"Image": models, code, and papers
Case Studies on X-Ray Imaging, MRI and Nuclear Imaging
Jun 07, 2023
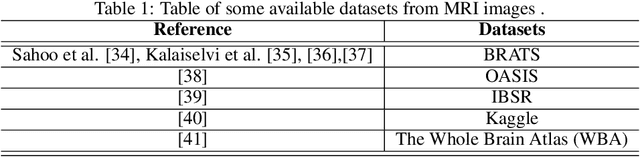
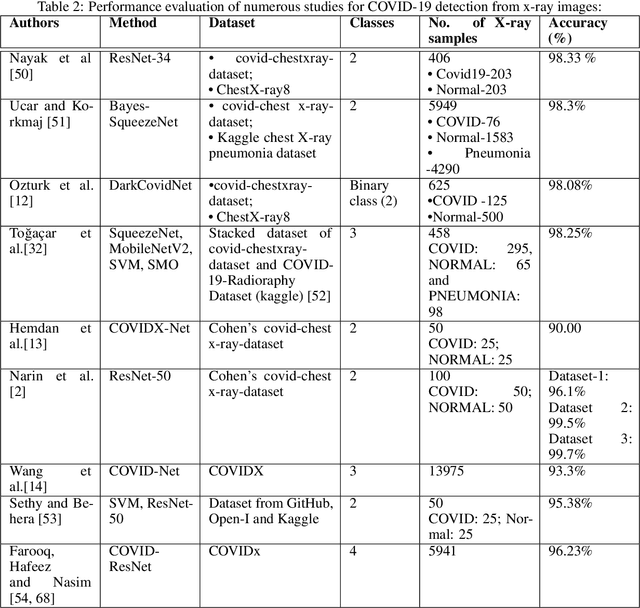
The field of medical imaging is an essential aspect of the medical sciences, involving various forms of radiation to capture images of the internal tissues and organs of the body. These images provide vital information for clinical diagnosis, and in this chapter, we will explore the use of X-ray, MRI, and nuclear imaging in detecting severe illnesses. However, manual evaluation and storage of these images can be a challenging and time-consuming process. To address this issue, artificial intelligence (AI)-based techniques, particularly deep learning (DL), have become increasingly popular for systematic feature extraction and classification from imaging modalities, thereby aiding doctors in making rapid and accurate diagnoses. In this review study, we will focus on how AI-based approaches, particularly the use of Convolutional Neural Networks (CNN), can assist in disease detection through medical imaging technology. CNN is a commonly used approach for image analysis due to its ability to extract features from raw input images, and as such, will be the primary area of discussion in this study. Therefore, we have considered CNN as our discussion area in this study to diagnose ailments using medical imaging technology.
UniControl: A Unified Diffusion Model for Controllable Visual Generation In the Wild
May 18, 2023
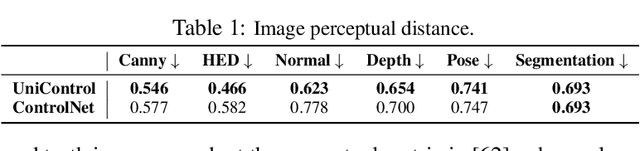
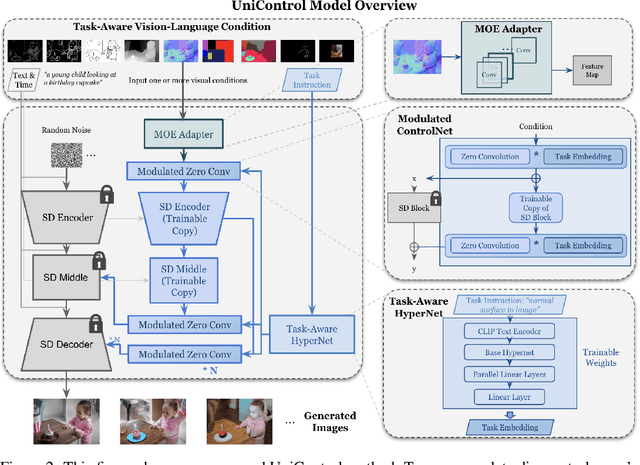

Achieving machine autonomy and human control often represent divergent objectives in the design of interactive AI systems. Visual generative foundation models such as Stable Diffusion show promise in navigating these goals, especially when prompted with arbitrary languages. However, they often fall short in generating images with spatial, structural, or geometric controls. The integration of such controls, which can accommodate various visual conditions in a single unified model, remains an unaddressed challenge. In response, we introduce UniControl, a new generative foundation model that consolidates a wide array of controllable condition-to-image (C2I) tasks within a singular framework, while still allowing for arbitrary language prompts. UniControl enables pixel-level-precise image generation, where visual conditions primarily influence the generated structures and language prompts guide the style and context. To equip UniControl with the capacity to handle diverse visual conditions, we augment pretrained text-to-image diffusion models and introduce a task-aware HyperNet to modulate the diffusion models, enabling the adaptation to different C2I tasks simultaneously. Trained on nine unique C2I tasks, UniControl demonstrates impressive zero-shot generation abilities with unseen visual conditions. Experimental results show that UniControl often surpasses the performance of single-task-controlled methods of comparable model sizes. This control versatility positions UniControl as a significant advancement in the realm of controllable visual generation.
OpenShape: Scaling Up 3D Shape Representation Towards Open-World Understanding
May 18, 2023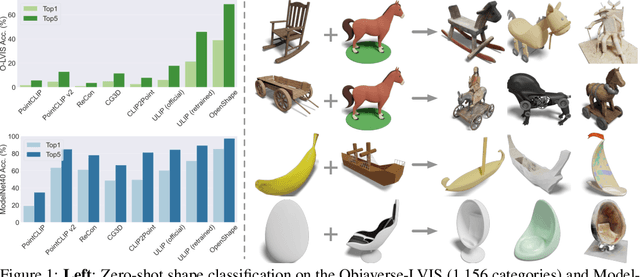
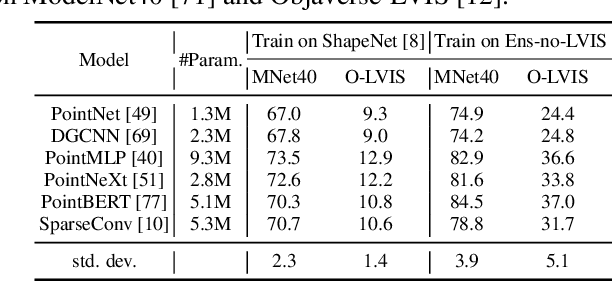
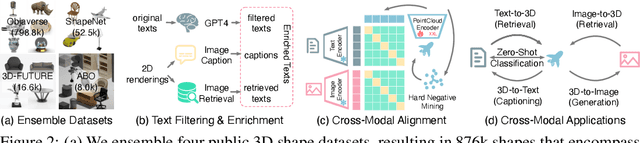
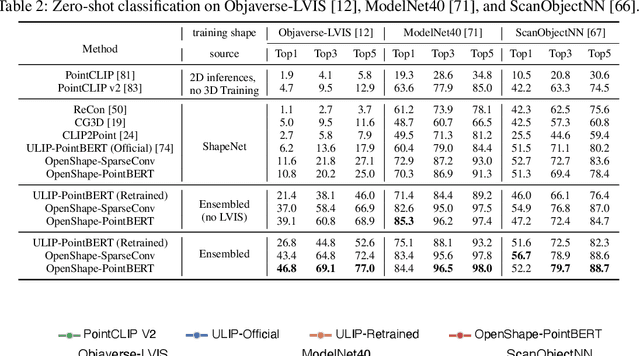
We introduce OpenShape, a method for learning multi-modal joint representations of text, image, and point clouds. We adopt the commonly used multi-modal contrastive learning framework for representation alignment, but with a specific focus on scaling up 3D representations to enable open-world 3D shape understanding. To achieve this, we scale up training data by ensembling multiple 3D datasets and propose several strategies to automatically filter and enrich noisy text descriptions. We also explore and compare strategies for scaling 3D backbone networks and introduce a novel hard negative mining module for more efficient training. We evaluate OpenShape on zero-shot 3D classification benchmarks and demonstrate its superior capabilities for open-world recognition. Specifically, OpenShape achieves a zero-shot accuracy of 46.8% on the 1,156-category Objaverse-LVIS benchmark, compared to less than 10% for existing methods. OpenShape also achieves an accuracy of 85.3% on ModelNet40, outperforming previous zero-shot baseline methods by 20% and performing on par with some fully-supervised methods. Furthermore, we show that our learned embeddings encode a wide range of visual and semantic concepts (e.g., subcategories, color, shape, style) and facilitate fine-grained text-3D and image-3D interactions. Due to their alignment with CLIP embeddings, our learned shape representations can also be integrated with off-the-shelf CLIP-based models for various applications, such as point cloud captioning and point cloud-conditioned image generation.
Fast and Painless Image Reconstruction in Deep Image Prior Subspaces
Feb 20, 2023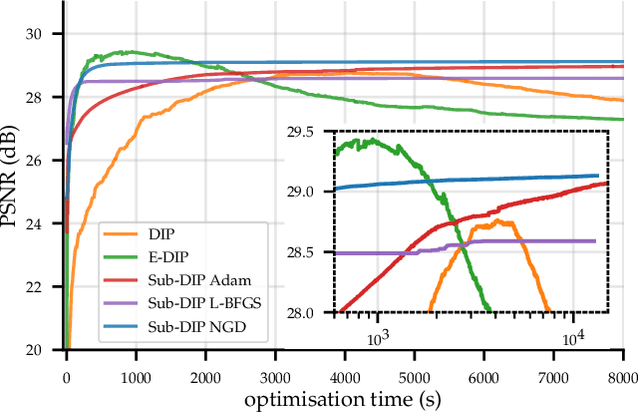
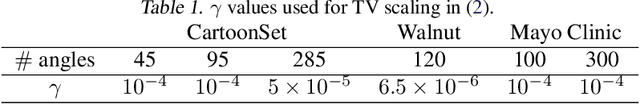
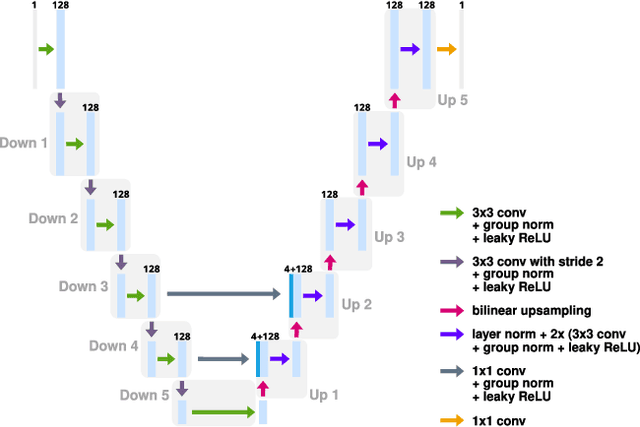
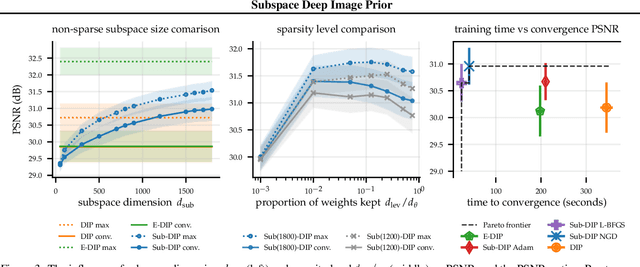
The deep image prior (DIP) is a state-of-the-art unsupervised approach for solving linear inverse problems in imaging. We address two key issues that have held back practical deployment of the DIP: the long computing time needed to train a separate deep network per reconstruction, and the susceptibility to overfitting due to a lack of robust early stopping strategies in the unsupervised setting. To this end, we restrict DIP optimisation to a sparse linear subspace of the full parameter space. We construct the subspace from the principal eigenspace of a set of parameter vectors sampled at equally spaced intervals during DIP pre-training on synthetic task-agnostic data. The low-dimensionality of the resulting subspace reduces DIP's capacity to fit noise and allows the use of fast second order optimisation methods, e.g., natural gradient descent or L-BFGS. Experiments across tomographic tasks of different geometry, ill-posedness and stopping criteria consistently show that second order optimisation in a subspace is Pareto-optimal in terms of optimisation time to reconstruction fidelity trade-off.
SAM3D: Zero-Shot 3D Object Detection via Segment Anything Model
Jun 04, 2023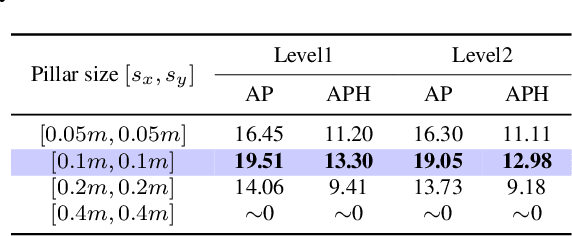
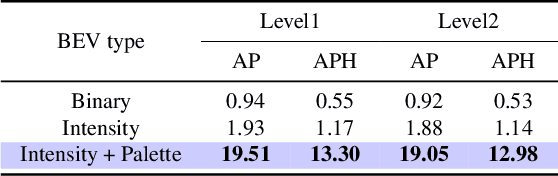
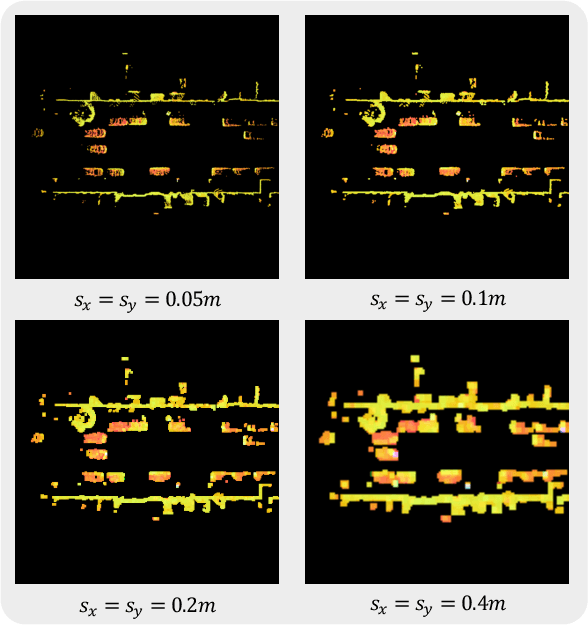
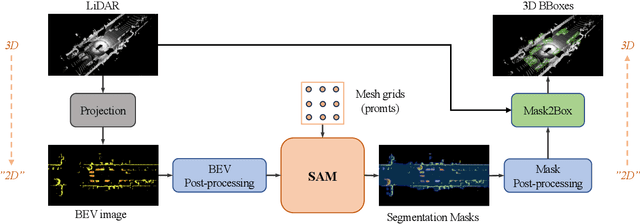


With the development of large language models, many remarkable linguistic systems like ChatGPT have thrived and achieved astonishing success on many tasks, showing the incredible power of foundation models. In the spirit of unleashing the capability of foundation models on vision tasks, the Segment Anything Model (SAM), a vision foundation model for image segmentation, has been proposed recently and presents strong zero-shot ability on many downstream 2D tasks. However, whether SAM can be adapted to 3D vision tasks has yet to be explored, especially 3D object detection. With this inspiration, we explore adapting the zero-shot ability of SAM to 3D object detection in this paper. We propose a SAM-powered BEV processing pipeline to detect objects and get promising results on the large-scale Waymo open dataset. As an early attempt, our method takes a step toward 3D object detection with vision foundation models and presents the opportunity to unleash their power on 3D vision tasks. The code is released at https://github.com/DYZhang09/SAM3D.
$IC^3$: Image Captioning by Committee Consensus
Feb 02, 2023
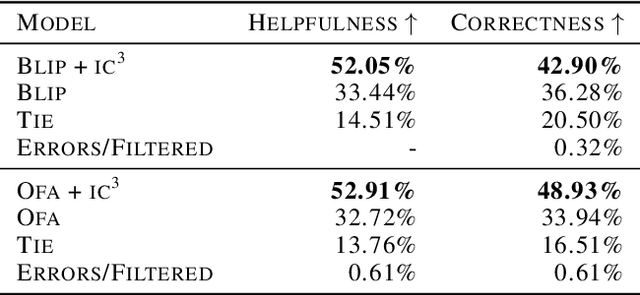
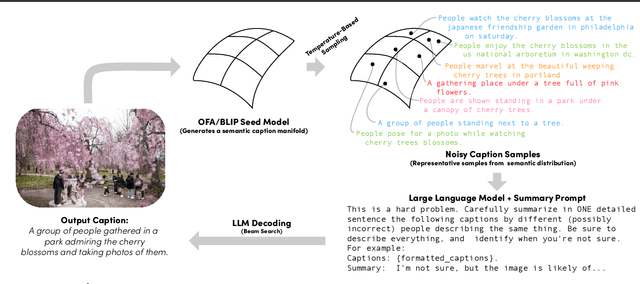
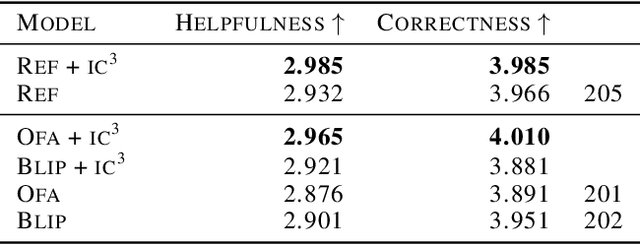
If you ask a human to describe an image, they might do so in a thousand different ways. Traditionally, image captioning models are trained to approximate the reference distribution of image captions, however, doing so encourages captions that are viewpoint-impoverished. Such captions often focus on only a subset of the possible details, while ignoring potentially useful information in the scene. In this work, we introduce a simple, yet novel, method: "Image Captioning by Committee Consensus" ($IC^3$), designed to generate a single caption that captures high-level details from several viewpoints. Notably, humans rate captions produced by $IC^3$ at least as helpful as baseline SOTA models more than two thirds of the time, and $IC^3$ captions can improve the performance of SOTA automated recall systems by up to 84%, indicating significant material improvements over existing SOTA approaches for visual description. Our code is publicly available at https://github.com/DavidMChan/caption-by-committee
Report of the Medical Image De-Identification (MIDI) Task Group -- Best Practices and Recommendations
Mar 18, 2023This report addresses the technical aspects of de-identification of medical images of human subjects and biospecimens, such that re-identification risk of ethical, moral, and legal concern is sufficiently reduced to allow unrestricted public sharing for any purpose, regardless of the jurisdiction of the source and distribution sites. All medical images, regardless of the mode of acquisition, are considered, though the primary emphasis is on those with accompanying data elements, especially those encoded in formats in which the data elements are embedded, particularly Digital Imaging and Communications in Medicine (DICOM). These images include image-like objects such as Segmentations, Parametric Maps, and Radiotherapy (RT) Dose objects. The scope also includes related non-image objects, such as RT Structure Sets, Plans and Dose Volume Histograms, Structured Reports, and Presentation States. Only de-identification of publicly released data is considered, and alternative approaches to privacy preservation, such as federated learning for artificial intelligence (AI) model development, are out of scope, as are issues of privacy leakage from AI model sharing. Only technical issues of public sharing are addressed.
Deep Representation Learning of Tissue Metabolome and Computed Tomography Images Annotates Non-invasive Classification and Prognosis Prediction of NSCLC
May 26, 2023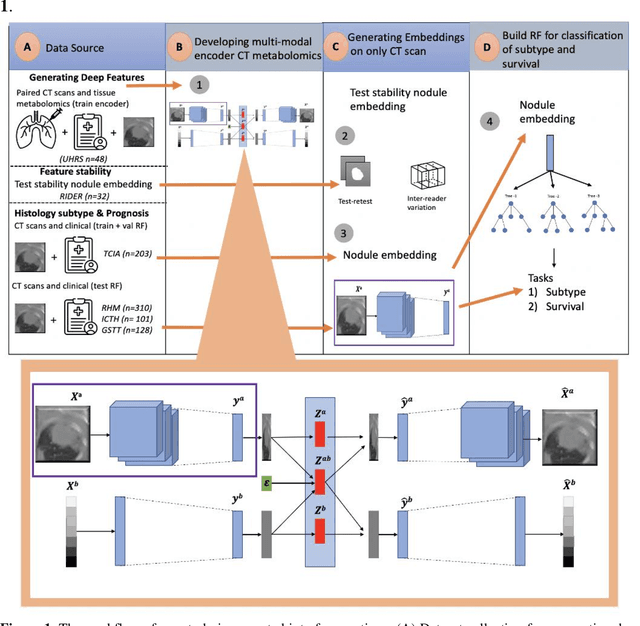
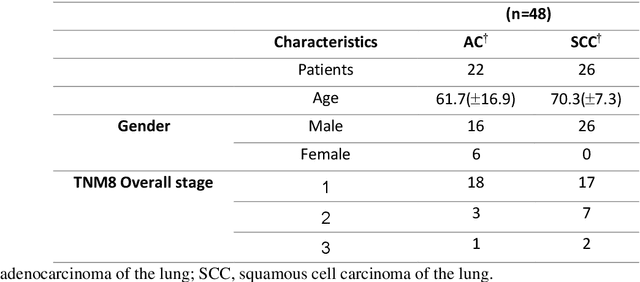

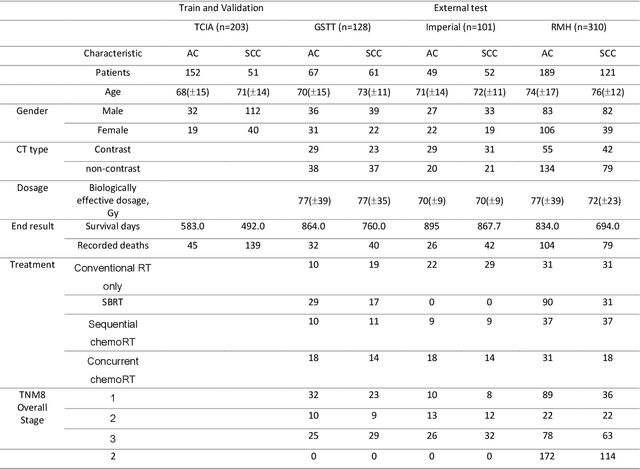
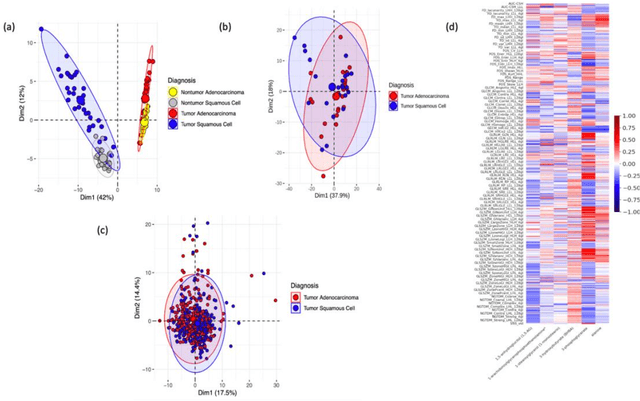
The rich chemical information from tissue metabolomics provides a powerful means to elaborate tissue physiology or tumor characteristics at cellular and tumor microenvironment levels. However, the process of obtaining such information requires invasive biopsies, is costly, and can delay clinical patient management. Conversely, computed tomography (CT) is a clinical standard of care but does not intuitively harbor histological or prognostic information. Furthermore, the ability to embed metabolome information into CT to subsequently use the learned representation for classification or prognosis has yet to be described. This study develops a deep learning-based framework -- tissue-metabolomic-radiomic-CT (TMR-CT) by combining 48 paired CT images and tumor/normal tissue metabolite intensities to generate ten image embeddings to infer metabolite-derived representation from CT alone. In clinical NSCLC settings, we ascertain whether TMR-CT achieves state-of-the-art results in solving histology classification/prognosis tasks in an unseen international CT dataset of 742 patients. TMR-CT non-invasively determines histological classes - adenocarcinoma/ squamous cell carcinoma with an F1-score=0.78 and further asserts patients' prognosis with a c-index=0.72, surpassing the performance of radiomics models and clinical features. Additionally, our work shows the potential to generate informative biology-inspired CT-led features to explore connections between hard-to-obtain tissue metabolic profiles and routine lesion-derived image data.
Interactive Data Synthesis for Systematic Vision Adaptation via LLMs-AIGCs Collaboration
May 22, 2023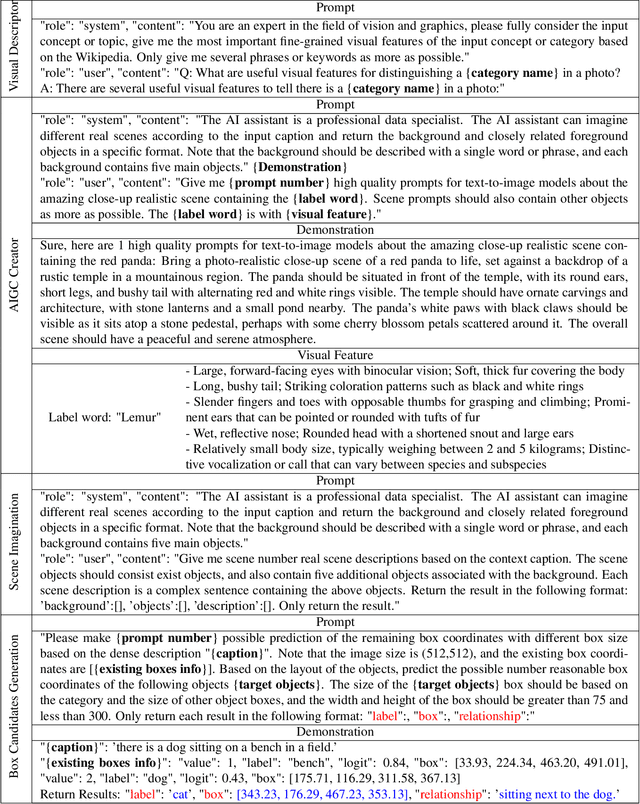
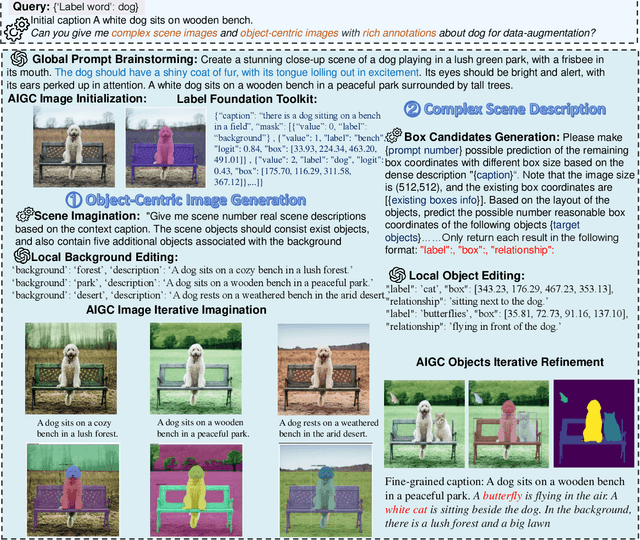
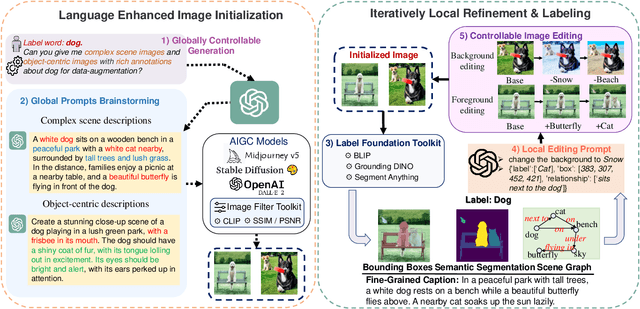
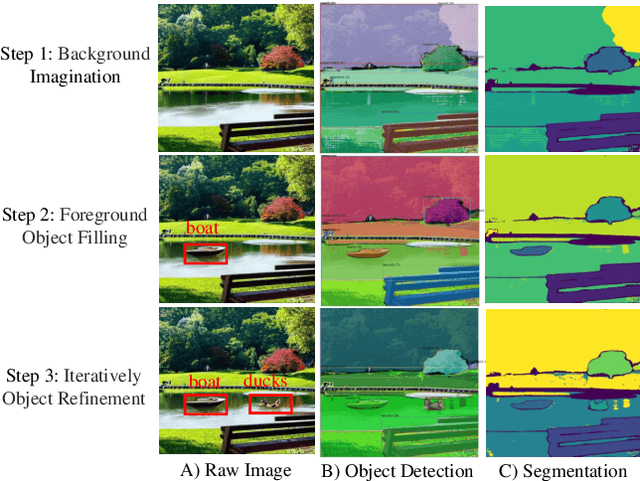

Recent text-to-image generation models have shown promising results in generating high-fidelity photo-realistic images. In parallel, the problem of data scarcity has brought a growing interest in employing AIGC technology for high-quality data expansion. However, this paradigm requires well-designed prompt engineering that cost-less data expansion and labeling remain under-explored. Inspired by LLM's powerful capability in task guidance, we propose a new paradigm of annotated data expansion named as ChatGenImage. The core idea behind it is to leverage the complementary strengths of diverse models to establish a highly effective and user-friendly pipeline for interactive data augmentation. In this work, we extensively study how LLMs communicate with AIGC model to achieve more controllable image generation and make the first attempt to collaborate them for automatic data augmentation for a variety of downstream tasks. Finally, we present fascinating results obtained from our ChatGenImage framework and demonstrate the powerful potential of our synthetic data for systematic vision adaptation. Our codes are available at https://github.com/Yuqifan1117/Labal-Anything-Pipeline.
Cycle Consistency-based Uncertainty Quantification of Neural Networks in Inverse Imaging Problems
May 22, 2023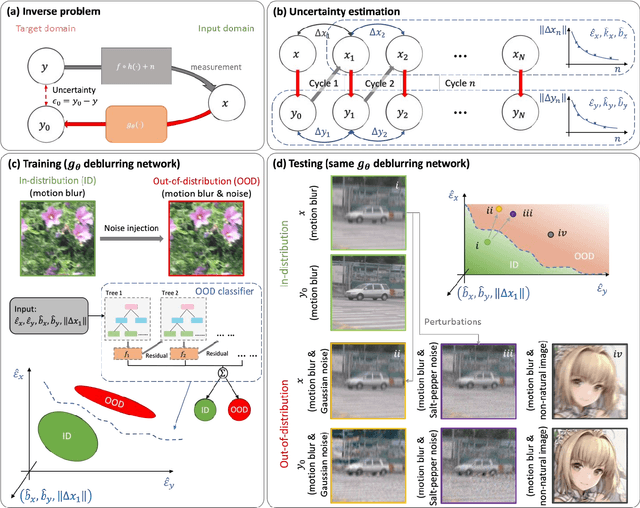
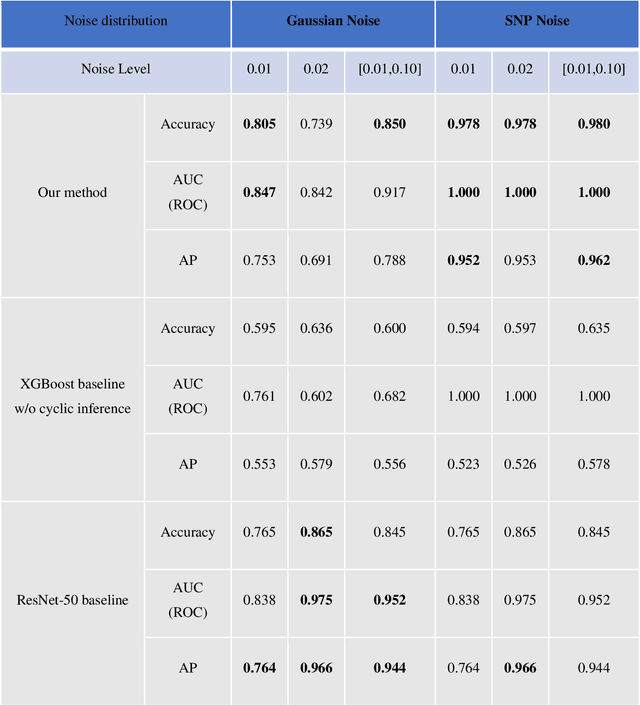
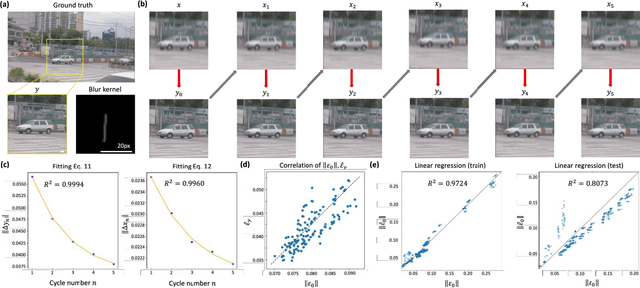
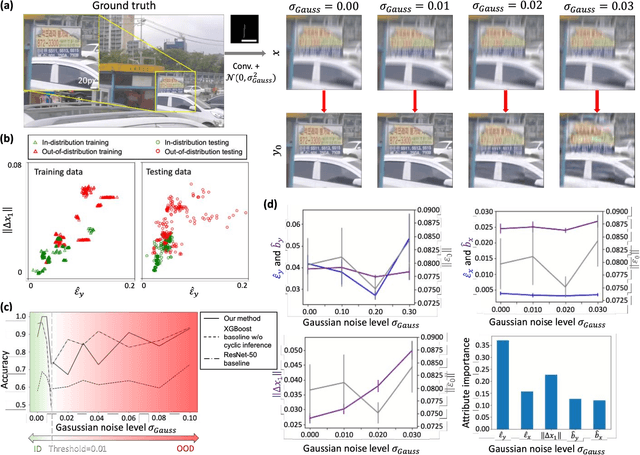
Uncertainty estimation is critical for numerous applications of deep neural networks and draws growing attention from researchers. Here, we demonstrate an uncertainty quantification approach for deep neural networks used in inverse problems based on cycle consistency. We build forward-backward cycles using the physical forward model available and a trained deep neural network solving the inverse problem at hand, and accordingly derive uncertainty estimators through regression analysis on the consistency of these forward-backward cycles. We theoretically analyze cycle consistency metrics and derive their relationship with respect to uncertainty, bias, and robustness of the neural network inference. To demonstrate the effectiveness of these cycle consistency-based uncertainty estimators, we classified corrupted and out-of-distribution input image data using some of the widely used image deblurring and super-resolution neural networks as testbeds. The blind testing of our method outperformed other models in identifying unseen input data corruption and distribution shifts. This work provides a simple-to-implement and rapid uncertainty quantification method that can be universally applied to various neural networks used for solving inverse problems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge