseases or tissue damage. Early detection of vascular abnormalities might have an important role during treatment and also raise the patient's awarenes. Current detection methods for vascular imaging are high-cost, invasive, and mostly radiation-based. In this study, a low-cost and portable microcomputer-based tool has been developed as a near-infrared (NIR) superficial vascular imaging device. The device uses NIR light-emitting diode (LED) light at 850 nm along with other electronic and optical components. It operates as a non-contact and safe infrared (IR) imaging method in real-time. Image and video analysis are carried out using OpenCV (Open-Source Computer Vision), a library of programming functions mainly used in computer vision. Various tests were carried out to optimize the imaging system and set up a suitable external environment. To test the performance of the device, the images taken from three diabetic volunteers, who are expected to have abnormalities in the vascular structure due to the possibility of deformation caused by high glucose levels in the blood, were compared with the images taken from two non-diabetic volunteers. As a result, tortuosity was observed successfully in the superficial vascular structures, where the results need to be interpreted by the medical experts in the field to understand the underlying reasons. Although this study is an engineering study and does not have an intention to diagnose any diseases, the developed system here might assist healthcare personnel in early diagnosis and treatment follow-up for vascular structures and may enable further opportunities.
Affordance Diffusion: Synthesizing Hand-Object Interactions
Mar 21, 2023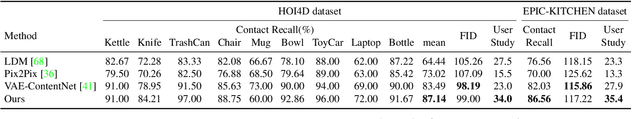
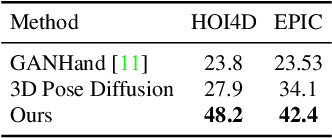
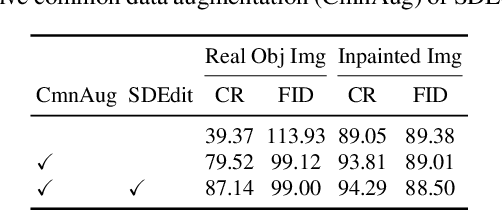
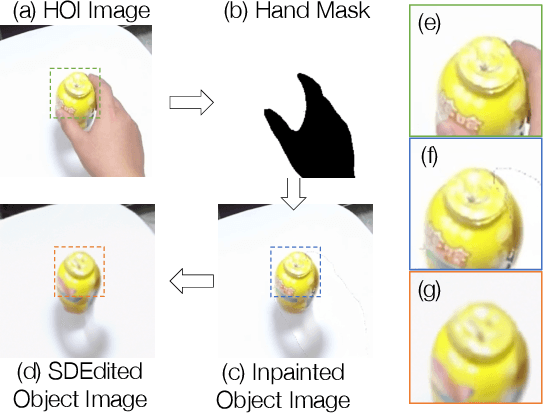

Recent successes in image synthesis are powered by large-scale diffusion models. However, most methods are currently limited to either text- or image-conditioned generation for synthesizing an entire image, texture transfer or inserting objects into a user-specified region. In contrast, in this work we focus on synthesizing complex interactions (ie, an articulated hand) with a given object. Given an RGB image of an object, we aim to hallucinate plausible images of a human hand interacting with it. We propose a two-step generative approach: a LayoutNet that samples an articulation-agnostic hand-object-interaction layout, and a ContentNet that synthesizes images of a hand grasping the object given the predicted layout. Both are built on top of a large-scale pretrained diffusion model to make use of its latent representation. Compared to baselines, the proposed method is shown to generalize better to novel objects and perform surprisingly well on out-of-distribution in-the-wild scenes of portable-sized objects. The resulting system allows us to predict descriptive affordance information, such as hand articulation and approaching orientation. Project page: https://judyye.github.io/affordiffusion-www
Few Shot Semantic Segmentation: a review of methodologies and open challenges
Apr 12, 2023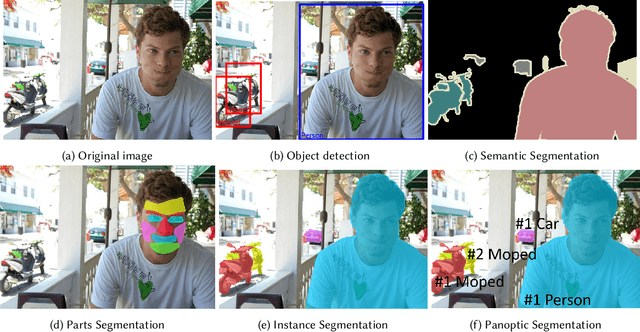
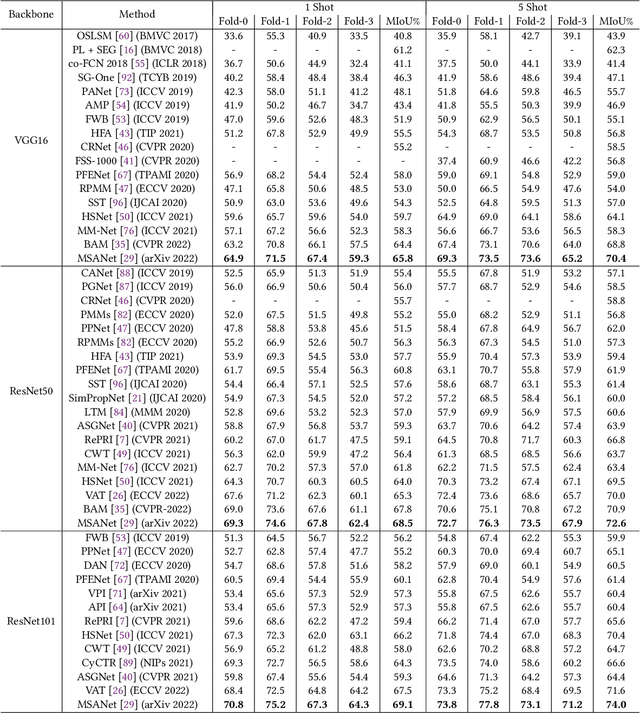
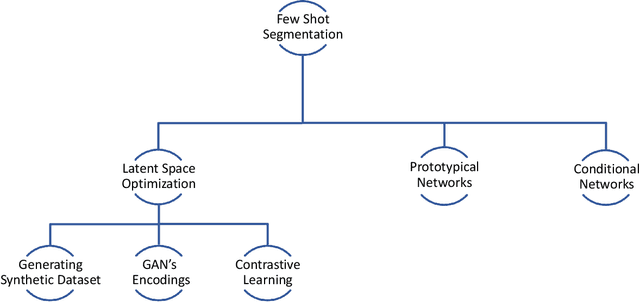
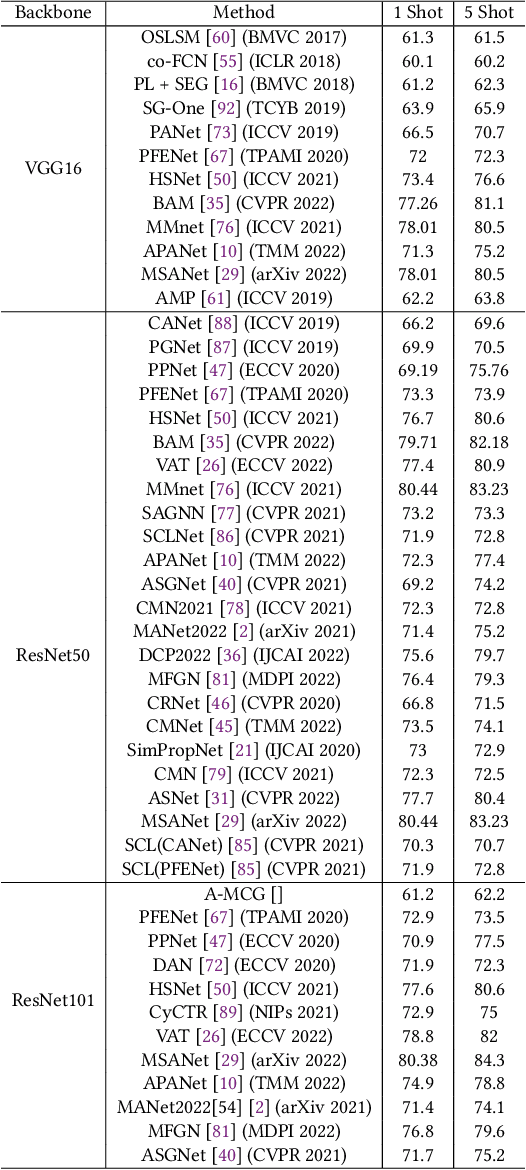
Semantic segmentation assigns category labels to each pixel in an image, enabling breakthroughs in fields such as autonomous driving and robotics. Deep Neural Networks have achieved high accuracies in semantic segmentation but require large training datasets. Some domains have difficulties building such datasets due to rarity, privacy concerns, and the need for skilled annotators. Few-Shot Learning (FSL) has emerged as a new research stream that allows models to learn new tasks from a few samples. This contribution provides an overview of FSL in semantic segmentation (FSS), proposes a new taxonomy, and describes current limitations and outlooks.
HADA: A Graph-based Amalgamation Framework in Image-text Retrieval
Jan 11, 2023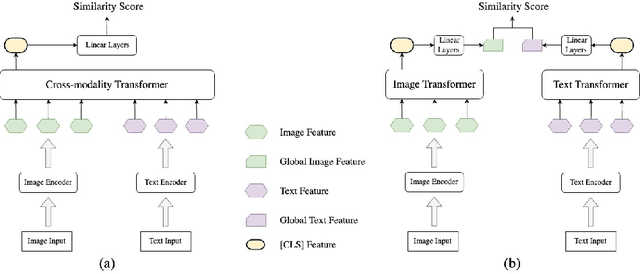
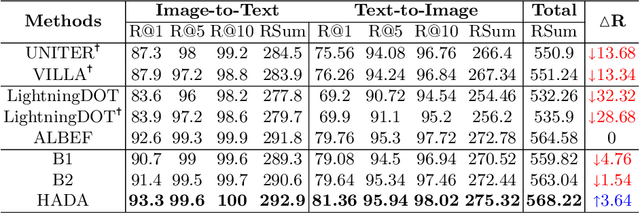
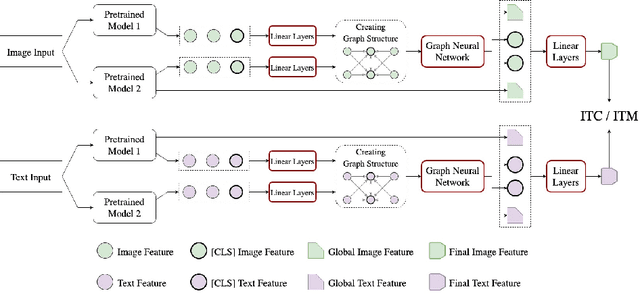
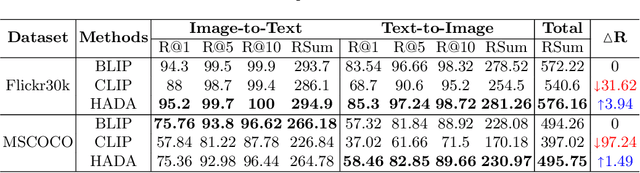
Many models have been proposed for vision and language tasks, especially the image-text retrieval task. All state-of-the-art (SOTA) models in this challenge contained hundreds of millions of parameters. They also were pretrained on a large external dataset that has been proven to make a big improvement in overall performance. It is not easy to propose a new model with a novel architecture and intensively train it on a massive dataset with many GPUs to surpass many SOTA models, which are already available to use on the Internet. In this paper, we proposed a compact graph-based framework, named HADA, which can combine pretrained models to produce a better result, rather than building from scratch. First, we created a graph structure in which the nodes were the features extracted from the pretrained models and the edges connecting them. The graph structure was employed to capture and fuse the information from every pretrained model with each other. Then a graph neural network was applied to update the connection between the nodes to get the representative embedding vector for an image and text. Finally, we used the cosine similarity to match images with their relevant texts and vice versa to ensure a low inference time. Our experiments showed that, although HADA contained a tiny number of trainable parameters, it could increase baseline performance by more than 3.6% in terms of evaluation metrics in the Flickr30k dataset. Additionally, the proposed model did not train on any external dataset and did not require many GPUs but only 1 to train due to its small number of parameters. The source code is available at https://github.com/m2man/HADA.
NASDM: Nuclei-Aware Semantic Histopathology Image Generation Using Diffusion Models
Mar 20, 2023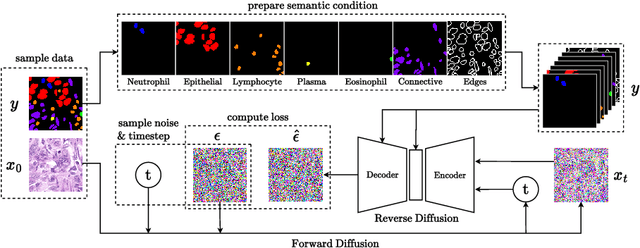
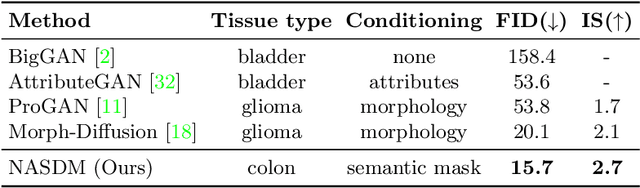

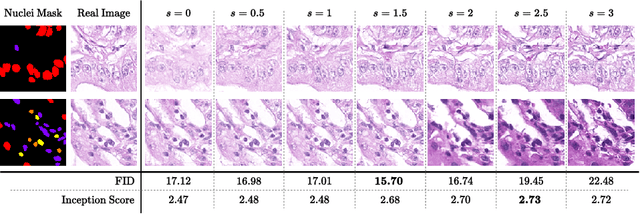
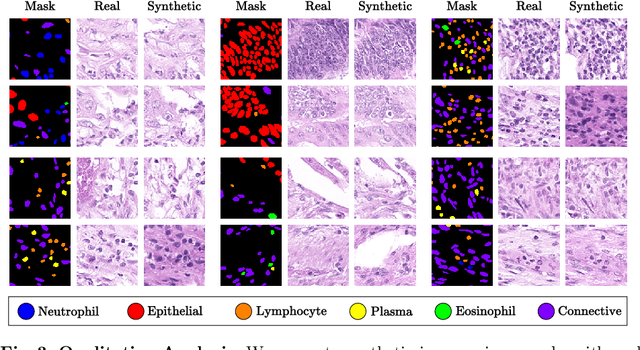
In recent years, computational pathology has seen tremendous progress driven by deep learning methods in segmentation and classification tasks aiding prognostic and diagnostic settings. Nuclei segmentation, for instance, is an important task for diagnosing different cancers. However, training deep learning models for nuclei segmentation requires large amounts of annotated data, which is expensive to collect and label. This necessitates explorations into generative modeling of histopathological images. In this work, we use recent advances in conditional diffusion modeling to formulate a first-of-its-kind nuclei-aware semantic tissue generation framework (NASDM) which can synthesize realistic tissue samples given a semantic instance mask of up to six different nuclei types, enabling pixel-perfect nuclei localization in generated samples. These synthetic images are useful in applications in pathology pedagogy, validation of models, and supplementation of existing nuclei segmentation datasets. We demonstrate that NASDM is able to synthesize high-quality histopathology images of the colon with superior quality and semantic controllability over existing generative methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge