"Image": models, code, and papers
Supervised Image Segmentation for High Dynamic Range Imaging
Dec 06, 2022
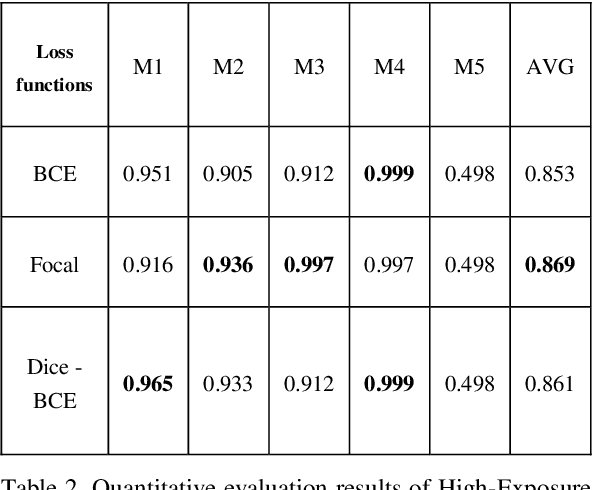
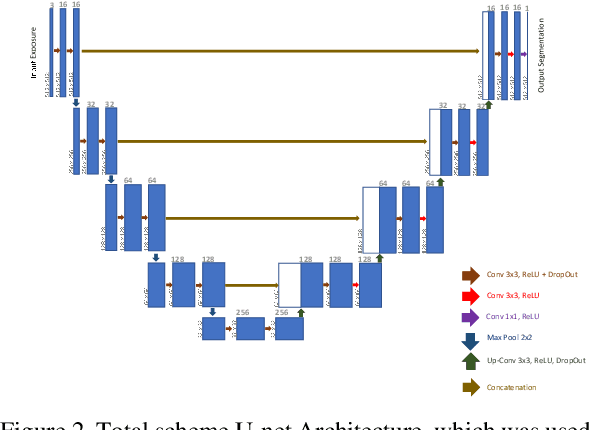
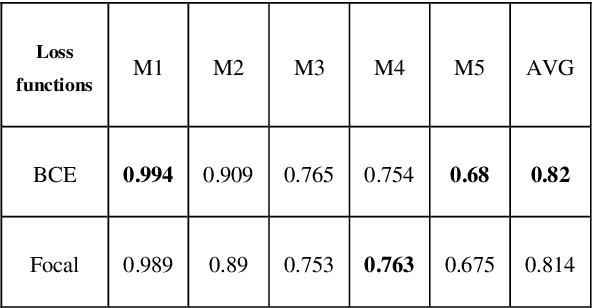
Regular cameras and cell phones are able to capture limited luminosity. Thus, in terms of quality, most of the produced images from such devices are not similar to the real world. They are overly dark or too bright, and the details are not perfectly visible. Various methods, which fall under the name of High Dynamic Range (HDR) Imaging, can be utilised to cope with this problem. Their objective is to produce an image with more details. However, unfortunately, most methods for generating an HDR image from Multi-Exposure images only concentrate on how to combine different exposures and do not have any focus on choosing the best details of each image. Therefore, it is strived in this research to extract the most visible areas of each image with the help of image segmentation. Two methods of producing the Ground Truth were considered, as manual threshold and Otsu threshold, and a neural network will be used to train segment these areas. Finally, it will be shown that the neural network is able to segment the visible parts of pictures acceptably.
Cyclic Generative Adversarial Networks With Congruent Image-Report Generation For Explainable Medical Image Analysis
Nov 16, 2022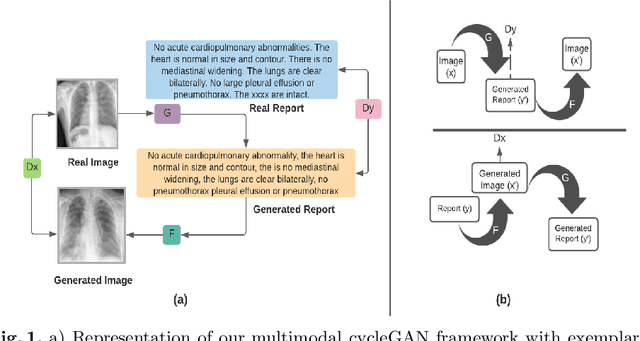
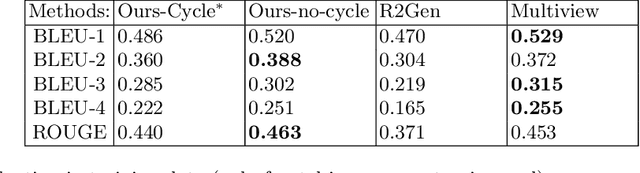
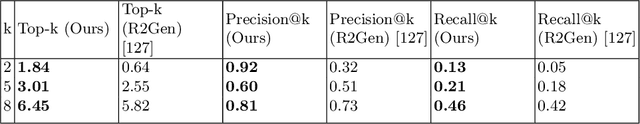
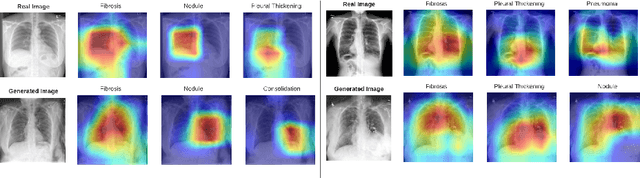
We present a novel framework for explainable labeling and interpretation of medical images. Medical images require specialized professionals for interpretation, and are explained (typically) via elaborate textual reports. Different from prior methods that focus on medical report generation from images or vice-versa, we novelly generate congruent image--report pairs employing a cyclic-Generative Adversarial Network (cycleGAN); thereby, the generated report will adequately explain a medical image, while a report-generated image that effectively characterizes the text visually should (sufficiently) resemble the original. The aim of the work is to generate trustworthy and faithful explanations for the outputs of a model diagnosing chest x-ray images by pointing a human user to similar cases in support of a diagnostic decision. Apart from enabling transparent medical image labeling and interpretation, we achieve report and image-based labeling comparable to prior methods, including state-of-the-art performance in some cases as evidenced by experiments on the Indiana Chest X-ray dataset
Score-Based Diffusion Models as Principled Priors for Inverse Imaging
Apr 23, 2023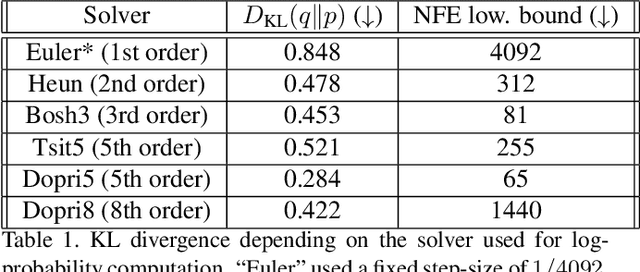
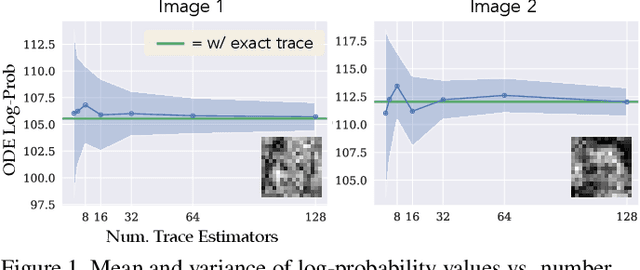
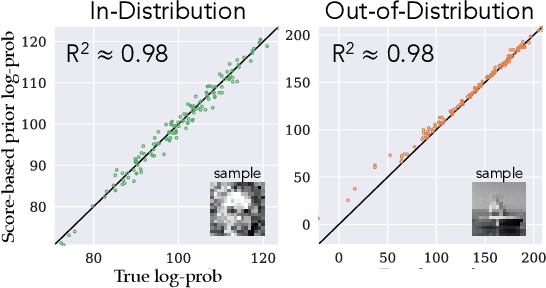
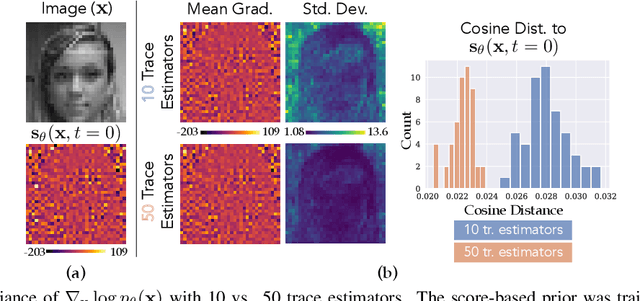
It is important in computational imaging to understand the uncertainty of images reconstructed from imperfect measurements. We propose turning score-based diffusion models into principled priors (``score-based priors'') for analyzing a posterior of images given measurements. Previously, probabilistic priors were limited to handcrafted regularizers and simple distributions. In this work, we empirically validate the theoretically-proven probability function of a score-based diffusion model. We show how to sample from resulting posteriors by using this probability function for variational inference. Our results, including experiments on denoising, deblurring, and interferometric imaging, suggest that score-based priors enable principled inference with a sophisticated, data-driven image prior.
If At First You Don't Succeed: Test Time Re-ranking for Zero-shot, Cross-domain Retrieval
Mar 30, 2023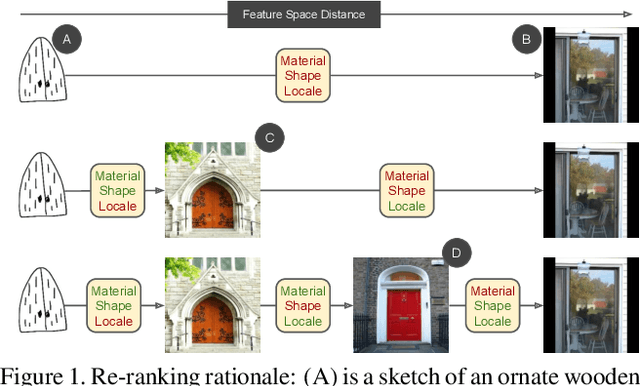
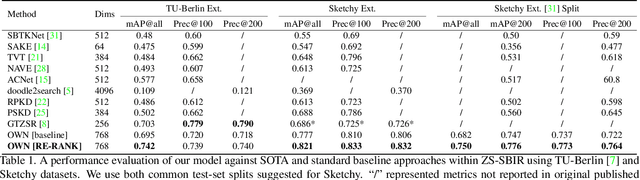
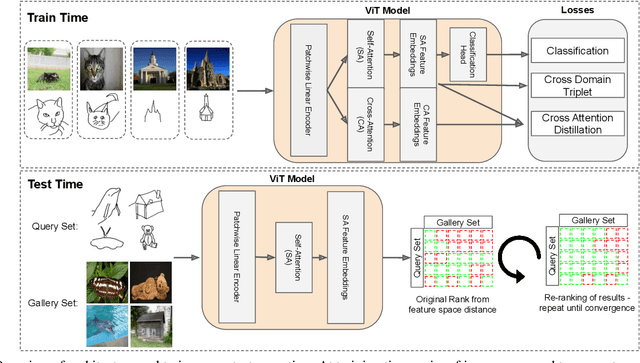
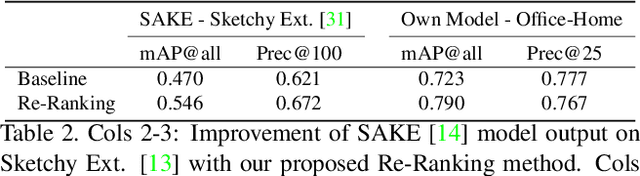
In this paper we propose a novel method for zero-shot, cross-domain image retrieval in which we make two key contributions. The first is a test-time re-ranking procedure that enables query-gallery pairs, without meaningful shared visual features, to be matched by incorporating gallery-gallery ranks into an iterative re-ranking process. The second is the use of cross-attention at training time and knowledge distillation to encourage cross-attention-like features to be extracted at test time from a single image. When combined with the Vision Transformer architecture and zero-shot retrieval losses, our approach yields state-of-the-art results on the Sketchy and TU-Berlin sketch-based image retrieval benchmarks. However, unlike many previous methods, none of the components in our approach are engineered specifically towards the sketch-based image retrieval task - it can be generally applied to any cross-domain, zero-shot retrieval task. We therefore also show results on zero-shot cartoon-to-photo retrieval using the Office-Home dataset.
Data-driven modelling of brain activity using neural networks, Diffusion Maps, and the Koopman operator
Apr 24, 2023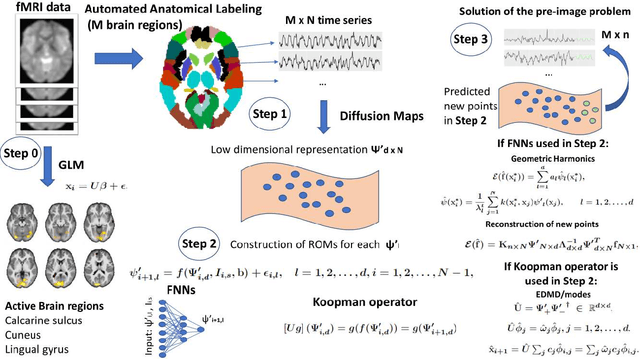
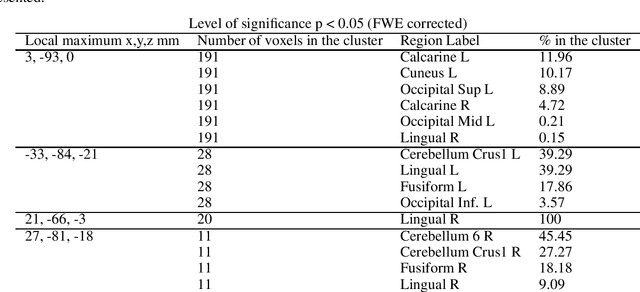
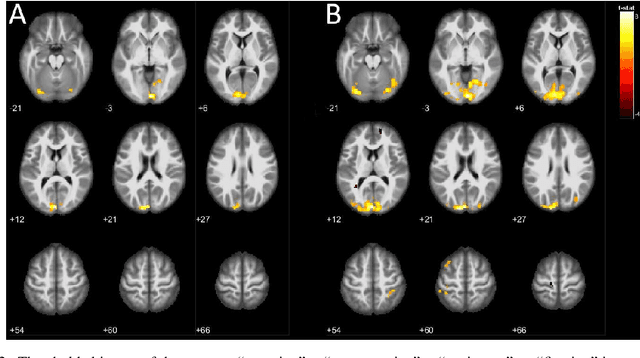
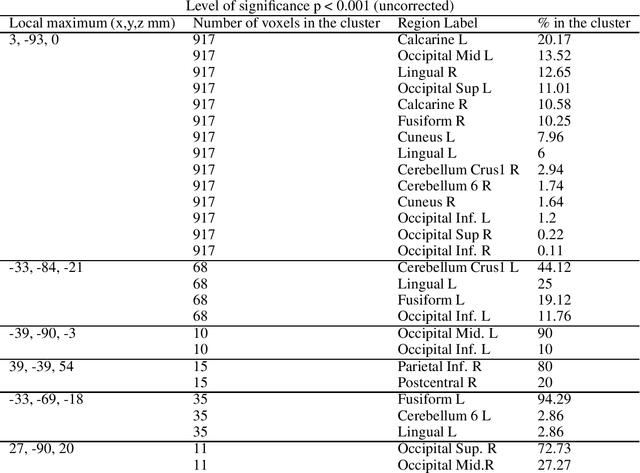
We propose a machine-learning approach to model long-term out-of-sample dynamics of brain activity from task-dependent fMRI data. Our approach is a three stage one. First, we exploit Diffusion maps (DMs) to discover a set of variables that parametrize the low-dimensional manifold on which the emergent high-dimensional fMRI time series evolve. Then, we construct reduced-order-models (ROMs) on the embedded manifold via two techniques: Feedforward Neural Networks (FNNs) and the Koopman operator. Finally, for predicting the out-of-sample long-term dynamics of brain activity in the ambient fMRI space, we solve the pre-image problem coupling DMs with Geometric Harmonics (GH) when using FNNs and the Koopman modes per se. For our illustrations, we have assessed the performance of the two proposed schemes using a benchmark fMRI dataset with recordings during a visuo-motor task. The results suggest that just a few (for the particular task, five) non-linear coordinates of the high-dimensional fMRI time series provide a good basis for modelling and out-of-sample prediction of the brain activity. Furthermore, we show that the proposed approaches outperform the one-step ahead predictions of the naive random walk model, which, in contrast to our scheme, relies on the knowledge of the signals in the previous time step. Importantly, we show that the proposed Koopman operator approach provides, for any practical purposes, equivalent results to the FNN-GH approach, thus bypassing the need to train a non-linear map and to use GH to extrapolate predictions in the ambient fMRI space; one can use instead the low-frequency truncation of the DMs function space of L^2-integrable functions, to predict the entire list of coordinate functions in the fMRI space and to solve the pre-image problem.
MProtoNet: A Case-Based Interpretable Model for Brain Tumor Classification with 3D Multi-parametric Magnetic Resonance Imaging
Apr 14, 2023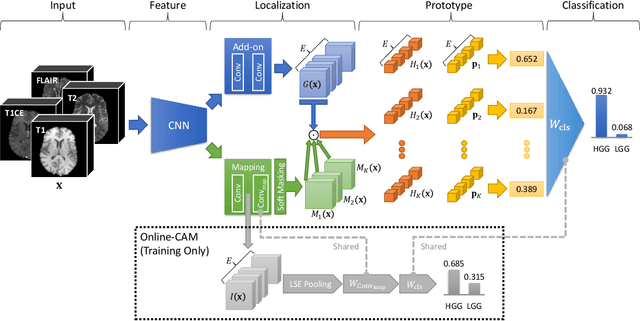
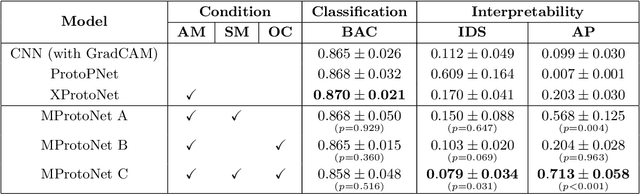
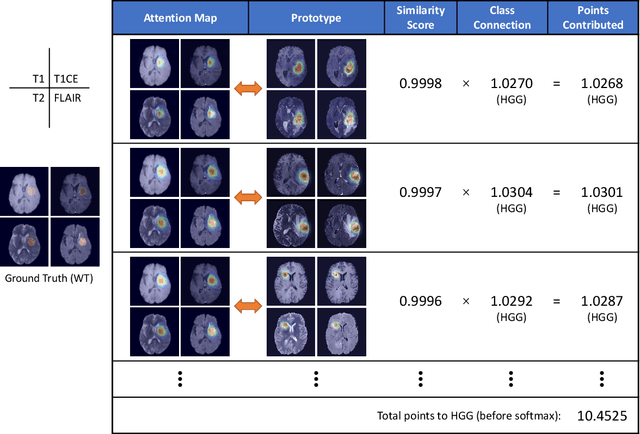
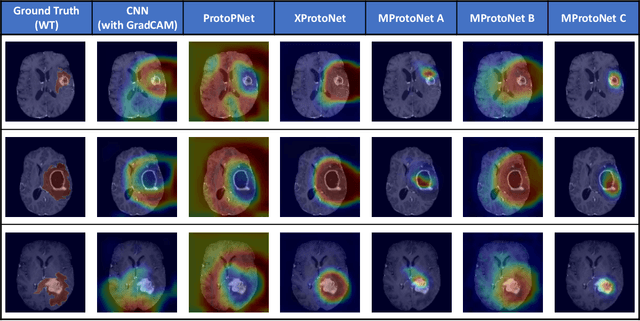
Recent applications of deep convolutional neural networks in medical imaging raise concerns about their interpretability. While most explainable deep learning applications use post hoc methods (such as GradCAM) to generate feature attribution maps, there is a new type of case-based reasoning models, namely ProtoPNet and its variants, which identify prototypes during training and compare input image patches with those prototypes. We propose the first medical prototype network (MProtoNet) to extend ProtoPNet to brain tumor classification with 3D multi-parametric magnetic resonance imaging (mpMRI) data. To address different requirements between 2D natural images and 3D mpMRIs especially in terms of localizing attention regions, a new attention module with soft masking and online-CAM loss is introduced. Soft masking helps sharpen attention maps, while online-CAM loss directly utilizes image-level labels when training the attention module. MProtoNet achieves statistically significant improvements in interpretability metrics of both correctness and localization coherence (with a best activation precision of $0.713\pm0.058$) without human-annotated labels during training, when compared with GradCAM and several ProtoPNet variants. The source code is available at https://github.com/aywi/mprotonet.
KS-GNNExplainer: Global Model Interpretation Through Instance Explanations On Histopathology images
Apr 14, 2023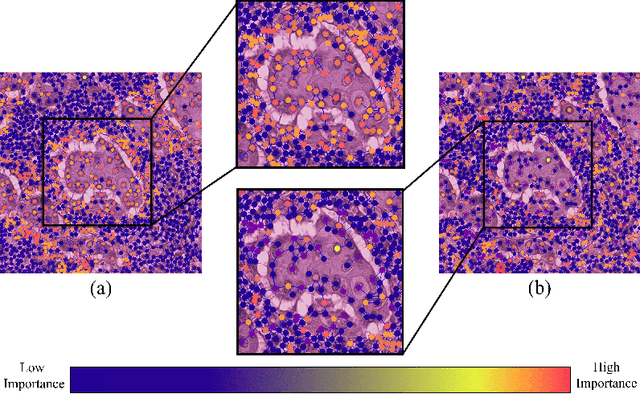
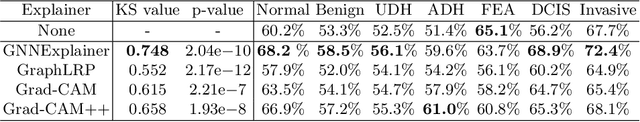

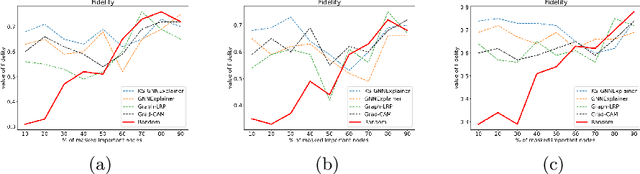
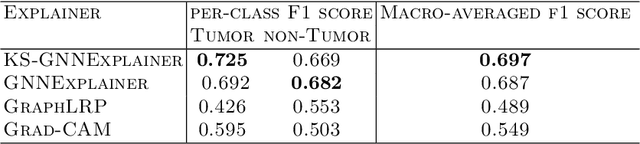
Instance-level graph neural network explainers have proven beneficial for explaining such networks on histopathology images. However, there has been few methods that provide model explanations, which are common patterns among samples within the same class. We envision that graph-based histopathological image analysis can benefit significantly from such explanations. On the other hand, current model-level explainers are based on graph generation methods that are not applicable in this domain because of no corresponding image for their generated graphs in real world. Therefore, such explanations are communicable to the experts. To follow this vision, we developed KS-GNNExplainer, the first instance-level graph neural network explainer that leverages current instance-level approaches in an effective manner to provide more informative and reliable explainable outputs, which are crucial for applied AI in the health domain. Our experiments on various datasets, and based on both quantitative and qualitative measures, demonstrate that the proposed explainer is capable of being a global pattern extractor, which is a fundamental limitation of current instance-level approaches in this domain.
Diffusion Models for Time Series Applications: A Survey
May 01, 2023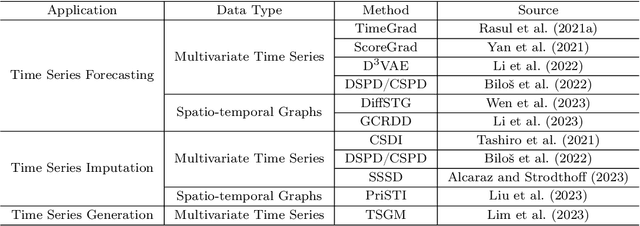
Diffusion models, a family of generative models based on deep learning, have become increasingly prominent in cutting-edge machine learning research. With a distinguished performance in generating samples that resemble the observed data, diffusion models are widely used in image, video, and text synthesis nowadays. In recent years, the concept of diffusion has been extended to time series applications, and many powerful models have been developed. Considering the deficiency of a methodical summary and discourse on these models, we provide this survey as an elementary resource for new researchers in this area and also an inspiration to motivate future research. For better understanding, we include an introduction about the basics of diffusion models. Except for this, we primarily focus on diffusion-based methods for time series forecasting, imputation, and generation, and present them respectively in three individual sections. We also compare different methods for the same application and highlight their connections if applicable. Lastly, we conclude the common limitation of diffusion-based methods and highlight potential future research directions.
Deep-Learning-based Vasculature Extraction for Single-Scan Optical Coherence Tomography Angiography
Apr 20, 2023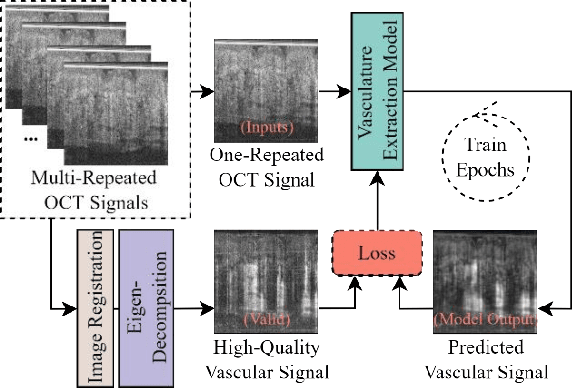
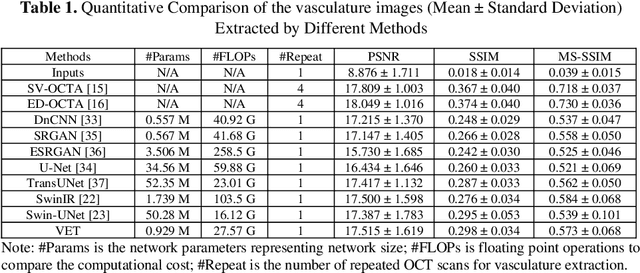
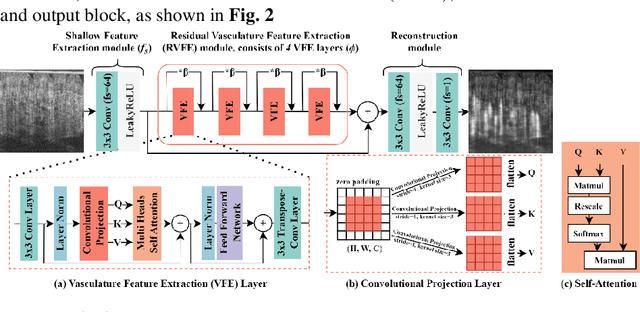
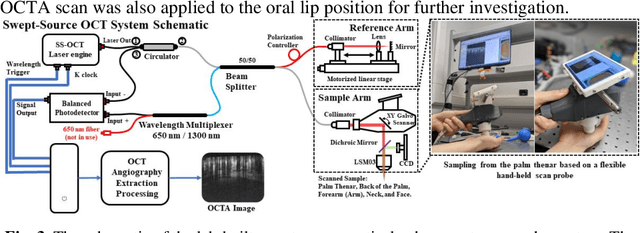
Optical coherence tomography angiography (OCTA) is a non-invasive imaging modality that extends the functionality of OCT by extracting moving red blood cell signals from surrounding static biological tissues. OCTA has emerged as a valuable tool for analyzing skin microvasculature, enabling more accurate diagnosis and treatment monitoring. Most existing OCTA extraction algorithms, such as speckle variance (SV)- and eigen-decomposition (ED)-OCTA, implement a larger number of repeated (NR) OCT scans at the same position to produce high-quality angiography images. However, a higher NR requires a longer data acquisition time, leading to more unpredictable motion artifacts. In this study, we propose a vasculature extraction pipeline that uses only one-repeated OCT scan to generate OCTA images. The pipeline is based on the proposed Vasculature Extraction Transformer (VET), which leverages convolutional projection to better learn the spatial relationships between image patches. In comparison to OCTA images obtained via the SV-OCTA (PSNR: 17.809) and ED-OCTA (PSNR: 18.049) using four-repeated OCT scans, OCTA images extracted by VET exhibit moderate quality (PSNR: 17.515) and higher image contrast while reducing the required data acquisition time from ~8 s to ~2 s. Based on visual observations, the proposed VET outperforms SV and ED algorithms when using neck and face OCTA data in areas that are challenging to scan. This study represents that the VET has the capacity to extract vascularture images from a fast one-repeated OCT scan, facilitating accurate diagnosis for patients.
OSTA: One-shot Task-adaptive Channel Selection for Semantic Segmentation of Multichannel Images
May 08, 2023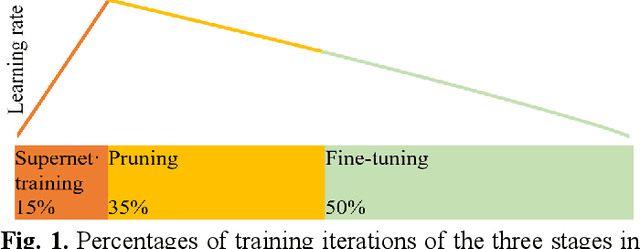
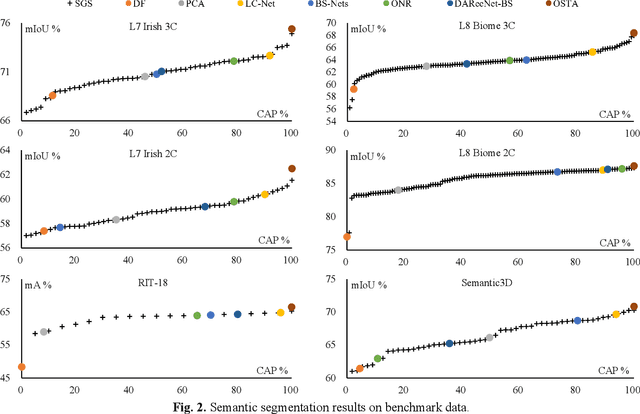
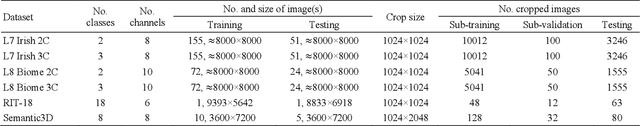
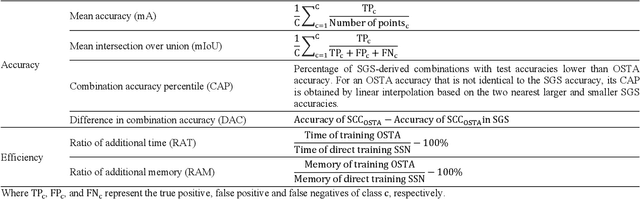
Semantic segmentation of multichannel images is a fundamental task for many applications. Selecting an appropriate channel combination from the original multichannel image can improve the accuracy of semantic segmentation and reduce the cost of data storage, processing and future acquisition. Existing channel selection methods typically use a reasonable selection procedure to determine a desirable channel combination, and then train a semantic segmentation network using that combination. In this study, the concept of pruning from a supernet is used for the first time to integrate the selection of channel combination and the training of a semantic segmentation network. Based on this concept, a One-Shot Task-Adaptive (OSTA) channel selection method is proposed for the semantic segmentation of multichannel images. OSTA has three stages, namely the supernet training stage, the pruning stage and the fine-tuning stage. The outcomes of six groups of experiments (L7Irish3C, L7Irish2C, L8Biome3C, L8Biome2C, RIT-18 and Semantic3D) demonstrated the effectiveness and efficiency of OSTA. OSTA achieved the highest segmentation accuracies in all tests (62.49% (mIoU), 75.40% (mIoU), 68.38% (mIoU), 87.63% (mIoU), 66.53% (mA) and 70.86% (mIoU), respectively). It even exceeded the highest accuracies of exhaustive tests (61.54% (mIoU), 74.91% (mIoU), 67.94% (mIoU), 87.32% (mIoU), 65.32% (mA) and 70.27% (mIoU), respectively), where all possible channel combinations were tested. All of this can be accomplished within a predictable and relatively efficient timeframe, ranging from 101.71% to 298.1% times the time required to train the segmentation network alone. In addition, there were interesting findings that were deemed valuable for several fields.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge